Abstract
Sustainable and eco-efficient alternatives for the production of platform chemicals, fuels and chemical building blocks require the development of stable, reusable and recyclable biocatalysts. Here we present a novel concept for the biocatalytic production of 1,5-diaminopentane (DAP, trivial name: cadaverine) using catalytically active inclusion bodies (CatIBs) of the constitutive L-lysine decarboxylase from E. coli (EcLDCc-CatIBs) to process L-lysine-containing culture supernatants from Corynebacterium glutamicum. EcLDCc-CatIBs can easily be produced in E. coli followed by a simple purification protocol yielding up to 43% dry CatIBs per dry cell weight. The stability and recyclability of EcLDCc-CatIBs was demonstrated in (repetitive) batch experiments starting from L-lysine concentrations of 0.1 M and 1 M. EcLDC-CatIBs exhibited great stability under reaction conditions with an estimated half-life of about 54 h. High conversions to DAP of 87–100% were obtained in 30–60 ml batch reactions using approx. 180–300 mg EcLDCc-CatIBs, respectively. This resulted in DAP titres of up to 88.4 g l−1 and space-time yields of up to 660 gDAP l−1 d−1 per gram dry EcLDCc-CatIBs. The new process for DAP production can therefore compete with the currently best fermentative process as described in the literature.
Similar content being viewed by others
Introduction
An interpolation from the current state of the petrochemical industry and fossil-based energy supply to the next century predicts the exhaustion of fossil carbon sources, which can be attributed to an alarmingly rapid exploitation of limited natural deposits1. In particular, this applies to crude oil, due to the steadily growing demand2,3. Consequently, society will face a notable future price increase for fossil resources, which has already focused public interest on sustainable and eco-efficient alternatives. This has thus encouraged the biotechnology industry to develop processes for the sustainable production of platform chemicals, biofuels4,5,6, and in particular bio-based polymers4,7. Increasing knowledge about the prokaryotic metabolism and ongoing developments in systems engineering pave the way for the development of microbial hosts also enabling the economic production of intermediates and bulk chemicals. However, to meet economic demand, it is necessary to develop innovative concepts and improved bioprocesses.
Biotechnological workhorses such as Escherichia coli, Saccharomyces cerevisiae and Corynebacterium glutamicum have been engineered to produce ω-amino acids8, aromatic monomers9, diamines10,11,12, dicarboxylic acids13,14,15, diols16 and hydroxy acids17, respectively. From this broad spectrum of building blocks for biopolymer production, the linear aliphatic diamine 1,5-diaminopentane (DAP) is probably one of the most attractive options. One reason is its ability to produce fully bio-based polyamides, such as PA 5.4 and PA 5.10, based on DAP and dicarboxylic acids such as succinate13,18 and sebacic acid19, respectively. PA 5.10, in particular, exhibits material properties comparable or even superior to the widely used petroleum-based polyamide PA 620.
One option for the biotechnological production of DAP is the use of engineered, well-established L-lysine producers, especially C. glutamicum10,21,22,23,24 and E. coli12,25,26,27,28,29,30. C. glutamicum DAP-producer strains are usually created by the introduction of one of the L-lysine decarboxylase (LDC) genes from E. coli (cadA21 or ldcC10 encoding the acid-inducible enzyme CadA, and the constitutive LDCc, respectively), thus enabling the intracellular decarboxylation of the L-lysine (1) to DAP (2) (Fig. 1). Both enzymes are very similar (sequence similarity 84%)31,32, require the cofactor pyridoxal-5′-phosphate (PLP), and appear as decamers composed of five dimers, as was deduced from cryo-electron microscopy33 and X-ray crystallography34.
Several constraints need to be tackled for fermentative microbial production, such as the tolerance of the microbial system with respect to DAP35, the avoidance of by-products such as N-acetyl-1,5-diaminopentane36 and the management of product export37, which is no longer possible via the well-engineered lysine exporter LysE38.
Another option for DAP production is the bioconversion of L-lysine by the addition of LDC to L-lysine containing culture supernatants. Here, immobilisation of the LDC enables easy separation from the reaction medium and recycling of the biocatalyst to decrease process costs. Different concepts have been employed for the immobilisation of LDC, e.g. using whole recombinant E. coli cells25,27,29,39, immobilised recombinant E. coli cells in alginate beads40,41, as well as immobilised LDC on poly(3-hydroxybutyrate) (P(3HB) biopolymer42 or crosslinked enzyme aggregates (CLEAS) of LDC43. Generally, the bioconversions were performed in buffer and only in a few cases directly in culture supernatants of L-lysine producers25,29.
Catalytically active inclusion bodies (CatIBs) represent biologically produced, cell-free and carrier-free immobilisates that can easily be produced in E. coli cells44,45,46,47. They are a simple and cheap alternative to common immobilisates, which require a case-to-case optimisation of several, often expensive and laborious, steps, including chromatographic purification of the enzyme followed by covalent or non-covalent immobilisation concepts in the presence or absence of carriers48,49. The production of immobilised enzymes directly in vivo could reduce the production costs of the biocatalyst to the level of crude cell extracts50, since the insoluble cell fraction can be directly used for biotransformations. Furthermore, CatIB-based biotransformations are free of genetically modified organisms (GMO-free), since any remaining vital recombinant E. coli cells can be efficiently inactivated and separated during the production process51.
Active inclusion bodies can be formed either naturally by self-aggregation of the enzyme46,52 or by fusion to a tag containing an aggregation-prone part, e.g. cellulose binding domains53,54,55,56, pyruvate oxidase (PoxB) of Paenibacillus polymyxa57, the viral capsid protein VP1, the human Aβ-amyloid peptide58 or various self-assembling peptides59,60. For a detailed overview of the state of the art in this field we refer to a recent review47. We previously evaluated the tetramerisation domain of the cell-surface protein tetrabrachion (known as TDoT) from Staphylothermus marinus61 for its potential to induce CatIB formation. The TDoT domain has a rope-like structure forming a stable parallel tetrameric coiled coil61. Previous studies have demonstrated that the fusion of the TDoT domain to various enzymes of different complexity resulted in all cases in the formation of CatIBs, which implies that the TDoT-domain is a promising new fusion tag to induce the formation of active inclusion bodies62.
We here report on an innovative immobilisation approach using CatIBs of the constitutive L-lysine decarboxylase (EcLDCc) and the application of this GMO-free approach to produce DAP in L-lysine-containing culture supernatants. In this study, we demonstrate that this approach also works for the complex PLP-dependent decameric EcLDCc. This enzyme was chosen instead of the frequently used CadA mainly because of the broader pH optimum of EcLDCc (pH 6.2–8) compared to CadA (pH 5.7)63. This property makes EcLDCc advantageous for application in L-lysine-containing culture supernatants of the respective C. glutamicum producer strains, which exhibit pH values in the range of 6 to 8.564,65.
The respective EcLDCc-CatIBs were produced in E. coli and successfully applied in culture supernatants of a C. glutamicum lysine producer with lysine concentrations of up to 1 M to demonstrate the applicability of this approach on the preparative scale. Under optimised conditions, 74.7–88.4 g l−1 DAP was produced with 10 g l−1 dry EcLDCc-CatIBs with a space-time yield of 296–660 gDAP l−1 d−1 per gram dry EcLDC-CatIBs (see Table 1).
Results and Discussion
Production of EcLDCc-CatIBs
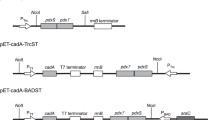
The gene encoding the constitutive EcLDCc was introduced into a pET28 vector already containing the gene sequences encoding the coiled-coil domain TDoT and an additional 3xGGGS linker region as described elsewhere62. Based on the quaternary structure of EcLDCc, the TDoT domain was fused to the C-terminus, since the N-terminus is located within the protein structure33.
EcLDCc-CatIBs could be easily produced in E. coli BL21(DE3) using an auto-induction medium (see Sect. 5.3). The formation of EcLDCc-CatIBs in the respective recombinant E. coli cells was demonstrated with inverted epifluorescence microscopy66,67 showing the CatIBs as bright spots at the cell poles (Fig. 2), which is typical of the deposition of recombinant proteins as inclusion bodies in E. coli68.
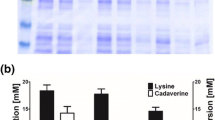
A previously developed protocol62 was further optimised for the purification of EcLDCc-CatIBs (see Fig. 3, left). After two washing steps with water followed by centrifugation, the pellet containing the CatIBs and some cell membrane fractions was lyophilised, yielding 130 ± 37 mg dry CatIBs per gram of wet cells (approx. 13% of the wet cell weight, corresponding to 43% dry CatIBs based on dry cell weight). The protein content of the pellet was about 68%, which is comparable to previous results obtained with other CatIB enzymes62. The production process was monitored by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE, Fig. 3, right) showing that the EcLDCc-TDoT fusion is predominantly present in the pellet. Due to the simple purification protocol (see Methods), further cellular proteins co-purified with the CatIBs were expected, as was also reported for other inclusion body formulations69.
Left: Production and purification of EcLDCc-CatIBs produced in E. coli BL21(DE3). Right: SDS-PAGE analysis of the EcLDCc-CatIB preparation (calculated molecular weight: 87.8 kDa, arrow); CCE = crude cell extract, which was centrifuged to separate the supernatant (S1) from the pellet (P1). The pellet P1 was washed once with MilliQ water by resuspension and subsequent centrifugation, resulting in S2 and P2; the protein concentration was measured using the Bradford assay (see Methods). For SDS-PAGE, samples were diluted with water to a protein concentration of 1 mg ml−1 by the following dilution factors: 4 for CCE, 2 for S1 and P1, 4.5 for P2; 1 for S2; M = Marker. For details see Methods.
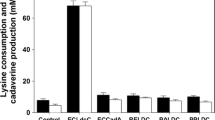
The activity of the EcLDCc-CatIBs was demonstrated in potassium phosphate buffer (KPi buffer) and cultivation medium (CGXII)64 (Supplementary Fig. S2). Additionally, the CatIBs were compared to an E. coli whole cell biocatalyst containing the overproduced soluble LDCc. The results demonstrate that EcLDCc-CatIBs can compete with the whole cell biocatalyst (for details see Supplementary Fig. S7a,b). Subsequently, the EcLDCc-CatIBs were characterised in CGXII medium and used for a case study under technical conditions in culture supernatants of a C. glutamicum L-lysine producer strain.
Characterisation of EcLDCc-CatIBs
Activity in phosphate buffer
In a first step, the EcLDCc-CatIBs were characterised in KPi buffer to determine the pH optimum in the pH range of 7–9 and the minimal requirement for PLP.
As already mentioned in the Introduction, the soluble wild-type EcLDCc is active in a relatively broad pH range exhibiting maximal activity between pH 6.2 and pH 8, whereas at pH 8.8 the activity was shown to decrease to 30%63. As demonstrated in Fig. 4, EcLDCc-CatIBs showed considerable activity between pH 7.5–9.0 with a clear activity maximum at pH 8. Furthermore, addition of the cofactor PLP was decisive in achieving optimal enzyme activity. Generally, the activity increased by 5–15% in the presence of the cofactor (Fig. 4). Strikingly, at pH 9 the positive PLP effect was approximately 35%. In additional studies, a PLP concentration of 0.05 mM was found to be sufficient for maximal activity of the EcLDCc-CatIBs in buffer (see Supplementary Fig. S3). A similar positive effect of PLP on the LDC activity was recently reported for the second isoenzyme in E. coli, the acid-inducible CadA overproduced in recombinant E. coli, which was used as a whole-cell biocatalyst. In this case, full conversion of 1 M L-lysine to DAP was observed in the presence of 0.025 mM PLP, whereas without additional PLP only 20% conversion was achieved29.
Activity in CGXII minimal medium
To verify the applicability of EcLDCc-CatIBs at the preparative scale, DAP production was tested in CGXII cultivation medium providing an experimental setup close to requirements on the technical scale.
First, the optimal pH was determined between pH 7.0–9.0 in fresh CGXII medium with 0.1 mM PLP and 10 mM L-lysine. EcLDCc-CatIBs revealed the highest conversion between pH 8–9, showing a maximum at 8.5 (see Supplementary Fig. S4), which closely corresponds to the pH optimum in KPi buffer (Fig. 4). To ensure comparability, all subsequent experiments were performed at pH 8 providing excellent conditions for EcLDCc-CatIBs in CGXII medium as well as KPi buffer.
In technical processes, L-lysine concentrations of up to 120 g l−1 (820 mM) are expected22. Therefore, EcLDCc-CatIBs were tested in (repetitive) batch reactions with substrate concentrations of up to 1 M L-lysine. In initial studies with 10–100 mM L-lysine, a concentration of 2 mg ml−1 EcLDCc-CatIBs was shown to be sufficient to completely convert 100 mM L-lysine to DAP in 4 h (Fig. 5). Notably, the estimated activity increased from approx. 0.3 U mg−1 (10 mM L-lysine) to approx. 0.8 U mg−1 (100 mM L-lysine), giving rise to the conclusion that the maximum velocity (Vmax) of the CatIBs requires a L-lysine concentration of 100 mM or higher. Under the applied conditions, EcLDCc-CatIBs exhibited half-maximum activity at approx. 23 mM L-lysine. This value is much higher compared to the KM value of 0.84 mM published for the soluble enzyme70, which was determined under different reaction conditions (soluble EcLDCc in 0.5 M sodium acetate buffer, pH 5.5), which makes a comparison of KM values meaningless. However, the enormously increased KM for the CatIBs could indicate a form of mass transport limitation of the substrate or product in the environment of the precipitated protein structure of the CatIBs. The highest enzymatic productivity of 4.9 gDAP gLDC-CatIBs−1 (48 mmolDAP gLDC-CatIBs−1) was achieved with 100 mM L-lysine in these first studies.
Application of EcLDC-CatIBs for the production of DAP
Subsequently, EcLDC-CatIBs were characterised in CGXII medium containing L-lysine produced by a C. glutamicum DM1945 strain71.
In the first trial, EcLDCc-CatIBs were directly added to the cultivation medium to enable the simultaneous production of L-lysine followed by decarboxylation to DAP in one pot. Surprisingly, only low yields of DAP were obtained, although PLP was added to the cultivation medium. This result could be due to the degradation of PLP by photolysis or oxidation72 or consumption of the cofactor by C. glutamicum, which was earlier reported by Kind et al., who studied the positive effect of adding PLP to the cultivation broth of a C. glutamicum DAP producer strain10. A further reason could be the low apparent affinity of EcLDCc-CatIBs to L-lysine (KM approx. 23 mM) as discussed above, which results in low conversion rates at substrate concentrations <100 mM.
In order to circumvent this issue, the lysine-producing cultivation of C. glutamicum DM1945 was first completed in CGXII medium. The cell-free culture supernatant was further supplemented with L-lysine to 0.1 M and 0.1 mM PLP.
The initially performed determination of the stability of EcLDC-CatIBs in this reaction system, as well as their continued application in repetitive batch mode, shows that EcLDCc-CatIBs are fully stable for at least 24 h and can be recycled several times (for details see Supplementary Fig. S5). In repetitive batch mode, the CatIBs were reused after centrifugation and resuspension for five batch cycles, resulting in a productivity of 19.4 gDAP gLDC-CatIBs−1 (190 mmolDAP gLDC-CatIBs−1), which is 4 times higher compared to a single batch (Fig. 5).
Based on these promising results, a repetitive batch on the 60 mL scale with 0.1 M L-lysine was set up in a pH-controlled environment, which was necessary since the reaction products CO2 and DAP shift the pH. The nine-batch cycles took either 4 or 15 hours. The results of the repetitive batch experiment revealed a constant high conversion of 84–98% during the first 46 h (Fig. 6). The first two repetitive batches (each lasting 4 h) showed almost full conversions of 90–98%. Also the 3rd batch reaction (performed for 15 h) yielded full conversion. After 54 h reaction time (batch 8) the half-life of the EcLDC-CatIBs has almost been achieved, since the conversion decreased to 54%. After 69 h reaction time, the 9th batch (lasting 15 h) only reached 76% conversion, demonstrating that 15 h reaction time was not sufficient to compensate the progressive inactivation. By means of the repetitive batch approach, the enzymatic productivity was increased to 25 gDAP/gLDC-CatIBs (Table 1), which is 7.5 times higher compared to a single 60 ml batch reaction (e.g. 2nd batch: 3.33 gDAP gLDC-CatIBs−1), yielding a final DAP concentration of 8.47 g l−1, a specific space-time yield (STY) of 16 gDAP l−1 d−1 per gram EcLDCc-CatIBs, and a total turnover number (ttn) for PLP of up to 994. This experiment showed that the EcLDCc-CatIBs could be reused for several cycles for at least 69 hours under the applied conditions.
Repetitive batches for the production of DAP with EcLDCc-CatIBs with pH-control. Experimental conditions: 3 mg ml−1 lyophilised EcLDCc-CatIBs, 0.1 M L-lysine, 0.1 mM PLP in 60 ml cell-free culture supernatant (CGXII medium, pH 8). Two 4 h batches (batch 1, 2, 4, 5, 7, 8) were followed by 15 h overnight batches (3, 6, 9) on three subsequent days.
To apply the EcLDCc-CatIBs under the requirements on a technical scale, where L-lysine concentrations of up to 1 M are converted to DAP22,29, the application was next tested in a batch reaction (30 ml) with 1 M L-lysine, which resulted in 87% conversion after approx. 24 h (Fig. 7). The specific activity of 0.75 U mg−1, deduced from conversions ≤10%, was comparable to the reaction velocity observed with 100 mM L-lysine (see above), which indicates that there is no substrate inhibition for EcLDCc-CatIBs up to 1 M L-lysine. Although the enzymatic productivity was reduced to 30% (8.8 gDAP g−1CatIBs) compared to the previous repetitive batch experiments with 100 mM L-lysine, due to the higher concentration of CatIBs, the STY was increased 180-fold to 296 gDAP l−1 d−1 per gram EcLDC-CatIBs. As demonstrated in Fig. 7, the reaction slowed down after a process time of 9 h and between 9 h and 24 h conversion only increased by about 10%. The analysis of the reasons for the slowdown of the reaction requires further investigation. One possibility could be inactivation of the enzyme by negative effects caused by the pH-adjustment with NaOH and HCl or due to high concentrations of DAP, which could be targeted by reaction engineering. If the high DAP concentration is the reason for deactivation of the enzyme, stopping the reaction after 9 h would be a good option (Fig. 7). This would result in a more than twofold higher specific space-time yield of 660 gDAP l−1 d−1 per gram EcLDC-CatIBs. Accordingly, the ttn for the cofactor PLP was increased by a factor of 10 relative to the repetitive batch with 0.1 M L-lysine (see Table 1). Consequently, a 10 times higher product concentration was reached in one third of the time (24 h) needed for the repetitive batch mode using 0.1 M L-lysine.
Conversion curve for the production of DAP with EcLDCc-CatIBs in a 30 ml batch reactor with pH control. Experimental conditions: 10 mg ml−1 lyophilised EcLDCc-CatIBs, 1 M L-lysine, 0.1 mM PLP, in 30 ml cell-free culture supernatant (CGXII medium, pH 8). For the dosage profile with NaOH and HCl, respectively, to keep the pH constant see Supplementary Fig. S6.
Conclusions
The development of cheap, stable, reusable and recyclable biocatalysts is necessary for the prospective creation of competitive sustainable and eco-efficient production processes for platform chemicals, fuels, and polymer building blocks. One promising approach to fulfil industrial demands in terms of productivity, yield, and product titre is the application of whole-cell biotransformation50 e.g. using resting or metabolically active microbial cells. However, the drawbacks of this approach are productivity issues, e.g. due to undesired side reactions, negative interactions of substrates and products with the microorganism as well as difficulties in downstream processing caused by lysed cells under process conditions. Furthermore, the application of genetically modified organisms (GMO) requires conformity with the respective safety standards. In contrast to whole cells, the preparation of catalytically active inclusion bodies (CatIBs) requires only a few additional steps (cell disruption, solid/liquid separation, washing with water) and thus represents a versatile and cheap GMO-free immobilisation method.
Here we demonstrate the application of CatIBs for the production of 1,5-diaminopentane (DAP) using the constitutive decameric and PLP-dependent lysine decarboxylase from E. coli (EcLDCc). Currently, this represents the structurally most complex enzyme in our CatIB toolbox62.
EcLDCc-CatIBs can be produced with high yields (about 13% dry CatIBs based on the wet cell weight, equivalent to 43% dry CatIBs based on dry cell weight) at low cost comparable to crude cell extract50. A two-step process was applied whereby L-lysine is produced first through a C. glutamicum producer strain and the culture supernatant is subsequently treated with EcLDCc-CatIBs to produce DAP. Maximal conversion rates were obtained with L-lysine concentrations of 0.1–1 M. As was found for soluble EcLDCc73 and whole cell catalysts27,29, the addition of PLP was decisive for optimal CatIB activity. After optimisation of the reaction conditions, a study on a preparative scale demonstrated that EcLDCc-CatIBs are recyclable and stable biocatalysts for DAP production directly applicable in L-lysine-containing culture supernatant. The EcLDCc-CatIBs were successfully reused by simple centrifugation and resuspension steps. Starting from 1 M L-lysine, a maximal DAP concentration of 74.7–88.4 g l−1 and a specific STY of up to 296–660 gDAP l−1 d−1 per gram EcLDC-CatIBs were obtained (see Table 1). This result compares well with the currently best fermentative process using C. glutamicum, which also achieved a final titre of 88 g l−1 DAP after 50 hours of a combined batch/fed-batch fermentation, but a STY of only 52.8 g l−1 d−1 (2.2 g l−1 h−1)23 (see Table 1). In order to fulfil the technically relevant demands, the usage of EcLDCc CatIBs in batch mode at high substrate concentrations proved to be appropriate in order to obtain high STY.
Methods
Materials
All chemicals were purchased from Sigma-Aldrich, Roth, KMF and Merck. Enzymes for molecular biology were purchased from Thermo Scientific.
Cloning
See Supplementary “Cloning & sequences.”
Protein production, cell disruption and protein purification
EcLDCc-CatIBs were produced in E. coli BL21(DE3) as recently described elsewhere62. Here, a temperature of 15 °C during protein production was decisive for the formation of active EcLDC-CatIBs. Cell disruption was performed with a high-pressure homogeniser (EmulsiFlex-C5, Avestin Europe GmbH, Mannheim, Germany) at 1000 bar using a cooled 10% (w/v) suspension of E. coli cells in cell lysis buffer (50 mM sodium phosphate, 100 mM NaCl, pH 8). To ensure thorough cell disruption, the suspension was passed three times through the high-pressure homogeniser under constant cooling. SDS-PAGE (see below) was used to analyse the distribution of the recombinant protein in the E. coli cells and during CatIB isolation. After cell disruption, the crude cell extract, and the soluble and insoluble protein fraction were separated by centrifugation at 15,000 × g for 30 min. The pellet was washed by suspension in MilliQ water in the initial volume followed by centrifugation. The obtained pellet was frozen overnight at −20 °C and a 10% (w/v) suspension in MilliQ water was prepared for lyophilisation (Christ ALPHA 1–3 LD Plus, Martin Christ Gefriertrocknungsanlagen GmbH, Osterode, Germany). In a mortar the dried pellet was ground to a fine powder, which was weighed and stored at −20 °C for further use.
SDS-PAGE and protein assay
SDS-PAGE analysis was performed using the NuPAGE® Kit (ThermoFisher Scientific), consisting of LDS Sample Buffer (4×) and NuPAGE® Reducing Agent (10×) with a final protein content of 1 mg ml−1. Previously, the soluble protein concentration had been measured using the Bradford assay74 and bovine serum albumin as a standard. Samples were applied to a NuPAGE™ 4–12% Bis-Tris protein gel, 1.0 mm, with 15 wells together with a protein marker (PageRuler Plus Prestained Protein ladder, ThermoFisher Scientific). Gel electrophoresis was performed in NuPAGE® MES SDS running buffer (1×) at 200 V, 100 mA and 15 W.
The protein content of the lyophilised CatIBs was determined by absorption at 280 nm. For this purpose, a defined amount (1–2 mg) of freeze-dried CatIBs was dissolved in 6 M aqueous guanidine hydrochloride solution (1 ml) and incubated at 30 °C for 30 min under constant shaking at 1000 rpm in a thermomixer (Thermomixer comfort, Eppendorf, Germany). The absorption of the protein solution was measured at 280 nm with a spectrophotometer (Shimadzu UV-1800/UV-1600). The protein content was estimated using the molar extinction coefficient (EcLDCc-CatIBs: ε = 109,210 l mol−1 cm−1) as calculated based on the amino acid composition using the ProtParam Tool (http://web.expasy.org/protparam).
Live cell imaging
See Supplementary.
pH optimum of EcLDC-CatIBs in KPi buffer and activity assay
In order to analyse the pH optimum, a reaction with 0.5 mg ml−1 lyophilised EcLDCc-CatIBs in a reaction tube (2 ml safe-lock tube, Eppendorf, Germany) in 1 ml KPi buffer (50 mM, pH 7.0, 7.5, 8.0, 8.5, 9.0) containing 10 mM L-lysine, 0.1 mM PLP was performed for 20 min at 30 °C and 1000 rpm in a thermomixer (Thermomixer comfort, Eppendorf, Germany). After different time intervals (5, 10 and 20 min), 20 µl of the samples was taken from one vial. The reaction was stopped by incubation at 90 °C for 2 min and subsequent centrifugation for 2 min at 15,800 × g. The samples were then diluted to 1:50 in KPi buffer (50 mM, pH 7.0). The specific activity was calculated based on the DAP formed within the linear range, which was determined by HPLC analysis (see below).
One unit (U) of specific activity is defined as the amount of enzyme (in mg, calculated on the basis of protein content) which catalyses the formation of 1 µmol DAP per minute from the respective L-lysine concentration under the applied reaction conditions. The formation of DAP was monitored using HPLC as described below.
Characterisation of EcLDCc-CatIBs in CGXII medium supplemented with different L-lysine concentrations
To characterise the performance of EcLDCc-CatIBs for the transformation of different L-lysine concentrations from 10–100 mM, reactions of 2 mg ml−1 lyophilised EcLDCc-CatIBs each were performed in reaction tubes (2 ml safe-lock tube, Eppendorf, Germany) in 1 ml CGXII medium64 adjusted to pH 8.0 and containing 10, 20, 50, and 100 mM L-lysine and 0.1 mM PLP for 4 h at 30 °C and 1000 rpm in a thermomixer (Thermomixer comfort, Eppendorf, Germany). After different time intervals (6, 12, 24, 36, 60, 120, 180, and 246 min), 20 µl samples were taken and stopped by 1:5 dilution with methanol. The reaction mixture was then 1:10 diluted in KPi buffer (50 mM, pH 8.0) and subsequently centrifuged for 2 min at 15,800 × g. The samples thus obtained were diluted in an appropriate manner to obtain a final DAP concentration suitable for HPLC analysis (see below) between 10 and 100 µM. The specific activity was calculated as described above.
Bioreactor cultivation of C. glutamicum
See Supplementary.
Application of EcLDCc-CatIBs in a (repetitive) batch
EcLDCc-CatIBs were characterised in a cell-free culture supernatant (30 or 60 ml), with an adjusted pH of 8.0 and 0.1 mM PLP. Decarboxylation reactions of 100 mM and 1 M L-lysine were performed in repetetive batch and batch experiments, respectively. For the repetitive batch experiment, 3 mg ml−1 EcLDCc-CatIBs were used in 60 ml reaction solution. For the single batch reaction, starting from 1 M L-lysine, 10 mg ml−1EcLDCc-CatIBs were added to 30 ml reaction solution. The experiments were performed under pH-control by dosing NaOH (2 M) and HCl (5%), respectively, using a 665 Dosimat, 632 pH meter equipped with a 614 Impulsomat from Metrohm, Germany. For the dosage profile of NaOH and HCl during the batch reaction starting from 1 M L-lysine see Supplementary Fig. S6. Reactions were performed in a doubled-walled 3-neck reactor vessel with two nozzles for the cooling supply to keep the temperature constant at 30 °C. The reaction mixture was stirred with a magnetic stirrer. For the repetitive batch approach, consecutive batch experiments were performed for either 4 h or 15 h. After a batch of 4 h or 15 h, the reaction mixture was transferred to a centrifugal beaker and centrifuged at 30,966 × g for 2 min. The pellet was suspended in the fresh reaction solution described above and transferred back into the doubled-walled flask reactor vessel. 20 µl samples were taken from the supernatant of the respective batch (60 ml approach) and the reaction was stopped by 1:5 dilution with methanol. The reaction mixture was then diluted 1:100 in KPi buffer (50 mM, pH 8.0). 5 µl samples were taken from the 30 ml batch experiment after different time intervals (6, 12, 18, 30, 45, 60, 90, 120, 180, 240, 300, 354, 426, 480, 543, and 1434 min) and the reaction was stopped by 1:20 dilution with methanol. The reaction mixture was then diluted (1:250) in KPi buffer (50 mM, pH 8.0). All samples were subsequently centrifuged for 2 min at 15,800 × g. The amount of DAP formed was determined by HPLC analysis (see below).The specific activity was calculated as described above.
Quantification of L-lysine and DAP by HPLC
To determine the DAP concentration in cell-free and CatIB-free reaction solutions, a HPLC-system (Agilent 1100 Infinity, Agilent Technologies, Santa Clara, USA) was used, equipped with a fluorescence detector (excitation: 230 nm; emission: 460 nm) and a C18 KinetexEvo column (Phenomenex, Torrence, USA). Prior to injection, samples were diluted 1:2 (v/v) with 100 µM α-aminobutyric acid as the internal standard (Sigma-Aldrich, St. Louis; USA). Analysis of DAP and L-lysine was performed by a method for amino acid quantification71 including a pre-column derivatisation step at 18 °C using 5 µl ortho-phthaldialdehyde (OPA, Sigma-Aldrich) and 5 µl sample (6 mixing iterations). The mobile phase A was composed of 7.12 g l−1 Na2HPO4, 6.24 g l−1 NaH2PO4 and 0.8% (v/v) THF in water, and the mobile phase B contained 50% (v/v) methanol, 45% (v/v) acetonitrile, and 5% (v/v) water. For chromatographic separation, a linear gradient was applied with a flow of 1 ml min−1 starting with 0% B, 0–2 min 0–38% B, 2–6 min 38–42% B, 6–7 min 42–70% B, 7–9 min 70–100% B, 9–13 min 100-0% B. Approximate retention times were 8 min for α-aminobutyric acid, 10 min for L-lysine, and 11 min for DAP. In order to correct for possible effects of the analytical matrix on derivatisation efficiency, α-aminobutyric acid (Sigma-Aldrich, St. Louis; USA) was used as an internal standard. The DAP concentration was derived from the linear calibration of five reference solutions (10 µM to 100 µM), included in each measurement run (for the calibration curve see Supplementary Fig. S8).
Data availability
The datasets generated during the current study are available from the corresponding author on reasonable request.
References
Kriegler, E. et al. Fossil-fueled development (SSP5): An energy and resource intensive scenario for the 21st century. Glob. Environ. Chang. 42, 297–315 (2017).
U.S. Energy Information Administration. International Energy Outlook 2016. 0484, (2016).
Magnusson, L. U., Gummesson, B., Joksimović, P., Farewell, A. & Nyström, T. Identical, independent, and opposing roles of ppGpp and DksA in Escherichia coli. J. Bacteriol. 189, 5193–5202 (2007).
Chung, H. et al. Bio-based production of monomers and polymers by metabolically engineered microorganisms. Curr. Opin. Biotechnol. 36, 73–84 (2015).
Kremer, F., Blank, L. M., Jones, P. R. & Akhtar, M. K. A comparison of the microbial production and combustion characteristics of three alcohol biofuels: Ethanol, 1-butanol, and 1-octanol. Front. Bioeng. Biotechnol. 3, 112 (2015).
Wendisch, V. F. Microbial production of amino acids and derived chemicals: Synthetic biology approaches to strain development. Curr. Opin. Biotechnol. 30, 51–58 (2014).
Erickson, B., Nelson & Winters, P. Perspective on opportunities in industrial biotechnology in renewable chemicals. Biotechnol. J. 7, 176–185 (2012).
Park, S. J. et al. Synthesis of nylon 4 from gamma-aminobutyrate (GABA) produced by recombinant Escherichia coli. Bioprocess Biosyst. Eng. 36, 885–892 (2013).
Meng, D. C. et al. Production of poly(3-hydroxypropionate) and poly(3-hydroxybutyrate-co-3-hydroxypropionate) from glucose by engineering Escherichia coli. Metab. Eng. 29, 189–195 (2015).
Kind, S., Jeong, W. K., Schröder, H. & Wittmann, C. Systems-wide metabolic pathway engineering in Corynebacterium glutamicum for bio-based production of diaminopentane. Metab. Eng. 12, 341–351 (2010).
Jiang, Y. et al. Microbial production of short chain diols. Microb. Cell Fact. 13, 165 (2014).
Ma, W. et al. Advances in cadaverine bacterial production and its applications. Engineering 3, 308–317 (2017).
Litsanov, B., Brocker, M. & Bott, M. Toward homosuccinate fermentation: Metabolic engineering of Corynebacterium glutamicum for anaerobic production of succinate from glucose and formate. Appl. Environ. Microbiol. 78, 3325–3337 (2012).
Raab, A. M., Gebhardt, G., Bolotina, N., Weuster-Botz, D. & Lang, C. Metabolic engineering of Saccharomyces cerevisiae for the biotechnological production of succinic acid. Metab. Eng. 12, 518–525 (2010).
Thakker, C., Martínez, I., San, K.-Y. & Bennett, G. N. Succinate production in Escherichia coli. Biotechnol. J. 7, 213–24 (2012).
Tai, Y.-S. et al. Engineering nonphosphorylative metabolism to generate lignocellulose-derived products. Nat. Chem. Biol. 12, 247–253 (2016).
Honjo, H., Tsuruno, K., Tatsuke, T., Sato, M. & Hanai, T. Dual synthetic pathway for 3-hydroxypropionic acid production in engineered Escherichia coli. J. Biosci. Bioeng. 120, 199–204 (2015).
Okino, S. et al. An efficient succinic acid production process in a metabolically engineered Corynebacterium glutamicum strain. Appl. Microbiol. Biotechnol. 81, 459–464 (2008).
Ogunniyi, D. S. Castor oil: A vital industrial raw material. Bioresour. Technol. 97, 1086–1091 (2006).
Thielen, M. Bio-polyamides for automotive applications. Bioplastics Mag. 5, 10–11 (2010).
Mimitsuka, T., Sawai, H., Hatsu, M. & Yamada, K. Metabolic engineering of Corynebacterium glutamicum for cadaverine fermentation. Biosci. Biotechnol. Biochem. 71, 2130–2135 (2007).
Becker, J., Zelder, O., Häfner, S., Schröder, H. & Wittmann, C. From zero to hero—Design-based systems metabolic engineering of Corynebacterium glutamicum for L-lysine production. Metab. Eng. 13, 159–168 (2011).
Kind, S. et al. From zero to hero - Production of bio-based nylon from renewable resources using engineered Corynebacterium glutamicum. Metab. Eng. 25, 113–123 (2014).
Eggeling, L. & Bott, M. A giant market and a powerful metabolism: l-lysine provided by Corynebacterium glutamicum. Appl. Microbiol. Biotechnol. 99, 3387–3394 (2015).
Oh, Y. H. et al. Development of engineered Escherichia coli whole-cell biocatalysts for high-level conversion of L-lysine into cadaverine. J. Ind. Microbiol. Biotechnol. 42, 1481–1491 (2015).
Liu, P., Zhu, X., Tan, Z., Zhang, X. & Ma, Y. in Advances in Biochemical Engineering/Biotechnology 123, 107–140 (2015).
Kim, J.-H. et al. Functional study of lysine decarboxylases from Klebsiella pneumoniae in Escherichia coli and application of whole cell bioconversion for cadaverine production. J. Microbiol. Biotechnol. 26, 1586–1592 (2016).
Li, S., Xu, N., Liu, L. & Chen, J. Engineering of carboligase activity reaction in Candida glabrata for acetoin production. Metab. Eng. 22, 32–9 (2014).
Kim, H. J. et al. Optimization of direct lysine decarboxylase biotransformation for cadaverine production with whole-cell biocatalysts at high lysine concentration. J. Microbiol. Biotechnol. 25, 1108–1113 (2015).
Qian, Z. G., Xia, X. X. & Lee, S. Y. Metabolic engineering of Escherichia coli for the production of cadaverine: A five carbon diamine. Biotechnol. Bioeng. 108, 93–103 (2011).
Yamamoto, Y., Miwa, Y., Miyoshi, K., Furuyama, J. & Ohmori, H. The Escherichia coli ldcC gene encodes another lysine decarboxylase, probably a constitutive enzyme. Genes Genet. Syst. 72, 167–172 (1997).
Snider, J. et al. Formation of a distinctive complex between the inducible bacterial lysine decarboxylase and a novel AAA+ ATPase. J. Biol. Chem. 281, 1532–1546 (2006).
Kandiah, E. et al. Structural insights into the Escherichia coli lysine decarboxylases and molecular determinants of interaction with the AAA+ ATPase RavA. Sci. Rep. 6, 24601 (2016).
Kanjee, U. et al. Linkage between the bacterial acid stress and stringent responses: the structure of the inducible lysine decarboxylase. EMBO J. 30, 931–944 (2011).
Ma, W. et al. Enhanced cadaverine production from L-lysine using recombinant Escherichia coli co-overexpressing CadA and CadB. Biotechnol. Lett. 37, 799–806 (2015).
Kind, S., Jeong, W. K., Schröder, H., Zelder, O. & Wittmann, C. Identification and elimination of the competing N-acetyldiaminopentane pathway for improved production of diaminopentane by Corynebacterium glutamicum. Appl. Environ. Microbiol. 76, 5175–80 (2010).
Li, M. et al. Improving the secretion of cadaverine in Corynebacterium glutamicum by cadaverine-lysine antiporter. J. Ind. Microbiol. Biotechnol. 41, 701–9 (2014).
Vrljic, M., Sahm, H. & Eggeling, L. A new type of transporter with a new type of cellular function: L-lysine export from Corynebacterium glutamicum. Mol. Microbiol. 22, 815–26 (1996).
Li, N., Chou, H., Yu, L. & Xu, Y. Cadaverine production by heterologous expression of Klebsiella oxytoca lysine decarboxylase. Biotechnol. Bioprocess Eng. 19, 965–972 (2014).
Bhatia, S. K. et al. Biotransformation of lysine into cadaverine using barium alginate-immobilized Escherichia coli overexpressing CadA. Bioprocess Biosyst. Eng. 38, 2315–2322 (2015).
Kim, J.-H. et al. Development of a continuous L-lysine bioconversion system for cadaverine production. J. Ind. Eng. Chem. 46, 44–48 (2016).
Seo, H. M. et al. In situ immobilization of lysine decarboxylase on a biopolymer by fusion with phasin: Immobilization of CadA on intracellular PHA. Process Biochem. 51, 1413–1419 (2016).
Park, S. H., Soetyono, F. & Kim, H. K. Cadaverine production by using cross-linked enzyme aggregate of Escherichia coli lysine decarboxylase. J. Microbiol. Biotechnol. 27, 289–296 (2017).
Li, S., Lin, K., Pang, H., Wu, Y. & Xu, J. Production, characterization, and application of an organic solvent-tolerant lipase present in active inclusion bodies. Appl. Biochem. Biotechnol. 169, 612–623 (2013).
Nahálka, J., Vikartovská, A. & Hrabárová, E. A crosslinked inclusion body process for sialic acid synthesis. J. Biotechnol. 134, 146–153 (2008).
Worrall, D. M. & Goss, N. H. The formation of biologically active beta-galactosidase inclusion bodies in Escherichia coli. Aust. J. Biotechnol. 3, 28–32 (1989).
Krauss, U., Jäger, V. D., Diener, M., Pohl, M. & Jaeger, K.-E. Catalytically-active inclusion bodies—Carrier-free protein immobilizates for application in biotechnology and biomedicine. J. Biotechnol. 258, 136–147 (2017).
Sheldon, R. A. Enzyme Immobilization: The Quest for Optimum Performance. Adv. Synth. Catal. 349, 1289–1307 (2007).
Barbosa, O. et al. Strategies for the one-step immobilization–purification of enzymes as industrial biocatalysts. Biotechnol. Adv. 33, 435–456 (2015).
Tufvesson, P., Lima-Ramos, J., Nordblad, M. & Woodley, J. M. Guidelines and cost analysis for catalyst production in biocatalytic processes. Org. Process Res. Dev. 15, 266–274 (2011).
Rodríguez-Carmona, E., Cano-Garrido, O., Seras-Franzoso, J., Villaverde, A. & García-Fruitós, E. Isolation of cell-free bacterial inclusion bodies. Microb. Cell Fact. 9, 71 (2010).
Tokatlidis, K., Dhurjati, P., Millet, J., Béguin, P. & Aubert, J.-P. High activity of inclusion bodies formed in Escherichia coli overproducing Clostridium thermocellum endoglucanase D. FEBS Lett. 282, 205–208 (1991).
Nahálka, J. & Pätoprstý, V. Enzymatic synthesis of sialylation substrates powered by a novel polyphosphate kinase (PPK3). Org. Biomol. Chem. 7, 1778–1780 (2009).
Choi, S. L. et al. Generation of catalytic protein particles in Escherichia coli cells using the cellulose-binding domain from Cellulomonas fimi as a fusion partner. Biotechnol. Bioprocess Eng. 16, 1173–1179 (2011).
Hrabárová, E., Achbergerová, L. & Nahálka, J. In Insoluble Protein: Methods and protocols (ed. García-Fruitós, E.) 1258, 411–422 (Springer New York, 2015).
Nahálka, J., Mislovicová, D. & Kavcová, H. Targeting lectin activity into inclusion bodies for the characterisation of glycoproteins. Mol. Biosyst. 5, 819–21 (2009).
Park, S. Y., Park, S. H. & Choi, S. K. Active inclusion body formation using Paenibacillus polymyxa PoxB as a fusion partner in Escherichia coli. Anal. Biochem. 426, 63–65 (2012).
García-Fruitós, E. et al. Aggregation as bacterial inclusion bodies does not imply inactivation of enzymes and fluorescent proteins. Microb. Cell Fact. 4, 27 (2005).
Wang, X., Zhou, B., Hu, W., Zhao, Q. & Lin, Z. Formation of active inclusion bodies induced by hydrophobic self-assembling peptide GFIL8. Microb. Cell Fact. 14, 88 (2015).
Zhou, B., Xing, L., Wu, W., Zhang, X.-E. & Lin, Z. Small surfactant-like peptides can drive soluble proteins into active aggregates. Microb. Cell Fact. 11, 10 (2012).
Peters, J. et al. Tetrabrachion: A filamentous archaebacterial surface protein assembly of unusual structure and extreme stability. J. Mol. Biol. 245, 385–401 (1995).
Diener, M., Kopka, B., Pohl, M., Jaeger, K.-E. & Krauss, U. Fusion of a coiled-coil domain facilitates the high-level production of catalytically active enzyme inclusion bodies. ChemCatChem 8, 142–152 (2016).
Kikuchi, Y., Kojima, H., Tanaka, T., Takatsuka, Y. & Kamio, Y. Characterization of a second lysine decarboxylase isolated from. Escherichia coli. J. Bacteriol. 179, 4486–4492 (1997).
Keilhauer, C., Eggeling, L. & Sahm, H. Isoleucine synthesis in Corynebacterium glutamicum: molecular analysis of the ilvB-ilvN-ilvC operon. J. Bacteriol. 175, 5595–603 (1993).
Follmann, M. et al. Functional genomics of pH homeostasis in Corynebacterium glutamicum revealed novel links between pH response, oxidative stress, iron homeostasis and methionine synthesis. BMC Genomics 10, 621 (2009).
Grünberger, A. et al. Microfluidic picoliter bioreactor for microbial single-cell analysis: Fabrication, system setup, and operation. J. Vis. Exp. 1–11 https://doi.org/10.3791/50560 (2013).
Grünberger, A. et al. Spatiotemporal microbial single-cell analysis using a high-throughput microfluidics cultivation platform. Cytom. Part A 87, 1101–1115 (2015).
Rokney, A. et al. E. coli transports aggregated proteins to the poles by a specific and energy-dependent process. J. Mol. Biol. 392, 589–601 (2009).
Ventura, S. & Villaverde, A. Protein quality in bacterial inclusion bodies. Trends Biotechnol. 24, 179–185 (2006).
Krithika, G., Arunachalam, J., Priyanka, H. & Indulekha, K. The two forms of lysine decarboxylase; Kinetics and effect of expression in relation to acid tolerance response in E. coli. J. Exp. Sci. 1, 10–21 (2010).
Limberg, M. H. et al. Metabolic profile of 1,5-diaminopentane producing Corynebacterium glutamicum under scale-down conditions: Blueprint for robustness to bioreactor inhomogeneities. Biotechnol. Bioeng. 114, 560–575 (2017).
Reiber, H. Photochemical reactions of vitamin B6 compounds, isolation and properties of products. Biochem. Biophys. Acta 279, 310–315 (1972).
Holde, V., Sabo, D. L. & Fischer, E. H. Chemical properties of Escherichia coli lysine decarboxylase including a segment of its pyridoxal 5′-phosphate binding site. Biochemistry 13, 670–676 (1974).
Bradford, M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 248–254 (1976).
Acknowledgements
The scientific activities of the Bioeconomy Science Center were financially supported by the Ministry of Innovation, Science and Research within the framework of the NRW Strategieprojekt BioSC (No. 313/323‐400‐002 13). BioSC is a research cluster of RWTH Aachen University, the universities of Düsseldorf and Bonn, and Forschungszentrum Jülich. The authors thank the Federal German Ministry of Education and Science for providing funding for the cluster project “Systematic consideration of inhomogeneity at the large scale” (grant no. 031A220A), which is embedded in the ERA-IB framework.
Author information
Authors and Affiliations
Contributions
D.H. performed the construction of plasmids under the direction of R.K. and U.K., who performed structure-based planning of the genetic construct. V.D.J. and U.K. supported CatIB preparation. U.M. performed production, purification, and characterisation of the CatIBs and analysed the data with the assistance of R.K. and M.H.L., and the coordination and supervision of M.P. A.G. performed live-cell imaging with the assistance of V.D.J. and R.K. M.H.L. and R.K. performed microbial L-lysine production under the coordination and supervision of M.O. R.K., M.H.L. and M.P. wrote the manuscript with input from A.G., U.K. and M.O.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kloss, R., Limberg, M.H., Mackfeld, U. et al. Catalytically active inclusion bodies of L-lysine decarboxylase from E. coli for 1,5-diaminopentane production. Sci Rep 8, 5856 (2018). https://doi.org/10.1038/s41598-018-24070-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-24070-2
- Springer Nature Limited
This article is cited by
-
Construction and characterization of BsGDH-CatIB variants and application as robust and highly active redox cofactor regeneration module for biocatalysis
Microbial Cell Factories (2022)
-
Tailor-made active enzyme aggregates in vivo for efficient degradation of biomass polysaccharide into reducing sugar
Biomass Conversion and Biorefinery (2022)
-
Construction and comprehensive characterization of an EcLDCc-CatIB set—varying linkers and aggregation inducing tags
Microbial Cell Factories (2021)
-
Thermostable adenosine 5′-monophosphate phosphorylase from Thermococcus kodakarensis forms catalytically active inclusion bodies
Scientific Reports (2021)
-
Coiled-coil inspired functional inclusion bodies
Microbial Cell Factories (2020)