Abstract
Long non-coding RNAs (lncRNAs) can regulate gene expression directly or indirectly through interacting with microRNAs (miRNAs). However, the role of differentially expressed mRNAs, lncRNAs and miRNAs, and especially their related competitive endogenous RNAs (ceRNA) network in head and neck squamous cell carcinoma (HNSCC), is not fully comprehended. In this paper, the lncRNA, miRNA, and mRNA expression profiles of 546 HNSCC patients, including 502 tumor and 44 adjacent non-tumor tissues, from The Cancer Genome Atlas (TCGA) were analyzed. 82 miRNAs, 1197 mRNAs and 1041 lncRNAs were found to be differentially expressed in HNSCC samples (fold change ≥ 2; P < 0.01). Further bioinformatics analysis was performed to construct a lncRNA-miRNA-mRNA ceRNA network of HNSCC, which includes 8 miRNAs, 71 lncRNAs and 16 mRNAs. Through survival analysis based on the expression profiles of RNAs in the ceRNA network, we detected 1 mRNA, 1 miRNA and 13 lncRNA to have a significant impact on the overall survival of HNSCC patients (P < 0.05). Finally, some lncRNAs, which are more important for survival, were also predicted. Our research provides data to further understand the molecular mechanisms implicated in HNSCC.
Similar content being viewed by others
Introduction
Head and neck squamous cell carcinoma (HNSCC) is the sixth most commonly reported malignancy worldwide1. There are about 500,000 new HNSCC patients diagnosed worldwide each year, and the death rate of new cases is about 20%2,3. Although comprehensive treatment, including surgical resection, radiotherapy, and chemotherapy, continues to develop, the overall 5-year survival rate of HNSCC patients has not been significantly improved over the past few decades and has not exceeded 50%. Thus, HNSCC is considered a malignant tumor with low survival rate4. Therefore, understanding the molecular nature of HNSCC carcinogenesis is a pivotal step for meliorating early diagnosis, predicting prognosis, and developing effective therapeutics5.
In recent years, the regulatory network composed of long non-coding RNAs (lncRNAs), microRNAs (miRNAs) and messenger RNAs (mRNAs) has gained a great interest in the study of molecular biological mechanisms involved in the process of tumor occurrence and progression. lncRNA refers to a non-coding RNA transcript with a length greater than 200 nucleotides, which is located in the nucleus and cytoplasm of eukaryotic cells6. A large number of clinical and experimental studies have shown that some lncRNAs play important roles in the emergence and development of malignant tumors7.
miRNA is an endogenous single stranded RNA molecule of 22 nucleotides length that does not encode for a protein. miRNA inhibits the expression of a target gene by complementation binding of its seed region to the microRNA response elements (MREs) on the mRNA8. The regulation network between miRNA and target genes is involved in a variety of biological processes, including tumor occurrence and metastasis9,10. In recent years, research on the role of miRNAs and their implication in the regulation of initiation and development of HNSCC has advanced, and a large number of papers have been published. In 2016, Irani provided a comprehensive literature review of the role of miRNAs in head and neck cancer metastasis11. These studies have laid a good foundation for understanding the mechanism of lncRNA/miRNA/mRNA interactions.
In 2011, Salmena and colleagues presented the competing endogenous RNA (ceRNA) hypothesis which states that the pool of mRNAs, lncRNAs, and other non-coding RNAs share common MREs with miRNAs, which can act as natural miRNA “sponges” and inhibit miRNA function through competitive binding to MREs on the target mRNA12. This theory is based on the fact that lncRNAs can regulate expression of target regulatory genes not only by direct interaction, but also indirectly through competitive binding with miRNAs, as they contain MREs which can bind to the core seed sequence of miRNA. At present, the ceRNA regulation theory has been proven to be implicated in the development of cancer by some related studies. For example, Guo et al. found that lncRNA-BGL3 was a target of miR-17, miR-93, miR-20a, miR-20b, miR-106a and miR-106b, microRNAs that repress mRNA of phosphatase and tensin homolog (PTEN). Further experiments showed that lncRNA-BGL3 operated as a competitive endogenous RNA for binding these microRNAs to cross-regulate PTEN expression13. The lncRNA gene, HLUC, acts as a ceRNA of PRKACB in liver cancer by competing with miR-372, thereby reducing miR-372’s inhibitory effect on CREB and causing up-regulation of CREB14. With the development of bioinformatics technology, more researchers have adopted data analysis and mining methods to study ceRNA networks. In addition, some representative databases, such as miRTarBase15, miRDB16, TargetScan17 and StarBase18 provide data and useful resources for studying ceRNA networks. The Cancer Genome Atlas (TCGA) research team has published a comprehensive resource for multidimensional molecular spectroscopy that stores a large number of tumor samples. These datasets make it feasible to build a ceRNA regulatory network in cancer.
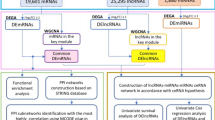
In recent years, numerous studies have confirmed that the lncRNA-miRNA-mRNA ceRNA regulatory network is implicated in the development of gastric, liver, breast, pancreatic and bladder cancer19,20,21,22,23. However, similar studies on HNSCC are limited, and there is still a lack of comprehensive analysis of lncRNAs and miRNAs related to HNSCC based on high-throughput sequencing and large-scale sample size. In this paper, we obtained RNA expression data and compared the expression profiles between 44 normal tissues and 502 tumor tissues of HNSCC. Following, we identified differentially expressed mRNAs, miRNAs and lncRNAs between the samples from HNSCC patients. Finally, 8 miRNAs, 16 mRNAs and 71 lncRNAs were selected to build the lncRNA-miRNA-mRNA ceRNA network. Based on the survival analysis of RNAs in the ceRNA network, we analyzed the lncRNAs that significantly affect the survival and prognosis of HNSCC patients.
Materials and Methods
Patients and TCGA data retrieval
The RNA sequence data of 546 samples with head and neck squamous cell carcinoma were retrieved from the TCGA data portal. The TCGA dataset (https://portal.gdc.cancer.gov/), being comprised of more than two petabytes of genomic data, is publically available, and this genomic information helps the cancer research community to improve the prevention, diagnosis, and treatment of cancer. This study is in accordance with the publication guidelines provided by TCGA (https://cancergenome.nih.gov/publications/publicationguidelines). Since the data comes from the TCGA database, no further approval was required from the Ethics Committee.
RNA sequence data processing
The RNA expression data (level 3) of 546 HNSCC samples were downloaded from the TCGA data portal (up to Aug 29, 2017). The RNA and miRNA sequence data from the 546 samples had been derived from the IlluminaHiSeq_RNASeq and the IlluminaHiSeq_miRNASeq sequencing platforms; all the data were freely available to download. The sequencing data of the 546 samples contained the corresponding RNA-seq and miRNA-seq data. 546 samples were divided into 2 cohorts: 502 tumor samples and 44 normal samples.
In this paper, we mainly used the program code written in Perl and R language to analyze and deal with RNA data.
Identification of differentially expressed mRNAs (DEmRNAs), miRNAs (DEmiRNAs) and long non-coding RNAs (DEIncRNAs)
We identified mRNAs and lncRNAs by using the Ensembl database (http://www.ensembl.org/index.html, version 89)24. In this study, lncRNAs and mRNAs that were not included in the database were excluded.
Before conducting differential expression analysis, we wrote the R language code to ensure that all unexpressed RNAs was filtered out. This was carried out by deleting all rows with a mean read of less than or equal to one. By using the edgeR25 software to further analyze the data, the differentially expressed mRNAs, lncRNAs and miRNAs were obtained. All P values use false discovery rate (FDR) to correct the statistical significance of the multiple test. Fold changes (log2 absolute) ≧2 and FDR adjusted to P < 0.01 were considered significant.
For the obtained differentially expressed mRNAs, lncRNAs, and miRNAs, we generated heat maps and volcano maps using the gplots and heatmap packages in the R platform.
Construction of a ceRNA regulatory network
The ceRNA control network of HNSCC was mainly established by the following steps.
First, the miRcode database26 was used to predict interactions between lncRNA with miRNAs. miRcode provides “whole transcriptome” human microRNA target predictions based on the comprehensive GENCODE gene annotation, including >10,000 long non-coding RNA genes26. miRNA targets can be predicted scientifically by entering the name of the relevant lncRNA and miRNA in the miRcode website platform (http://www.mircode.org/).
Next, we used miRTarBase15, miRDB16 and TargetScan17 databases to retrieve miRNA targeted mRNAs. miRTarBase is a database that has accumulated more than three hundred and sixty thousand miRNA-target interactions (MTIs), which are collected by manually surveying pertinent literature and mining the text systematically to filter research articles related to functional studies of miRNAs15. miRDB is an online database for miRNA target prediction and functional annotations. All the targets in miRDB were predicted by mirTarget, which was developed by analyzing thousands of miRNA-target interactions from high-throughput sequencing experiments16. TargetScan predicts biological targets of miRNAs by searching for the presence of conserved 8mer, 7mer, and 6mer sites that match the seed region of each miRNA17. In order to improve the validity of our search results, we only included miRNA-targeted mRNAs present in all three databases to construct the ceRNA network.
At last, we used Cytoscape 3.5.1 software to visually map the results. Cytoscape (http://www.cytoscape.org/) is an open source software platform for visualizing molecular interaction networks and biological pathways and integrating these networks with annotations, gene expression profiles and other state data.
Survival analysis
We use the R survival package (https://CRAN.R-project.org/package=survival, Version: 2.41-3) for survival analysis for all RNAs in the ceRNA network. The univariate Cox proportional hazards regression27 was carried out to identify the lncRNAs, mRNAs and miRNAs whose expression correlated with overall survival. For the overall survival rates, the log-rank test was used to compare the significant differences in univariate analysis between subgroups. Unless otherwise stated, a P value of less than 0.05 is considered statistically significant.
Data availability
The datasets analysed during the current study are available in the TCGA repository, https://portal.gdc.cancer.gov/.
Results
Differentially expressed mRNAs (DEmRNAs) in HNSCC
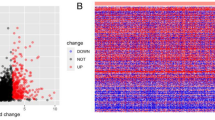
This study investigated the expression levels of RNAs in 502 tumor tissues (cohort T) and 44 normal tissues (cohort N). A differentially expressed gene is defined as a gene whose log2 value of the differential expression multiple value (logFC) is greater than 2 and the corrected P value (FDR) is less than 0.01. According to this standard, 869 (43.51%) up-regulated mRNAs and 1128 (56.49%) down–regulated mRNAs were identified by using the edgeR package. The first 25 up-regulated mRNAs, the first 25 down-regulated mRNAs, and their corresponding logFC, P-value, and FDR values, generated by edgeR, are outlined in Table 1. A complete list of DEmRNAs is provided in appendix 1. In Fig. 1, we show the distribution of all the differentially expressed mRNAs on the two dimensions of -log (FDR) and logFC through a volcano map. All the mRNA expression levels were standardized to the sample mean.
From Table 1 and Fig. 1, we can see that the difference of down-regulated mRNAs is more significant than the up-regulated mRNAs, but the logFC value between them is relatively close.
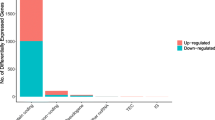
Meanwhile, in order to better understand the mechanisms involved in the development of head and neck squamous cell carcinoma, we used the ClusterProfiler package in the R language for KEGG analysis and mapping, to further analyze the functional characteristics of differentially expressed RNA. The 23 significantly enriched KEGG pathways were listed in Table 2, and the first 12 pathways with the most significant p-values were shown in Fig. 2.
Differentially expressed lncRNAs (DElncRNAs) in HNSCC
We identified 1041 lncRNAs to have statistically significant differential expression (logFC > 2 and FDR < 0.01)in HNSCC samples compared to normal tissues, including 755 up-regulated lncRNAs (72.5%) and 286 down-regulated lncRNAs (27.5%). The first 25 up-regulated lncRNAs and the first 25 down-regulated lncRNAs are outlined in Table 3. The distribution of all the differentially expressed lncRNA on -log (FDR) and logFC are depicted in the volcano map in Fig. 3.
As demonstrated in Table 3 and Fig. 3, most of the differentially expressed lncRNAs are up-regulated (72.5%), but the statistical significance in the down-regulated lncRNAs is more prominent.
Differentially expressed miRNAs (DEmiRNAs) in HNSCC
To construct the lncRNA-miRNA-mRNA ceRNA regulatory network, we also compared miRNA differential expression profiles in tumor tissues and normal tissues. As a result, a total of 82 differentially expressed miRNAs were identified in samples from HNSCC patients, including 44 up-regulated miRNAs (53.6%) and 38 down-regulated miRNAs (46.4%). The first 25 up-regulated miRNAs and the first 25 down-regulated miRNAs are listed in Table, and the volcano map of the related differentially expressed miRNAs is shown in Fig. 4.
Table 4 and Fig. 4 show that, although the number of up-regulated miRNAs is slightly higher than the number of down-regulated miRNAs, the significance of the down-regulated miRNAs remains prominent.
Construction of a ceRNA regulatory network in HNSCC
In order to better understand the role of differentially expressed lncRNAs in HNSCC and to further elucidate the interaction between these differentially expressed lncRNAs and differentially expressed miRNAs, we constructed a lncRNA-miRNA-mRNA related ceRNA regulatory network of HNSCC.
First, we used the 1041 differentially expressed lncRNAs retrieved from the miRcode database26, and applied the Perl program to identify 173 pairs of interacting lncRNAs and miRNAs. From the 82 retrieved DEmiRNAs, we predicted that 8 of them could interact with 71 differentially expressed lncRNAs. Following, we found that these 8 DEmiRNAs targeted 411 mRNAs in all three databases (miRTarBase, miRDB and TargetScan). The 411 targeted mRNAs were cross-checked with the 1997 DEmiRNAs retrieved from the miRcode database. The results showed that 16 mRNAs were involved in the construction of the ceRNA regulatory network (Fig. 5). Finally, we constructed the ceRNA regulatory network of head and neck Squamous cell carcinoma by incorporating 71 DElncRNAs, 16 DEmRNAs and 8 DEmiRNAs, as shown in Fig. 6.
Venn diagram of mRNAs involved in ceRNA regulation network. As shown in the figure, the number of mRNA expressed in the red area is only the difference expression, rather than the target number. The blue area presents only the target number instead of the number of mRNA expressed differently, while the purple area in the middle represents the number of mRNA which is both the differential expression and the target.
DElncRNA mediated ceRNA regulatory network in head and neck squamous cell carcinoma. The nodes highlighted in red indicate expression up-regulation, and the nodes highlighted in blue indicate expression down-regulation. IncRNAs, miRNAs and mRNAs are represented by diamonds, rounded rectangles, and ellipses, respectively.
Correlation analysis of survival and the expression of lncRNAs, miRNAs and mRNAs in the ceRNA network
To identify the potential lncRNAs, miRNAs and mRNAs with prognostic characteristics, the expression levels of 71 lncRNAs, 8 miRNAs and 16 mRNAs in the ceRNA network were profiled using the univariate Cox proportional hazards regression model. As a result, only one miRNA, one mRNA and thirteen lncRNAs were found to be significantly associated with overall survival (P < 0.05). Through the Kaplan-Meier curve analysis, we found that the expression levels of the STC2 mRNA and hsa-mir-206 miRNA were negatively correlated with the overall survival rate (P < 0.05; Fig. 7). In addition, 13 lncRNAs were closely related to survival. 8 differentially expressed IncRNA, including ABCA9-AS1, AL163952.1, AL356056.1, ANO1-AS2, AP002478.1, HOTTIP, LINC00052, and LINC00460, were negatively correlated with overall survival (P < 0.05). On the contrary, 5 differentially expressed IncRNA, including AL161645.1, HCG22, MIAT, MUC19, and ZFY-AS1, were positively correlated with overall survival time (P < 0.05). In Fig. 8, we show the Kaplan-Meier curves of 6 lncRNAs with the most significant P values. In the meantime, in order to better understand the relationship between the expression of these 13 lncRNAs and the patient’s survival time, we generated risk heat maps of these 13 lncRNAs in combination with clinical survival data (Fig. 9).
The constructed ceRNA network showed a possible interaction between DElncRNAs and mRNAs in head and neck squamous cell carcinoma. To confirm this finding, we performed regression analysis between the expression levels of 13 DElncRNAs significantly related to survival and all 16 DEmRNAs included in the network. It was shown that most lncRNA and mRNA in the network do not have direct linear correlation, except HOTTIP and HOXA10 (Fig. 10).
Discussion
In recent years, lncRNA related research has attracted the attention of various cancer fields. Accumulative evidence shows that long-chain non-coding RNAs play a very important regulatory role in tumorigenesis and tumor progression. Although research on lncRNAs for head and neck squamous cell carcinoma is limited, a recent study has reported few differentially expressed lncRNAs such as HOTAIR, NEAT1, UCA1, and MALAT1, in oral cancers28. The HOX transcript antisense intergenic RNA (HOTAIR) is one of the most widely studied lncRNAs29,30. Different studies indicate that HOTAIR plays a significant role in the metastatic process and may be a predictor of poor patient prognosis when is highly expressed31,32. A meta-analysis, performed by Troiano et al., revealed that HOTAIR’s high expression was related to advanced tumor stage, lymph-node metastasis and poor overall survival, which demonstrated the potential prognostic role of HOTAIR in HNSCC30. Tang et al. first demonstrated that HOTAIR, NEAT-1, UCA1 expression levels were up-regulated, while MEG-3 expression levels were down-regulated in oral squamous cell carcinoma (OSCC) and its corresponding adjacent tissues33. They also showed that only HOTAIR could be detected in the patient’s saliva, especially in patients with lymph node metastases33. Li et al. found that HOTAIR expression in laryngeal squamous cell carcinoma (LSCC) was significantly higher than that in para-cancerous tissues, it was correlated with patients’ poor prognosis, and was an independent prognostic factor of LSCC34.
In the ceRNA network constructed in this paper, we found that the high expression of the lncRNA HOTAIR is closely related to 4 differentially expressed miRNAs (hsa-mir-301b is highly expressed, whilst hsa-mir-211, hsa-mir-206 and hsa-mir-375 are low expressed). The highly homologous miRNA to hsa-mir-301b, hsa-miR-301a-3p, has been proven to act as an oncogene by directly regulating the anti-oncogene Smad4, thereby playing a role in the emergence and development of laryngeal squamous cell carcinoma35. Koshizuka et al. showed that miR-1 and miR-206 were down-regulated in HNSCC clinical specimens, which is in agreement with our results. Furthermore, they found that two growth factor receptors, the epidermal growth factor receptor (EGFR) and the hepatocyte growth factor receptor (c-MET), were directly regulated by both miR-206 and miR-1 in HNSCC cells36. Bruce et al. has reported that metadherin was the direct target of miR-375 and their control mechanism may represent a novel oncogenic pathway that drives human head and neck cancer (HNC) progression, possibly through the PI(3) K pathway37. Finally, mir-211 has also been reported to promote head and neck carcinoma progression by targeting TGFβRII38.
Considering all the identified lncRNAs, HOTTIP’s abnormal expression had the most significant impact on the survival of patients (P = 6.241e-04). Meanwhile, the only miRNA and mRNA showing significant differences in their expression levels (P < 0.05) were hsa-mir-206 and STC2, respectively. HOTTIP has been proven to be correlated with the progression and prognosis of tongue squamous cell carcinoma, pancreatic cancer, osteosarcoma, and hepatocellular carcinoma39,40,41,42. miR-206 has been closely linked to the diagnosis, proliferation, and prognosis of multiple cancers, including HNSCC, by many research reports43,44,45,46. Yang et al. revealed that STC2 controls HNSCC metastasis via the PI3K/AKT/Snail signaling pathway and that targeted therapy against STC2 may be a novel strategy to effectively treat patients with metastatic HNSCC47. In 2014, Ren et al. demonstrated that miR-206 was expressed at markedly low levels in a cohort of gastric tumors in comparison with their matching normal tissues, and in high amounts in gastric cancer cell lines48. Furthermore, they found that the miR-206’s anti-metastatic effect was probably mediated through targeting the metastasis regulatory gene STC2 and other mRNAs, which were drastically down-regulated by the exogenous miR-206’s stable expression in SCG-7901 cells48. However, in our study, STC2 was obviously down-regulated by the low expression of mir-206 in HNSCC patients.
In addition to HOTTIP, we also found several lncRNAs in the ceRNA network that were closely linked to the survival of HNSCC patients. 13 lncRNAs had a significant impact on survival, and the expression of some lncRNAs showed a more obvious change trend as the risk increased. From the survival analysis, only LINC00460 had been previously described as a therapeutic target and a novel prognostic biomarker for the diagnosis and treatment of nasopharyngeal carcinoma49. The remaining five lncRNAs have been identified for the first time to be closely related to the prognosis of HNSCC, which can serve as potential targets for future clinical treatments.
Conclusion
To conclude, our study has identified differentially expressed mRNAs, lncRNAs, and miRNAs in HNSCC patients. Importantly, a ceRNA network has been constructed to propose a novel regulatory mechanism for the development of HNSCC. The lncRNAs identified in our constructed ceRNA network may have an important impact on the survival and prognosis of HNSCC patients.
References
Jemal, A., Siegel, R., Xu, J. Q. & Ward, E. Cancer Statistics. Cancer 59, 1–24 (2010).
Torre, L. A. et al. Global cancer statistics, 2012. Ca A Cancer Journal for Clinicians 65, 87–108 (2015).
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2015. Ca A Cancer Journal for Clinicians 65, 5 (2014).
Nohata, N. et al. Tumor suppressive microRNA-375 regulates oncogene AEG-1/MTDH in head and neck squamous cell carcinoma (HNSCC). Journal of Human Genetics 56, 595 (2011).
Jamali, Z. et al. MicroRNAs as prognostic molecular signatures in human head and neck squamous cell carcinoma: A systematic review and meta-analysis. Oral Oncology 51, 321–331 (2015).
Ponting, C. P., Oliver, P. L. & Reik, W. Evolution and functions of long noncoding RNAs. Cell 136, 629–641 (2009).
Yan, X. et al. Comprehensive Genomic Characterization of Long Non-coding RNAs across Human Cancers. Cancer Cell 28, 529–540 (2015).
Ye, W. et al. The effect of central loops in miRNA:MRE duplexes on the efficiency of miRNA-mediated gene regulation. Plos One 3, e1719 (2008).
Amirkhah, R., Schmitz, U., Linnebacher, M., Wolkenhauer, O. & Farazmand, A. MicroRNA–mRNA interactions in colorectal cancer and their role in tumor progression. Genes Chromosomes & Cancer 54, 129–141 (2015).
Wang, C., Hu, D. Z. & Liu, J. Z. Identification of critical TF-miRNA-mRNA regulation loops for colorectal cancer metastasis. Genetics & Molecular Research Gmr 14, 5485–5495 (2015).
Irani, S. miRNAs Signature in Head and Neck Squamous Cell Carcinoma Metastasis: A Literature Review. Journal of Dentistry 17, 71–83 (2016).
Salmena, L., Poliseno, L., Tay, Y., Kats, L. & Pandolfi, P. P. A ceRNA hypothesis: the Rosetta stone of a hidden RNA language? Cell 146, 353–358 (2011).
Guo, G. et al. A long noncoding RNA critically regulates Bcr-Abl-mediated cellular transformation by acting as a competitive endogenous RNA. Oncogene 34, 1768–1779 (2015).
Wang, J. et al. CREB up-regulates long non-coding RNA, HULC expression through interaction with microRNA-372 in liver cancer. Nucleic Acids Research 38, 5366–5383 (2010).
Chou, C. H. et al. miRTarBase 2016: updates to the experimentally validated miRNA-target interactions database. Nucleic Acids Research 44, D239 (2016).
Wong, N. & Wang, X. miRDB: an online resource for microRNA target prediction and functional annotations. Nucleic Acids Research 43, D146 (2015).
Fromm, B. et al. A Uniform System for the Annotation of Vertebrate microRNA Genes and the Evolution of the Human microRNAome. Annual Review of Genetics 49, 213–242 (2015).
Li, J. H., Liu, S., Zhou, H., Qu, L. H. & Yang, J. H. star Basev2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Research 42, 92–97 (2014).
Xia, T. et al. Long noncoding RNA associated-competing endogenous RNAs in gastric cancer. Scientific Reports 4, 6088 (2014).
Zhang, J., Fan, D., Jian, Z., Chen, G. G. & Lai, P. B. S. Cancer Specific Long Noncoding RNAs Show Differential Expression Patterns and Competing Endogenous RNA Potential in Hepatocellular Carcinoma. Plos One 10, e0141042 (2015).
Zhou, X., Liu, J. & Wang, W. Construction and investigation of breast-cancer-specific ceRNA network based on the mRNA and miRNA expression data. Iet Systems Biology 8, 96–103 (2014).
Zhou, M. et al. Construction and analysis of dysregulated lncRNA-associated ceRNA network identified novel lncRNA biomarkers for early diagnosis of human pancreatic cancer. Oncotarget 7, 56383 (2016).
Wang, H. et al. Comprehensive analysis of aberrantly expressed profiles of lncRNAs and miRNAs with associated ceRNA network in muscle-invasive bladder cancer. Oncotarget 7 (2016).
Aken, B. L. et al. The Ensembl gene annotation system. Database the Journal of Biological Databases & Curation 2016, baw093 (2016).
Robinson, M. D., Mccarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139 (2010).
Ashwini, J., Marks, D. S. & Erik, L. miRcode: a map of putative microRNA target sites in the long non-coding transcriptome. Bioinformatics 28, 2062 (2012).
Kirch, W. Cox Proportional Hazards Regression. (Springer Netherlands, 2008).
Gomes, C. C., de Sousa, S. F., Calin, G. A. & Gomez, R. S. The emerging role of long noncoding RNAs in oral cancer. Oral Surg Oral Med Oral Pathol Oral Radiol, 235–241 (2017).
Loewen, G., Jayawickramarajah, J., Zhuo, Y. & Shan, B. Functions of lncRNA HOTAIR in lung cancer. Journal of Hematology & Oncology 7, 90 (2014).
Troiano, G. et al. Expression of the long non-coding RNA HOTAIR as a prognostic factor in squamous cell carcinoma of the head and neck: a systematic review and meta-analysis. Oncotarget 8, 73029–73036 (2017).
Xing, B. et al. Knockdown of long non-coding RNA HOTAIR increases miR-454-3p by targeting Stat3 and Atg12 to inhibit chondrosarcoma growth. Cell Death & Disease 8, e2605 (2017).
Liu, X. et al. The long non-coding RNA HOTAIR indicates a poor prognosis and promotes metastasis in non-small cell lung cancer. Bmc Cancer 13, 464–464 (2013).
Tang, H., Wu, Z., Zhang, J. & Su, B. Salivary lncRNA as a potential marker for oral squamous cell carcinoma diagnosis. Molecular Medicine Reports 7, 761–766 (2013).
Li, D. et al. Long intergenic noncoding RNA HOTAIR is overexpressed and regulates PTEN methylation in laryngeal squamous cell carcinoma. American Journal of Pathology 182, 64–70 (2013).
Lu, Y. et al. Hsa-miR-301a-3p Acts as an Oncogene in Laryngeal Squamous Cell Carcinoma via Target Regulation of Smad4. Journal of Cancer 6, 1260–1275 (2015).
Koshizuka, K. et al. Dual-receptor (EGFR and c-MET) inhibition by tumor-suppressive miR-1 and miR-206 in head and neck squamous cell carcinoma. Journal of Human Genetics 62, 113 (2017).
Bruce, J. P. et al. Abstract 3967: Identification of metadherin as a novel target of miR-375 in head and neck cancer. Cancer Research 71, 3967–3967 (2011).
Chu, T. H. et al. miR-211 promotes the progression of head and neck carcinomas by targeting TGFβRII. Cancer Letters 337, 115–124 (2013).
Zhang, H. et al. Long non-coding RNA HOTTIP is correlated with progression and prognosis in tongue squamous cell carcinoma. Tumour Biology the Journal of the International Society for Oncodevelopmental Biology & Medicine 36, 8805 (2015).
Cheng, Y., Jutooru, I., Chadalapaka, G., Corton, J. C. & Safe, S. The long non-coding RNA HOTTIP enhances pancreatic cancer cell proliferation, survival and migration. Oncotarget 6, 10840–10852 (2015).
Fan, L., Lei, C., Donghua, H., Fang, W. & Qiugen, W. Long non-coding RNA HOTTIP is up-regulated and associated with poor prognosis in patients with osteosarcoma. International Journal of Clinical & Experimental Pathology 8, 11414 (2015).
Quagliata, L. et al. lncRNA HOTTIP/HOXA13 expression is associated with disease progression and predicts outcome in hepatocellular carcinoma patients. Hepatology 59, 911 (2014).
Kondo, N., Toyama, T., Sugiura, H., Fujii, Y. & Yamashita, H. miR-206 Expression is down-regulated in estrogen receptor alpha-positive human breast cancer. Cancer Research 68, 5004 (2008).
Zhang, T. et al. Down-regulation of MiR-206 promotes proliferation and invasion of laryngeal cancer by regulating VEGF expression. Anticancer Research 31, 3859–3863 (2011).
Zhang, L. et al. miR-206 inhibits gastric cancer proliferation in part by repressing cyclinD2. Cancer Letters 332, 94 (2013).
Wang, S. H. et al. Long non-coding RNA Malat1 promotes gallbladder cancer development by acting as a molecular sponge to regulate miR-206. Oncotarget 7, 37857 (2016).
Yang, S. et al. STC2 promotes head and neck squamous cell carcinoma metastasis through modulating the PI3K/AKT/Snail signaling. Oncotarget 8, 5976 (2017).
Ren, J. et al. MicroRNA-206 suppresses gastric cancer cell growth and metastasis. Cell & Bioscience 4, 26 (2014).
Kong, Y. G. et al. LncRNA-LINC00460 facilitates nasopharyngeal carcinoma tumorigenesis through sponging miR-149-5p to up-regulate IL6. Gene 639, 77 (2018).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (61572298, 61772322, 61602283), the Shandong Provincial Natural Science Foundation of China (ZR2011HZ004, ZR2017BC073, ZR2016FB10) and the Project of Shandong Province Higher Educational Science and Technology Program (J10LC21 and J16LE07).
Author information
Authors and Affiliations
Contributions
X.F. and M.Y. wrote the main manuscript text, H.L. and C.L. prepared all figures. C.X., G.Y. and H.Z. designed the experiments, and all authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fang, XN., Yin, M., Li, H. et al. Comprehensive analysis of competitive endogenous RNAs network associated with head and neck squamous cell carcinoma. Sci Rep 8, 10544 (2018). https://doi.org/10.1038/s41598-018-28957-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-28957-y
- Springer Nature Limited
This article is cited by
-
Prediction of Potential Biomarkers in Early-Stage Nasopharyngeal Carcinoma Based on Platelet RNA Sequencing
Molecular Biotechnology (2023)
-
Comprehensive analysis of the LncRNAs, MiRNAs, and MRNAs acting within the competing endogenous RNA network of LGG
Genetica (2022)
-
Development and analysis of long non-coding RNA-associated competing endogenous RNA network for osteosarcoma metastasis
Hereditas (2021)
-
A ceRNA-associated risk model predicts the poor prognosis for head and neck squamous cell carcinoma patients
Scientific Reports (2021)
-
Transcriptomics and Epigenomics in head and neck cancer: available repositories and molecular signatures
Cancers of the Head & Neck (2020)