Abstract
Pin1, a polypeptide proline isomerase parvulin, plays a key role in Alzheimer’s disease (AD), common tumors and cancers. Two conservative histidine residues, His59 and His157, are important for maintaining the stability of the PPIase domain. Hence multiple spectral and computational techniques were performed to investigate the potential mechanism of two histidine residues. Thermal denaturation indicated that both residues His59 and His157 are not sensitive to the lower temperatures, while residue His59 is more sensitive to the higher temperatures than residue His157. Acidic denaturation suggested that influences of both residues His59 and His157 to acidic stability were the difference from Pin1-WT. ANS and RLS spectra hinted that there was no significant effect on hydrophobic change and aggregation by histidine mutations. The GndHCl-induced denaturation implied that residues His59 and His157 contributed the most to the chemical stability. MD simulations revealed that residues His59 and His157 mutations resulted in that the hydrogen bond network of the dual histidine motif was destroyed wholly. In summary, these histidine residues play an important role in maintaining the structural stability of the PPIase domain.
Similar content being viewed by others
Introduction
Pin1 (Protein interaction with NIMA1), which was discovered in 1996, is a peptidyl-prolyl cis-trans isomerase (PPIase)1. Indeed, Pin1 is associated with many biological processes including transcription regulation, cell growth, cell division, cell cycle, apoptosis and DNA damage repair2,3,4. Moreover, Pin1 interacts with a number of phosphoproteins, such as c-Jun, cyclin D1, p53, tau protein and β-catenin, to modulate their structures and functions3,5,6,7,8. Interestingly, the changes of Pin1 activity are closely related to the types of disease. For example, the up-regulation of Pin1 activity leads to various cancers, on the contrary, it causes Alzheimer’s disease2,3,5,9,10. The stability of Pin1 plays a key role in performing its normal physiological activities and functions, hence researching the factors that affect the stability of Pin1 will be conducive to the diagnosis and treatment of Pin1-related diseases10,11.
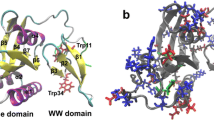
Pin1 contains 163 amino acids and consists of two domains, the WW and PPIase domain, connected by a long flexible loop12,13 (Fig. 1A). The WW domain, which contains two highly conserved tryptophan residues, possesses a substrate identification pocket surrounded by the residues Ser16, Arg17, Tyr23 and Trp3411,14. The PPIase domain is a catalytic domain to with the function isomerize the peptides interacting with the residues Lys63, Arg68, Arg69, Cys113 and Ser1547,14. Moreover, two highly conserved histidine residues His59 and His157, within the active site, form a catalytic tetrads with the residues Cys113 and Thr15215,16 (Fig. 1B). Although their sequences are conservative in the PPIase domain, the role of these histidine residues remains unclear in Pin1.
The three-dimensional structure of Pin1. (A) The Overall crystal structure of Pin1. Yellow arrows, green lines and red helixes represent β-sheet, random coil and α-helix, respectively. The model of white sticks represents residues His59 and His157, respectively. (B) The dual-histidine motif of Pin1. The model of white, blue-green and pink sticks represents histidine, cysteine and threonine residues, respectively. The black line represents hydrogen bonds.
Moreover, mutational analyses have already reported that several key amino acid residues are of great significance in Pin1, such as the residues Lys13, Trp11 and Ser32, they have played an important role in maintaining the structure and function of WW domain11,17. Similarly, the residues Val55, Cys57, Trp73, Leu61, Gly120, Ala137 and Gly155 have illustrated that they are essential to the structural stability of the PPIase domain6,11,17. Interestingly, the residues Leu61, Lys63, Ser67, Arg74 and Ala85 have possessed the function of phosphate binding6,17, and the residues Ser111 and Cys113 have owned the role of catalysis6,17. In addition, previous research has indicated that residues His59 and His157 are very vital in sustaining the domain structure or stability of Pin115,16. In the present research, the site-directed mutation of histidine to arginine was constructed, and the spectral methodologies and molecular dynamics (MD) simulations were performed to investigate the structural stability of the PPIase domain. In conclusion, the present work will contribute to further understand the mechanism by which dual histidine motif maintain the stability of the PPIase domain.
Results
Thermal stability of Pin1-WT, H59R and H157R
The fluorescence spectra can reflect accurately the information of tertiary structure and hydrophobic change concerning the microenvironments around the chromophore18,19. Generally, the spectra with λex = 295 nm (λex = excitation wavelength) reveal the microenvironments around the tryptophan residues20,21. As shown in Fig. 2A, the fluorescence intensity of Pin1-WT decreased gradually with the increase of temperature, data of mutants not shown, which indicated that the polarity around the tryptophan residues gradually increased in the unfolded process. Ordinarily, the F350/F335 ratio was used to reflect the changes in the λmax of the fluorescence spectra (λmax = max emission wavelength)11,22. The increase of F350/F335 ratio meant red-shift of λmax, conversely, the decrease of the ratio indicated blue-shift of λmax. The shifts of F350/F335 ratio (Fig. 2B) had significant differences between Pin1-WT and mutants, suggesting that histidine mutations have an impact on the thermal stability of the protein structure. In addition, the intermediate states were obviously observed at about 60 °C by normalized data of Pin1-WT, H59R and H157R, which suggested that they involve a three-state transition during thermal denaturation process20,21. Therefore, the thermal unfolding of Pin1-WT and mutants was fitted to two-step denaturation, from 20 to 60 °C and 60 to 95 °C, in order to calculate the unfolded fraction of each protein on the basis of the fluorescence intensity at 340 nm using Eq. (2) (Fig. 2C).
The thermal unfolding curves of Pin1-WT, H59R and H157R. (A) The representative fluorescence spectra of Pin1-WT. (B) The structural changes in the F350/F335 ratio for the spectra of Pin1-WT, H59R and H157R. (C) The thermal unfolding curves of Pin1-WT, H59R and H157R on the basis of fluorescence spectra with λex = 295 nm. (D) The representative far-UV CD spectra of Pin1-WT. (E) The relative intensity of far-UV CD spectra of Pin1-WT, H59R and H157R at 208 nm. (F) The thermal unfolding curves of Pin1-WT, H59R and H157R on the basis of far-UV CD spectra.
Tm, a criterion for the thermostability, defined as the midpoint of the denaturation process11,23, was summarized in Table 1. On the one hand, the Tm values of Pin1-WT, H59R and H157R were similar when the temperature was between 20 and 60 °C, indicating that the influence of histidine mutations to structural stability was not sensitive below 60 °C. On the other hand, the Tm value of H59R was the lowest and the others were similar, when the temperature was between 60 and 95 °C, suggesting that residues His59 was more sensitive to high temperature. Previous research has illustrated that histidine residues His59 and His157 play a significant role in structural stability, and His157 is not key in Pin1 function as His5915. Therefore, we deduced that the influences of residues His59 and His157 to the thermostability were similar Pin1-WT when the temperature was below 60 °C, while the influence of residue His59 to the thermostability was more sensitive when the temperature is above 60 °C.
The far-UV CD spectra can provide accurately the information of secondary structure11,19,23. The representative far-UV CD signal of Pin1-WT showed in Fig. 2D. The CD signal revealed that heat treatment changed the secondary structure of Pin1, resulting in the decreases of α-helix structure and increases of β-sheet and turn structure, respectively (Fig. 2D and Table S1). The unfolded fractions of Pin1-WT and mutants were plotted in Fig. 2E on the basis of the signal intensity at 208 nm. The free energies of unfolding \((\Delta {{G}}^{{{H}}_{{2}}{O}})\) were obtained by using the Eqs (3) and (4) (Fig. 2F). The one-step denaturation was applied to the unfolding of Pin1-WT and mutants by far-UV CD spectra and summarized in Table 1. Tm value of Pin1-WT was about 54 °C, while the histidine mutations resulted in that Tm values decrease by 3 °C, suggesting that histidine residues play an important role in maintaining the secondary structural stability of Pin1. The free energies of unfolding \((\Delta {{G}}^{{{H}}_{{2}}{O}})\) for Pin1-WT, H59R and H157R were 3.10, 2.83, 3.05 kcal mol−1, respectively, suggesting that H59R was relatively sensitive to heat and less thermostability than the others.
Acidic stability of Pin1-WT, H59R and H157R
The Fig. 3A showed the fluorescence spectra of Pin-WT by acidic treatment, data of mutants not shown. We observed that the fluorescence intensity decreased gradually in the acidic denaturation process with the decrease of acidic pH. The F350/F335 ratio was performed in the study of acid stability for Pin1-WT and mutants22,24. As shown in Fig. 3B, the F350/F335 ratio changed barely from pH 7.0 to pH 3.0, however, the ratio increased dramatically from pH 3.0 to pH 1.0. These phenomena elaborated that the acidic denaturation of Pin1-WT and mutants was a three-state transition and existed an intermediate, which are in accordance with the previous studies24. Therefore, the acidic denaturation of Pin1-WT and mutants was fitted to two-step denaturation, from pH 7.0 to pH 3.0 and pH 3.0 to pH 1.0. In order to calculate the unfolded fraction of protein on the basis of the fluorescence intensity at 340 nm, Eq. (2) was applied to fit data (Fig. 3C and Table 2).
The acidic unfolding curves of Pin1-WT, H59R and H157R. (A) The representative fluorescence spectra of Pin1-WT. (B) The structural changes in the F350/F335 ratio for the spectra of Pin1-WT, H59R and H157R. (C) The acidic unfolding curves of Pin1-WT, H59R and H157R on the basis of fluorescence spectra with λex = 295 nm. (D) The representative far-UV CD spectra of Pin1-WT. (E) The relative intensity of far-UV CD spectra of Pin1-WT, H59R and H157R at 208 nm. (F) The acidic unfolding curves of Pin1-WT, H59R and H157R on the basis of far-UV CD spectra.
pKa, the midpoint pH value in the unfolding process, is a criterion for the acidic stability of the protein structure25,26. For one thing, the pKa value of H59R was the highest, and the others was similar, when pH was between 7.0 and 3.0, suggesting that residue His59 was relatively sensitive to acid. For another, the opposite results were observed that the pKa value of H157R was the highest when pH is between 3.0 and 1.0. These results indicated that influences of both residues His59 and His157 to acidic stability were difference from Pin1-WT.
The representative far-UV CD signal of Pin-WT, data of mutants not shown, was plotted in Fig. 3D. The CD signal showed two typical negative bands at 208 and 217 nm in a native state. However, the acidic treatment changed greatly the shape of CD spectra, resulting in that the negative band at 217 nm faded away with the decrease of acidic pH11,24. In addition, the signal changes at 208 nm of Pin-WT and mutants decreased gradually from pH 7.0 to pH 1.0, suggesting the decrease of α-helix structures (Fig. 3E and Table S2). The unfolded fractions were showed in Fig. 3F on the basis of the signal intensity at 208 nm. The one-step denaturation was performed to the unfolding of Pin1-WT and mutants by far-UV CD spectra11. As seen in Table 2, the pKa value of Pin-WT was the lowest, suggesting mutants were more sensitive to acid. These results further illustrated that the histidine residues are very important to the secondary stability of Pin1.
The ANS fluorescence spectra were carried out to assess the exposure of the hydrophobic regions in Pin1-WT and mutants22,24. The representative ANS spectra of Pin-WT, data of mutants not shown, was plotted in Fig. 4A. Similarly, the fluorescence intensity at 488 nm was showed in Fig. 4B. These facts illustrated that ANS intensity was very weak from pH 7.0 to pH 5.0, while it increased dramatically from pH 5.0 to pH 3.0 and remained constant from pH 3.0 to pH 1.0. To calculate the hydrophobic parameters by fitting data, pKb is the pH of one half of full exposure by ANS spectra (Table 2). The pKb value of Pin1-WT was in accordance with these mutants at about pH 4.5, implying that the histidine residues had a little effect on the acidic pH-induced hydrophobic change.
The ANS and RLS spectra of Pin1-WT, H59R and H157R by acidic pH treatments. (A) The representative ANS fluorescence spectra of Pin1-WT. (B) Relative ANS intensity at 488 nm of Pin1-WT, H59R and H157R. (C) The representative RLS fluorescence signal of Pin1-WT. (D) Relative RLS intensity at 440 nm of Pin1-WT, H59R and H157R.
The RLS fluorescence spectra are usually implemented to evaluate the aggregation and size of Pin1-WT and mutants in unfolding process19,24. The representative RLS spectra of Pin-WT and the fluorescence intensities were shown in Fig. 4C,D, respectively. pKc, which is defined as the pH of the maximum RLS fluorescence intensity at 450 nm, was summarized in Table 2. The pKc value of Pin1-WT was in accordance with these mutants at about pH 3.3, indicating that the histidine residues to the acidic pH-induced aggregation were not a significant influence.
Chemical stability of Pin1-WT, H59R and H157R
The representative fluorescence spectra of Pin-WT, data of mutants not shown, was plotted in Fig. 5A by GndHCl treatment. The λmax of native Pin1-WT and mutants were located at about 342 nm, while the GndHCl denaturation shifted λmax to about 350 nm, indicating that the tryptophan residues in Pin1-WT and mutants were fully exposed after GndHCl denaturation. The F350/F335 ratio was used in GndHCl induced denaturation for Pin1-WT and mutants11. As shown in Fig. 5B, the F350/F335 ratio has hardly changed in the GndHCl of low concentration (<2.0 mol/l), however, the ratio increased significantly with the increase of GndHCl concentration. The unfolded fraction of each protein was showed in Fig. 5C. Cm, the unfolding transition midpoint concentration, is a measure for the chemical stability of protein structure11,19. Obviously, the unfolding curves and Cm values of these histidine mutants were significantly different from Pin1-WT, suggesting that the residues His59 and His157 contribute to the chemical stability of Pin1 (Table 3).
The chemical unfolding curves of Pin1-WT, H59R and H157R. (A) The representative fluorescence spectra of Pin1-WT. (B) The structural changes in the F350/F335 ratio for the spectra of Pin1-WT, H59R and H157R. (C) The chemical unfolding curves of Pin1-WT, H59R and H157R on the basis of fluorescence spectra with λex = 295 nm. (D) The representative far-UV CD spectra of Pin1-WT. (E) The relative intensity of far-UV CD spectra of Pin1-WT, H59R and H157R at 222 nm. (F) The chemical unfolding curves of Pin1-WT, H59R and H157R on the basis of far-UV CD spectra.
The representative far-UV CD signal of Pin-WT, data of mutants not shown, was plotted in Fig. 5D by GndHCl treatment. With the increasing concentration of GdnHCl, the CD spectra were greatly changed and negative bands at 222 nm faded away, indicating the decreases of α-helix structure and increases of β-sheet and turn structure (Table S2). Moreover, the relative CD intensity at 222 nm decreased to the minimum at 7.0 M GdnHCl, implying complete denaturation of Pin1-WT and mutants (Fig. 5E). The unfolding curves of H59R and H157R were similar and their Cm values were less than Pin1-WT, meaning that histidine mutants were sensitive to the GdnHCl (Fig. 5F and Table 3). These phenomena were consistent with thermal and acidic denaturation by CD spectra, indicating that the histidine residues play a significant role in maintaining the secondary stability.
The free energies of unfolding (ΔG0) and rate constants of unfolding (Kobs) in Pin1-WT and mutants were calculated by Eqs (5) and (6), respectively7,20. The ΔG0 of H59R and H157R were similar and greater than Pin1-WT as revealed by fluorescence and far-UV CD spectra, suggesting that these histidine mutations is conducive to unfolding for Pin1 (Fig. 6A,C and Table 3). The Kobs, which reflect the rate of unfolding, also illustrated that histidine mutations have an influence on the unfolding rate (Fig. 6B,D and Table 3).
Chemical-induced conformational changes. (A,B) Represent free energy of unfolding (ΔG0) and unfolding kinetics rate constants (log Kui) against GdnHCl on the basis of fluorescence spectra, respectively. (C,D) Represent free energy of unfolding (ΔG0) and unfolding kinetics rate constants (log Kui) against GdnHCl on the basis of far-UV CD spectra, respectively. Intercepts at y axis give ΔG0 and log Kobs, respectively.
Overall dynamics behavior of the Pin1-WT, H59R and H157R
It is well known that structural stability and flexibility can be altered by mutation methods27,28. Therefore, Cα-RMSD and Cα-RMSF were plotted in Fig. 7 to assess structural stability and flexibility, respectively. In Fig. 7A, the global structure Cα-RMSD values for Pin1-WT and mutants reached equilibrium after 5 ns with an average Cα-RMSD value of 3 Å. Previous researches have demonstrated that the movement of the WW and PPIase domain was relatively independent, so we calculated Cα-RMSD values of the WW and PPIase domain, respectively28. As shown in Fig. 7B, the WW domain of Pin1-WT and mutants shared similar RMSD values, indicating that the effect of histidine mutations was very weak for WW domain. However, there were obvious differences in the PPIase domain between Pin1-WT and mutants, with 3.54, 3.67, 3.85 Å of the average RMSD values (Table S4), respectively. The results further indicated that residues His59 and His157 have an effect on the stability of PPIase domain due to these histidine residues were located in this region (As seen in Fig. 1A). Bailey et al. works revealed that the dual histidine motif (His59 and His157) has contribution to structural stability rather than function in Pin115. Combined with the results above, our work also implied the similar result about the influence of residues His59 and His157 to the PPIase domain of structural stability in the PPIase domain.
Usually, Cα-RMSF value provides the information for the structural flexibility and mobility of each protein22,27. Compared with Pin-WT, H59R and H157R may have an influence on the overall structural flexibility and mobility, with the average RMSF values increased by 0.59 and 0.92 Å, respectively, (Fig. 7D and Table S4). As seen in Fig. 7E, the significant difference of Cα-RMSF values occurred at residues 15–25 in the WW domain, which consist of a flexible loop between β1 and β2. In Fig. 7F, H59R had an impact on the RMSF values of the residues 67–72, which are located at a functional loop to bind substrates. However, H157R affected both the RMSF values of residues 95–103 and 125–145, which are located at α1/α2 and α4 structure (Fig. 1A). Generally, the active center of Pin1 in the PPIase domain is composed of residues Lys63, Arg68, Arg69, Cys113 and Ser15412,13. Therefore, we conjectured that the residue His59 might affect the structural flexibility and mobility of active center in Pin1 due to its spatial position is close to the key catalytic residues Cys11315,29. Moreover, the hydrophobic pocket are mainly composed of residues His59, Ser114, Ala116, Leu122, Gln129, Gln131, Phe134 and His157 in the PPIase domain14,30. We guessed that H157 affects possibly the structural flexibility and mobility of the hydrophobic pocket.
Analysis of distance and Rg in the Pin1-WT, H59R and H157R
Generally, the function of the WW domain is mainly to identify the substrate, while the PPIase domain is the active center to catalyze substrate11,30. It is very meaningful to evaluate the distance between the WW and PPIase domain for analyzing the structure and function of Pin1. In Fig. 8A, the distance values of H59R and H157R were significantly higher than Pin1-WT after 5 ns MD simulations. Similarly, the average distance values of H59R (21.51 Å) and H157R (22.63 Å) were about the twofold increase, compared with Pin-WT (10.94 Å). Combined with the above results of RLS spectra, the RLS intensity of Pin1-WT at 450 nm was less than H59R and H157R, indicating that their sizes increase in the unfolding process. Therefore, we speculated that the histidine mutations may cause that the WW domain moves away from the PPIase domain in Pin1.
The radius of gyration (Rg), which is defined as a radial distance of atoms from the center of mass, usually estimate the compactness of protein28 (Fig. 8B–D). Obviously, the Rg value of global structure for H157R was greater than the others, suggesting that residue H157 may have an influence on the compactness of Pin1 (Fig. 8B and Table S4). The further analysis has shown that Rg values of the WW domain for Pin1-WT and mutants did not change significantly (Fig. 8C and Table S4). However, the Rg values of the PPIase domain were more sensitive to mutants, indicating that residues His59 and His157 mutations may result in a decrease in the compactness of the PPIase domain. (Fig. 8D and Table S4).
Analysis of hydrogen bonds of the dual-histidine motif in the Pin1-WT, H59R and H157R
Previous studies have shown that a dual histidine motif is composed of residues His59 and His157 and they form a hydrogen bond network with residues Cys113 and Thr15215,16 (As seen in Fig. 1B). Therefore, the hydrogen bonds from the hydrogen bond network were calculated by molecular dynamic (MD) simulation method. Meanwhile, the backbone hydrogen bonds also were calculated by VMD1.9.2 program to estimate overall structural stability31. As seen in Table S4, compared with Pin1-WT, the average numbers of the backbone hydrogen bonds of H59R and H157R were decreased about two hydrogen bonds. Interestingly, the two missing hydrogen bonds located possibly at dual histidine motif owing to residues His59 and His157 mutations resulted in that the hydrogen bond network has been destroyed.
We calculated the distances changes in the hydrogen bond network between partial atom pairs along with the 20 ns simulation time32. The Cys113: SG forms a hydrogen bond with His59: NE2, His59: ND1 forms a hydrogen bond with His157: ND1, and His157:NE2 forms a hydrogen bond with Thr152: OG1. In crystal structure of Pin1 (PDB code: 1PIN), the distance between Cys113: SG and His59: NE2 is 3.3 Å, it was consistent with the average distance of Pin1-WT (3.4 Å) and less than H59R (4.2 Å) and H157R (3.8 Å) along with MD simulation (Fig. 1B, Fig. 9 and Table S5). Moreover, the hydrogen bond occupancy between Cys113: SG and His59:NE2 was very short in these histidine mutants (Table S5). For the distance between His59:ND1 and His157:ND1, the crystal distance was observed about 2.9 Å, however, the average distance was 3.5 Å (WT-Pin1), 4.1 Å (H59R), 4.8 Å (H157R) along with MD simulation, respectively (Fig. 9 and Table S5). Interestingly, the occupancy of H59R between His59:ND1 and His157:ND1 was less than the others. Moreover, the distance between His157:NE2 and Thr152:OG1 was about 2.7 Å in the crystal structure, whereas the average distance and occupancy of H157R (4.8 Å and 19.1%) were significantly different from Pin-WT (2.9 Å and 96.6%) and H59R (3.5 Å and 63.1%), respectively (Table S5). These results illustrated that the hydrogen bond network was destroyed in the dual histidine motif by residues His59 and His157 mutations.
The distance between dual-histidine motif in Pin1-WT, H59R and H157R as a function of 20 ns MD simulation. (A,B) Represent the distance and model of the dual-histidine motif in Pin1-WT, respectively. (C,D) Represent the distance and model of the dual-histidine motif in H59R, respectively. (E,F) Represent the distance and model of the dual-histidine motif in H157R, respectively. The black line represents the cut-off distance of hydrogen bond as 3.5 Å.
Discussion and Conclusion
Pin1, a peptidyl-prolyl cis-trans isomerase, plays a key role in tumorigenesis and Alzheimer’s disease (AD)3,33. Several clinical trials have reported that Pin1 overexpression is closely related to multiple malignant cancers7,8,34,35,36, such as the colon, ovarian, cervical, endometrial, breast, prostatic and liver cancer. Whereas Pin1 inactivation is a potential cause of Alzheimer’s disease9,33. Up to now Pin1 exhibits a unique characteristic to specifically isomerize cis/trans conformation for p-Ser/Pro or p-Thr/Pro motifs, facilitating kinds of dephosphorylation pathway3,37. Isomerization of these motifs occurs in a modular fashion: the WW domain firstly recognizes them and then catalyzes them through the catalytic PPIase domain38.
A dual histidine motif, which consists of residues His59 and His157, forms a hydrogen bond network with residues Cys113 and Thr152 in the active site of the PPIase domain39,40. Zhou et al. and Behrsin et al. illustrated that mutants H157A/L/N still have partially activities6,16. Notably, Bailey et al. revealed that mutants H59X (X = A/N/S) and H157X (X = L/A/N/F/S) support partly viability, suggesting that the dual histidine motif has a structural role instead of catalytic function15. Moreover, Bailey et al. indicated that residue His157 is less important for enzyme function than His5915. Tossavainen et al. also reported the residue His59 plays a key role in the correct folding of the PPIase domain of PrsA from Bacillus subtilis41.
Exploring the structure and function of the dual histidine motif is necessary to understand the mechanism of the structural stability in the active site of Pin1 PPIase domain. Therefore, the H59R and H157R mutants were constructed for the following reasons: (1) Both arginine and histidine are basic amino acids with similar chemical properties; (2) Arginine mutants have not been studied and reported so far. Interestingly, my colleague measured the enzyme activity of Pin1-WT, H59R and H157R according to Behrsin et al.16 research, data not published, and the result suggested that two mutants have still partially activity (about 50%).
In the present work, the structural stability of wild-type and histidine mutants in Pin1 were extensive studies by thermal, acidic, chemical stability and MD simulation. Thermal denaturation revealed that both His59 and His157 are not sensitive to the lower temperatures, while His59 is more sensitive to the higher temperatures than His157, suggesting that His59 is possibly more important to maintain structural stability than His157. Wang et al. used site-directed mutagenesis to construct tryptophan mutants, including W11L, W34L and W73L, and illustrated that similar phenomenon about mutations reduce thermal stability11. Similarly, Bailey et al. also suggested that mutant H59L is less stable at higher temperatures, at least 50 °C, than H157L16. Acidic denaturation implied that influences of both residues His59 and His157 to acidic stability were difference from Pin1-WT. According to the crystal structure14,29, residue His59 is spatially closer to Cys113, which is considered as a critical catalytic residue in Pin1, than His157, indicating that it may play a role in catalysis. This may be one of the reasons that their acid stability is different. Moreover, the ANS and RLS spectra indicated that the histidine residues had little effect on the acidic pH-induced hydrophobic change and aggregation. One of the possible reasons is that a single amino acid mutation has little effect on the overall hydrophobicity and aggregation of the protein. Obviously, chemical denaturation suggested that both residues His59 and His157 contributed the most to the chemical stability of Pin1. Previous laboratory studies have indicated that key residue mutations cause a decrease in chemical stability and are more prone to unfolding11.
MD simulation can provide an atomic-level information for protein structures, which is difficult to obtain from traditional experiments. RMSD values suggested the histidine mutations mainly affect the structural stability of the PPIase domain rather than the WW domain. Similarly, RMSF values implied that residues His59 and His157 primarily affect the flexibility of the active center in the PPIase domain. It is well known that residues His59 and His157 are located in adjacent antiparallel β chains of the PPIase domain, so their mutations do not affect the WW domain15,29. Meanwhile, these results further indicated that residues His59 and His157 play an important role in stabilizing the PPIase domain. Obviously, our results have shown that the hydrogen bond network in the dual histidine motif was destroyed by residue His59 and His157 mutations. Chang et al. reported that mutant C113D destroyed the hydrogen bond network, leading to the instability of the catalytic tetrads (Cys113 -His59-His157-Thr152 motif)42. Barman et al. illustrated that the hydrogen bond network was rearranged by the protonation of residue Cys113, with the switching of the tautomeric states of the dual histidine motif43. Similarly, Wang et al. demonstrated that residues Cys113 and Ser138 modulate the hydrogen bond network dynamics due to the allosteric breakage of the hydrogen bond within the dual histidine motif in the PPIase domain39,40. These results suggested that the hydrogen bond network plays a very important role in stabilizing the PPIase domain, and a single residue mutation, including residues Cys113, His59, His157, and Thr152, has an effect on its stability. All in all, this work is based on the research of Bailey et al.15 to reveal deeply the mechanism by which residues His59 and His157 maintain the stability of the PPIase domain.
Methods
Preparation of Pin1-WT and mutants
The recombinant plasmids pET-19b-Pin1, pET-19b-H59R and pET-19b-H157R were expressed in E. coli BL21, which were induced with a specific condition (0.5 nM, 30 °C, 4 h)11. The interest proteins were purified by Ni-NAT Sepharose on an AKTA FPLC30. His-tag was excised by enterokinase and then separated through Amicon Ultra-15 centrifugal filter. The purity and concentration were detected by SDS-PAGE and Bradford assay, respectively44.
Measurements of fluorescence spectra
The fluorescence spectra were measured on a F-4500 fluorescence spectrophotometer (Hitachi) with λex = 295 nm (310–400 nm), which were set as 5 nm excitation and 10 nm emission slit11,30. The ANS (8-anilino-1-naphthalenesulfonic acid) fluorescence spectra were recorded with λex = 380 nm (400–600 nm), setting as 5 nm excitation and 5 nm emission slit, respectively22,45. The RLS (Resonance light scattering) fluorescence spectra were recorded with Δλ = 0 nm (400–600 nm), setting as 2.5 nm excitation and 2.5 nm emission slit, respectively24. The concentration of Pin1-WT and mutants set to 5 μM.
To avoid the inner filter effect, the fluorescence intensities were corrected using the following relationship46,47:
where Fcor and Fobs are the corrected fluorescence intensity and observed fluorescence intensity, respectively. Aex and Aem represent the absorption of the excitation and the emission wavelength, respectively.
Measurements of far-UV circular dichroism (CD) spectra
The far-UV CD spectra were recorded between 190 to 260 nm in 2 mm cuvettes by an Aviv Model 400 circular dichroism spectrophotometer (AVIV)28,48. The solvent spectrum was subtracted and the CDNN software was used to calculate the content of the secondary structure11. The concentration of Pin1-WT and mutants set to 10 μM.
Thermal denaturation analysis of Pin1-WT and mutants
Fluorescence and far-UV CD measurements were implemented to assess the impacts of the histidine mutations on the thermal stability of Pin111,20,21. Therefore, for fluorescence experiments, the thermal stability of Pin1-WT and mutants were performed from 20 to 95 °C at an interval of 5 °C using peltier temperature controller. The samples (5 μM) of Pin1-WT and mutants were incubated a given conditions (20 mM Tris-HCl, pH 7.0) for 5 min. For far-UV CD experiments, the samples (10 μM) were incubated for 5 min from 20 to 80 °C, and the ellipticity at 208 nm was recorded.
In the unfolding studies, the fraction of Pin1-WT and mutants at each temperature was calculated using the following equation11,19:
where fu is the fraction of unfolded Pin1-WT and mutants at a given temperature, Fobs is the fluorescence intensity or CD signal intensity at the temperature, Fn and Fd are the fluorescence intensity or CD signal intensity of the native and denatured protein, respectively. Tm, a temperature of the denaturation process at the midpoint, was calculated by the plotting fu against temperature11,23,49.
Based on a two-state model, the apparent Gibbs free energy (ΔGu) was calculated using the following equation at each temperature19,50:
where R and T are the ideal gas constant and specific temperature, respectively. Keq is the fraction of unfolded protein at each temperature. The apparent Gibbs free energy (ΔGu) is related to the free energy of unfolding \((\Delta {{G}}^{{{H}}_{{2}}{O}})\) by the following equation23,51:
where m is the experimental measure of the dependence of ΔGu on temperature.
Acidic denaturation analysis of Pin1-WT and mutants
The acidic-pH induced denaturation of Pin1-WT and mutants were evaluated by fluorescence and far-UV CD measurements24. The solution of Pin1-WT and mutants were added to the different acidic pH buffer. The ANS fluorescence spectra were carried out to assess the extent of hydrophobic surface exposure in Pin1-WT and mutants, and the fluorescence intensity at 488 nm was recorded24. The RLS fluorescence spectra were implemented to assess the aggregation of Pin1-WT and mutants, and the signal intensity at 450 nm was recorded to gauss curves in origin 8.5 software.
Chemical denaturation analysis of Pin1-WT and mutants
The chemical denaturation of Pin1-WT and mutants were assessed by fluorescence and far-UV CD experiments. The samples were incubated at GdnHCl solution of different concentrations for 2 h at room temperature. The unfolding constants for Pin1-WT and mutants were calculated according to the following equation19,50:
where Kui is unfolding constant, Pt is the molecular concentration of protein when all the protein is folded, fu is the fraction of unfolded, n is the number of the chain formed after denaturation.
The free energy (ΔGui) of folding at any given concentration was evaluated from the following equation19,50:
where R and T are the ideal gas constant and specific temperature, respectively. Free energy of protein unfolding (ΔG0) was obtained from a plot of ΔGui (y) as a function of GdnHCl concentrations (x) where y intercept equals to the ΔG0 value. Similarly, the values of unfolding kinetics rate constants (Kobs) was determined from a half-Chevron plot of Kui (y) as a function of GdnHCl concentrations (x) where y intercept equals to the Kobs value.
Molecular dynamics simulation
The crystal structure of Pin1 was downloaded from Protein Data Base (PDB code: 1PIN)14,52. The water molecules, ions and ligands were removed and missing residues were added by the Modeller 9.16 software53. Two mutants H59R and H157R were constructed by using Pymol 1.X program54.
The MD simulations of Pin1-WT and mutants were performed by Gromacs 4.6.5 with Gromacs 96–53a6 force filed55,56,57. Before the formal simulation, the energy minimization was carried out for all systems22,27. Firstly, steepest descent method was adopted to energy minimization of 1000 steps and then heated to 300 K in vacuum27. Secondly, all the systems were immersed in a cubic box and solvated with SPC water patterns using a solute-box distance of 1.2 nm27. To neutralize the system, Na+ and Cl− ions were inserted in the solute-box58,59. Thirdly, all systems were further performed energy minimization of 100 ps in the NVT ensemble, and subsequently equilibrated for 100 ps in the NPT ensemble30. All systems were performed to 20 ns simulation in the NPT ensemble. The temperature and pressure were controlled at 300 K and 1 atm by the methods of Velocity-Rescaling and Parrinello-Rahman, respectively60. The short-range interactions for cutoff and switch distance were set to 12 Å and 10 Å, respectively61. The long-range electrostatic interaction was described by Particle mesh Ewald (PME) method62. LINCS (Linear Constraint Algorithm) method was used to constrain the hydrogen bonds63.
References
Lu, K. P., Hanes, S. D. & Hunter, T. A human peptidyl–prolyl isomerase essential for regulation of mitosis. Nature 380, 544–547 (1996).
Driver, J. A., Zhou, X. Z. & Lu, K. P. Regulation of protein conformation by Pin1 offers novel disease mechanisms and therapeutic approaches in Alzheimer’s disease. Discovery. Medicine 17, 93–99 (2014).
A Driver, J. & Lu, K. P. Pin1: a new genetic link between Alzheimer’s disease, cancer and aging. Current aging science 3, 158–165 (2010).
Wulf, G. M. et al. Role of Pin1 in the regulation of p53 stability and p21 transactivation, and cell cycle checkpoints in response to DNA damage. Journal of Biological Chemistry 227, 47976–47979 (2002).
Lu, P. J. et al. The prolyl isomerase Pin1 restores the function of Alzheimer-associated phosphorylated tau protein. Nature 399, 784–788 (1999).
Zhou, X. Z. et al. Pin1-dependent prolyl isomerization regulates dephosphorylation of Cdc25C and tau proteins. Molecular cell 6, 873–883 (2000).
Wulf, G. M. et al. Pin1 is overexpressed in breast cancer and cooperates with Ras signaling in increasing the transcriptional activity of c-Jun towards cyclin D1. The EMBO journal 20, 3459–3472 (2001).
Yeh, E. S. & Means, A. R. PIN1, the cell cycle and cancer. Nature Reviews Cancer 7, 381–388 (2007).
Driver, J. A., Zhou, X. Z. & Lu, K. P. Pin1 dysregulation helps to explain the inverse association between cancer and Alzheimer’s disease. Biochim Biophys Acta. 1850, 2069–2076 (2015).
Wang, J. Z. et al. Aluminum(III) interferes with the structure and the activity of the peptidyl-prolyl cis-trans isomerase (Pin1): a new mechanism contributing to the pathogenesis of Alzheimer’s disease and cancers? Journal of Inorganic Biochemistry 126, 111–117 (2013).
Wang, J. Z. et al. The structural and functional role of the three tryptophan residues in Pin1. Journal of Photochemistry & Photobiology B Biology 146, 58–67 (2015).
Guo, C. et al. Structure-based design of novel human Pin1 inhibitors (I). Bioorganic & medicinal chemistry letters 19, 5613–5616 (2009).
Dong, L. et al. Structure-based design of novel human Pin1 inhibitors (II). Bioorganic & Medicinal Chemistry Letters 20, 2210–2214 (2010).
Urusova, D. V. et al. Epigallocatechin-gallate suppresses tumorigenesis by directly targeting Pin1. Cancer prevention research 4, 1366–1377 (2011).
Bailey, M. L. et al. The dual histidine motif in the active site of Pin1 has a structural rather than catalytic role. Biochemistry 47, 11481–11489 (2008).
Behrsin, C. D. et al. Functionally important residues in the peptidyl-prolyl isomerase Pin1 revealed by unigenic evolution. Journal of Molecular Biology 365, 1143–1162 (2007).
Wu, X. et al. The Ess1 prolyl isomerase is linked to chromatin remodeling complexes and the general transcription machinery. The EMBO Journal 19, 3727–3738 (2014).
Yang, H. et al. Binding modes of environmental endocrine disruptors to human serum albumin: insights from STD-NMR, ITC, spectroscopic and molecular docking studies. Scientific Reports 7, 11126 (2017).
Devesh, K., Suman, K. & Kayastha, A. M. Thermal, Chemical and pH Induced Denaturation of a Multimeric β-Galactosidase Reveals Multiple Unfolding Pathways. Plos One 7, 304–309 (2012).
Wang, J. Z. et al. Spectroscopic studies on the irreversible heat-induced structural transition of Pin1. Spectrochimica Acta Part A Molecular & Biomolecular Spectroscopy 78, 142–147 (2011).
Wang, J. Z. et al. Stability of Pin1 as revealed by thermal and spectroscopic studies. Journal of Molecular Structure 975, 310–316 (2010).
Wang, W. et al. The acidic pH-induced structural changes in apo-CP43 by spectral methodologies and molecular dynamics simulations. Journal of Molecular Structure 1152, 177–188 (2018).
Zhu, G. F. et al. Temperature induced structural transitions from native to unfolded aggregated states of tobacco etch virus protease. Journal of Molecular Structure 1082, 80–90 (2015).
Wang, J. Z. et al. The acidic pH-induced structural changes in Pin1 as revealed by spectral methodologies. Spectrochim Acta A Mol Biomol Spectrosc 98, 199–206 (2012).
Kojima, M. et al. pH-dependent unfolding of aspergillopepsin II studied by small-angle X-ray scattering. Biochemistry 39, 1364–1372 (2000).
Dubey, V. K. & Jagannadham, M. V. Differences in the unfolding of procerain induced by pH, guanidine hydrochloride, urea, and temperature. Biochemistry 42, 12287–11297 (2003).
Zhao, M. L. et al. In silico structure prediction and inhibition mechanism studies of AtHDA14 as revealed by homology modeling, docking, molecular dynamics simulation. Computational Biology & Chemistry 75, 120–130 (2018).
Zhu, G. F. et al. Interaction between ginkgolic acid and human serum albumin by spectroscopy and molecular modeling methods. Journal of Solution Chemistry 43, 1232–1249 (2014).
Ranganathan, R. et al. Structural and functional analysis of the mitotic rotamase Pin1 suggests substrate recognition is phosphorylation dependent. Cell 89, 875–886 (1997).
Xi, L. et al. Interaction between Pin1 and its natural product inhibitor epigallocatechin-3-gallate by spectroscopy and molecular dynamics simulations. Spectrochimica Acta Part A Molecular & Biomolecular. Spectroscopy 169, 134–143 (2016).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. Journal of Molecular Graphics 14, 33–38 (1996).
Wang, Y. et al. Insight into the structural stability of wild type and mutants of the tobacco etch virus protease with molecular dynamics simulations. Journal of Molecular Modeling 19, 4865–4875 (2013).
Lee, T. H., Pastorino, L. & Lu, K. P. Peptidyl-prolyl cis–trans isomerase Pin1 in ageing, cancer and Alzheimer disease. Expert reviews in molecular medicine 13, 1–26 (2011).
Bao, L. et al. Prevalent overexpression of prolyl isomerase Pin1 in human cancers. The American journal of pathology 164, 1727–1737 (2004).
Kim, G., Bhattarai, P. Y. & Choi, H. S. Peptidyl-prolyl cis/trans isomerase NIMA-interacting 1 as a molecular target in breast cancer: a therapeutic perspective of gynecological cancer. Archives of pharmacal research 42, 128–139 (2019).
Wu, K. J. et al. Small Molecule Pin1 Inhibitor Blocking NF‐κB Signaling in Prostate Cancer Cells. Chemistry–An Asian Journal 13, 275–279 (2018).
Cheng, C. W. & Tse, E. PIN1 in cell cycle control and cancer. Frontiers in pharmacology 9, 1–10 (2018).
Pu, W. et al. Targeting Pin1 by inhibitor API-1 regulates microRNA biogenesis and suppresses hepatocellular carcinoma development. Hepatology 68, 547–560 (2018).
Wang, J. et al. Allosteric breakage of the hydrogen bond within the dual-histidine motif in the active site of human Pin1 ppiase. Biochemistry 54, 5242–5253 (2015).
Wang, J. et al. Dynamic allostery modulates catalytic activity by modifying the hydrogen bonding network in the catalytic site of human pin1. Molecules 22, 992 (2017).
Tossavainen, H. et al. NMR solution structure and characterization of substrate binding site of the PPIase domain of PrsA protein from Bacillus subtilis. Febs Letters 580, 1822–1826 (2006).
Chang, S. et al. Molecular dynamics simulations of wild type and mutant of Pin1 peptidyl-prolyl isomerase. Molecular Simulation 42, 1281–1291 (2016).
Barman, A. & Hamelberg, D. Cysteine-mediated dynamic hydrogen-bonding network in the active site of Pin1. Biochemistry 53, 3839–3850 (2014).
Zor, T. & Selinger, Z. Linearization of the Bradford protein assay increases its sensitivity: theoretical and experimental studies. Analytical biochemistry 236, 302–308 (1996).
Poggetto, E. D., Chiti, F. & Bemporad, F. The Folding process of Human Profilin-1, a novel protein associated with familial amyotrophic lateral sclerosis. Scientific Reports 5, 12332 (2015).
Parker, C. A. & Rees, W. T. Correction of fluorescence spectra and measurement of fluorescence quantum efficiency. Analyst 85, 16–27 (1960).
Lakowicz, J. R. Principles of Fluorescence Spectroscopy. Springer Science & Business Media (2013).
Du., W. et al. Spectroscopic studies on the interaction of bovine serum albumin with ginkgolic acid: Binding characteristics and structural analysis. Journal of Luminescence 132, 1207–1214 (2012).
Xiao, Q. J. et al. Heat-induced unfolding of apo-CP43 studied by fluorescence spectroscopy and CD spectroscopy. Photosynthesis Research 126, 427–435 (2015).
Greenfield, N. J. Determination of the folding of proteins as a function of denaturants, osmolytes or ligands using circular dichroism. Nature Protocols 1, 2733–2741 (2006).
Rabbani, G., Ahmad, E., Zaidi, N. & Khan, R. H. pH-Dependent Conformational Transitions in Conalbumin (Ovotransferrin), a Metalloproteinase from Hen Egg White. Cell Biochemistry & Biophysics 61, 551–560 (2011).
HM, B. et al. The Protein Data Bank. Genetica 106, 149–158 (2000).
Eswar, N. et al. Protein structure modeling with MODELLER. Structural proteomics: high-throughput methods 1137, 145–159 (2008).
DeLano, W. L. Pymol: An open-source molecular graphics tool. CCP4 Newsletter On Protein Crystallography 40, 82–92 (2002).
Berendsen, H. J., van der Spoel, D. & van Drunen, R. GROMACS: a message-passing parallel molecular dynamics implementation. Computer physics communications 91, 43–56 (1995).
Hess, B. et al. GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. Journal of chemical theory and computation 4, 435–447 (2008).
Lindahl, E., Hess, B. & Spoel, D. V. D. GROMACS 3.0: a package for molecular simulation and trajectory analysis. Molecular modeling annual 7, 306–317 (2001).
Wang, X. et al. In silico identification of small molecules as novel LXR agonists for the treatment of cardiovascular disease and cancer. Journal of Molecular Modeling 24, 57 (2018).
Li, J. et al. Structure and energy based quantitative missense variant effect analysis provides insights into drug resistance mechanisms of anaplastic lymphoma kinase mutations. Scientific Reports 8, 10664 (2018).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. Journal of Chemical Physics 126, 104010 (2007).
Berendsen, H. J. C. et al. Molecular dynamics with coupling to an external bath. Journal of Chemical Physics 81, 3684–3690 (1984).
Essmann, U. et al. A smooth particle mesh Ewald method. Journal of Chemical Physics 103, 8577–8593 (1995).
Hess, B. et al. LINCS: a linear constraint solver for molecular simulations. Journal of computational chemistry 18, 1463–1472 (1997).
Acknowledgements
The authors thank Hai-Jun Cao from the institute of blood transfusion (IBT) for their kind help with the far-UV CD measurements. This work was financially supported by the National Key Technology R&D program of China (2009BAK61B04, 2006BAF07B01) and Science & Technology Foundation of Sichuan Province (2011JTD0026).
Author information
Authors and Affiliations
Contributions
L.D. conceived the idea and supervised the project. W.W. performed molecular dynamics simulation experiments and wrote the manuscript. L.X. carried out spectral experiments. W.W., X.X. and X.L. analyzed the data. X.L., Q.Z., L.D. and W.Y. edited the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, W., Xi, L., Xiong, X. et al. Insight into the structural stability of wild-type and histidine mutants in Pin1 by experimental and computational methods. Sci Rep 9, 8413 (2019). https://doi.org/10.1038/s41598-019-44926-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-44926-5
- Springer Nature Limited