Abstract
RIF1 plays a key role in inhibiting DNA end resection and promoting NHEJ mediated DNA double stand break repair in G1. However, whether SUMOlyation may regulate RIF1 functions is still largely unknown. Here, we report that RIF1 is SUMOlyated in response to DNA damage. We identified PIAS4 as the primary SUMO E3 ligase required for the SUMOylation of RIF1 protein. Mammalian cells compromised of PIAS4 expression, show impaired RIF1 SUMOylation and defective for the disassembly of DNA damage responsive RIF1 foci. Mechanistically, we show that PIAS4 knockdown abrogates UHRF1-dependent ubiquitination of RIF1, compromising RIF1 protein turnover. We detected intense RPA foci that colocalize with RIF1 foci in PIAS4 knockdown cells. These data highlight an important role of PIAS4-dependent regulation of RIF1, likely mediated by SUMOylation, in the disassembly of RIF1 DNA damage response (DDR) foci. We propose that unresolved RIF1 protein at sites of DNA damage in PIAS4-depleted cells largely accumulates in S phase, and subsequently leads to DNA double strand breaks. Therefore, PIAS4 promotes genomic stability by regulating the timely removal of RIF1 from sites of DNA damage.
Similar content being viewed by others
Introduction
DNA damage activates a wide range of responses including altered gene expression, cell cycle arrest and activation of DNA repair1. To preserve genome integrity after genotoxic insult, eukaryotic cells have developed a highly conserved surveillance mechanism, collectively termed the DNA damage response (DDR) pathway2,3. In response to DNA double strand breaks (DSBs), components of DDR signaling drive two main repair pathways, NHEJ and HR4,5. In G1 cells, in the absence of sister chromatid and inadequate CDK activity, nucleolytic resection of 5′ end is inhibited, which in turn promotes the 53BP1-mediated NHEJ break processing6. However, in S and G2 phases, CDK phosphorylation of BRCA1/CtIP drives the 5′–3′ DNA end resection which facilitates the HR process to repair the DNA DSBs7.
PTMs involve (but not limited to) phosphorylation, methylation, acetylation, SUMOylation and Ubiquitination. In the latter two PTMs, Ubiquitin and SUMO polypeptides are covalently attached to target protein via isopeptide linkage8,9. The extent of SUMO modifications of the target proteins depends on the number of SUMO conjugation. Some of the target proteins have a single SUMO attached, while in others, multiple Lys residues on the target are individually linked to SUMO10,11.
Coordinated PIAS1 and PIAS4 mediated protein SUMOylation and ubiquitination facilitate the distribution of DDR components (MDC1, BRCA1 and 53BP1) at the sites of DNA breaks and promote the repair process12. SUMOylation deficient mouse embryos die early due to defective chromosomal segregation, suggesting a key role for SUMO in maintaining genomic integrity13,14. It has been established that SUMO conjugates, SUMO-conjugating enzymes UBC9 (UBE2I) and SUMO E3 ligases, PIAS1 (protein inhibitor of activated STAT 1) and PIAS4 (PIASy), are recruited at sites of DSB, which in turn promote DSB signaling and repair12,15. PIAS4 mediates SUMO-2 conjugation of Topoisomerase-II on mitotic chromosomes16. SUMO2 modification of Rev1 by PIAS4 regulates p53-dependent cancer cell death in response to oxidative stress17. Elegant works from different laboratories indicates that PIAS1 and PIAS4 function in parallel but overlapping SUMO-conjugation pathways to facilitate the DNA break repair12,15. Previous studies have also detected SUMOylated 53BP1 in His purified SUMO2 conjugates and unlike BRCA1 and MDC1, SUMOylated 53BP1 was not increased after RNF4 knockdown18. Earlier studies have revealed a function for SUMO and ubiquitin in the recruitment and disassembly of DNA repair foci to prevent genomic instability19,20,21,22.
Identification of RIF1 at the sites of DNA breaks was reported previously23,24,25. However, its broader function in the regulation of key DNA repair process has only recently been evidenced. RIF1 has been identified as an effector of 53BP1, which modulates the DNA DSBs repair by regulating NHEJ in G1 cells. In contrast, during S/G2 phase of cell cycle, BRCA1-CtIP mediated DNA end resection prevents NHEJ through the removal of 53BP1-RIF1 from DSBs26,27,28,29,30,31. Several earlier reports have demonstrated novel functions of RIF1 in the maintenance of genomic stability, replication timing, nuclear architecture, class switch recombination and immunological functions32,33,34,35,36. RIF1 is a large nuclear protein. It’s molecular and biochemical basis of action and its upstream regulation is still unclear. BLM and RIF1 interact physically and are recruited at the stalled replication fork with similar kinetics37. In addition, BLM SUMOylation is required for RAD51 localization at damaged replication forks and repair by HR38,39.
In this study we report that RIF1 is regulated by SUMOylation in response to DNA damage. We identified PIAS4 as the main SUMO E3 ligase required for RIF1 SUMOylation. PIAS4 deficient mammalian cells showed impaired RIF1 SUMOylation and defective disassembly of RIF1 DDR foci after recovery from DNA damage. These RIF1 foci resulted in increased replication stress and DNA double strand breaks. Moreover, we noticed multiple RIF1 and 53BP1 nuclear bodies in PIAS4 depleted cells. Overall, we have identified RIF1 as a novel PIAS4 target protein required for the maintenance of genomic integrity.
Results
RIF1 SUMOylation is increased in response to DNA double strand breaks
The increasing importance of SUMOylation in the regulation of DDR response and protein dynamics at DNA breaks prompted us to investigate the role of RIF1 SUMOylation in the regulation of RIF1 functions. To detect RIF1 SUMOylation in vivo, we have used a U2OS cell line stably expressing 10 His SUMO240,41. DMSO or bleocin treated cells were lysed and His purified SUMO2 protein samples were immunoblotted to detect SUMOylated RIF1 protein. We noted a distinct slower migrating form representing SUMOylated RIF1. Interestingly, SUMOylated RIF1 protein signal was further increased in response to bleocin and MG132 treatment (Figs 1A and S1A). We further tested the specificity of DNA damaging agents (listed in Table 1), triggering RIF1 SUMOylation and we noticed an enhanced SUMOylated signal in response to all genotoxic agents included in this study (Fig. 1B). We noticed a better RIF1 SUMOylation signal in response to bleocin and MMS treatment. Similar to RIF1, SUMOylation of 53BP1 was also increased in response to different forms of DNA damaging agents (Figs 1C and S1B). His purified SUMO2 conjugates were blotted with SUMO2/3 antibody to determine the total level of SUMO2 in different protein samples (Figs 1A, S1A, 1D and S1B lower panels).
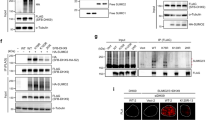
RIF1 is SUMOylated protein. (A) U2OS cells stably expressing 10 His-SUMO2 were either DMOS treated or treated with indicated doses of bleocin and MG132. Cells were harvested and SUMO2 protein conjugates were purified from denaturing lysates. Whole cell lysate (WCE) and His purified SUMO2 conjugates were immunoblotted with antibodies directed against RIF1 and SUMO2/3. Additionally, parental U2OS cells were included in all experiments, as a negative control. (B) U2OS cells stably expressing 10 His-SUMO2 were either DMOS treated or treated with the indicated DNA damaging agents. His purified SUMO2 conjugates and WCE were immunoblotted with RIF1 antibody. (C) Protein samples from Fig. 1B were immunoblotted with antibody directed against 53BP1. (D) Total SUMO2/3 in His purified SUMO2 conjugates were detected using SUMO2/3-specific antibody. (E) Colocalization between RIF1 and SUMO2 foci. U2OS cell were transiently transfected with RFP-SUMO2 construct and treated with bleocin. Cells were immunostained with RIF1 antibody and subsequently stained with Alexa Fluor 488 secondary antibody. Horizontal bar represents 10 μm. (F) U2OS cells transfected with RFP-SUMO2, RFP-SUMO3 and RFP empty vector constructs. Cell were mock treated or treated with low dose (2.5 µg/ml) of bleocin. RFP-tagged proteins and associated protein conjugates were purified using RFP-Trap® assay and immunoblotted with RIF1 antibody.
RIF1 is recruited at sites of DNA damage and colocalize with phosphorylated H2AX (pS139)24,42. Earlier reports have also shown that 53BP1 and BRCA1 colocalize differentially with SUMO isoforms at the local site of DNA damage12,15. We found that the majority of RIF1 foci colocalize with SUMO2 (Fig. 1E). In contrast, only a small fraction of RIF1 foci colocalize with SUMO1 (Fig. S1C). To confirm our cell biology data, RFP fused SUMO2 and SUMO3 were analyzed for their ability to bind RIF1. Interestingly, we detected RIF1 in RFP-SUMO2 and SUMO3 purified protein complexes (Fig. 1F). Taken together our results indicate that RIF1 is a SUMO modified protein and SUMOylation of RIF1 is increased in response to a broad range of DNA damaging agents.
RIF1 SUMOylation is increased in G1 cells
To determine the cell cycle specificity of the RIF1 SUMOylation, cell synchronization experiments were performed (Fig. 2A). Flow cytometry data confirms the enrichment of cells at different cell cycle phases (Fig. 2B). Interestingly, a clear enhancement of SUMOylated RIF1 signal was observed in G1 cells, which was further increased in response to bleocin treatment (Fig. 2C). RIF1 SUMOylation was decreased in S and G2 cells, confirming the specificity of RIF1 SUMOylation in G1. Similar to RIF1 SUMOylation, DNA damage induced 53BP1 SUMOylation was also enriched in G1 cells (Fig. 2D). His purified SUMO2 conjugates were immunoblotted with SUMO2/3 antibody (Fig. 2E), which demonstrated that the levels of purified protein complexes are similar in different samples.
Cell cycle dependent RIF1 SUMOylation is increased in G1 cells. (A) Schematic diagram of the cell cycle synchronization and release experiment. (B) Graph represents the percentages of G1, S and G2/M cell populations determined by flow cytometry. (C) Timed release of G1 synchronized His10-SUMO2 expressing U2OS cells from double thymidine block followed by treatment with bleocin and/or MG132. His purified SUMO2 conjugates and WCE were immunoblotted with RIF1 antibody. (D) His purified protein samples from the Fig. 3C were immunoblotted with 53BP1 antibody. (E) His purified protein samples and WCE were immunoblotted with SUMO2/3 and actin antibody respectively.
SUMO E3 ligase PIAS4 regulates RIF1 SUMOylation
Increasing evidences suggest that members of the protein inhibitor of activated signal transducer and activator of transcription (PIAS) family of SUMO E3 ligases play crucial roles in the maintenance of genomic integrity. To identify the SUMO E3 ligase required for RIF1 SUMOylation, we first depleted cells of PIAS1 and PIAS4 by using specific RNAi sequences (listed in Table 2). Interestingly, we noted a substantial reduction of RIF1 SUMOylation specifically upon PIAS4 depletion (Fig. 3A), whereas PIAS1 depletion affected only partially the extent of RIF1 SUMOylation (Figs 3A and S3A). This indicates a predominant role of PIAS4 in regulating RIF1 SUMOylation. Consistent with an earlier report18, our data also suggest that SUMO2 modification of 53BP1 is significantly increased in response to DNA double strand breaks (Figs 1C, 2B and S1B). Here, we further demonstrated that PIAS4 is also required for SUMO-2 modification of 53BP1. PIAS4 depletion significantly reduced the level of SUMOylated 53BP1 and residual SUMOylation signal was abolished after PIAS1 and PIAS4 co-depletion (Fig. 3B). Total SUMO2 proteins in purified complexes were determined by SUMO2/3 (Figs 3C and S3A) or 6xHis (Fig. S3B) immunoblotting.
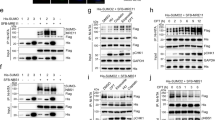
SUMO E3 ligase PIAS4 regulates RIF1 SUMOylation. (A) U2OS cells stably expressing 10 His-SUMO2 were either transfected with control shRNA or PIAS1 and PIAS4-specific shRNAs, and were treated with bleocin or DMSO. His-SUMO2 conjugates were purified and immunoblotted with RIF1 antibody. Protein samples from WCE were immunoblotted with RIF1 antibody. (B) Protein samples from the above experiment were analyzed for 53BP1 SUMOylation. WCE were probed with 53BP1, PIAS1, PIAS4 and y-H2AX antibodies. (C) His purified SUMO2 conjugates from (A) were immunoblotted with SUMO2/3 antibody to detect total SUMO2/3. (D) U2OS cells stably expressing His10-SUMO2 were either transfected with control siRNA or with two independent siRNAs targeting 53BP1. 3 days later, cells were treated with bleocin or DMSO as control. His purified SUMO2 conjugates and WCE were immunoblotted with RIF1 antibody. WCE was immunoblotted with 53BP1 antibody. His purified SUMO2 conjugates were immunoblotted with anti-6XHis tag antibody to establish equivalent His pull-down efficiency in all conditions. (E) U2OS cells were transiently transfected with FLAG-PIAS4 and synchronized at G1, S and G2. Flag-IP samples were probed with RIF1 antibody to determine the RIF1-PIAS4 bindings in different cell cycle stages. Whole cell lysates were immunoblotted with RIF1 and Flag antibody to determine the total proteins.
Given that phosphorylation of 53BP1 was reported to be important in the regulation of RIF1 functions in NHEJ27, we asked if 53BP1 is directly required for RIF1 SUMOylation. However, we did not observe any substantial reduction of SUMOylated RIF1 signal in 53BP1-depleted cells (Fig. 3D), suggesting that RIF1 SUMOylation is not regulated by 53BP1. Whole cell lysates and His purified SUMO2 conjugates were immunoblotted for His6 (Fig. 3D lower panel) to determine the total level of proteins.
To understand the molecular basis of PIAS4 dependent RIF1 SUMOylation, we analyzed the extent of RIF1-PIAS4 interaction in cells and asked whether that may be cell cycle-regulated. U2OS cells were transiently transfected with FLAG-PIAS4 plasmid DNA and cells were enriched in different cell cycle phases, using the same synchronization protocol as depicted in Fig. 2A. Clearly, we observed an increased interaction between RIF1 and PIAS4 in G1-enriched cells (Fig. 3E) as well as in S phase enriched cells. In contrast, a reduced RIF1-PIAS4 binding was detected in G2-enriched synchronized cells. This result provided evidence that support our earlier findings that RIF1 is mainly SUMOylated in G1 cells (Fig. 2C). Additionally, the increased PIAS4-RIF1 interaction in S phase cells, which was further enhanced in response to DNA damage, suggest other potential roles of PIAS4 in regulating RIF1 in replicating cells.
PIAS4 is required for the resolution of RIF1 foci, in response to DNA DSBs
To investigate the importance of PIAS4-dependent RIF1 SUMOylation, we examined the recruitment of RIF1 to γH2AX-marked sites of damage in PIAS4-depleted cells. We observed a substantial increase in γH2AX foci that colocalized with Rif1 foci upon bleocin treatment (Fig. S4A). RIF1 foci also colocalize with pATM foci. (Fig. S4B). Despite an initial delay in the recruitment of RIF1 and γH2AX to DDR sites in PIAS4-depleted cells (Fig. 4C), we did not observe any drastic change in RIF1 or γH2AX foci formation (Fig. 4B-II and 4C), suggesting that the recruitment of RIF1 to sites of DNA damage is not completely dependent on PIAS4 functions. This prompted us to investigate if the dynamics of RIF1/DDR foci following recovery from DNA damage may be dependent on PIAS4. Therefore, we examined the resolution of RIF1 and γ-H2AX foci following recovery of cells from DNA damage (Fig. 4A and B). As expected, in control cells, the kinetics of RIF1/γH2AX foci shows a time-dependent resolution. Surprisingly, a striking contrast in the resolution of RIF1 foci was observed in PIAS4-depleted cells (Fig. 4BII and C). We observed a four-fold increase in the signal intensity of RIF1 foci remaining in PIAS4-depleted cells compared to control cells at 10 hours after recovery from bleocin (Fig. 4D), suggesting that PIAS4 depletion significantly impaired the clearance of RIF1 from DNA damage sites. PIAS4 depletion was tested by immunoblotting (Fig. 4E). Similarly, we noticed an increased RIF1, RPA70 and pCHK1 proteins, indicating an overall increase in DNA damage in PIAS4 depleted cells (Fig. 4F). Taken together, our data suggest that PIAS4 function is required for the disassembly of RIF1 DDR foci from the sites of DNA damage.
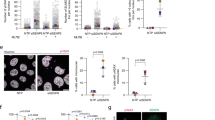
PIAS4 is required for the resolution of DNA damage induced RIF1 foci. (A) Schematic representation of experiment to assess the extent of resolution of DDR foci following recovery from DNA damage. (B) Representative images of U2OS cells were transfected with (i) control (Luc) siRNA and PIAS4 siRNA. Cells were treated with bleocin for 1 hour and recovered in fresh medium to allow for recovery from DNA damage. At the indicated timepoint following incubation in fresh media, cells were permeabilized and fixed with formaldehyde, followed by co-immunostaining with RIF1 and γ-H2AX antibodies. (C) Intensities of nuclear RIF1 signals were quantified using ImageJ and expressed as ratios of the maximal signal intensity detected within the 10 hours of recovery. Graph shows the intensity ratios of cells transfected with PIAS4-specific siRNA or control (Luc) siRNA at different timepoints following recovery. (D) As in (B) and (C), the relative signal intensities were detected using (i) RIF1 and γ-H2AX antibodies and quantified using ImageJ. Relative signal intensity ratios for the 10 hours timepoint were plotted, comparing control (Luc) siRNA and PIAS4 siRNA transfected cells. Data represent mean and SD from two independent experiments (*) P < 0.001. (E) WCE were immunoblotted to determine the knockdown level PIAS4 in experiment A to D. (F) Control (Luc) and PIAS4 siRNA transfected cells were treated with bleocin and harvested at the indicated timepoints following recovery in fresh media. WCE were immunoblotted using antibodies against RIF1, RPA70, pChk1 and PIAS4.
PIAS4 prevents DNA double stand breaks in S phase cells and Ultrafine Bridges
The persistent nature of the RIF1 foci in PIAS4-depleted cells led us to further question the fate and consequence of these RIF1 foci. To investigate that, we extended the recovery time and examined the resolution of RIF1 foci at later time points. Consistently, most of the RIF1 foci were effectively resolved in cells having normal PIAS4 expression. However, we still observed intense RIF1 foci in PIAS4 depleted cells despite extended recovery (16 and 24 hours) (Fig. 5A). Interestingly, in the absence of PIAS4, we observed persistent RIF1 foci in S phase, as marked by EdU positivity (Fig. 5A). A careful quantification of EdU stained cells revealed that a substantial proportion of replicating cells harbor RIF1 foci in the absence of PIAS4 (Fig. 5B). Interestingly we noted distinct RPA70 foci formation in the absence of PIAS4 (Fig. 5C). A careful quantification revealed that RPA foci are clearly increased in PIAS4 depleted cells, compared to control cells, at the 16 hours of time point following recovery from bleocin. Secondly, a large proportion of these foci colocalized with the RIF1 foci and to some extent with γ-H2AX foci (Fig. 5D). Consistently, we observed a significant increase in RIF1 foci in PIAS4-depleted cells, compared with the control cells. Therefore, it is conceivable that one of the major consequences of PIAS4 absence is unresolvable RIF1 DDR foci, which persist longer and posing a potential threat to genomic integrity.
PIAS4 depletion resulted in persistent RIF1 foci and replication stress. (A) U2OS cells were transfected with control siRNA or PIAS4 siRNA and treated with bleocin (1 hr) or DMSO as control. Cells were recovered in fresh media for 16 hours or 24 hours. Formaldehyde fixed cells were immunostained with RIF1 antibody and EdU positive S phase cells were detected using Click-iT® assay kit. (B) Percentage of Edu positive cells displaying RIF1 foci at 16 hrs. or 24 hrs. following drug recovery, either in presence (Luc si) or in the absence of PIAS4 (PIAS4 si). (C) Control (Luc) siRNA and PIAS4 siRNA transfected U2OS cells were treated with bleocin (1 hr) and recovered in fresh medium for 16 hours. Cells were immunostained for RIF1, RPA70 and γ-H2AX. Inset shows colocalized RIF1, RPA70 and γ -H2AX foci. (D) After the recovery, number RIF1, γ-H2AX and RPA foci per cell were counted. (E) U2OS cells were transfected with control (Luc) siRNA or PIAS4 siRNA and immunostained with RIF1 antibody. Mitotic cells were analyzed for the presence of RIF1-positive ultrafine anaphase bridges (UFBs). (F) Percentage of mitotic cells with UFBs in control or PIAS4 siRNA transfected cells. Data represent at least two independent experiments; values represent mean ± SD.
Recently, RIF1 has been reported to be recruited at Ultrafine Bridges (UFBs), which is a major consequence of abnormal DNA structure, carried-over from S-phase into mitosis43. During anaphase RIF1 promotes the resolution of UFBs and prevents these structures or chromosomal lesions from being transmitted to G1 cells in the form of nuclear bodies44. Consistent with this observation, we noticed RIF1 at the sites of UFBs in mitosis (Fig. 5E). We further investigated the involvement of PIAS4 in the regulation of the UFBs. Surprisingly, the frequency of UFBs formation was considerably increased in PIAS4 depleted cells (Fig. 5F). These observations strongly suggest that cells lacking PIAS4 activity are susceptible to replication stress, frequent UFBs formation and subsequently lead to genomic instability.
PIAS4 prevents unusual RIF1 and 53BP1 nuclear bodies
Unresolved replication intermediates generated during late stage of replication, can lead to the formation of nuclear bodies in G1 cells44,45,46. In normal proliferating cells, the p53 binding protein 53BP1 has been identified as a key component of these nuclear bodies45,46,47. Upon induction of low dose of replication stress, 53BP1 differentially colocalize to MDC1, pATM, RNF168, Ubiquitin conjugating enzyme (FK1), BLM and few other proteins involved in DNA replication stress. We consistently noticed intense and enlarged RIF1 foci resembling the previously described 53BP1 nuclear bodies (Fig. 6A). The previously reported accumulation of 53BP1 and other DNA Repair proteins in nuclear bodies (NBs), prompted us to examine the colocalization of RIF1 with NBs. Indeed, a majority of RIF1 colocalize with 53BP1-positive NBs (Fig. 6A). To determine the cell cycle specificity, cells were immunostained with RIF1 and CyclinB1 antibodies (listed in Table 3). We noticed that RIF1 NBs were mainly present in Cyclin B1 negative cells, indicating that these specific foci are absent in G2 cells (Fig. 6B). We further confirmed our observation using EdU labeling of S phase cells and noticed that RIF1 NBs were present predominantly in EdU negative cells (Fig. 6C). Together, these data suggest that, similar to previously identified 53BP1, RIF1 also exist as nuclear bodies in G1 cells.
PIAS4 is required to suppress multiple RIF1/53BP1 Nuclear Bodies. (A) Nuclear bodies containing RIF1 and 53BP1 were detected using specific antibodies. (B) U2OS cells were co-immunostained with RIF1 and Cyclin B1 antibodies. (C) U2OS cells were incubated with EdU for 60 minutes and immunostained with RIF1 antibody. (D) U2OS cells transfected with either control siRNA or PIAS4 siRNA were immunostained with RIF1 and 53BP1 antibodies. (E) Percentage of EdU positive cells containing different numbers of RIF1 nuclear bodies either in presence of in the absence of PIAS4. (F) Percentage of Edu positive cells containing different numbers of 53BP1 nuclear bodies either in presence of in the absence of PIAS4. Data represent at least two independent experiments; values represent mean ± SD.
Given that nuclear bodies are a potential consequence of aberrant UFBs resolution45 and our data suggests that PIAS4 plays a critical role in suppressing abnormal UFBs formation (Fig. 5E and F), we predicted that PIAS4 might have important role in preventing the formation of NBs. Indeed, we found that PIAS4 depleted cells displayed a remarkable increase in the number of intense NBs, containing both RIF1 and 53BP1 (Fig. 6D). Considering the differential number of nuclear bodies in each cell, we grouped cells according to the number of NBs present in different cells (Fig. 6E). Interestingly, compared to control siRNA treated cells, the proportion of cells with more than 5 nuclear bodies were increased in PIAS4 depleted cells (Fig. 6E and F). Together, these data suggest an important role of PIAS4 in the suppression of RIF1 and 53BP1 nuclear bodies.
PIAS4 is required for RIF1 Ubiquitination
Epigenetic regulator UHRF1 (Ubiquitin-like, with PHD and RING finger domains 1) has been shown to play important role in the maintenance of genomic integrity. Recently, it has been shown that UHRF1 is recruited by BRCA1 in S phase to ubiquitinate RIF1 and promotes its dissociation from 53BP1 at DNA DSB sites48. Here, we show that RIF1 foci persist in EdU positive cells in the absence of SUMO E3 ligase PIAS4. These results prompted us to question if RIF1 ubiquitination and protein turnover at sites of DNA breaks has been compromised, specifically when PIAS4-dependent SUMOylation of RIF1 is impaired. We hypothesized that PIAS4 mediated RIF1 SUMOylation is required for RIF1 ubiquitination and dissociation from the site of DNA damage. To analyze the RIF1 Ubiquitination in vivo, U2OS cell stably expressing 10-His Ubiquitin were treated with bleocin. Consistently, we detected RIF1 ubiquitination that was largely dependent on UHRF1. Remarkably, we also noticed a substantial reduction in the extent of RIF1 Ubiquitination when PIAS4 is depleted (Fig. 7A). The level of ubiquitinated RIF1 signal intensity in UHRF1 depleted cells was comparable to that in PIAS4-depleted cells (Fig. 7B). Whole cell lysates were analyzed to detect the protein levels of PIAS4, UHRF1 and γ-H2AX in damaged and untreated cells (Fig. 7C). His purified Ubiquitin protein conjugates were immunoblotted to determine the total level of ubiquitin (Fig. 7D).
PIAS4 regulates RIF1 Ubiquitination. (A) U2OS cells stably expressing His10-Ubiquitin were transfected with control (Luc) siRNA or siRNAs specific to UHRF1 or PIAS4. Cells were treated with bleocin or DMSO as control and harvested for His purification. His purified Ubiquitin conjugates were immunoblotted with RIF1 antibody, and anti-His and anti-Ub to detect total His/Ub pulldown in all conditions. (B) The intense ubiquitinated RIF1 protein bands were quantified by imageJ software and relative image signal intensity of different protein samples were plotted on Y-axis. (C) Protein samples from WCE were immunoblotted with indicated antibodies to detect the expression of RIF1 (Fig. 7A lower panel), PIAS4, UHRF1 and y-H2AX (Fig. 7C). (D) His ubiquitin conjugates were immunoblotted with ubiquitin specific antibody to determine the level of total ubiquitin in different protein samples.
C-Terminal region of RIF1 is SUMOylated
Biochemical analysis of BLM protein complex revealed Rif1 as an important component. BLM binding of Rif1 is absolutely dependent on its conserved C-terminal domain and is independent of the N-terminal Heat repeat region of Rif137. A domain structure of RIF1 is shown in Fig. 8A. To determine the potential sites of RIF1 SUMOylation, we obtained different deletion fragments of RIF125 and established stable cell lines of GFP-fused RIF1 fragments expressing, aa 406–2446 (CFB129), aa 1355–2446 (CFB130), aa 1701–2446 (CFB 131) and aa 1924–2446 (CFB 91) in 10-His-SUMO2 U2OS background. His purified SUMO2 conjugates were immunoblotted either with antibodies raised against RIF1 or GFP. We demonstrated that CFB131 was the most efficiently SUMOylated fragment compared to other deletion fragments (Fig. 8B). Bioinformatic tool and online available database suggest that a majority of consensus SUMOylation sites are clustered in C-terminal region of the protein. Four potential SUMO acceptor Lysine sites (K1883, K1889, K2093 and K2097) are found in CFB131. In addition, a previous study using mass spectrometry analysis shown that another Lysine residue, K1880, is SUMOylated49. Therefore, we mutated all five potential SUMO acceptor lysine residues in the CFB131 fragment. Interestingly, we found that RIF1 SUMOylation was significantly reduced in Lysine deficient (5K0) mutant (Figs 8C & S8), strongly suggesting that one or more of these five lysine sites could be SUMOylated. Overall, our results indicate that PIAS4 mediated RIF1 SUMOylation play important role in the maintenance of Genomic stability (Fig. 9).
RIF1 C-terminal region is efficient for SUMOylation. (A) Schematic presentation of Linear structure of RIF1 protein. (B) U2OS cells stably co-expressing His10-SUMO2 and indicated GFP fused RIF1 fragments were either DMSO treated or treated with Bleocin. Cells were harvested and His purification was performed. His purified SUMO2 conjugates were resolved in 3–8% Tris-acetate gel and immunoblotted with anti-RIF1 (upper panel) and ant-GFP (lower panel) and SUMO2/3 antibody (lowest pane). (C) U2OS cells stably expressing His10-SUMO2 were transiently transfected with GFP fused RIF1 fragment (CFB131), Lysine deficient mutant RIF1 fragment (5K0) and an empty vector. After indicated treatment cells were harvested and His purification was performed with the Lysed samples. WCE and His purified SUMO2 conjugates were separated in 4–12% Bis-Tris gel and immunoblotted with RIF1 antibody.
MODEL: SUMO E3 Ligase PIAS4 maintains genomic stability. SUMO E3 ligases PIAS4 and PIAS1 are central to DDR signaling. In normal conditions, DNA damage induced 53BP1 and RIF1 foci colocalize with y-H2AX. SUMOylated 53BP1 and RIF1 facilitate efficient recruitment, break repair and removal of RIF1 from the sites of DNA breaks. Timely resolution of RIF1 coupled to the cessation of DDR signals and y-H2AX foci allows cells to resume normal cell cycle progression, thus maintaining overall genomic stability. In the absence of PIAS4, RIF1 SUMOylation is impaired, resulting in decreased RIF1 ubiquitination and it's dissociation from the sites of DNA dmage. Unresolved RIF1 foci accumulate even as cells enter S phase. Consequently, enhanced UFBs are also detected in mitotic cells depleted of PIAS4 that potentially leads to the formation of multiple RIF1 nuclear bodies in the subsequent G1 cell cycle phase. Taken together, our findings support a model in which PIAS4 is needed to promote genomic integrity by promoting the timely clearance of RIF1 from sites of repaired DNA breaks.
Discussion
Coordinated protein SUMOylation and ubiquitination are key PTMs required for the recruitment and repair of DSBs in a timely manner11,50,51. SUMO E3 Ligases PIAS1 and PIAS4 have been shown to play important roles in the SUMO modification of target proteins12,15. We identified PIAS4 as a critical regulator of RIF1 SUMOylation. Earlier studies have shown that the DNA damage response mediator proteins BRCA1, 53BP1 and MDC1 are SUMOylated in response to genotoxic stress12,18,20. In this study we have identified RIF1 as a SUMOylated protein. RIF1 SUMOylation is increased in G1 cells and enriched in response to different DNA damaging agents, suggesting a broader role of RIF1 SUMOylation to counteract genotoxic stress. In addition to RIF1, we also demonstrated that 53BP1 protein is SUMO2 modified. Cell cycle synchronization study further highlights that both RIF1 and 53BP1 are strongly SUMOylated in G1 and extent of SUMOylation was increased in response to bleocin treatment. Our data suggesting that DNA damage induced RIF1 foci strongly colocalize with SUMO2 and to some extent with SUMO3, but not with SUMO1. Together our data strongly suggest that RIF1 is predominantly modified by SUMO2/3.
Earlier findings suggest that PIAS4 regulates both SUMO1 and SUMO2/3 accrual in laser tracks12. Here we identified an additional DDR protein RIF1, as a PIAS4 target for SUMOylation. Consistent with earlier findings12, we noticed only a partial loss of 53BP1 SUMOylation after PIAS1 knockdown and a substantial loss was observed after PIAS4 knockdown, suggesting that PIAS4 is the main SUMO E3 ligase required for the 53BP1 SUMOylation. Although it is a possibility that PIAS4 may regulate the function of RIF1 via 53BP1, given that many reports have demonstrated that RIF1’s function in NHEJ is regulated by 53BP126,27,28,29,30,31,32,52, our data clearly show RIF1 SUMOylation is not affected by 53BP1 depletion. This strongly suggests that PIAS4 promote the SUMOylation of 53BP1 and RIF1 independently, and argues for a direct role of PIAS4-dependent SUMOylation of RIF1. Coimmunoprecipitation experiment suggest a stronger binding of RIF1-PIAS4 in G1 cells compared to G2 cells, and supports our finding that RIF1 is mainly SUMOylated by PIAS4 in G1. In addition, we also noticed a DNA damage responsive interaction of RIF1 and PIAS4 in S phase cells, indicating other potential role of PIAS4 in regulating RIF1 in replicating cells, a subject of future investigation.
In addition to known consequences of PIAS4 depletion, our results suggest new roles in the regulation of DNA damage induced RIF1 foci resolution. RIF1 is recruited at the sites of DNA damage and colocalize with different DDR signaling proteins including γ-H2AX. In accordance with the role of PIAS4 in DDR signaling, we observed a clear delay in the accumulation of DNA damage induced RIF1 and γ-H2AX foci in PIAS4-depleted cells. Importantly, we found that RIF1 foci was persistent even at 16–24 hours of DNA damage recovery when PIAS4 was depleted. Therefore, we speculate that the persistence of RIF1 at sites of DNA damage in the absence of PIAS4, indicate that DNA damage induced PIAS4 dependent RIF1 SUMOylation is essentially required for the complete removal of RIF1 from the sites of DNA breaks post repair.
The peculiar nature of the unresolved RIF1 foci prompted us to analyze these cells at longer time points, followed by recovery from acute DNA damage. We observed a large number of S phase cells harboring RIF1 foci. The distinct RPA70 foci colocalization with RIF1 strongly suggesting that SUMO E3 ligase activity of the PIAS4 is required for the removal of RIF1 from the site of DNA breaks. This raises important questions as to how cells deal with unresolved RIF1 during replication and if these unresolved foci may pose impediments to DNA replication which can risk the genomic stability. In line with this, we demonstrated that PIAS4 activity is needed to prevent the occurrence of UFBs, which are known to result from replication problems and are associated with the loci that are either under-replicated and/or associated with abnormal DNA structures. Lack of PIAS4 increases the incidence of UFBs (>3 fold). We speculate that lack of RIF1 removal from chromatin may impede normal replication and result in abnormal replication of genomic regions that are visualized as UFBs in mitosis. In PIAS4 depleted cells, we also detected an increase in RIF1/53BP1 nuclear bodies. We described here for the first time that RIF1 is present in 53BP1-positive nuclear bodies, which are thought to be chromatin lesions that are transmitted from mitosis to G145,46. Accordingly, aberrant UFB dissolution in mitosis may cause the accumulation of nuclear bodies in the subsequent G1 phase. Our data suggest that the increase in nuclear bodies that comprise of RIF1 and 53BP1 in PIAS4 depleted cells may result from the increase in UFBs and, most likely replication problems. Interestingly, recently it has been shown that RIF1 plays a 53BP1 independent role in the maintenance of genomic stability through its recruitment at UFBs and its resolution44. Currently we cannot exclude the possibility that PIAS4 mediated RIF1 function may be directly required for the resolution of UFBs. Therefore, we propose that PIAS4 activity is required to suppress the formation of 53BP1 and RIF1 nuclear bodies.
Recently UHRF1 has been identified as downstream target of BRCA1. In S phase cells UHRF1 ubiquitinates RIF1 and promotes its dissociation from 53BP1, which in turn promotes HR48. Our study suggests that PIAS4 mediated SUMO2/3 modification of RIF1 is a primary event, required for efficient break repair and removal of RIF1 from the sites of DNA damage by promoting the ubiquitination of RIF1. Further studies are required to elucidate the mechanism of how SUMOylation of RIF1 may directly influence the ubiquitination of RIF1. In addition, further studies are required to investigate the role of PIAS4 in the regulation of UHRF1 activity.
Our data suggest that one or more of the five lysine residues located in the C-terminal region of the protein, required for RIF1 SUMOylation. Indeed, one of the site (K1880) has already been reported to be SUMOylated in vivo49. Lysine1880 is located in the motif ETKEEKPEETP, contains plenty of glutamic acids in the region and two prolines, indicating an unstructured region in the protein most likely located at the surface as a result of the charged side chains. Overall, our study proposes a model in which PIAS4 dependent regulation of RIF1 SUMOylation is required for efficient DDR signaling and resolution of RIF1 from sites of DNA damage in a timely manner. PIAS4 deficiency therefore results in genomic instability, as evidenced by the rise of DNA DSBs, accumulation of UFBs in mitotic cells and RIF1 nuclear bodies in G1 cells (Fig. 9).
Method
Plasmid DNA
YFP-SUMO1, RFP-SUMO2 and RFP-SUMO3 plasmid DNA constructs are a kind gift from Prof. Ron Hay Lab. FLAG-PIAS4 (add gene #15208).
Lentiviral transduction
The source of shRNA clones are listed in Table 4. One million cells were seeded in a 15-cm dish and the next day, cells were either infected with shRNA viruses directed against PIAS1 and PIAS4 or control non-targeting shRNA viruses at MOI 2. After changing media on the third day, cells were incubated for another 3 to 4 days before the experiment.
siRNA transfection
The siRNA duplexes were purchased from Dharmacon (listed in Table 2). 1.8 million cells were seeded in a 15 cm dish and reverse transfection was performed according to manufacturer’s instructions. 18 hours after the transfection, fresh growth medium was added. 72 hrs after the transfection, the indicated drug treatments were performed and cells were harvested.
Cell culture and cell cycle analysis
U2OS cells and U2OS cells stably expressing His SUMO2 were grown in DMEM high glucose medium added with 10% FBS and 100 U/ml penicillin plus 100 µg/ml streptomycin (Life Technologies) at 37 °C with 5% CO2. To arrest cells at the G1/S cell cycle stage, cells were treated with 2 mM thymidine for 19 hrs and then released for 9 hours, followed by a second thymidine block for 18 hrs. To release G1-arrested cells, they were washed with PBS and prewarmed cell culture medium. Cells were collected after 4 hours and 8 hours of G1 release to obtain S-phase or S/G2 enriched population. After washing with PBS, cells were transferred into 70% ethanol and incubated for 30 minutes. Subsequently cells were incubated with Ribonuclease A and stained with propidium iodide18 for 20 minutes and analyzed by flow cytometry.
Microscopy
Cells for immunofluorescence microscopy were cultured on square glass slides (22 × 22 mm) in 6 well plates. After treatment with MG132 (10 µM) for 5 hrs. and/or Bleocin (5 µg/ml) for 4 hours, U20S cell were washed 3 times in PBST (phosphate buffer saline containing 0.1% tween-20 detergent). Cells were incubated with ice- cold 2% formaldehyde (w/v) for 20 minutes and washed 3 times with PBST. Fixed cells were blocked with 5% BSA in PBST for overnight at 4-degree C. After blocking, cells were stained with appropriate antibodies (listed in Table 3) in PBST containing 5% BSA. Cells were washed 3 times in PBST and then co-immunostained with anti-rabbit or anti-mouse Alexa Fluor® 488 and anti-rabbit or anti-mouse Alexa Fluor® 594 for 30 minutes. Cell were washed 3 times with PBST and counter stained with DAPI. After mounting, imaging was performed with Zeiss AxioImager Z1 upright (EBL) Olympus microscope.
Purification of SUMO-2 and Ubiquitin conjugates
His-SUMO-2 and Ubiquitin conjugates were purified essentially, as described previously (Vyas et al., 2013). U2-OS cells and stably expressing 10 His SUMO2 or 10 His-Ub, were washed and collected in ice-cold PBS. Small aliquots of cells were lysed in 1% SDS, 1% NP-40, 100 mM Tris/Hcl (pH 7.5), 150 mM Nacl to determine the protein concentration. Guanidinium lysis buffer (6 M guanidinium-HCl, 0.1 M Na2HPO4/NaH2PO4, 0.01 M Tris/HCl, pH 8.0, and competing imidazole) was added to the cell pellet to lyse the cells, after which, the cells were sonicated to reduce the viscosity. His-SUMO-2 conjugates were enriched on nickel-nitrilotriacetic acid-agarose beads (Qiagen) after which the beads were washed using wash buffers A to D. Wash buffer A: 6 M guanidinium-HCl, 0.1 M Na2HPO4/NaH2PO4, 0.01 M Tris/HCl (pH 8.0), 10 mM β-mercaptoethanol, 0.3% Triton X-100. Wash buffer B: 8 M urea, 0.1 M Na2HPO4/NaH2PO4, 0.01 M Tris/HCl (pH 8.0), 10 mM b-mercaptoethanol, 0.3% Triton X-100. Wash buffer C: 8 M urea, 0.1 M Na2HPO4/NaH2PO4, 0.01 M Tris/HCl (pH 6.3), 10 mM β-mercaptoethanol, 0.3% Triton X-100. Wash buffer D: 8 M urea, 0.1 M Na2HPO4/NaH2PO4, 0.01 M Tris/HCl, pH 6.3, 10 mM β-mercaptoethanol, 0.1% Triton X-100. Samples were eluted in 7 M urea, 0.1 M NaH2PO4/Na2HPO4, 0.01 M Tris/HCl (pH 7.0), 500 mM imidazole.
RFP-TRAP Assay
Asynchronously growing U2OS cells were washed with PBS, trypsinized and resuspended in ice-cold PBS. 10 million cells were transferred into Protein LoBind Eppendorf tubes and spun down for 5′ at 1500 rpm. Cells were lysed in 0.3 ml EBC buffer (50 mM Tris pH 7.3, 150 mM NaCl, 0.5% NP-40, 1 mM MgCl2. 50 mM Idoacetamide) and sonicated. The Lysed protein samples were added with 500 U Benzonase (in 700 µl EBC buffer). Protein lysates were incubated for 60 min under rotation. NaCl concentration was made to 300 mM, by adding 30 µl 5 M NaCl to 1 ml lysate. spin down lysate for 10′ full speed in Eppendorf centrifuge. Equilibriate RFP-Beads (Chromotek RTA-20) in wash buffer. Transfer lysate to Eppendorf tube with 50 µl RFP-Trap A beads suspension. Incubate 1.5 hr under rotation. Wash 3x with EBC buffer without Iodoacetamides. Elute with 4X LDS buffer.
Immunoblotting
Whole cell extracts (WCE) were lysed in RIPA buffer supplemented with competing amount of protease inhibitor. WCE or purified protein samples were first separated on Novex 4–12% gradient gels (Life Technologies) using MOPS buffer or on Novex 3–8% gradient gels (Life Technologies) using Tris-Acetate buffer and then transferred onto 0.45 NC Nitrocellulose blotting membrane (GE Healthcare; 10600003) using a submarine system (Life Technologies). Membranes were stained with Ponceau S to visualize total protein and blocked with PBST containing 5% milk powder and 0.01% Tween-20 before incubating with primary antibodies as indicated.
Recovery Assay
U2OS cells were seeded at 50 k cell density on cover slips. Next day cells were either treated with Luc si RNA or PIAS4 si RNA. After 72 hours of siRNA transfection, cells were either mock treated or treated with 5 µM of bleocin. After one hour of bleocin treatment cells were washed 2 times with PBS, one time with pre-warmed media and started to recover in new media. Cells on coverslips were permeabilized and fixed at different time points. U2OS cells were grown in 6 well plates transfected either with Luc siRNA or PIAS4 siRNA. As described above, after 72 hours of transfection, recovery experiment was performed. Cells were lysed in RIPA buffer supplemented with competing concentration of protease inhibitor and immunoblotted with indicated antibodies.
Edu staining
To determine the no. of S-phase cells, U2OS cells were added with 10 µM of the Edu dye for 30 minutes. Premetallized and formaldehyde fixed cells were stained with RIF1 primary antibody and respective secondary antibodies according to Invitrogen- Click IT Edu Alexa Fluor 488 kit’s instructions.
Cell Imaging and data analysis
All microscopic imaging was done in at 63X oil immersion objective Lens in Olympus Zeiss AxioImager Z1 upright fluorescence microscope. The image processing and quantification was primarily done with the help of FIJI and ImageJ software.
Change history
22 March 2018
A correction to this article has been published and is linked from the HTML and PDF versions of this paper. The error has been fixed in the paper.
References
Zhou, B. B. & Elledge, S. J. The DNA damage response: putting checkpoints in perspective. Nature 408, 433–439, https://doi.org/10.1038/35044005 (2000).
Wyman, C. & Kanaar, R. DNA double-strand break repair: all’s well that ends well. Annu Rev Genet 40, 363–383, https://doi.org/10.1146/annurev.genet.40.110405.090451 (2006).
Mailand, N. et al. RNF8 ubiquitylates histones at DNA double-strand breaks and promotes assembly of repair proteins. Cell 131, 887–900, https://doi.org/10.1016/j.cell.2007.09.040 (2007).
Mattiroli, F. et al. RNF168 ubiquitinates K13-15 on H2A/H2AX to drive DNA damage signaling. Cell 150, 1182–1195, https://doi.org/10.1016/j.cell.2012.08.005 S0092-8674(12)01000-8 (2012).
Chapman, J. R., Sossick, A. J., Boulton, S. J. & Jackson, S. P. BRCA1-associated exclusion of 53BP1 from DNA damage sites underlies temporal control of DNA repair. J Cell Sci 125, 3529–3534, https://doi.org/10.1242/jcs.105353 (2012).
Lieber, M. R. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu Rev Biochem 79, 181–211, https://doi.org/10.1146/annurev.biochem.052308.093131 (2010).
Heyer, W. D., Ehmsen, K. T. & Liu, J. Regulation of homologous recombination in eukaryotes. Annu Rev Genet 44, 113–139, https://doi.org/10.1146/annurev-genet-051710-150955 (2010).
Kerscher, O., Felberbaum, R. & Hochstrasser, M. Modification of proteins by ubiquitin and ubiquitin-like proteins. Annu Rev Cell Dev Biol 22, 159–180, https://doi.org/10.1146/annurev.cellbio.22.010605.093503 (2006).
Dorval, V. & Fraser, P. E. Small ubiquitin-like modifier (SUMO) modification of natively unfolded proteins tau and alpha-synuclein. J Biol Chem 281, 9919–9924, https://doi.org/10.1074/jbc.M510127200 (2006).
Bergink, S. & Jentsch, S. Principles of ubiquitin and SUMO modifications in DNA repair. Nature 458, 461–467, https://doi.org/10.1038/nature07963 nature07963 (2009).
Jackson, S. P. & Durocher, D. Regulation of DNA damage responses by ubiquitin and SUMO. Mol Cell 49, 795–807, https://doi.org/10.1016/j.molcel.2013.01.017 S1097-2765(13)00050-6 (2013).
Galanty, Y. et al. Mammalian SUMO E3-ligases PIAS1 and PIAS4 promote responses to DNA double-strand breaks. Nature 462, 935–939, https://doi.org/10.1038/nature08657 (2009).
Nacerddine, K. et al. The SUMO pathway is essential for nuclear integrity and chromosome segregation in mice. Dev Cell 9, 769–779, https://doi.org/10.1016/j.devcel.2005.10.007 (2005).
Wang, L. et al. SUMO2 is essential while SUMO3 is dispensable for mouse embryonic development. EMBO reports 15, 878–885, https://doi.org/10.15252/embr.201438534 (2014).
Morris, J. R. et al. The SUMO modification pathway is involved in the BRCA1 response to genotoxic stress. Nature 462, 886–890, https://doi.org/10.1038/nature08593 (2009).
Azuma, Y., Arnaoutov, A., Anan, T. & Dasso, M. PIASy mediates SUMO-2 conjugation of Topoisomerase-II on mitotic chromosomes. EMBO J 24, 2172–2182, 7600700, https://doi.org/10.1038/sj.emboj.7600700 (2005).
Shim, H. S., Wei, M., Brandhorst, S. & Longo, V. D. Starvation promotes REV1 SUMOylation and p53-dependent sensitization of melanoma and breast cancer cells. Cancer Res 75, 1056–1067, https://doi.org/10.1158/0008-5472.CAN-14-22490008-5472.CAN-14-2249 (2015).
Vyas, R. et al. RNF4 is required for DNA double-strand break repair in vivo. Cell Death Differ 20, 490–502, https://doi.org/10.1038/cdd.2012.145 cdd2012145 (2013).
Bergink, S. et al. Role of Cdc48/p97 as a SUMO-targeted segregase curbing Rad51-Rad52 interaction. Nat Cell Biol 15, 526–532, https://doi.org/10.1038/ncb2729 (2013).
Luo, K., Zhang, H., Wang, L., Yuan, J. & Lou, Z. Sumoylation of MDC1 is important for proper DNA damage response. EMBO J 31, 3008–3019, https://doi.org/10.1038/emboj.2012.158 emboj2012158 (2012).
Kohler, J. B., Jorgensen, M. L., Beinoraite, G., Thorsen, M. & Thon, G. Concerted action of the ubiquitin-fusion degradation protein 1 (Ufd1) and Sumo-targeted ubiquitin ligases (STUbLs) in the DNA-damage response. PLoS One 8, e80442, https://doi.org/10.1371/journal.pone.0080442 (2013).
Nie, M. et al. Dual recruitment of Cdc48 (p97)-Ufd1-Npl4 ubiquitin-selective segregase by small ubiquitin-like modifier protein (SUMO) and ubiquitin in SUMO-targeted ubiquitin ligase-mediated genome stability functions. J Biol Chem 287, 29610–29619, https://doi.org/10.1074/jbc.M112.379768 (2012).
d’Adda di Fagagna, F. et al. A DNA damage checkpoint response in telomere-initiated senescence. Nature 426, 194–198, https://doi.org/10.1038/nature02118 (2003).
Silverman, J., Takai, H., Buonomo, S. B., Eisenhaber, F. & de Lange, T. Human Rif1, ortholog of a yeast telomeric protein, is regulated by ATM and 53BP1 and functions in the S-phase checkpoint. Genes Dev 18, 2108–2119, https://doi.org/10.1101/gad.1216004 18/17/2108 (2004).
Xu, L. & Blackburn, E. H. Human Rif1 protein binds aberrant telomeres and aligns along anaphase midzone microtubules. J Cell Biol 167, 819–830, https://doi.org/10.1083/jcb.200408181 (2004).
Chapman, J. R. et al. RIF1 is essential for 53BP1-dependent nonhomologous end joining and suppression of DNA double-strand break resection. Molecular cell 49, 858–871, https://doi.org/10.1016/j.molcel.2013.01.002 (2013).
Di Virgilio, M. et al. Rif1 prevents resection of DNA breaks and promotes immunoglobulin class switching. Science 339, 711–715, https://doi.org/10.1126/science.1230624 (2013).
Escribano-Diaz, C. et al. A cell cycle-dependent regulatory circuit composed of 53BP1-RIF1 and BRCA1-CtIP controls DNA repair pathway choice. Molecular cell 49, 872–883, https://doi.org/10.1016/j.molcel.2013.01.001 (2013).
Zimmermann, M., Lottersberger, F., Buonomo, S. B., Sfeir, A. & de Lange, T. 53BP1 regulates DSB repair using Rif1 to control 5′ end resection. Science 339, 700–704, https://doi.org/10.1126/science.1231573 (2013).
Feng, L., Fong, K. W., Wang, J., Wang, W. & Chen, J. RIF1 counteracts BRCA1-mediated end resection during DNA repair. The Journal of biological chemistry 288, 11135–11143, https://doi.org/10.1074/jbc.M113.457440 (2013).
Callen, E. et al. 53BP1 mediates productive and mutagenic DNA repair through distinct phosphoprotein interactions. Cell 153, 1266–1280, https://doi.org/10.1016/j.cell.2013.05.023 S0092-8674(13)00590-4 (2013).
Kumar, R. & Cheok, C. F. RIF1: a novel regulatory factor for DNA replication and DNA damage response signaling. DNA Repair (Amst) 15, 54–59, https://doi.org/10.1016/j.dnarep.2013.12.004 S1568-7864(13)00291-7 (2014).
Panier, S. & Boulton, S. J. Double-strand break repair: 53BP1 comes into focus. Nat Rev Mol Cell Biol 15, 7–18, https://doi.org/10.1038/nrm3719 nrm3719 (2014).
Manis, J. P. et al. 53BP1 links DNA damage-response pathways to immunoglobulin heavy chain class-switch recombination. Nat Immunol 5, 481–487, https://doi.org/10.1038/ni1067 ni1067 (2004).
Ward, I. M. et al. 53BP1 is required for class switch recombination. J Cell Biol 165, 459–464, https://doi.org/10.1083/jcb.200403021 jcb.200403021 (2004).
Foti, R. et al. Nuclear Architecture Organized by Rif1 Underpins the Replication-Timing Program. Mol Cell, https://doi.org/10.1016/j.molcel.2015.12.001 (2015).
Xu, D. et al. Rif1 provides a new DNA-binding interface for the Bloom syndrome complex to maintain normal replication. EMBO J 29, 3140–3155, https://doi.org/10.1038/emboj.2010.186 emboj2010186 (2010).
Ouyang, K. J. et al. SUMO modification regulates BLM and RAD51 interaction at damaged replication forks. PLoS Biol 7, e1000252, https://doi.org/10.1371/journal.pbio.1000252 (2009).
Ouyang, K. J., Yagle, M. K., Matunis, M. J. & Ellis, N. A. BLM SUMOylation regulates ssDNA accumulation at stalled replication forks. Front Genet 4, 167, https://doi.org/10.3389/fgene.2013.00167 (2013).
Gonzalez-Prieto, R., Cuijpers, S. A., Kumar, R., Hendriks, I. A. & Vertegaal, A. C. c-Myc is targeted to the proteasome for degradation in a SUMOylation-dependent manner, regulated by PIAS1, SENP7 and RNF4. Cell Cycle 14, 1859–1872, https://doi.org/10.1080/15384101.2015.1040965 (2015).
Schimmel, J. et al. Uncovering SUMOylation dynamics during cell-cycle progression reveals FoxM1 as a key mitotic SUMO target protein. Mol Cell 53, 1053–1066, https://doi.org/10.1016/j.molcel.2014.02.001 S1097-2765(14)00115-4 (2014).
Buonomo, S. B., Wu, Y., Ferguson, D. & de Lange, T. Mammalian Rif1 contributes to replication stress survival and homology-directed repair. J Cell Biol 187, 385–398, https://doi.org/10.1083/jcb.200902039 jcb.200902039 (2009).
Liu, Y., Nielsen, C. F., Yao, Q. & Hickson, I. D. The origins and processing of ultra fine anaphase DNA bridges. Curr Opin Genet Dev 26, 1–5, https://doi.org/10.1016/j.gde.2014.03.003 S0959-437X(14)00009-4 (2014).
Hengeveld, R. C. et al. Rif1 Is Required for Resolution of Ultrafine DNA Bridges in Anaphase to Ensure Genomic Stability. Dev Cell 34, 466–474, https://doi.org/10.1016/j.devcel.2015.06.014 S1534-5807(15)00421-9 (2015).
Lukas, C. et al. 53BP1 nuclear bodies form around DNA lesions generated by mitotic transmission of chromosomes under replication stress. Nature cell biology 13, 243–253, https://doi.org/10.1038/ncb2201 (2011).
Harrigan, J. A. et al. Replication stress induces 53BP1-containing OPT domains in G1 cells. J Cell Biol 193, 97–108, https://doi.org/10.1083/jcb.201011083 jcb.201011083 (2011).
Doil, C. et al. RNF168 binds and amplifies ubiquitin conjugates on damaged chromosomes to allow accumulation of repair proteins. Cell 136, 435–446, https://doi.org/10.1016/j.cell.2008.12.041 S0092-8674(09)00004-X (2009).
Zhang, H. et al. A cell cycle-dependent BRCA1-UHRF1 cascade regulates DNA double-strand break repair pathway choice. Nat Commun 7, 10201, https://doi.org/10.1038/ncomms10201 ncomms10201 (2016).
Hendriks, I. A. et al. Uncovering global SUMOylation signaling networks in a site-specific manner. Nature structural & molecular biology 21, 927–936, https://doi.org/10.1038/nsmb.2890 (2014).
Dantuma, N. P. & Pfeiffer, A. Real Estate in the DNA Damage Response: Ubiquitin and SUMO Ligases Home in on DNA Double-Strand Breaks. Front Genet 7, 58, https://doi.org/10.3389/fgene.2016.00058 (2016).
Dou, H., Huang, C., Van Nguyen, T., Lu, L. S. & Yeh, E. T. SUMOylation and de-SUMOylation in response to DNA damage. FEBS Lett 585, 2891–2896, https://doi.org/10.1016/j.febslet.2011.04.002 S0014-5793(11)00248-1 (2011).
Chapman, J. R., Taylor, M. R. & Boulton, S. J. Playing the end game: DNA double-strand break repair pathway choice. Mol Cell 47, 497–510, https://doi.org/10.1016/j.molcel.2012.07.029 S1097-2765(12)00656-9 (2012).
Acknowledgements
We thank Dr. Alfred Vertegaal (LUMC, Leiden, Netherlands) for cell lines and reagents. We thank Prof. Ron Hay (University of Dundee, UK) for SUMO constructs. We thank Prof. Noel F. Lowndes (NUI Galway, Ireland) for critical reading of this manuscript and valuable suggestions to improve the manuscript. We thank to Gopinath Meenakshi Sundaram (IMB, A-STAR Institute) for critical reading of the manuscript. We thank Obed Akwasi Aning for his help in image quantification. This work is supported by IFOM Principle Investigator award to CF and IFOM postdoctoral fellowship to RK. We sincerely apologize to all authors whose work is not cited here due to space constraints.
Author information
Authors and Affiliations
Contributions
R. Kumar and C.F. conceived the project. R. Kumar and C.F. designed and performed experiments. R. Kumar and C.F. analyzed data. R. Kumar wrote the manuscript. R. Kumar and C.F. worked on the final draft.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
A correction to this article is available online at https://doi.org/10.1038/s41598-018-23385-4.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kumar, R., Cheok, C.F. Dynamics of RIF1 SUMOylation is regulated by PIAS4 in the maintenance of Genomic Stability. Sci Rep 7, 17367 (2017). https://doi.org/10.1038/s41598-017-16934-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-16934-w
- Springer Nature Limited
This article is cited by
-
Rif1 interacts with non-canonical polycomb repressive complex PRC1.6 to regulate mouse embryonic stem cells fate potential
Cell Regeneration (2022)
-
Identification of protein inhibitor of activated STAT 4, a novel host interacting partner that involved in bovine viral diarrhea virus growth
Virology Journal (2020)
-
Rif1 S-acylation mediates DNA double-strand break repair at the inner nuclear membrane
Nature Communications (2019)