Abstract
The inducible Di-Cre system was used to delete the putative ubiquitin-conjugating enzyme 13 gene (ubc13) of Plasmodium falciparum to study its role in ubiquitylation and the functional consequence during the parasite asexual blood stage. Deletion resulted in a significant reduction of parasite growth in vitro, reduced ubiquitylation of the Lys63 residue of ubiquitin attached to protein substrates, and an increased sensitivity of the parasite to both the mutagen, methyl methanesulfonate and the antimalarial drug dihydroartemisinin (DHA), but not chloroquine. The parasite was also sensitive to the UBC13 inhibitor NSC697923. The data suggest that this gene does code for an ubiquitin conjugating enzyme responsible for K63 ubiquitylation, which is important in DNA repair pathways as was previously demonstrated in other organisms. The increased parasite sensitivity to DHA in the absence of ubc13 function indicates that DHA may act primarily through this pathway and that inhibitors of UBC13 may both enhance the efficacy of this antimalarial drug and directly inhibit parasite growth.
Similar content being viewed by others
Introduction
Plasmodium falciparum, the parasite responsible for the most severe malaria and one of the leading causes of infant mortality in Africa, has a complex multi-stage life cycle, with sexual reproduction in the Anopheles mosquito and asexual proliferative stages within hepatocytes and red blood cells (RBCs) of a human host. During the asexual blood phase of its life cycle, every ~ 48 h the parasite invades new RBCs, grows and multiplies within them, developing through ring, trophozoite and schizont stages to produce extracellular merozoites that invade new RBCs. Because of its high replication rate and the accompanying rapid cell transformations, it is likely that the reuse of resources, the control of protein quality, and accurate DNA synthesis machinery are important features of progression through the life cycle1.
The Ubiquitin–Proteasome System (UPS) is a highly regulated cellular mechanism of protein turnover with critical roles in protein degradation, signal transduction, cell cycle progression, transcriptional regulation and DNA synthesis and repair2. Protein ubiquitylation is a cascade of reactions carried out by three classes of enzymes: ubiquitin-activating enzymes (E1s), ubiquitin-conjugating enzymes (E2s) and ubiquitin ligases (E3s). E1 uses AMP to activate the C-terminus of ubiquitin, forming a covalent thioester to an E1 cysteine, and then in a transthiolation reaction transfers the activated ubiquitin to a cysteine side chain of an E2. Either directly, or facilitated by the involvement of an E3, the ubiquitin is transferred again and covalently conjugated to specific protein substrates, usually through a lysine residue. However, some E2s can transfer ubiquitin directly to the substrate N-terminal amino group or to Lys, Cys, Ser and Thr side chains, thereby playing a role in where and how a target is modified to form diverse ubiquitin linkages3,4. Polyubiquitin (poly-Ub) chains are often added to substrates; these chains are formed by ubiquitylation of ubiquitin through one of its seven lysine residues (K6, K11, K27, K29, K33, K48, and K63)5. The nature of this linkage in poly-Ub chains is very important for both proteasome-dependent and proteasome-independent functions. For example, in well-studied model cells, poly-Ub chains containing K48 and K11 linkages target substrate proteins for degradation by the 26S proteasome, while poly-Ub chains containing K63 linkages have other roles in a variety of cellular processes, including DNA repair, kinase activation, and vesicle trafficking5,6.
In the P. falciparum genome, many genes are annotated as coding for UPS–related proteins7,8, including ubiquitin-activating enzyme, PfUba1, (PF3D7_1225800) and at least eight potential E2s. One of these E2 genes (PF3D7_0527100) encodes putative ubiquitin-conjugating enzyme 13 (UBC13), which catalyses the formation of K63-linked polyubiquitin chains, as shown in a variety of organisms, including human (Homo sapiens [Hs]), rodent (Mus musculus), yeast (Saccharomyces cerevisiae [Sc]), fly (Drosophila melanogaster), plant (Arabidopsis thaliana), worm (Caenorhabditis elegans) and trypanosome (Leishmania spp.)9,10,11,12,13,14,15,16. UBC13 functions as part of a complex with an E2-related protein that lacks the active site cysteine, and is called Ubs variant (UEV). Different UEVs such as MMS2 or UEV1A function together with UBC13 to catalyze polyubiquitylation in different cellular processes. In P. falciparum, there is one E2-like protein (encoded by PF3D7_0305700) lacking the cysteine and with high similarity to ScMMS2. The mutation of ubc13 and mms2 genes in S. cerevisiae results in greater sensitivity than wild type cells to UV exposure and also greater sensitivity to treatment with the alkylating agent methyl methanesulfonate (MMS) that induces DNA damage. ubc13 and mms2 genes are epistatic and belong to the same DNA-repair pathway12. In Arabidopsis, mutations of UBC13 result in increased sensitivity to stress conditions, such as high salt, oxidative stress, and treatment with an abscisic acid (ABA) analogue; UBC13 (and two other E2s, UBC7 and UBC14) are important in the response of plants to multiple stress conditions17. HsUBC13 functions together with its two cofactors, Uev1A (UBE2V1) and Mms2 (UBEV2), to promote the formation of K63 poly-Ub chains, which leads to either NF-κB activation or recruitment of repair proteins to DNA lesions, respectively9,18,19. UBC13 also regulates other cellular processes, including nuclear localization of the tumour suppressor p53 protein and MAPK activation20,21. However, neither UBC13 nor UEV alone can promote K63-linked poly-Ub chain formation22,23.
PfUBC13 (PF3D7_0527100) is a 152-residue protein that was first identified as a substrate of the P. falciparum Protein Kinase 9 (PfPK9), an essential kinase of P. falciparum that does not appear to be a member of any known eukaryotic kinase family24,25,26. It was reported that Ser106 of PfUBC13 is phosphorylated by PfPK9, resulting in suppression of UBC13 activity27. However, in contrast, inhibition of PfPK9 decreased PfUBC13 activity and resulted in the reduction of K63-linked polyubiquitylation24. Both of these studies suggest that phosphorylation of PfUBC13 is important in controlling its function. The structures of recombinant putative PfUBC13 monomer and PfUBC13 in complex with Pf MMS2/UEV1 (PF3D7_0305700) have been determined (https://www.rcsb.org/structure/2r0j and https://www.rcsb.org/structure/3e95, respectively). NSC697923 (2-[(4-methylphenyl)sulfonyl]-5-nitrofuran) is a small molecule covalent inhibitor of UBC1328, which also inhibits the growth of P. falciparum, in vitro29, although its mode of action against the parasite has not been established. Since the putative Pfubc13 is essential for P. falciparum asexual blood stage development26, it is possible that this protein might be a new target for anti-malarial drug development.
In this study, we aimed to identify the role of UBC13 in P. falciparum using an inducible DiCre-recombinase-based gene knockout approach. Following Pfubc13 inducible gene knockout (iKO) there was decreased K63 linked poly-ubiquitylation and a growth defect. Furthermore the ubc13-iKO parasite showed higher sensitivity to MMS and to dihydroartemisinin (DHA), an antimalarial drug that also affects DNA repair30,31. Parasite treatment with NSC697923 caused a growth defect similar to that of the ubc13-iKO. These results suggest that Pfubc13 is involved in K63 polyubiquitylation, is essential for the proper growth of P. falciparum and is important for the correct function of the DNA repair system. Inhibitors of UBC13 are likely to inhibit parasite growth and also enhance the activity of DHA and similar artemisinin-derived compounds against the parasite.
Results
Comparison of the putative P. falciparum UBC13 with the human enzyme
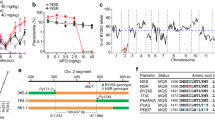
An alignment of the HsUBC13 and putative PfUBC13 revealed that the 152-residue protein coded by PF3D7_0527100 is orthologous to HsUBC13 with 68% amino acid sequence identity (Fig. 1a). Most of the sequence differences are located away from the active site cysteine (Fig. 1b).
Structural comparison of HsUBC13 and the putative PfUBC13. (a) Homo sapiens (Hs)UBC13 and Plasmodium falciparum (Pf)UBC13 amino acid sequences are aligned, with identical residues in the two sequences highlighted in black. The active site cysteine residue that binds activated ubiquitin covalently and reacts with NSC697923 in human UBC13 is highlighted in yellow and marked with an asterisk. (b) Three- dimensional structures with the active site cysteine highlighted in yellow and marked with an asterisk. In the PfUBC13 structure, amino acid residues where the two sequences differ are highlighted in blue, or in cyan if the substitution is conservative.
Generation of an inducible ubc13 knockout in P. falciparum
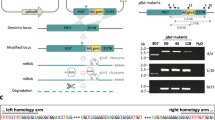
We used CRISPR-Cas9 to modify the 3’ end of the putative ubc13 gene (PF3D7_0527100), inserting a loxP element in the fourth and final intron, inserting a recodonised final exon and placing a loxP site after the stop codon in the ubc13 sequence (Supplementary Fig. 1a). The changes were made in the II-3 parasite line, which expresses functional DiCre recombinase activity following rapamycin addition32. After transfection, the parasite population was screened by PCR for DNA integration using diagnostic primer pairs P1/P2, P1/P4, P3/P4, P1/P6 and P5/P4 (Supplementary Fig. 1b, Supplementary Table S1) and products from both modified and unmodified locus were detected (Supplementary Fig. 1c). Parasites were cloned by limiting dilution and integration was examined in individual clones; for recombinant clones, primers P1/P6 and P5/P4, produced bands of 589 and 550 bp, respectively, diagnostic of integration and no product was obtained from primer pairs P1/P2, and P3/P4 indicating the absence of unmodified locus (Supplementary Fig. 1c). The parasite clone 2E was used in subsequent experiments.
Induced excision of ubc13 shows that the gene is essential for the growth and survival of P. falciparum asexual blood stage
To truncate Pfubc13 and abolish expression of functional protein, we used a synchronized population of cloned parasites and at 4 h after invasion treated it with rapamycin or with DMSO alone (the control) for 24 h. Excision of the floxed DNA was monitored by PCR amplification with primer pair P1 and P4 (Supplementary Fig. 1d). For the control (DMSO-treated) sample a 1280 bp band was amplified, indicating no excision, and from the rapamycin-treated sample a 975 bp band was observed, indicating quantitative excision of the floxed DNA (Supplementary Fig. 1e). This result indicates that by the late trophozoite stage at 28 h after invasion, only a truncated ubc13 gene remained following rapamycin-treatment.
We next investigated whether functional ubc13 is essential for parasite growth and survival. Parasites were treated at 2 h post invasion with rapamycin or DMSO for 24 h and then further incubated for either one (48 h) or two (96 h) cell cycles and analysed by flow cytometry (FACS) and examination of Giemsa-stained thin blood smears by microscopy. At 48 h the parasitemia had increased in both samples (Fig. 2a) but was significantly less (p < 0.008) following rapamycin treatment, with fewer new ring-stage forms resulting from merozoite release and reinvasion (Fig. 2b). There was a clear difference in the parasitemia of the control and rapamycin-treated parasites at the second cycle (p < 0.0002) (Fig. 2a). At 96 h most of the parasites were at the ring stage of development in the control cultures, whereas the rapamycin-treated parasites were largely at the schizont stage, and looking very unhealthy on Giemsa-stained smears. Little increase in parasitemia had occurred overall (Fig. 2b).
Effect of an inducible Pfubc13 knockout in P. falciparum on parasite growth and ubiquitin ubiquitylation (a) Growth over two cycles of intraerythrocytic development for parasites treated with either DMSO (control) or rapamycin to induce Di-Cre mediated excision within ubc13. A synchronised parasite population (0.2% parasitemia) was treated with either DMSO or rapamycin for 24 h; parasitemia was measured by FACS analysis. Data are shown as mean ± standard error of the mean (SEM) from triplicate experiments performed in duplicate. (b) Analysis of parasites by light microscopy of Giemsa-stained thin smears, from parasites treated with either DMSO or rapamycin. At 48 h post invasion (PI), schizonts and ring stages were visible in DMSO treated cultures and there were fewer ring stages in the rapamycin treated cultures. By 96 h PI, second cycle ring stages were visible in DMSO-treated cultures and morphologically abnormal late stage parasites were predominant in rapamycin-treated cultures. Scale bar is 5 µm. (c) Parasitemia over up to 5 cycles of intraerythrocytic growth. The control (DMSO-treated) parasites were monitored for only two cycles; starting parasitemia, 0.3%. The rapamycin-treated parasites were monitored over five cycles; starting parasitema, 0.3 and 1%. The data shown are mean ± SEM from duplicate experiments. (d) The parasite populations were monitored by PCR amplification with primers P1/P4 (see Supplementary Fig. 1) at cycle 3 and cycle 5; no parasites without excision were detected in the rapamycin treated sample. (e) Analysis of schizont lysates by immunoblotting with antibodies specific for either Lys48 or Lys63 ubiquitylation of ubiquitin. Equal amounts of protein from rapamycin-treated or –untreated parasites at 44 h PI were resolved by SDS-PAGE, transferred to nitrocellulose and then probed with antibodies specific for either K48 or K63 ubiquitin linkages, or antibodies specific for BiP as a loading control. Full length gels and immunoblots are presented in Supplementary Figs. 3 and 4.
Further development of the rapamycin-treated parasite was monitored for up to five cycles (Fig. 2c). Parasites were still detectable in the culture although the parasitemia gradually declined. Analysis by PCR of DNA collected at cycles 3 and 5 confirmed that parasites contained the truncated gene, rather than representing parasites in which the gene was still intact (Fig. 2d). After the second cycle the rapamycin-treated parasites lost synchrony of development and some visible abnormalities were present in late stages, such as pyknotic and deformed cells (Supplementary Fig. 2). These results indicate that ubc13 is not absolutely essential for completion of the first growth cycle following rapamycin treatment, perhaps because of residual active UBC13 protein. However, for the second and subsequent cycles the absence of growth and decrease in parasitemia, coupled with the abnormal morphology of many parasites, indicate that UBC13 is essential for asexual blood stage growth of P. falciparum.
ubc13 deletion affects K63 polyubiquitylation in the parasite
We examined whether or not deletion of the putative ubc13 affected polyubiquitylation in the parasite. We used two antibodies, one specific for ubiquitin ubiquitylation at lysine-48 (K48) and the other for ubiquitin ubiquitylation at lysine-63 (K63). Parasites were treated with rapamycin or DMSO as described above and then extracts were prepared from schizonts for western blotting with the two antibodies (Fig. 2e). In a comparison of treated and control parasites, there was no clear difference or a slight increase in the pattern of K48-linked ubiquitylation, but the level of K63-linked ubiquitylation was clearly diminished in the rapamycin-treated cells. In the control-treated cell lysate, six discrete bands were clearly visible, with molecular masses in the range of 32 to ~ 150 kDa and the intensity of at least five of these was reduced. These results indicate that the putative UBC13 is largely responsible for synthesis of K63-linked ubiquitin ubiquitylation in blood stage parasites of P. falciparum, and suggest that the PF3D7_0527100 gene product is a functional homologue of UBC13, which is responsible for K63-linked ubiquitylation in human cells.
ubc13 deletion increases parasite sensitivity to DNA damage induced by treatment with methyl methanesulfonate
In human cells UBC13 plays an important role in DNA damage repair19,22 and we were interested to examine whether this protein has a similar function in P. falciparum. Therefore, we induced DNA damage in the parasite following rapamycin treatment, by incubation with MMS, a mutagen that alkylates DNA bases and induces single and double-strand breaks, and which has been shown to cause DNA damage in P. falciparum30,33. Parasites were exposed for different periods to either 2.25 mM or 4.5 mM MMS, washed to remove the drug and then cultured further to 48 h after initial invasion, at which point parasite survival was measured. The rapamycin-induced ubc13-iKO parasite was more sensitive to MMS than the DMSO-treated parasite. This was particularly evident at 2.25 mM MMS, since at the higher MMS concentration 40 min or more exposure resulted in no survival of either parasite population (Fig. 3a).
Deletion of Pfubc13 renders parasites more susceptible to the mutagen, methyl methanesulfonate (MMS) and dihydroartemisinin (DHA), but not chloroquine (CQ). (a) Synchronised parasite populations (5 to 8 h post invasion [PI], 1% parasitemia and 3% hematocrit) were pre-treated with DMSO (control) or rapamycin to induce ubc13 excision for 24 h, and then treated with 2.25 mM or 4.5 mM MMS for 0, 20, 40, 60 and 100 min. Parasite survival was assessed at 48 h. The data shown are mean ± SEM from duplicate experiments. (b) Long term survival of parasites treated with MMS. Synchronized parasite populations (5 h PI, 0.5% parasitemia and 3% hematocrit) were treated with rapamycin or DMSO and with or without 500 µM MMS. The cells were then washed to remove the drugs 48 h after the start of the experiment and the incubation was continued in complete medium. Giemsa-stained thin blood smears were examined at the time points indicated to determine percentage parasitemia. The data shown are mean ± SEM from triplicate experiments. (c) Parasites were treated with rapamycin or DMSO and increasing concentrations of either DHA or CQ. The percentage parasite survival was measured, allowing the EC50 of the drugs in each condition to be calculated. The data shown are mean ± SEM from triplicate experiments The DHA EC50 was 1.69 nM (95% CI: 1.41 to 1.99) and 4.1 nM (95% CI: 3.63 to 4.63) for the rapamycin and DMSO treated parasites, respectively, which is significantly different (P < 0.005, t-test), whereas there was no significant difference in the CQ EC50 (18.51 [95% CI: 16.05 to 20.20] and 20.19 [95% CI: 18.53 to 21.48]) for the two treatments.
The effect of adding rapamycin and a lower concentration of MMS together in the first intracellular cycle was also investigated. Parasites were treated with and without 20 nM rapamycin and with and without 500 µM MMS for 48 h and then parasitemia was monitored for up to four cycles of growth. The parasites not treated with rapamycin were severely inhibited by MMS but started to recover in cycle 3 at 144 h. In contrast the rapamycin and MMS treated parasites were killed and no recovery over the period of the experiment was observed (Fig. 3b).
Pfubc13 deletion increases parasite sensitivity to dihydroartemisin (DHA) but not chloroquine (CQ)
DHA generates oxidative free radicals that can cause DNA damage30. In contrast, CQ is not known to act in this way. Therefore, we examined parasite sensitivity to DHA and CQ, in mock treated or rapamycin-treated ubc13 inducible knockout parasites. The results (Fig. 3c) revealed that the ubc13-iKO parasite is more sensitive to DHA than the control parasite, with a significantly decreased DHA IC50 (1.69 nM and 4.1 nM for the rapamycin and DMSO treated parasites; P < 0.005). In contrast, the CQ IC50 was not affected by ubc13 deletion (18.51 and 20.19 for the two treatments). These results suggest that PfUBC13 functions, at least in part, in the DNA damage repair system.
Treatment of parasites with NSC697923 mimics the induced ubc13 knockout
NSC697937 is a UBC13 inhibitor in human cells and forms a covalent adduct with the active site cysteine28. Since NSC697937 has been shown to inhibit the asexual blood stage development of P. falciparum 29, we wished to compare the outcome of rapamycin and NSC697923 treatment on the growth of blood stage parasites. Assessment of the percent parasitemia by FACS at the start of the experiment (2 h post invasion) and at time points corresponding to the first cycle (50 hpi) and second cycle (98 hpi) showed that the treated parasites showed significantly reduced parasite numbers (Fig. 4a; DMSO treatment compared to either rapamycin or NSC697937 treatments, p < 0.0001). Giemsa stained thin blood smears were prepared to examine parasite development and morphology after treatment. Both NSC697923 and rapamycin treatment resulted in a growth defect. Both treatments resulted in slowed growth; for example, at 49 h post-invasion, treated parasites with either treatment were largely at the schizont stage, whereas the control parasites were predominantly at the ring stage (Fig. 4b).
Both NSC697923 and rapamycin treatment affect parasite growth. (a) Parasite growth over two cycles of intraerythrocytic development for DMSO-, rapamycin- or NSC697923-treated parasites. Synchronised parasite populations (0.2% parasitemia) were treated with either DMSO (control), rapamycin (to induce excision within ubc13) or NSC697923 (to inhibit UBC13), and parasitemia was measured for two cycles by FACS analysis. The data shown are mean ± SEM from triplicate experiments. The parasitemia in both rapamycin and NSC697923-treated parasite culture was significantly lower than that of the control at both cycle 1 and cycle 2. (** p < 0.001) (b) Analysis by microscopy of Giemsa-stained parasites at 24, 44, 49 and 69 h post-invasion. All parasites develop to schizonts but further development of both rapamycin- and NSC697923-treated parasites is delayed: at 49 and 69 h PI control parasites have formed new ring stage and trophozoite stages, whereas both rapamycin and NSC697923-treated parasites are largely schizont and early ring stages at these two time points. Scale bar is 5 µm.
Discussion
Human UBC13 (HsUBC13) specifically catalyzes the formation of K63-linked polyubiquitin chains, which are important in, for example, cytoplasmic NF-κB signaling and the DNA repair system. In P. falciparum, the role of UBC13 is unclear although PF3D7_0527100 appears to be an essential gene26 and the protein’s function is controlled by the activity of PK924,27. In this study, we generated a transgenic parasite to allow the inducible knockout of this gene using the DiCre recombinase strategy, and examined the resultant phenotype.
The structural alignment of the PfUBC13 and HsUBC13, and the fact that deletion of Pfubc13 affected K63-linked ubiquitylation in the parasite, indicate that the protein’s catalytic function mirrors that of UBC13 in higher eukaryotes. These data, together with crystal structure information for PfUBC13 (https://www.rcsb.org/structure/2r0j) and PfUBC13 complexed with PfUEV1a (https://www.rcsb.org/structure/3e95) indicate that PF3D7_0527100 is a homolog of UBC13 in higher eukaryotes. This is the first demonstration of the potential role of PfUBC13 in K63-linked polyubiquitylation in the parasite because the previous studies focused on PK9 and PK9 inhibitors24,27. The six visible K63 ubiquitylated substrates ranged in size from 32 to ~ 150 kDa with no evidence of heterogeneity, suggesting that the ubiquitylation is either a chain of just two or a small discrete number of ubiquitin units. These substrates remain to be identified, although they are likely present in the experimentally determined trophozoite/schizont ubiquitomes29. Interestingly, K63-polyubiquitylation of histone H2A34 and proliferating cell nuclear antigen (PCNA)35 is important in DNA repair processes, and both these proteins are present in the P. falciparum ubiquitome. In future work it may be possible to identify the proteins directly following their affinity purification with K63 polyubiquitin-specific antibody. Their identification may illuminate further the biological processes in which this modification is involved in the parasite. The slight increase in K48 ubiquitylation in rapamycin treated cells may reflect a loss of proteostasis, leading to enhanced ubiquitylation of proteins destined for degradation in the proteasome.
Rapamycin-induced Pfubc13 deletion affected parasite growth severely. After one cycle the number of new ring stage parasites was reduced significantly but not completely; and this effect was amplified at the second cycle and beyond, although morphologically intact parasites were still present in the culture at cycle 5. These parasites did not have an intact ubc13 gene and some visible abnormalities were present in late stages, such as pyknotic and deformed cells. Some parasites could develop from ring to schizont stage, and apparently survive to the next cycle although growth was severely retarded. Parasite largely failed to develop into viable late stages, which is when DNA replication, nuclear division and merozoite differentiation occur, suggesting a role of Pfubc13 in maintaining normal DNA replication.
K63-linked polyubiquitylation has been identified to play a role in DNA repair, signal transduction, and kinase activation in eukaryotes5,6,34, while K48-linked polyubiquitylation targets protein substrates to the proteasome. Therefore, we chose to examine the importance of PfUBC13 in DNA repair processes for P. falciparum. In Plasmodium spp., DNA damage has been studied experimentally and can result from exposure to UV irradiation; ionizing radiation like X-rays and gamma rays; chemical mutagens such as alkylating agents (e.g., methyl methanesulfonate and cisplatin) and also exposure to oxidative free radicals33,36. The damage includes inter-strand cross-linking, DNA strand breaks, and inhibition of DNA synthesis and replication, leading to cell cycle arrest, and parasite death37. Genome integrity is maintained in the parasite by repair mechanisms, for example, homologous recombinant repair (HR), mismatch repair (MMR), nucleotide excision repair (NER) and base excision repair (BER). Canonical non-homologous end joining (C-NHEJ) repair is absent in Plasmodium, but an alternative inefficient NHEJ pathway has been described38,39.
We chose to use the DNA alkylating agent, MMS, which predominantly modifies guanine to N7-methyl guanine (7meG) and O6-methyl guanine (O6meG), to study the effect of ubc13 deletion on DNA repair. These lesions can lead to the collapse of the replication fork and subsequent induction of DNA double-strand breaks40 that are commonly repaired by BER, NER and MMR processes37,41. Up-regulation of genes that function directly in DNA repair mechanisms such as PfRAD51 and PfRAD54, and changes in histone modification were reported after MMS treatment of P. falciparum30,42. These changes in chromatin modulate the access of repair factors to the damage site resulting in DNA repair43. The up-regulation of Pfubc13 transcription in the MMS-treated parasite has not been reported30. However, our results show that the Pfubc13-inducible knockout parasite has greater sensitivity to MMS treatment, and fails to survive under conditions in which the intact parasite recovers. These results are consistent with a major role for UBC13 and K63-polyubiquitylation in DNA repair in P. falciparum.
DHA is the active metabolite of artemisinin and related compounds, and its mode of action against the malaria parasite is complex44,45. One property, following activation, is its ability to alkylate macromolecules, and recent studies have suggested that one cellular process where this is important is in DNA damage and repair30,31. The Pfubc13-inducible knockout parasite had increased sensitivity to DHA, with a significantly decreased IC50, and showing a similar response profile to that with MMS, while CQ did not show this effect. This suggests that DHA is acting at least in part through mechanisms where repair is dependent on UBC13 and K63 ubiquitylation.
NSC697923 is an inhibitor of HsUBC13 function, reacting with the sulfhydryl group of the active site Cys87 to form a 5-nitrofuran adduct28. It has activity against P. falciparum growth in vitro29, and therefore it was of interest to compare mock and rapamycin-treated Pfubc13- inducible knockout parasites to parasites treated with this drug. Both treatments produced a similar result with a similar delay in parasite development. Although NSC697923 is not a potent inhibitor, the results suggest that if the primary target of NSC697923 in the parasite is UBC13 then better compounds with higher affinity, specificity, and a good therapeutic index, could be potential antimalarial compounds. Furthermore, we would expect that inhibitors of PfUBC13 could be used in combination to enhance the efficacy of artemisinin drugs.
In conclusion, our results demonstrate that the PF3D7_0527100 gene product is PfUBC13, a functional homologue of HsUBC13, which is responsible for K63-linked ubiquitylation and is involved in DNA repair systems. Deletion of this gene results in a P. falciparum growth defect and increases the sensitivity of the parasite to agents that cause DNA damage, in particular MMS and DHA.
Materials and methods
Bioinformatics
HsUBC13 and PfUBC13 sequences were obtained from the Uniprot (https://www.uniprot.org/uniprot/P61088) and PlasmoDB (https://plasmodb.org/plasmo/app/record/gene/PF3D7_0527100) databases, respectively. The amino acid sequences were aligned using the Protein BLAST alignment tool (https://blast.ncbi.nlm.nih.gov). Three-dimensional structures were generated with Phyre246.
In vitro culture and synchronisation of P. falciparum
The inducible DiCre recombinase-expressing P. falciparum line II-332 was cultured in RPMI 1640 medium supplemented with 1% (w/v) Albumax at 37˚C in gassed (5% CO2, 5% O2, 90% N2) flasks. The parasite population was synchronized by schizont centrifugation onto a 63% Percoll cushion. Purified schizonts were cultured for 2 to 3 h to allow merozoite invasion of fresh red blood cells and ring stage formation, and then residual schizonts were lysed by treatment with 5% D-sorbitol for 10 min. RBCs were purchased from the United Kingdom National Health Service Blood and Transplant (NHSBT).
Generation of an inducible Pfubc13 knockout parasite
To generate an inducible knockout of ubc13, we used CRISPR-Cas9 to insert two loxP sites flanking the last exon of the gene and the rapamycin-inducible active DiCre-recombinase system to excise the intervening DNA so that a truncated inactive protein would be produced, using the methodology described previously29,32. The Guide RNA sequence was designed to direct Cas9 to introduce a double-stranded break in Exon IV (Supplementary Fig. 1 and Supplementary Table 1). A pair of complementary oligonucleotides was phosphorylated with T4 polynucleotide kinase, annealed, and ligate into the plasmid pDC2-Cas9-hDHFRyFCU that had been digested with BbsI restriction enzyme32. For the repair plasmid, synthetic DNA (GeneArt, Life Technologies) was used to replace Intron IV with a synthetic intron containing a 34 bp loxP element (loxPint, 103 bp)47 followed by a recodonized Exon IV (225 bp) and a second loxP site (34 bp) after the stop codon. This sequence was placed between two homology regions, HR1 (262 bp) and HR2 (352 bp) to facilitate homologous recombination and cloned in the pMK vector with EcoRI and XhoI restriction sites at the 5’ and 3’ end, respectively. The repair plasmid (50 μg) was digested with EcoRI and XhoI, ethanol precipitated together with 20 μg plasmid to express the guide RNA, and redissolved in 10 μl sterile TE (10 mM Tris–HCl 1 mM EDTA pH 8.0).
For transfection, purified P. falciparum II-3 schizonts (10–40 µl) were mixed with DNA dissolved in 100 µl of AMAXA primary cell solution P3 and electroporated using an AMAXA 4D nucleofector with program FP158. Following electroporation, schizonts were transferred to 2 ml culture medium containing 300 µl of erythrocytes, and incubated with shaking at 37 °C for 30 min. A further 8 ml of culture medium was added and then 24 to 48 h post-transfection at 37 °C, 2.5 to 10 nM WR99210 was added to provide selection. The parasite population was screened for integration by PCR, using pairs of primers (Supplementary Fig. 1 and Supplementary Table 1) and then individual lines were selected following cloning by limiting dilution and screening by PCR.
Rapamycin-induced DiCre mediated loxP excision
Synchronized ring-stage parasites (2 h PI) were treated with a final concentration of 20 nM rapamycin or with DMSO for 24 h. Samples were taken for DNA analysis by PCR at this time or for protein analysis by Western blotting at 44 h PI.
Parasite growth and survival study
One hundred microliters synchronised ring-stage parasites (2 h PI, 0.2% parasitemia; 3% hematocrit), treated with 20 nM rapamycin or 0.01% DMSO, were placed in wells of a round-bottom 96-well plate at time zero. At the start and after specific periods (48 and 96 h) under normal culture conditions, parasite samples were fixed with 100 µl 4% paraformaldehyde + 0.1% glutaraldehyde for 1 h at room temperature (RT). The plates were centrifuged and fixative was replaced with 50 µl PBS and the parasites were stained with 2 × SyBR Green I in the dark for 30 min. One microlitre of stained cells was diluted into 3 ml PBS in a FACS tube and analysed using a BD LSRFortessa flow cytometer, with a 488 nm UV laser and fluorescence at 530 nm. Duplicate samples on triplicate plates were analysed for each time point. Data were analysed with one-way ANOVA analysis (Dunnett’s multiple comparison test) using Graphpad software. At various time points thin blood smears were prepared, stained with Giemsa’s reagent and examined by microscopy to evaluate parasitemia and parasite morphology.
Western blotting of parasite lysates
Parasites (3–4 h PI) were treated with 20 nM rapamycin or 0.01% DMSO for 40 h, then schizonts were purified by centrifugation over a 63% Percoll cushion, suspended in PBS containing 0.15% (w/v) saponin to lyse erythrocyte membranes, and collected by centrifugation at 5,000 g for 5 min. The cell pellet was dissolved in 10 volumes of lysis buffer (150 mM NaCl, 1% Triton X-100, 0.1% SDS, 1 μl/ml benzonase [Roche], and 1X complete protease inhibitors [Roche]). After incubation on ice for 20 min, samples were centrifuged at 17,000 g for 30 min. The protein content of the supernatant was measured using a BCA Protein Assay (Pierce), and 10 μg total protein was resolved on a 4 to 12% Tris–acetate PAGE gel (Invitrogen), transferred to nitrocellulose, and probed with specific antibodies. Protein ubiquitylation was detected with antibodies to ubiquitin Lys48- and Lys63-specific linkages (Merck-Millipore rabbit mAbs 05–1307 and 05–1308, respectively). Antibodies to BiP29 were used as a loading control.
Parasite survival following treatment with the mutagen, methyl methanesulfonate
Parasites pretreated with rapamycin or DMSO for 24 h were treated with 2.25 mM and 4.5 mM MMS for 0, 20, 40, 60 and 100 min, washed twice with incomplete RPMI 1640 medium and finally returned to culture in complete RPMI 1640 medium. At 48 h after the start of the experiment, thin blood smears were prepared, stained with Giemsa’s reagent and examined by microscopy to calculate percent parasite survival.
To examine parasite recovery from MMS induced-DNA damaged, ring stage parasites 4 to 6 h PI were adjusted to 0.5% parasitemia, 3% hematocrit and then treated for 48 h with 20 nM rapamycin or DMSO and with or without 500 μM MMS. The medium was changed every 2 days, and the percentage parasitemia calculated every day from examination of Giemsa-stained thin blood smears.
Parasite drug sensitivity
Synchronised ring stage parasites (0.05–1% parasitemia; 3% hematocrit) were treated with either DMSO alone or 20 nM rapamycin, together with serial dilutions (from 200 nM) of DHA and CQ as 100 µl final volume of culture in a black 96-well plate. Following incubation under normal culture conditions for 48 or 96 h, parasite survival was determined by using the malaria SYBR Green I-based fluorescence assay (MSF), as described previously48. Briefly, 100 µl of SYBR Green I in lysis buffer (0.2 µl of SYBR Green I/ml of lysis buffer [20 mM Tris–HCl, 5 mM EDTA, 0.008% saponin, and 0.08% Triton X100]) was added to each well, then the plate was mixed and incubated in the dark at RT for 1 h. The fluorescence was measured with excitation and emission at 485 and 530 nm, respectively48,49,50. The data were analysed and the EC50 for each compound was calculated using GraphPad Prism software.
Data availability
All plasmids and transgenic parasites described in this study are available upon request.
References
Aminake, M. N., Arndt, H. D. & Pradel, G. The proteasome of malaria parasites: A multi-stage drug target for chemotherapeutic intervention?. Int. J. Parasitol. Drugs Drug Resist. 2, 1–10. https://doi.org/10.1016/j.ijpddr.2011.12.001 (2012).
Bhoj, V. G. & Chen, Z. J. Ubiquitylation in innate and adaptive immunity. Nature 458, 430–437. https://doi.org/10.1038/nature07959 (2009).
McGinty, R. K., Henrici, R. C. & Tan, S. Crystal structure of the PRC1 ubiquitylation module bound to the nucleosome. Nature 514, 591–596. https://doi.org/10.1038/nature13890 (2014).
Stewart, M. D., Ritterhoff, T., Klevit, R. E. & Brzovic, P. S. E2 enzymes: More than just middle men. Cell Res. 26, 423–440. https://doi.org/10.1038/cr.2016.35 (2016).
Adhikari, A. & Chen, Z. J. Diversity of polyubiquitin chains. Dev. Cell 16, 485–486. https://doi.org/10.1016/j.devcel.2009.04.001 (2009).
Ye, Y. & Rape, M. Building ubiquitin chains: E2 enzymes at work. Nat. Rev. Mol. Cell Biol. 10, 755–764. https://doi.org/10.1038/nrm2780 (2009).
Ponts, N. et al. Deciphering the ubiquitin-mediated pathway in apicomplexan parasites: A potential strategy to interfere with parasite virulence. PLoS ONE 3, e2386. https://doi.org/10.1371/journal.pone.0002386 (2008).
Chung, D. W., Ponts, N., Prudhomme, J., Rodrigues, E. M. & Le Roch, K. G. Characterization of the ubiquitylating components of the human malaria parasite’s protein degradation pathway. PLoS ONE 7, e43477. https://doi.org/10.1371/journal.pone.0043477 (2012).
Andersen, P. L. et al. Distinct regulation of Ubc13 functions by the two ubiquitin-conjugating enzyme variants Mms2 and Uev1A. J. Cell. Biol. 170, 745–755. https://doi.org/10.1083/jcb.200502113 (2005).
Ashley, C., Pastushok, L., McKenna, S., Ellison, M. J. & Xiao, W. Roles of mouse UBC13 in DNA postreplication repair and Lys63-linked ubiquitination. Gene 285, 183–191. https://doi.org/10.1016/s0378-1119(02)00409-2 (2002).
Bai, Z., Li, Z. & Xiao, W. Drosophila bendless catalyzes K63-linked polyubiquitination and is involved in the response to DNA damage. Mutat. Res. 808, 39–47. https://doi.org/10.1016/j.mrfmmm.2018.02.003 (2018).
Brusky, J., Zhu, Y. & Xiao, W. UBC13, a DNA-damage-inducible gene, is a member of the error-free postreplication repair pathway in Saccharomyces cerevisiae. Curr. Genet. 37, 168–174. https://doi.org/10.1007/s002940050515 (2000).
Burge, R. J., Damianou, A., Wilkinson, A. J., Rodenko, B. & Mottram, J. C. Leishmania differentiation requires ubiquitin conjugation mediated by a UBC2-UEV1 E2 complex. PLoS Pathog. 16, e1008784. https://doi.org/10.1371/journal.ppat.1008784 (2020).
Gudgen, M., Chandrasekaran, A., Frazier, T. & Boyd, L. Interactions within the ubiquitin pathway of Caenorhabditis elegans. Biochem. Biophys. Res. Commun. 325, 479–486. https://doi.org/10.1016/j.bbrc.2004.10.047 (2004).
Sato, M., Konuma, R., Sato, K., Tomura, K. & Sato, K. Fertilization-induced K63-linked ubiquitylation mediates clearance of maternal membrane proteins. Development 141, 1324–1331. https://doi.org/10.1242/dev.103044 (2014).
Wen, R., Newton, L., Li, G., Wang, H. & Xiao, W. Arabidopsis thaliana UBC13: Implication of error-free DNA damage tolerance and Lys63-linked polyubiquitylation in plants. Plant Mol. Biol. 61, 241–253. https://doi.org/10.1007/s11103-006-0007-x (2006).
Feng, H., Wang, S., Dong, D., Zhou, R. & Wang, H. Arabidopsis ubiquitin-conjugating enzymes UBC7, UBC13, and UBC14 are required in plant responses to multiple stress conditions. Plants (Basel) https://doi.org/10.3390/plants9060723 (2020).
Wenzel, D. M., Stoll, K. E. & Klevit, R. E. E2s: Structurally economical and functionally replete. Biochem. J. 433, 31–42. https://doi.org/10.1042/BJ20100985 (2011).
Pulvino, M. et al. Inhibition of proliferation and survival of diffuse large B-cell lymphoma cells by a small-molecule inhibitor of the ubiquitin-conjugating enzyme Ubc13-Uev1A. Blood 120, 1668–1677. https://doi.org/10.1182/blood-2012-02-406074 (2012).
Laine, A. et al. Regulation of p53 localization and activity by Ubc13. Mol. Cell Biol. 26, 8901–8913. https://doi.org/10.1128/MCB.01156-06 (2006).
Yamamoto, M. et al. Key function for the Ubc13 E2 ubiquitin-conjugating enzyme in immune receptor signaling. Nat. Immunol. 7, 962–970. https://doi.org/10.1038/ni1367 (2006).
McKenna, S. et al. Noncovalent interaction between ubiquitin and the human DNA repair protein Mms2 is required for Ubc13-mediated polyubiquitination. J. Biol. Chem. 276, 40120–40126. https://doi.org/10.1074/jbc.M102858200 (2001).
Hofmann, R. M. & Pickart, C. M. Noncanonical MMS2-encoded ubiquitin-conjugating enzyme functions in assembly of novel polyubiquitin chains for DNA repair. Cell 96, 645–653. https://doi.org/10.1016/s0092-8674(00)80575-9 (1999).
Raphemot, R. et al. Plasmodium PK9 inhibitors promote growth of liver-stage parasites. Cell Chem. Biol. 26, 411–419. https://doi.org/10.1016/j.chembiol.2018.11.003 (2019).
Solyakov, L. et al. Global kinomic and phospho-proteomic analyses of the human malaria parasite Plasmodium falciparum. Nat. Commun. 2, 565. https://doi.org/10.1038/ncomms1558 (2011).
Zhang, M. et al. Uncovering the essential genes of the human malaria parasite Plasmodium falciparum by saturation mutagenesis. Science 360, 7847. https://doi.org/10.1126/science.aap7847 (2018).
Philip, N. & Haystead, T. A. Characterization of a UBC13 kinase in Plasmodium falciparum. Proc. Natl. Acad. Sci. U.S.A. 104, 7845–7850. https://doi.org/10.1073/pnas.0611601104 (2007).
Hodge, C. D. et al. Covalent inhibition of Ubc13 affects ubiquitin signaling and reveals active site elements important for targeting. ACS Chem. Biol. 10, 1718–1728. https://doi.org/10.1021/acschembio.5b00222 (2015).
Green, J. L. et al. Ubiquitin activation is essential for schizont maturation in Plasmodium falciparum blood-stage development. PLoS Pathog. 16, e1008640. https://doi.org/10.1371/journal.ppat.1008640 (2020).
Gupta, D. K., Patra, A. T., Zhu, L., Gupta, A. P. & Bozdech, Z. DNA damage regulation and its role in drug-related phenotypes in the malaria parasites. Sci. Rep. 6, 23603. https://doi.org/10.1038/srep23603 (2016).
Xiong, A. et al. K13-Mediated Reduced Susceptibility to Artemisinin in Plasmodium falciparum Is Overlaid on a Trait of Enhanced DNA Damage Repair. Cell Rep. 32, 107996. https://doi.org/10.1016/j.celrep.2020.107996 (2020).
Knuepfer, E., Napiorkowska, M., van Ooij, C. & Holder, A. A. Generating conditional gene knockouts in Plasmodium—a toolkit to produce stable DiCre recombinase-expressing parasite lines using CRISPR/Cas9. Sci. Rep. 7, 3881. https://doi.org/10.1038/s41598-017-03984-3 (2017).
Gopalakrishnan, A. M. & Kumar, N. in Encyclopedia of Malaria (eds Marcel Hommel & Peter G. Kremsner) Ch. Chapter 50–2, 1–8 (Springer New York, 2013).
Hodge, C. D., Spyracopoulos, L. & Glover, J. N. Ubc13: the Lys63 ubiquitin chain building machine. Oncotarget 7, 64471–64504. https://doi.org/10.18632/oncotarget.10948 (2016).
Bi, X. mechanism of DNA damage tolerance. J. Biol. Chem. 6, 48–56 (2005).
Lee, A. H., Symington, L. S. & Fidock, D. A. DNA repair mechanisms and their biological roles in the malaria parasite Plasmodium falciparum. Microbiol. Mol. Biol. Rev. 78, 469–486. https://doi.org/10.1128/MMBR.00059-13 (2014).
Fu, D., Calvo, J. A. & Samson, L. D. Balancing repair and tolerance of DNA damage caused by alkylating agents. Nat. Rev. Cancer 12, 104–120. https://doi.org/10.1038/nrc3185 (2012).
Gardner, M. J. et al. Genome sequence of the human malaria parasite Plasmodium falciparum. Nature 419, 498–511. https://doi.org/10.1038/nature01097 (2002).
Singer, M. et al. Zinc finger nuclease-based double-strand breaks attenuate malaria parasites and reveal rare microhomology-mediated end joining. Genome Biol. 16, 249. https://doi.org/10.1186/s13059-015-0811-1 (2015).
Kondo, N., Takahashi, A., Ono, K. & Ohnishi, T. DNA damage induced by alkylating agents and repair pathways. J. Nucleic Acids 2010, 543531. https://doi.org/10.4061/2010/543531 (2010).
Drablos, F. et al. Alkylation damage in DNA and RNA–repair mechanisms and medical significance. DNA Repair (Amst) 3, 1389–1407. https://doi.org/10.1016/j.dnarep.2004.05.004 (2004).
Gopalakrishnan, A. M. & Kumar, N. Opposing roles for two molecular forms of replication protein A in Rad51-Rad54-mediated DNA recombination in Plasmodium falciparum. MBio 4, e00252-e1213. https://doi.org/10.1128/mBio.00252-13 (2013).
Dinant, C., Houtsmuller, A. B. & Vermeulen, W. Chromatin structure and DNA damage repair. Epigenet. Chromatin 1, 9. https://doi.org/10.1186/1756-8935-1-9 (2008).
O’Neill, P. M., Barton, V. E. & Ward, S. A. The molecular mechanism of action of artemisinin–the debate continues. Molecules 15, 1705–1721. https://doi.org/10.3390/molecules15031705 (2010).
Sutherland, C. J., Henrici, R. C. & Artavanis-Tsakonas, K. Artemisinin susceptibility in the malaria parasite Plasmodium falciparum: propellers, adaptor proteins and the need for cellular healing. FEMS Microbiol. Rev. 45, 1–15. https://doi.org/10.1093/femsre/fuaa056 (2021).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858. https://doi.org/10.1038/nprot.2015.053 (2015).
Jones, M. L. et al. A versatile strategy for rapid conditional genome engineering using loxP sites in a small synthetic intron in Plasmodium falciparum. Sci. Rep. 6, 21800. https://doi.org/10.1038/srep21800 (2016).
Smilkstein, M., Sriwilaijaroen, N., Kelly, J. X., Wilairat, P. & Riscoe, M. Simple and inexpensive fluorescence-based technique for high-throughput antimalarial drug screening. Antimicrob. Agents Chemother. 48, 1803–1806. https://doi.org/10.1128/AAC.48.5.1803-1806.2004 (2004).
Co, E. M., Dennull, R. A., Reinbold, D. D., Waters, N. C. & Johnson, J. D. Assessment of malaria in vitro drug combination screening and mixed-strain infections using the malaria Sybr green I-based fluorescence assay. Antimicrob. Agents Chemother. 53, 2557–2563. https://doi.org/10.1128/AAC.01370-08 (2009).
Johnson, J. D. et al. Assessment and continued validation of the malaria SYBR green I-based fluorescence assay for use in malaria drug screening. Antimicrob. Agents Chemother. 51, 1926–1933. https://doi.org/10.1128/AAC.01607-06 (2007).
Acknowledgements
SW was funded by The Royal Golden Jubilee PhD. Programme (NSTDA), Thailand Research Fund (PHD/0210/2557) and a Thailand Research Fund-British Council Fund Newton Fund PhD Placement (http://www.britishcouncil.org/newton-fund). This research was funded in part by the National Center for Genetic Engineering and Biotechnology, and Thailand Research Fund (RSA5880064), and the National Science and Technology Development Agency (P1850116). Work in the Holder laboratory was funded by the Francis Crick Institute (FC001097), which receives its core funding from Cancer Research UK (FC001097), the UK Medical Research Council (FC001097), and the Wellcome Trust (FC001097). For the purpose of Open Access, the author has applied a CC BY public copyright licence to any Author Accepted Manuscript version arising from this submission.
Author information
Authors and Affiliations
Contributions
S.M., A.A.H., E.K., J.L.G. : conceptualised and designed experiments; S.M., P.P.: performed experiments; E.K., J.L.G., P.P. provided reagents and methodology; S.M., A.A.H., S.S., C.U. analysed the data; A.A.H, S.S., C.U. acquired funding and supervised the work; S.M., A.A.H wrote initial manuscript draft; all authors reviewed manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Maneekesorn, S., Knuepfer, E., Green, J.L. et al. Deletion of Plasmodium falciparum ubc13 increases parasite sensitivity to the mutagen, methyl methanesulfonate and dihydroartemisinin. Sci Rep 11, 21791 (2021). https://doi.org/10.1038/s41598-021-01267-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-01267-6
- Springer Nature Limited