Abstract
The phospholipid biosynthesis of the malaria parasite, Plasmodium falciparum is a key process for its survival and its inhibition is a validated antimalarial therapeutic approach. The second and rate-limiting step of the de novo phosphatidylcholine biosynthesis is catalysed by CTP: phosphocholine cytidylyltransferase (PfCCT), which has a key regulatory function within the pathway. Here, we investigate the functional impact of the key structural differences and their respective role in the structurally unique pseudo-heterodimer PfCCT protein in a heterologous cellular context using the thermosensitive CCT-mutant CHO-MT58 cell line. We found that a Plasmodium-specific lysine-rich insertion within the catalytic domain of PfCCT acts as a nuclear localization signal and its deletion decreases the nuclear propensity of the protein in the model cell line. We further showed that the putative membrane-binding domain also affected the nuclear localization of the protein. Moreover, activation of phosphatidylcholine biosynthesis by phospholipase C treatment induces the partial nuclear-to-cytoplasmic translocation of PfCCT. We additionally investigated the cellular function of several PfCCT truncated constructs in a CHO-MT58 based rescue assay. In absence of the endogenous CCT activity we observed that truncated constructs lacking the lysine-rich insertion, or the membrane-binding domain provided similar cell survival ratio as the full length PfCCT protein.
Similar content being viewed by others
Introduction
Malaria is a major global threat with around 50% of the world population being in danger of getting infected. The disease is caused by the parasites of the Plasmodium genus, most notably Plasmodium falciparum. Despite the immense effort to eradicate malaria worldwide, ca. 228 million cases of malaria have been reported in 2018 among which 405.000 cases were lethal1. The occurrence of drug-resistant parasite strains makes it even more difficult to eliminate the threat2. Therefore, the development of new compounds with different mechanisms of action is essential for effective therapy3. Currently, the artemisinin-based combination therapy (ACT) is the first-line of defence in many regions, although there are already parasite strains with delayed parasite clearance in many regions4. Essential biosynthetic pathways are in the focus of research to exploit the fundamental need of the Plasmodium parasite on processes such as DNA replication, nucleotide and phospholipid biosynthesis5,6.
During the intraerythrocytic life cycle, Plasmodium species produce up to 20 daughter parasite cells that egress from a single infected red blood cell in just 48 h6. This fast proliferation requires an excessive amount of membrane to be generated de novo. Therefore, the synthesis of phospholipid components is a process of key importance for the parasites, especially in the trophozoite and schizont stages. In eukaryotic cells, phosphatidylcholine (PC) is the major phospholipid component in most of the membrane structures7. Within the Plasmodium genus, PC is primarily generated in a three-step biosynthetic pathway called the Kennedy-pathway and that is partially supplemented by the methylation pathways of phosphoethanolamine and phosphatidylethanolamine8. Albitiazolium, a lead compound targeting the parasite PC biosynthesis, has already been successfully evaluated in human phase I and II clinical trials9, yet later in pediatric tests its high metabolic clearance was observed in children, which ultimately stopped its clinical development10. Nevertheless, the detailed analysis of the mode of action and pharmacokinetics of this drug could be utilized in future developments10.
The second and rate-limiting step of the Kennedy-pathway is catalysed by the enzyme CTP:phosphocholine cytidylyltransferase (CCT), which converts the precursor phosphocholine (PhoC) to cytidine-5′-diphosphocholine (CDPCho) by incorporating CTP and releasing pyrophosphate as a by-product. CCT proteins of Plasmodium species, including the Plasmodium falciparum CCT (PfCCT) are unique orthologues in terms of their architecture. While CCT enzymes regularly form a homodimer structure11, PfCCT is a pseudo-heterodimer that contains both monomer units as two repeat units within a single polypeptide chain as a result of a gene duplication event12. The pseudo-monomer repeat units correspond to the basic domain structure of CCT enzymes, including the catalytic (C) domain and the putative regulatory membrane-binding (M) domain (Fig. 1A)13. However, other regions such as the N-terminal nuclear localization signal (NLS) and the C-terminal phosphorylation site present in mammalian orthologs have not been identified in PfCCT (Fig. 1B)12,14. Within the active sites, the repeat units also contain the CTP-binding motif HxGH as well as the RTEG(I/V)ST(S/T) motif that overlaps with the N-terminus of the αE helix. The latter motif serves as a bridge between the C and M domains to assist membrane-binding induced regulatory changes15. Between the two pseudo-monomers (C1M1 and C2M2), a more than 200 amino acid long disordered inter-linker region is present, whose purpose has not yet been characterized (Fig. 1B). The precise molecular background of the PfCCT regulatory mechanism has not been revealed in detail, but it is expected to be reminiscent to the mode of action described for the mammalian CCTs16,17. In the inactive soluble form, the catalytic domain of the CCT is inhibited by an autoinhibitory (AI) helix in the M domain18,19. PC-depleted membranes have altered physicochemical properties, which leads to increased negative surface charge and membrane curvature, creating membrane packing defects17,20,21. The CCT protein is able to sense these alterations and the M domain undergoes significant conformational changes. Subsequently, the AI helix releases the catalytic domain and turns into a membrane-induced amphipathic helix (m-AH) that docks into the PC-depleted membrane surfaces15,21,22. Intriguingly, in PfCCT, the single continuous m-AH in mammalian CCTs23 is substituted by two separate amphipathic helices in the putative membrane binding domain, a shorter N-terminal and a longer C-terminal helices (m-AH-N and m-AH-C, respectively)24. Upon membrane tethering, the catalytic domain is released from autoinhibition, resulting in a dramatic increase of enzyme activity in mammalian CCTs15,25. Accordingly, an engineered CCT construct comprising the catalytic but not the membrane-binding domain possesses constitutive, lipid-independent enzyme activity26. Nevertheless, compared to its orthologs, PfCCT has a moderate enzyme activity in the absence of lipid activators and it only shows a sixfold increase in activity in the presence of lipid vesicles27. The reason and the purpose of this difference compared to other orthologous CCTs is yet to be explored.
Molecular architecture and regulation of the CCT protein. (A) The conceptual regulatory mechanism of CCT on the macromolecular level. In the inactive state, an autoinhibitory (AI) helix in the membrane-binding (M—grey) domain inhibits the catalytic domain (C—blue) in the inactive state (left). The decreased PC content alters the physicochemical properties of membranes. This induces a conformational change in the M domain, turning the AI helix into the membrane-induced amphipathic helix (m-AH) that docks into the PC-depleted membrane surface and thereby releases the inhibition (right)16. (B) The schematic representation of mammalian CCT proteins and the truncated protein constructs of PfCCT used in this study. N, C, M and P represent the N-terminal cap region, the catalytic domain, the membrane-binding domain and the region of phosphorylation, respectively. C1 and C2 constructs solely contain the catalytic domain of the first and second repeat unit of PfCCT, respectively. C1M1 and C2M2 also include the respective putative membrane binding domains of each repeat unit. ΔK constructs lack the respective lysine-rich loops (red segments noted as K).
Another major difference between Plasmodial and mammalian CCTs concerns the number of enzyme isoforms. The vast majority of CCT proteins in higher eukaryotic organisms have two major isoforms, CCTα/CCT1 and CCTβ/CCT2, which localize in different cellular compartments and translocate to different membrane surfaces28,29. The dominant CCTα/CCT1 isoform is a nuclear protein that translocate to the inner nuclear membrane22,30,31, while CCTβ/CCT2 is a cytosolic isoform that primarily docks into the surface of the endoplasmic reticulum in the cytoplasm32,33,34. Interestingly, in cells with excessive need for PC synthesis e.g. pulmonary epithelial cells, CCTα can relocate to the cytoplasm, supposedly to enhance the PC biosynthetic process35. Furthermore, CCTα is able to shuttle between the nucleus and cytoplasm upon changes in the cellular PC homeostasis33,36. External stimuli such as phospholipase treatment28, addition of lipid activators (e.g. farnesol, oleate)36,37 or choline deprivation38, can be used in experimental systems to modulate CCT function and investigate its subcellular behaviour. In Drosophila S2 cells, CCT1 was able to relocate from the nucleus to the surface of cytoplasmic lipid droplets, but not in the mammalian 3T3-L1 cell line and differentiating human preadipocytes22,37. In the lower eukaryotic organism, Saccharomyces cerevisiae the sole CCT isoform is found in the nucleus39,40. However, in the rodent malaria parasite, Plasmodium berghei, the only CCT was found to be diffusely localized in the cell, both in the nucleus and cytoplasm8. Plasmodium falciparum also possesses only one CCT isoform, which localization is currently only known in schizont stages and was shown to be mostly diffuse in the parasite41.
In the present study, we investigate the functional impact of individual PfCCT structural elements on subcellular localization and activity with several truncated constructs expressed in the CCT-mutant CHO-MT58 cell line (Fig. 1B). We found that the elimination of either the Plasmodium-specific lysine-rich insertion within the catalytic domain or the putative membrane-binding domain disrupted the nuclear localization of the protein. Furthermore, treatment of the cells with a translocation inducing enzyme, phospholipase C (PLC) resulted in a partial nuclear-to-cytosolic relocation of the full length PfCCT and the second repeat unit construct. Previously, we started to characterize PfCCT in a cellular context by its transient heterologous expression in the thermosensitive CCT-mutant CHO-MT58 cell line42. In this system, we have already demonstrated that PfCCT is able to rescue the CHO-MT58 cells from apoptosis at the restrictive temperature42. Here, we performed rescue experiments with a set of truncated PfCCT constructs encompassing the catalytic domain of the first or second repeat unit with optional presence of the local lysine-rich insertion and/or the adjacent membrane binding domain. All the truncated constructs retained their in vivo functionality by managing to rescue the cells with similar potential as the full length PfCCT protein. Our results thus indicate that the sole presence of the catalytic domain is enough to rescue the cells at the restrictive temperature.
Results
Identification of a potential nuclear localization signal in PfCCT
The sequence alignment of PfCCT with two mammalian orthologs, rat CCT (Rattus norvegicus, RnCCT) and human CCT (Homo sapiens, HsCCT) shows a highly conserved catalytic domain with one major exception (Fig. 2A). An 18 amino acid-long, lysine-rich insertion is present in the catalytically important L5 loop in both the first and second catalytic domains (C1 and C2) of PfCCT, respectively11. The L5 loop is responsible for the coordination of the quaternary ammonium moiety of choline with residues Y131/Y714, N133/N716 and Y158/Y741 in C1/C2 and contributes to the formation of the composite aromatic box cleft43,44. We generated a homology model of PfCCT to complete the recently resolved crystal structure (PDB: 4ZCS, Fig. 2B)44 by visualizing the 720–737 Lys-rich insertion that was deleted from the crystallized construct. The insertion appears as a highly flexible region, characterized by low QMEAN local quality scores with an average of 0.4. Therefore, the homology model shows only a representative position of this dynamic region. Notably, the deletion of the lysine-rich segment had no significant impact on the in vitro constitutive enzyme activity of a catalytic domain construct PfCCT528–79512.
In silico analysis of PfCCT-specific protein segments. (A) The protein sequence alignment of the rat (RnCCT), human (HsCCT) and the two CCT repeat units of P. falciparum CCT (PfCCT_1 and PfCCT_2). Red boxes and red letters indicate the identical and similar amino acid residues within the catalytic domain, respectively. Blue box shows the Plasmodium-specific lysine-rich loop. Green, blue and yellow letters represent acidic, basic and hydrophobic residues in the membrane binding domain, respectively. The highly conserved catalytic domain is highlighted above the sequences (red) and a white line indicates the position of L5 loop. The membrane-induced amphipathic helix of RnCCT (brown box) with the determined autoinhibitory helix (red line) is shown for reference. The two, previously hypothesized N- and C-terminal membrane-induced amphipathic helix of PfCCT M domain is highlighted under the sequence with yellow boxes24. Alignment was generated with ESpript 3.045. (B) Homology model of the active site of PfCCT with one potential representative conformation of the flexible lysine-rich loop in a close-up view. Catalytically important residues in the proximity of L5 loop are highlighted, based on the crystal structure of PfCCT (green, PDB: 4ZCS). The position of CDPCho (yellow) is shown in the active site. Blue colored line shows the position of the lysine residues on the main chain of the L5 loop (grey). (C) Helical representation of the putative membrane-induced amphipathic helices in the membrane-binding domain of the first and second repeat unit, respectively were made with HeliQuest46. <µH> represents the hydrophobic momentum of each helix. The abundance of basic residues (blue) supports the higher affinity of membrane-binding towards negatively charged membrane surfaces, whilst a few acidic (red) and the vast majority of hydrophobic residues (grey) found in the longer m-AH-C helices supposedly have a role in the detection of local H+ accumulation and support the docking into the membrane leaflets47.
However, other potential functions might be related to this segment. Firstly, the complex regulatory mechanism of CCT enzymes include several dynamic conformational changes, which could be altered by the lysine-rich loop. In addition, its presence may provide an explanation to the unique diffuse cellular localization of PfCCT. As the large molecular weight (~ 105 kDa) of PfCCT excludes the possibility of its passive nuclear transport, we hypothesize that the presence of a nuclear localization signal (NLS) in the protein is indispensable for nuclear accumulation. In silico predictions with cNLSmapper showed the possibility of a bipartite NLS in the lysine-rich loop with a score of 5.8, that indicates a weak NLS with the potential to partially relocate the protein to the nucleus. Based on its amino acid composition as well as the homology model of the PfCCT protein, the lysine-rich loop is expected to be disordered and solvent-accessible region, that facilitates its recognition by Plasmodial nuclear import proteins. Since the nuclear import machinery must be able to access the signal peptide to effectively carry its cargo into the nucleus, the lysine-rich loop can be valid candidate to have an NLS function. Additionally, the Lys-rich insert might be associate with negatively charged membrane surfaces, given its pronounced basic character.
Two additional NLS predictions were found between residues 264–293 and 823–851 with a score 5.7 and 5.2, respectively. These regions both overlap with the presumed C-terminal peptide of the membrane-induced amphipathic helix of the M domain (m-AH-C, Fig. 2C). It was previously reported that the M domain is required for the shuttle between the nucleus and the cytoplasm36, therefore we attempted to analyse the effect of the truncation of the M-domain on the localization of PfCCT.
To elucidate the impact of the proposed regions on the nuclear localization, confocal microscopy-based colocalization analysis was applied. Both the full length PfCCT and the second repeat unit construct, C2M2 has a mixed nuclear-cytoplasmic localization in CHO-MT58 cells (Fig. 3A,B). However, the lysine-rich loop truncated C2M2ΔK construct showed a predominantly cytosolic localization, which indicates the nuclear localization signal role of this insertion (Fig. 3C). Furthermore, the membrane-binding domain truncated second repeat unit construct, C2 also showed a decreased nuclear localization compared to the C2M2 construct and the full length PfCCT protein (Fig. 3D,H,M). We also investigated the consequences of a double truncated C2ΔK construct, which showed a decreased nuclear accumulation similar to the C2 and C2M2ΔK constructs (Fig. 3E,K).
Colocalization analysis of PfCCT construct in CHO-MT58 cells. Confocal immunofluorescence images of cells transfected with (A) the full length PfCCT, (B) the second repeat unit C2M2 construct, (C) the lysine-rich loop truncated C2M2ΔK construct, (D) the membrane-binding domain truncated C2 construct, (E) C2ΔK and (F) SV40 NLS-tagged C2M2 construct. (G–L) Confocal images of cells transfected with PfCCT constructs and treated with PLC. White lines indicate 50 µm scalebars. (K) Nuclear overlap coefficients were calculated with Fiji. The number of cells used for the calculation are indicated in the columns. *p < 0.001.
We wanted to further investigate the role of these segments on the translocation mechanism. For this, we used phospholipase C (PLC), an enzyme that hydrolyses PC in the cellular membrane structures and therefore simulates a PC-depleted membrane status in the cells48,49. The subcellular localization of the full length PfCCT showed a peculiar pattern upon treatment, as the protein distribution became predominantly cytosolic (Fig. 3G). We observed the same change in distribution in case of the C2M2 construct (Fig. 3H). To test whether the relocation could be associated with the nuclear localization signal of PfCCT, we created a simian virus 40 (SV40) NLS-tagged C2M2 construct as a positive control. Interestingly, this construct did not relocate to the cytosol which indicates that a stronger NLS keeps the protein in the nucleus even after PC depletion (Fig. 3F,L). We did not observe any changes in the localization of the C2M2ΔK, C2 and C2ΔK constructs upon PLC treatment (Fig. 3I–K).
Functional impact of the lysine-rich loop on the rescue potential
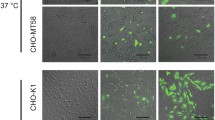
To evaluate the functionality of the different constructs in a cellular environment, we exploited the thermosensitive nature of the CCT-mutant CHO-MT58 cell line. Notably, these cells behave similarly to the parental CHO-K1 cell line at 37 °C, however, they are not viable at 40 °C, mainly due to thermal destabilizing effect of the CCT mutation50,51. Transient transfection of CHO-MT58 cells with the full length PfCCT protein was shown to be able to rescue the cells with a rescue potential of 55.9 ± 2.6% (Fig. 4), which is in good correspondence with the data reported42. We also used the inactive (IA) mutant PfCCT H45N H630N designed previously42 as a negative control that has a rescue potential of 11.7 ± 2.5%. In case of the constructs from the first repeat unit, C1M1 and C1 showed no significant difference in terms of cell survival, with a respective rescue potential of 46.2 ± 4.9% and 52.2 ± 10.9%. The second repeat constructs, C2M2 and C2 have similar results with a rescue potential of 45.6 ± 3.3% and 44.9 ± 7.5%, respectively. This suggests that the truncation of the putative M domain had no significant impact on the in vivo functionality of either the first or the second pseudo-monomer of the PfCCT protein. The deletion of lysine-rich loop also did not perturb the activity as reflected in a rescue potential of 52.2 ± 10.9% and 47.7 ± 6.7% for C1M1ΔK and C2M2ΔK is, respectively. Remarkably, the double truncated constructs C1ΔK and C2ΔK displayed a difference regarding their activity. While the second repeat unit construct C2ΔK had a rescue potential of 52.2 ± 14.1%, the first repeat unit construct C1ΔK had around half this value, 30.8 ± 9.3. Furthermore, the mammalian CCT control Homo sapiens CCTα (HsCCTα) was also found to be less potent in rescuing the cells as reflected by its rescue potential of 28.5 ± 7.1. While the underlying reason for these different rescue efficiencies remains unclear, notably, the rescue potential of all constructs tested is significantly (p < 0.01) higher than the inactive control PfCCT H45N H630N, indicating CCT enzyme activity dependent rescue. We hypothesize this variability is due to the different transfection efficiencies of constructs in this study.
Rescue potential of the different PfCCT constructs carried out in the thermosensitive CCT-mutant CHO-MT58 cell line system. Rescue potentials were determined as the quotient of surviving cells at 40 °C and 37 °C. Our results indicate that the sole presence of the catalytic domain is enough to rescue the cells at the restrictive temperature. All experiments were carried out in triplicates with averages ± S.D. being indicated. One-way ANOVA reveals no significant difference amongst the full length PfCCT and the truncated constructs (p > 0.05). All constructs are significantly higher than our inactive PfCCT mutant control (p < 0.01).
Discussion
Plasmodium falciparum CCT is in focus of research for a long time as a potential drug target due to its regulatory role in the phosphatidyl-choline de novo biosynthetic pathway. The duplication of catalytic and membrane-binding domains as well as the presence of Plasmodium-specific segments distinguishes PfCCT from its metazoan CCT homologs and the functional relevance of its many traits are yet unexplored. While structural and biochemical studies delineated the catalytic mechanism of PfCCT12,43,44, the precise regulatory mechanism including membrane binding and compartmentalization for this enzyme is less understood. Here, we used a mammalian cellular system based on the thermosensitive CCT-mutant CHO-MT58 to evaluate the function of the Plasmodium-specific structural elements of the PfCCT protein. The mammalian pathogens of the Plasmodium genus contain a typically lysine-rich insertion in the catalytic domain of their CCT protein that is absent in higher eukaryotic organisms (Fig. 5A). Intriguingly, other protists of the Apicomplexan phylum also possess similar insertions, albeit with fewer basic residues within. Noteworthy, this insertion is conserved in both repeat units of all Plasmodial CCTs, except the P. berghei CCT that contains an insertion that is less abundant in lysine residues in its C-terminal repeat unit and primarily contains non-charged asparagine residues (Fig. 5B). We investigated whether the positively charged cluster of lysine residues in PfCCT would allow a potential nuclear localization signal role for this segment.
Alignment of the lysine-rich loop from protists and Plasmodium species infecting mammals. (A) Phylogenetic tree of the CCT enzymes were put into context regarding the presence of an insertion in the catalytic domain. Orange background indicates the lack of insertion in higher eukaryotic organisms and Trypanosoma brucei. Babesia bovis evolved in a similar direction and has a smaller, less basic insertion highlighted by a grey background. Blue background represents that Toxoplasma gondii and many species of the Plasmodium genus have a longer, lysine-rich insertion. N-term and C-term indicates that whether the C- or the N-terminal repeat unit is included in the alignment of the protein sequences. The numbers indicate the mean number of substitutions per site. Phylogenetic tree was created with iToL tool52 and was manually annotated. (B) The lysine-rich character of the motif is conserved amongst Plasmodium species and is present in at least one of the two repeat units of the Plasmodial CCT enzymes. Blue and red letters highlight the basic and acidic residues, respectively. Numbers in the parenthesis show the length and the net charge of the insertion of each CCT protein and repeat unit. Visual representation of the alignment was generated with ESpript 3.0 and manually annotated45.
The full length PfCCT protein displays a diffuse localization pattern in both schizont stage parasites41 and the studied CHO-MT58 cell line (present study, Fig. 3). The large molecular mass of the pseudo-heterodimer PfCCT protein prevents its passive nuclear transport, thus the presence of a nuclear localization signal is required for nuclear targeting of this protein. Our confocal microscopy analysis shows that the lysine-rich loop has a nuclear localization signal function as the lysine-rich loop truncated constructs C2ΔK and C2M2ΔK were primarily found in the cytoplasm, in contrast to the full length PfCCT and the C2M2 constructs (Fig. 3). A possible explanation on why this lysine-rich motif is present in many of the Plasmodium species is the AT-rich genome and the tendency to generate indels, possibly due to slippage of DNA polymerase over the AT-rich, low complexity genomic regions, resulting in a biased use of AT-rich codons e.g. lysine or asparagine53,54. Nevertheless, it is intriguing that a similar insertion is visible in the CCT protein of Toxoplasma gondii, a taxonomically close relative of Plasmodium species, despite its 52% GC content genome55. Here, DNA polymerase slippage -driven emergence of the insert is less likely, which in turn raises the possibility for the functional relevance of this insert in parasites. Notably, a similar insertion with importin-dependent NLS function was also reported previously in the Plasmodium falciparum trimethyl-guanosine synthase enzyme56. Additionally, low complexity, Lys-rich repetitive regions of Plasmodial proteins are reported to modulate protein targeting into the periphery of infected red blood cells57. The reason why the presence of a Lys-rich insertion is advantageous for the parasite is still unclear and requires further characterization.
Additionally, we investigated how the PfCCT-expressing CHO-MT58 cells respond to PLC treatment that perturbs the phosphatidylcholine homeostasis. Our confocal microscopy analysis revealed a significant decrease in the nuclear fraction of both the full length PfCCT and the construct C2M2 following PLC treatment. In comparison, the PfCCT C2 construct that lacks the M domain displayed a decreased nuclear compartmentalization and the PLC treatment had no further effect on its localization. It was previously demonstrated that farnesol or oleate induced nuclear trafficking of exogenous CCT proteins in CHO-MT58 cells is dependent on amphipathic helix within M-domain36. Based on our results, the putative M domain of PfCCT also influences nuclear-to-cytoplasmic localization though its precise function is still unclear as we found no evidence of translocation occurring in either M domain truncated construct. A notable difference in the function of M domain was highlighted previously as it was proposed that it consists of two, shorter membrane-induced amphipathic helices24. Furthermore, this disparity is linked to a mere sixfold activation in the presence of oleate27 that is in sharp contrast to the robust, up to 100-fold activation increase in case of rat CCT, accounting for a less fine-tuned PfCCT regulatory mechanism58.
To evaluate the impact of the presence of the M domain on the exogenous PfCCT activity in a cellular environment, we transiently transfected the CHO-MT58 cells with several truncated constructs and assessed their effect on cell survival at the restrictive 40 °C temperature. Our findings suggest that neither the lysine-rich insert in the catalytically important L5 loop, nor the putative regulatory M domain is essential to rescue the cells. Nonetheless, C1ΔK showed a slightly lower capability of rescuing the thermosensitive CHO-MT58 cells, though it was still significantly more potent than our inactive PfCCT control. Moreover, we showed that HsCCTα was also capable of rescuing the cells and further shows the possibility of a previously established concept to test PfCCT-specific drugs by creating two, HsCCT- and PfCCT-expressing transgenic CHO-MT58 cell lines and use it as a cell-based test system for structure–function relation studies for drug development, targeting the PC biosynthetic pathway42.
We conclude that PfCCT has adapted to provide a unique robustness in its functionality and has a key role in supporting the excessive need for PC biosynthesis during the intraerythrocytic development. Its unique membrane-binding domain as well as the Lys-rich insertion may contribute to the regulation of the parasite PC homeostasis and intracellular transport throughout the widely diverse developmental stages. As confocal imaging did not reveal any distinct membrane compartmentalization following PLC treatment, it would be of further interest to investigate whether the enzyme could serve the increased need for PC biosynthesis in absence of the membrane-binding induced regulatory mechanism throughout the intraerythrocytic life cycle of parasites. A notable limitation of our results, combined from several independent experiments, originates from the mammalian cell line-based model system. The true functional relevance of PfCCT-specific structural elements has to be further assessed by in vivo experiments in parasites.
Materials and methods
Materials
Transient transfection was carried out with X-treme GENE HP transfection reagent (Merck KGaA, Darmstadt, Germany) according to the instructions of the manufacturer. Trypsin–EDTA (TE) solution and phosphate buffered saline (PBS) were used for general cell culture maintenance (Merck KGaA, Darmstadt, Germany). Plasmid pIRES-EGFP-puro was obtained from AddGene (#45567), provided by Michael McVoy (Virginia Commonwealth University). Enzymes Phusion Hot Start Flex DNA polymerase, NheI and XhoI restriction endonucleases and Instant Sticky End Ligase Master Mix are from New England Biolabs (Ipswich, MA, USA). Custom oligos were ordered from Sigma-Aldrich (Saint Louis, MO, USA). DNA purification kits for plasmid DNA and PCR product were obtained from Macherey–Nagel (Düren, Germany). Phospholipase C from C. perfringens (C. welchii) and every other reagent used here were obtained from Sigma-Aldrich (Saint Louis, MO, USA). Polyclonal anti-CCT was produced in rabbit after immunization with the second catalytic domain construct of PfCCT580–775, Δ720–737 and was isolated in the Department of Immunology, Eötvös Lóránd University, Hungary.
In silico analysis
Multiple sequence alignment was carried out by COBALT (NIH NCBI) with default parameters. To visually represent the alignment, ESpript 3.0 was used and manually annotated with the relevant secondary structure elements and domains. Homology model of PfCCT including the lysine-rich loop was created with SwissModel, based on the crystal structure of the PfCCT catalytic domain (PDB: 4ZCS)44 and was assessed using the Structure Assessment tool of SwissModel. Nuclear localization signal prediction was carried out with cNLSmapper for the entire protein with a cut-off score of 5.0. Helical representation of the different structural elements was made with HeliQuest using 3–11 helical structure46. Phylogenetic tree was generated with the Interactive Tree of Life (iTOL) tool52 after multiple sequence alignment of the different proteins using ClustalOmega.
Cloning of PfCCT constructs
The PfCCT cDNA sequence (PlasmoDB: PF3D7_1316600) was cloned into pIRES-EGFP-puro plasmid. The cDNA sequence was then used to design primer pairs for the respective monomer and their truncated constructs. Constructs C1, C1M1, C2, C2M2, SV40-NLS C2M2 and their respective lysine-rich loop deleted (ΔK) versions (present in Fig. 1B) were amplified with the following primers (Table 1).
Cell culture and rescue experiments
CHO-MT58 cells were kindly provided by Professor Dennis Vance (Department of Biochemistry, University of Alberta). Cells were cultured in an F12 Ham’s media supplemented with 10% foetal bovine serum (FBS) and 1% Penicillin–Streptomycin (Thermo Fisher Scientific, Waltham, MA, USA). CHO-MT58 cells were cultured at 37 °C and 40 °C in an incubator with a humidified, 5% CO2 atmosphere. Cells were prepared for rescue experiments seeding 100,000 cells in a 6-well-plate. At 80% confluence cells were transfected with 200 µl of transfection reagent mix containing 6 µl of X-treme GENE HP transfection reagent, 2 µg purified plasmid DNA in serum-free Opti-MEM medium. Media was always changed before the addition of the transfection mix. On the following day, culture was checked for transfection efficiency by live-cell imaging of the expressed green fluorescent protein (GFP) and if found proper, culture was split in half and incubated at 37 °C or 40 °C for an additional 3 days. Both cultures were then harvested. The supernatant was collected and mixed with the adherent cells resuspended by Trypsin–EDTA. To check the viability of cells, propidium-iodide (PI) was added in 1 µg/ml concentration. Flow cytometry was carried out with a FACS Attune Acoustic Focusing Flow Cytometer. After intact cells were gated based on Forward Scatter (FSC) and Side Scatter (SSC), GFP positivity and PI fluorescence were detected to determine the number of cells successfully transfected and the amount of dead cells. Rescue potential was calculated by the following formula:
Immunofluorescent staining
50,000 CHO-MT58 cells were passed to a coverslip and were transiently transfected at 80% confluence with 50 µl of transfection reagent mix containing 1.5 µl of X-treme GENE HP transfection reagent and 0.5 µg purified plasmid DNA in Opti-MEM serum-free medium. After 24 h, PLC + cells were treated with 10 mU/ml phospholipase C for 3 h at 37 °C. Cells were then fixed with 4% paraformaldehyde for 10 min and permeabilized with 0.1% Triton X-100 solution. A blocking solution containing 3% bovine serum albumin (BSA) and 5% FBS in PBS was added for an hour. After the blocking, polyclonal primary antibody anti-CCT was added in 1:1000 ratio for an hour. Secondary antibody anti-rabbit Alexa Fluor 633 from goat was added in 1:1000 ratio after the primary antibody treatment for an additional hour. Finally, the cells were stained with 0.1 µg/ml 4′,6-diamidino-2-phenylindole dihydrochloride (DAPI) in PBS. After every step, the coverslips were washed 3 times with PBS. Every step was carried out at room temperature. The stained slides were embedded in FluorSave antifading agent to preserve fluorescence. Confocal imaging was carried out on a Zeiss LSM 710 microscope using a Plan-APOCHROMAT 40 × oil immersion objective.
Colocalization analysis
8-bit images acquired by the confocal imaging were analysed in Fiji60 by the Coloc 2 plug-in. Threshold regression was carried out with the Costes method on images with determined regions of interest (ROI). Mander’s correlation coefficient was used to determine the nuclear overlap coefficient based on the DAPI and the corresponding anti-CCT signal using the following formula: \(M=\frac{\sum_{i}{S1}_{i,coloc}}{\sum_{i}{S1}_{i}}\), where S1i,coloc equals S1i (intensity of the anti-CCT) signal if DAPI is above the determined threshold.
References
World Health Organization (WHO). World Malaria Report 2019. (2019).
Menard, D. & Dondorp, A. Antimalarial drug resistance: a threat to malaria elimination. Cold Spring Harb. Perspect. Med. 7, a025619 (2017).
Kumar, S., Bhardwaj, T. R., Prasad, D. N. & Singh, R. K. Drug targets for resistant malaria: Historic to future perspectives. Biomed. Pharmacother. 104, 8–27 (2018).
Rosenthal, M. R. & Ng, C. L. Plasmodium falciparum artemisinin resistance: the effect of heme, protein damage, and parasite cell stress response. ACS Infect. Dis. 6, 1599–1614 (2020).
Alam, A. et al. Novel antimalarial drug targets: hope for new antimalarial drugs. Expert Rev. Clin. Pharmacol. 2, 469–489 (2009).
Matthews, H., Duffy, C. W. & Merrick, C. J. Checks and balances? DNA replication and the cell cycle in Plasmodium. Parasit. Vect. 11, 1–13 (2018).
Wein, S. et al. Contribution of the precursors and interplay of the pathways in the phospholipid metabolism of the malaria parasite. J. Lipid Res. 59, 1461–1471 (2018).
Déchamps, S. et al. The Kennedy phospholipid biosynthesis pathways are refractory to genetic disruption in Plasmodium berghei and therefore appear essential in blood stages. Mol. Biochem. Parasitol. 173, 69–80 (2010).
Wein, S. et al. Transport and pharmacodynamics of albitiazolium, an antimalarial drug candidate. Br. J. Pharmacol. 166, 2263–2276 (2012).
Wein, S. et al. High accumulation and in vivo recycling of the new antimalarial albitiazolium lead to rapid parasite death. Antimicrob. Agents Chemother. 61, 1–17 (2017).
Lee, J., Johnson, J., Ding, Z., Paetzel, M. & Cornell, R. B. Crystal structure of a mammalian CTP: phosphocholine cytidylyltransferase catalytic domain reveals novel active site residues within a highly conserved nucleotidyltransferase fold. J. Biol. Chem. 284, 33535–33548 (2009).
Nagy, G. N. et al. Evolutionary and mechanistic insights into substrate and product accommodation of CTP: phosphocholine cytidylyltransferase from Plasmodium falciparum. FEBS J. 280, 3132–3148 (2013).
Huang, H. K. H. et al. The membrane-binding domain of an amphitropic enzyme suppresses catalysis by contact with an amphipathic helix flanking its active site. J. Mol. Biol. 425, 1546–1564 (2013).
Dennis, M. K., Taneva, S. G. & Cornell, R. B. The intrinsically disordered nuclear localization signal and phosphorylation segments distinguish the membrane affinity of two cytidylyltransferase isoforms. J. Biol. Chem. 286, 12349–12360 (2011).
Taneva, S. G. et al. Interdomain communication in the phosphatidylcholine regulatory enzyme, CCTα, relies on a modular αE helix. J. Biol. Chem. 294, 15517–15530 (2019).
Cornell, R. B. Membrane lipids assist catalysis by CTP: phosphocholine cytidylyltransferase. J. Mol. Biol. 1862, 183135 (2020).
Cornell, R. B. Membrane lipid compositional sensing by the inducible amphipathic helix of CCT. Biochim. Biophys. Acta 1861, 847–861 (2016).
Lee, J., Taneva, S. G., Holland, B. W., Tieleman, D. P. & Cornell, R. B. Structural basis for autoinhibition of CTP: phosphocholine cytidylyltransferase (CCT), the regulatory enzyme in phosphatidylcholine synthesis, by its membrane-binding amphipathic helix. J. Biol. Chem. 289, 1742–1755 (2014).
Wang, Y. & Kent, C. Identification of an inhibitory domain of CTP:phosphocholine cytidylyltransferase. J. Biol. Chem. 270, 18948–18952 (1995).
Braker, J. D., Hodel, K. J., Mullins, D. R. & Friesen, J. A. Identification of hydrophobic amino acids required for lipid activation of C. elegans CTP: phosphocholine cytidylyltransferase. Arch. Biochem. Biophys. 492, 10–16 (2009).
Chong, S. S. Y., Taneva, S. G., Lee, J. M. C. & Cornell, R. B. The curvature sensitivity of a membrane-binding amphipathic helix can be modulated by the charge on a flanking region. Biochemistry 53, 450–461 (2014).
Haider, A. et al. pcyt1a regulates phosphatidylcholine homeostasis from the inner nuclear membrane in response to membrane stored curvature elastic stress. Dev. Cell 45, 481-495.e8 (2018).
Dunne, S. J., Cornell, R. B., Johnson, J. E., Glover, N. R. & Tracey, A. S. Structure of the membrane binding domain of CTP: phosphocholine cytidylyltransferase. Biochemistry 35, 11975–11984 (1996).
Larvor, M.-P. et al. Characterization of the lipid-binding domain of the Plasmodium falciparum CTP: phosphocholine cytidylyltransferase through synthetic-peptide studies. Biochem. J. 375, 653–661 (2003).
Knowles, D. G., Lee, J., Taneva, S. G. & Cornell, R. B. Remodeling of the interdomain allosteric linker upon membrane binding of CCTα pulls its active site close to the membrane surface. J. Biol. Chem. 294, 15531–15543 (2019).
Friesen, J. A., Campbell, H. A. & Kent, C. Enzymatic and cellular characterization of a catalytic fragment of CTP: phosphocholine cytidylyltransferase. J. Biol. Chem. 274, 13384–13389 (1999).
Yeo, H., Larvor, M., Ancelin, M. & Vial, H. J. Plasmodium falciparum CTP: phosphocholine cytidylyltransferase expressed in Escherichia coli: purification, characterization and lipid regulation. Biochem. J. 910, 903–910 (1997).
Clement, J. M. & Kent, C. CTP: phosphocholine cytidylyltransferase: insights into regulatory mechanisms and novel functions. Biochem Biophys Res Commun 257, 643–650 (1999).
Fagone, P. & Jackowski, S. Phosphatidylcholine and the CDP-choline cycle. Biochim. Biophys. Acta 1831, 523–532 (2013).
Cornell, R. B. & Ridgway, N. D. CTP:phosphocholine cytidylyltransferase: function, regulation, and structure of an amphitropic enzyme required for membrane biogenesis. Prog. Lipid Res. 59, 147–171 (2015).
Cornell, R. & Antonny, B. CCTα commands phospholipid homeostasis from the nucleus. Dev. Cell 45, 419–420 (2018).
Karim, M., Jackson, P. & Jackowski, S. Gene structure, expression and identification of a new CTP: phosphocholine cytidylyltransferase ß isoform. Biochim. Biophys. Acta 1633, 1–12 (2003).
Krahmer, N. et al. Phosphatidylcholine synthesis for lipid droplet expansion is mediated by localized activation of CTP: phosphocholine cytidylyltransferase. Cell Metab. 14, 504–515 (2011).
Lykidis, A., Baburina, I., Jackowski, S., Murti, K. & Jackowski, S. Distribution of CTP: phosphocholine cytidylyltransferase (CCT) isoforms. J. Biol. Chem. 274, 26992–27001 (1999).
Ridsdale, R., Tseu, I., Wang, J. & Post, M. CTP: phosphocholine cytidylyltransferase is a cytosolic protein in pulmonary epithelial cells and tissues. J. Biol. Chem. 276, 49148–49155 (2001).
Gehrig, K., Morton, C. C. & Ridgway, N. D. Nuclear export of the rate-limiting enzyme in phosphatidylcholine synthesis is mediated by its membrane binding domain. J. Lipid Res. 50, 966–976 (2009).
Aitchison, A. J., Arsenault, D. J. & Ridgway, N. D. Nuclear-localized CTP:phosphocholine cytidylyltransferase α regulates phosphatidylcholine synthesis required for lipid droplet biogenesis. Mol. Biol. Cell 26, 2927–2938 (2015).
Watkins, J. D. & Kent, C. Immunolocalization of membrane-associated CTP:phosphocholine cytidylyltransferase in phosphatidylcholine-deficient Chinese hamster ovary cells. J. Biol. Chem. 267, 5686–5692 (1992).
LaCount, D. J. et al. A protein interaction network of the malaria parasite Plasmodium falciparum. Nature 438, 103–107 (2005).
Mackinnon, M. A. et al. The Kap60-kap95 karyopherin complex directly regulates phosphatidylcholine synthesis. J. Biol. Chem. 284, 7376–7384 (2009).
Contet, A. et al. Plasmodium falciparum CTP:phosphocholine cytidylyltransferase possesses two functional catalytic domains and is inhibited by a CDP-choline analog selected from a virtual screening. FEBS Lett. 589, 992–1000 (2015).
Marton, L. et al. Heterologous expression of CTP: Phosphocholine cytidylyltransferase from Plasmodium falciparum rescues Chinese Hamster Ovary cells deficient in the Kennedy phosphatidylcholine biosynthesis pathway. Sci. Rep. 8, 4–11 (2018).
Nagy, G. N. et al. Composite aromatic boxes for enzymatic transformations of quaternary ammonium substrates. Angew. Chemie Int. Ed. 53, 13471–13476 (2014).
Guca, E. et al. Structural determinants of the catalytic mechanism of Plasmodium CCT, a key enzyme of malaria lipid biosynthesis. Sci. Rep. 8, 11215 (2018).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, 320–324 (2014).
Gautier, R., Douguet, D., Antonny, B. & Drin, G. HELIQUEST: a web server to screen sequences with specific α-helical properties. Bioinformatics 24, 2101–2102 (2008).
Johnson, J. E., Xie, M., Singh, L. M. R., Edge, R. & Cornell, R. B. Both acidic and basic amino acids in an amphitropic enzyme, CTP: phosphocholine cytidylyltransferase, dictate its selectivity for anionic membranes. J. Biol. Chem. 278, 514–522 (2003).
Sleight, R. & Kent, C. Regulation of phosphatidylcholine biosynthesis in mammalian. Cells 258, 831–835 (1983).
Sweitzer, T. D. & Kent, C. Expression of wild-type and mutant rat liver CTP: phosphocholine cytidylyltransferase in a cytidylyltransferase-deficient Chinese hamster ovary cell line. Arch. Biochem. Biophys. 311, 107–116 (1994).
Marton, L. et al. Molecular mechanism for the thermo-sensitive phenotype of CHO-MT58 cell line harbouring a mutant CTP: phosphocholine cytidylyltransferase. PLoS ONE 10, 1–17 (2015).
Esko, J. D., Wermuth, M. M. & Raetz, C. R. Thermolabile CDP-choline synthetase in an animal cell mutant defective in lecithin formation. J. Biol. Chem. 256, 7388–7393 (1981).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v4: recent updates and new developments. Nucleic Acids Res. 47, 256–259 (2019).
Hamilton, W. L. et al. Extreme mutation bias and high AT content in Plasmodium falciparum. Nucleic Acids Res. 45, 1259 (2017).
Paila, U., Kondam, R. & Ranjan, A. Genome bias influences amino acid choices: analysis of amino acid substitution and re-compilation of substitution matrices exclusive to an AT-biased genome. Nucleic Acids Res. 36, 6664–6675 (2008).
Lau, Y. L. et al. Deciphering the draft genome of Toxoplasma gondii RH strain. PLoS ONE 11, 1–15 (2016).
Babar, P. H., Dey, V., Jaiswar, P. & Patankar, S. An insertion in the methyltransferase domain of P. falciparum trimethylguanosine synthase harbors a classical nuclear localization signal. Mol. Biochem. Parasitol. 210, 58–70 (2016).
Davies, H. M., Thalassinos, K. & Osborne, A. R. Expansion of lysine-rich repeats in Plasmodium proteins generates novel localization sequences that target the periphery of the host erythrocyte. J. Biol. Chem. 291, 26188–26207 (2016).
Ding, Z. et al. A 22-mer segment in the structurally pliable regulatory domain of metazoan CTP: phosphocholine cytidylyltransferase facilitates both silencing and activating functions. J. Biol. Chem. 287, 38980–38991 (2012).
Xia, Y., Chu, W., Qi, Q. & Xun, L. New insights into the QuikChangeTM process guide the use of Phusion DNA polymerase for site-directed mutagenesis. Nucleic Acids Res. 43, e12–e12 (2015).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
Supported by the ÚNKP-18-2 New National Excellence Program of the Ministry of Human Capacities, the National Research, Development and Innovation Office of Hungary (K119493, 2017-1.3.1-VKE-2017-00002, 2017-1.3.1-VKE-2017-00013, VEKOP-2.3.2-16-2017-00013 to BGV, NKP-2018-1.2.1-NKP-2018-00005) and the BME-Biotechnology FIKP grant of EMMI (BME FIKP-BIO).
Author information
Authors and Affiliations
Contributions
R.I., L.M., G.N.N. and B.G.V. designed the experiments. R.I., L.M., H.L.P. and N.K. conducted the experiments and analysed the results. R.I., G.N.N. and B.G.V. wrote the final versions of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Izrael, R., Marton, L., Nagy, G.N. et al. Identification of a nuclear localization signal in the Plasmodium falciparum CTP: phosphocholine cytidylyltransferase enzyme. Sci Rep 10, 19739 (2020). https://doi.org/10.1038/s41598-020-76829-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-76829-1
- Springer Nature Limited