Abstract
It is particularly important to provide precise therapies and understand tumor heterogeneity based on the molecular typing of mutational landscape. However, the landscape of somatic mutations in different subtypes of advanced breast cancer (ABC) is largely unknown. We applied target-region capture deep sequencing to determine the frequency and spectrum of common cancer-related gene mutations in circulating tumor DNA (ctDNA) among different ABC subtypes and analyze their association with clinical features. In this retrospective study of 100 female advanced breast cancer patients, 96 (96.0%) had somatic genomic alterations in ctDNA, including copy number variants and point mutations. The results revealed that different subtypes of ABC have distinct features in terms of genetic alterations. Multivariate regression analyses revealed that the number of somatic mutations increased with the line of endocrine therapy and the fractions of trunk mutations was positive associated with the line of target therapy.
Similar content being viewed by others
Introduction
Breast cancer, one of the most common cancers worldwide, is a heterogeneous disease with a variety of outcomes and drug responses. The St. Gallen subtype classification, introduced in 2011, categorizes breast cancer into five basic therapeutic groups based on immunohistochemical staining. These subtypes include luminal A, luminal B HER2-neu negative, luminal B HER2-neu positive, HER2-neu non-luminal and basal-like. In most cases, the St. Gallen intrinsic subtype classification for breast cancer can effectively predict disease features, recurrence patterns and disease-free survival1. However, the outcomes and drug responses of patients with the same subtype are diverse. With the rapid development of genome sequencing technology, molecular characterization based on genomic alteration is widely accepted as a relevant source of cancer stratification. Therefore, it is particularly important to provide precise therapies and understand tumor heterogeneity based on the molecular typing of genomic alteration.
Most molecular studies of breast cancer have focused only on revealing the characteristics of primary cancer based on tissue sequencing2,3,4. As a result of therapeutic selective pressure and tumor evolution, the mutation landscape of advanced breast cancer (ABC) may shift and vary. We need to acquire tissue from the primary tumor and all of the metastatic tumors to understand the mutation characteristics of ABC. However, tissue biopsies are limited by the presence of spatial heterogeneity, which leads to tumor sampling bias5, 6. Most of the time, it is difficult to obtain metastatic tissue in clinical work. Circulating tumor DNA (ctDNA) has increasingly attracted attention for its convenient, easily accepted, and minimally invasive method of collection7. ctDNA analysis facilitates studies of tumor heterogeneity, for it is able to detect contributions from multiple tumor deposits. A recent study illustrated that ctDNA could reveal tumor heterogeneity in non-small-cell lung cancer8.
Here, we applied target-region-capture deep sequencing to detect somatic mutations in plasma ctDNA from ABC patients to understand the mutational characteristics of ABC and analyze the association of clinical features and therapeutic history with gene variations.
Results
Samples and clinical data
The main characteristics of these patients are shown in Table 1. A total of 100 female ABC patients were enrolled in the present study. The mean age at breast cancer diagnosis was 45.3 years and all of them are Chinese. One hundred plasma samples collected from these patients were assayed for somatic genomic alterations by target-capture NGS. A panel of 1021 genes was assayed in the present study (Supplementary Table S1).
Identification of somatic genomic alterations
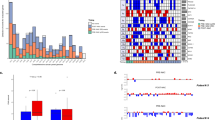
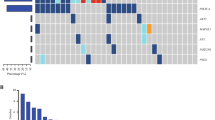
Somatic genomic alterations in ctDNA, including CNVs and point mutations, were identified in 96 of 100 patients (96.0%). (Figure 1)
The landscape of hotspot mutations in advanced breast cancer. Each of the 25 hotspot gene mutations has been identified in more than 5 patients listed to the left of the figure. The number of mutations in each gene among the 100 patients is shown (rows). Point mutations, copy number changes and both are colored red, blue and purple, respectively.
Target-capture sequencing of plasma ctDNA and matched WBC DNA was performed to detect somatic mutations in each sample, achieving a mean sequencing coverage of 1130×(317~3015×) for plasma ctDNA and 317×(69~541×) for WBC DNA. The sequencing and data analysis pipline used in this study were validated and the results showed that our test had 96.30% sensitivity at mutant allele frequency (MAF)≥0.5% with high specificity (99.9997%) and accuracy (99.9996%) for SNV detection. For CNV detection, the approach had 95.83% sensitivity for copy numbers at 1.25×(25.6% extra copies) with high specificity (99.77%) and accuracy (99.76%)9. In additional, the final candidate somatic variations were all manually verified in IGV browser.
The number of somatic mutations varied markedly between individual patients (mean 2.9, range 1–31). No difference in the number of somatic mutations was found between the four examined subtypes (p > 0.05). We examined the relationship between the number of somatic mutations and the age at diagnosis in the 100 patients. Across the entire sample and within each of the four subtype groups, no correlation was observed between the number of somatic mutations and the patient’s age. However, the mean number of somatic mutations was higher in patients aged 40–50 years than in patients aged more than 60 years (8.46 vs 3.88, respectively; p = 0.039). The results of multivariate analyses showed that the number of somatic mutations increased with the line of endocrine therapy (p = 0.007, Table 2). However, there were no differences between the number of mutations and the line of chemotherapy or target therapy (p > 0.05) (Fig. 2).
The correlation between the number of somatic mutations and clinical characteristics. (A).The relationship between the number of somatic mutations and the age at diagnosis in all 100 patients. (B).The correlation between the number of somatic mutations and the line of endocrine therapy in all 100 patients. Dots correspond to number of somatic mutations and whiskers correspond to its Standard Error. Red line correspond to the mean number of somatic mutations.
The fractions of trunk mutations were ranged from 0.3–80%. Multivariate regression analyses revealed that the fractions of trunk mutations was positive associated with the line of target therapy (p = 0.035, Table 3).
CNVs
We identified CNVs in 13 out of 100 (13.0%) patients by analyzing the sequencing data for plasma matched with blood cells from the same patient (Supplementary Table S2). Thirteen genes had copy number gains. Amplification of the ERBB2 gene, which encodes the HER2 protein, was predominant and was identified in 6 of 100 (6.0%) patients, all of whom belonged to the HER2+ group. Amplification of the FGFR1 gene was also identified in 5 of 100 (5.0%) patients, all of whom belonged to the HR+ group. Elevated levels of CDK12 were present in 4 of 100 patients (4.0%), all of whom were characterized by ERBB2 and CDK12 co-amplification. Moreover, amplification of the AURKA, IGFR1 and RPS6KB1 genes was captured in 2 of 100 patients (2.0%). In addition, ctDNA sequencing identified 7 other, less common CNVs in the study population; each of these CNVs was only detected in 1 patient (1.0%).
Point mutations
Point mutations in breast cancer-related genes were present in 96 of 100 patients (96.0%, Supplementary Table S3). TP53 and PIK3CA were the two most frequently mutated genes detected in the ctDNA of the 100 patients; these genes appeared in 43 (43.0%) and 32 (32.0%) patients, respectively. TP53 was the most frequently mutated gene, appearing in 43 (43.0%) patients. Thirty-nine kinds of mutations were detected, and 3 of the patients had R273H mutations. PIK3CA gene mutations were detected in 32 (32.0%) patients and included 17 point mutations and 1 CNV: H1047R mutation was the most frequently detected of these, appearing in 14 patients, while 4 patients had N345K mutations, 4 E542K, and 3 E545K. Twelve patients had ESR1 mutations, including 7 patients with Y537S mutations, 7 with D538G mutations and 4 with other kinds of point mutations. ERBB2 mutations were detected in 11 (11%) patients, all of whom belonged to the HER2-positive group.
Genomic characterization of the molecular subtypes
We analyzed the CNVs and point mutations of each patient in our cohort to delineate the biological characteristics of each molecular subtype of ABC.
There were 28 patients (28.0%) in the HR-positive HER2-negative group. We detected 8 gene CNVs in 7 of these 28 patients (25.0%). The frequency of mutations, including CNVs and point mutations, of eight genes was more than 10% in all 28 patients. The HR-positive HER2-positive group contained 37 patients (37.0%). We detected 6 genes whose mutation frequency was more than 10%, and 4 gene CNVs were detected in 3 patients (8.1%) from this group. Ten genes were had variations of more than 10% in the HR-negative HER2-positive group, which contained 31 patients (31.0%). Three patients (9.7%) out of 31 had CNVs. The triple-negative breast cancer (TNBC) group only contained four patients (4.0%), and 22 gene mutations were detected. ESR1 mutations were detected in 12 of the HR+ patients, who had a significantly higher mutation frequency than did the HR− patients (18.46% vs 0.00%, respectively; p = 0.007). NOTCH1 mutations were detected in 5 of the HER2- patients, which was more than the number of HER2+ patients that had such mutations (15.63% vs 2.94%, respectively; p = 0.033). Mutations in three genes, namely, NOTCH3, ESR1 and FGFR1, were detected in only the HR-positive group. PIK3CA mutations occurred more frequently in HR+ patients than in HR− patients (40.00% vs 17.14%, respectively; p = 0.025) (Fig. 3).
The frequency of common genes in four subtypes among 100 patients. (A) The frequency of common genes in different HER2 statues. Four gene mutations including TP53/PIK3CA/ERBB2/ESR1 detected more than 10 patients and NOTCH1 which differential detected between different HER2 statues patients listed to the bottom the figure. HER2 positive and negative patients are colored yellow and blue respectively. (B) The frequency of common genes in different HR statues patients. Four gene mutations including TP53/PIK3CA/ERBB2/ESR1 detected more than 10 patients listed to the bottom of the figure. HR positive and negative patients are colored yellow and blue respectively. Whiskers correspond to the percentage Standard Error of the gene frequency.
The associations between point mutations and clinical characteristics
Multiple logistic regression analysis indicated that pathological grade, tumor size at diagnosis and PR status were positively associated with PIK3CA mutations. However, menstruation status was inversely associated with PIK3CA mutations. Multiple regression analysis also revealed that ki-67, the presence of metastasis at diagnosis, the number of metastatic sites, and the line of endocrine therapy were associated with ESR1 mutations. HER2 status, number of metastatic sites and tumor size at diagnosis were negatively associated with NOTCH1 mutations in multiple regression analysis. (Table 4).
Discussion
Currently, breast cancer is divided into different molecular subtypes based on the St. Gallen subtype classification, which can effectively predict disease features and prognosis1. However, the outcomes and drug responses of patients with the same subtype are also diverse. Several studies revealed that one tumor often consists of multiple cell subpopulations and this heterogeneity has been proposed as one of the major reasons for the failure of drug treatment10, 11. Gene variations based subtyping has the potential to improve or even replace the current classification system12, 13. Several breast cancer studies have revealed mutations characteristic of primary cancer based on tissue sequencing2,3,4. Because of the spatial and temporal heterogeneity of breast cancer, the genetic alterations of ABC are different from those of early breast cancer. This study applied the next-generation sequencing of ctDNA to understand the mutant characterization of different subtypes of ABC and analyze the association between clinical features and gene somatic variation profiling. The results revealed that different subtypes of ABC have distinct features with respect to genetic alterations.
The somatic mutations in a cancer genome accumulate over a patient’s lifetime, and the number of somatic mutations may increase with the age2. A previous study found no correlation between the number of somatic mutations and age both in ER+ and ER− breast cancer patients2. Our data also support this conclusion, as there was no relationship between the number of somatic mutations and age in any of the four examined subtypes. However, the number of somatic mutations was higher in patients aged 40–50 years than in patients aged more than 60 years. The basis for this pronounced difference is unclear and may be correlated with therapy because multivariate analyses showed that the number of somatic mutations increased with the line of endocrine therapy. Recent evidences suggest that a high degree of cancer cell heterogeneity could lead to drug resistance10. The number of somatic mutations increased with the line of endocrine therapy maybe a result of endocrine therapy resistance. But we did not found the same phenomenon for target therapy and chemotherapy and further research is needed on this subject. We also found that the fraction of trunk mutations increased with the line of target therapy. This maybe the result of target therapy selection or maybe a result of drug resistance.
In general, TP53 and PIK3CA are the most frequently mutated genes in ABC, which is consistent with early breast cancer. TP53, the most frequently mutated gene in breast cancer, is more frequently mutated in HER2-positive breast cancer and TNBC14, 15. We did not find any significant difference in TP53 mutation frequency among the subtypes of ABC. No relationship was observed between clinical characteristics and TP53 mutations. PIK3CA is the second most frequently mutated gene, following TP53, currently known in breast cancer patients, occurring at a frequency of 20–45% in early breast cancer16,17,18,19,20. PIK3CA mutations are more prevalent in the HR+/HER2+ subtype of breast cancer19, 20. The results of the present study suggest that mutations are significantly more frequent in HR+ cancers than in HR− cancers19,20,21. The association of PIK3CA mutations with HR and HER2 status remains controversial. Some studies have supported a positive association between PIK3CA mutations and hormonal and HER2 status, while others have suggested no significant correlation21,22,23,24,25,26,27. Our data suggest that PIK3CA mutations are associated with progesterone receptor status, but there was no association with estrogen receptor or HER2 status. Multivariate analysis indicated that pathological grade and tumor size at diagnosis were positively associated with PIK3CA mutations, which indicates that cancers with PIK3CA mutations may be more invasive.
Mutations in the ER gene (ESR1) have been described in ABCs that had been exposed to previous therapy with aromatase inhibitors (AIs)28, 29. ESR1 mutations are only rarely detectable in primary breast cancer and are only found at an appreciable frequency after the development of hormone resistance30, 31. The frequency of ESR1 mutations ranges from 20% to 89% in HR-positive patients previously exposed to AIs28,29,30. Our study found that the mutation frequency of ESR1 was significantly higher in HR+ patients than in HR− patients, especially in the HR+/HER2− group. ESR1 mutations were positively related to ki-67 index, the presence or absence of metastasis at first diagnosis, the number of metastatic sites, and the line of endocrine therapy. These results support the previous view that ESR1 mutations are acquired from endocrine therapy28.
The NOTCH1 signaling pathway is associated with cell proliferation, cell differentiation, cell motility, cell-cell connections and cell polarity32. NOTCH1 was recently found to be related to cancer cell metastasis and the maintenance of cancer stem cells33. A recent meta-analysis found that the expression of NOTCH1 was enriched in the triple-negative subtype of breast cancer34. Previous studies indicated that NOTCH1 was inversely correlated with HR status, but there was no significant relationship between NOTCH1 and HER2 status34. Our study detected NOTCH1 gene mutations more frequently in the HR+/HER2− group, in contrast to previous research. Multivariate analysis in the present study found that NOTCH1 mutations were significantly negatively correlated with HER2 status. We also found that NOTCH1 mutations were negatively related to the number of metastatic sites. The mechanism is unclear and the result need further study and verify for our limited sample size in NOTCH1 mutant group.
ERBB2 point mutations and CNVs were only detected in HER2+ cancers, and the mutant frequency in the HER2+ group was 8.8%. Theoretically, all HER2+ patients should show an ERBB2 copy number gain in ctDNA. However, only 8.8% patients in the HER2+ group showed an ERBB2 copy number gain. Although the variation in the timing of primary tissue and metastatic plasma sampling may partly account for the discordance, possible methodological concerns were still explored. The ctDNA assay may not be sufficiently sensitive to accurately detect CNVs.
Despite the advantages delineated above, several limitations of the study should be noted. To begin, HER2 amplification was identified by ctDNA assay in only 6 of 68 (8.8%) HER2+ patients. Further studies should be conducted to improve the sensitivity of the ctDNA assay to detect CNVs. Second, the depth of coverage was low for most of the ctDNA samples; this limitation could be addressed in future studies by improving the sequencing depth. Third, the present study did not have sufficient power to arrive at statistically sound conclusions for the small number of patients with each subtype, especially for the TNBC group.
In conclusion, our data indicate that different subtypes of ABC have distinct features in terms of genetic alterations. Certain gene mutations may be related to clinical treatment, especially endocrine therapy.
Materials and Methods
Patients and sample collection
The present study is a retrospective study. All blood samples were obtained from female ABC patients who underwent therapy at Cancer Hospital, Chinese Academy of Medical Sciences, from March 2015 to September 2016. The study group comprised 100 patients with invasive ductal carcinoma, ranging from 22 to 72 years of age. Patients were eligible for enrollment if they were over 18 years of age and had a histologic/cytologic diagnosis of metastatic breast cancer. Patients were excluded if they had received chemotherapy within the past month. The study was reviewed and approved by the ethical committee at National Cancer Center/Cancer hospital, Chinese Academy of Medical Sciences and Peking Union Medical College. All methods were performed in accordance with the relevant guidelines and regulations, and informed consent was obtained from the patients and parents.
DNA extraction
Circulating DNA was isolated from 0.6–1.8 mL of plasma using the QIAamp Circulating Nucleic Acid Kit (Qiagen) and extracted using the DNeasy Blood and Tissue Kit. The DNA concentration was assessed using a Qubit fluorometer (Invitrogen, Carlsbad, CA USA) and the Qubit dsDNA HS (High Sensitivity) Assay Kit. The size distribution of the cell-free DNA (cfDNA) was assessed using an Agilent 2100 Bioanalyzer and the DNA HS kit (Agilent Technologies, Santa Clara, CA, USA).
Target capture and next-generation sequencing
Sequencing libraries were prepared for cfDNA using the KAPA DNA Library Preparation Kit (Kapa Biosystems, Wilmington, MA, USA), and gDNA sequencing libraries were prepared using the protocols recommended in the Illumina TruSeq DNA Library Preparation Kit (Illumina, San Diego, CA). For samples at or near the minimum input requirement, additional pre-capture PCR cycles were performed to generate sufficient PCR product for hybridization. Libraries were hybridized to custom-designed biotinylated oligonucleotide probes (Roche NimbleGen, Madison, WI, USA) covering ~1.1 Mbp of sequence. DNA sequencing was carried out with the HiSeq 3000 Sequencing System (Illumina, San Diego, CA) with 2 × 101-bp paired-end reads.
Sequencing data analysis
After removing the terminal adaptor sequences and low-quality data, the reads were mapped to the reference human genome. GATK (https://www.broadinstitute.org/gatk/, The Genome Analysis Toolkit) and MuTect were used to call small insertions and deletions (indels) and single nucleotide variants (SNVs) in the somatic DNA by filtering peripheral blood (PBL) sequencing data. In addition, we used the NoahCare Tool Kit using NCfilter (software developed by self, version 1.5.0) for fastq data QC, NCbamInfo (version 0.2.0) for alignment QC; NCanno (version 0.1.1) for annotation with multiple databases; and NChot (version 0.1.0) for hotspot region variant review and recall. Contra was used to detect copy number variants, and BreakDancer was used to detect cancer-associated structural variants. The final candidate variants were all manually verified using the Integrative Genomics Viewer (IGV) Browser.
Statistical analysis
The relationship between the molecular subtypes and gene variation profiles was studied by cross-tabulation with the chi-square test or Fisher’s exact test. Potential links between various parameters and the occurrence of a particular mutation were evaluated using logistic regression. Linear regression models were constructed examining the relationship between mutant allele fractions of trunk mutation or the number of total mutations and clinical characteristics of ABC. Mutations, which had the highest predicted cellular prevalence, were classified as trunk mutations. The Mann-Whitney U was used to compare the mean number of somatic mutations between different age groups. All statistical tests were two-sided, and p < 0.05 was considered to indicate statistical significance. All statistical analyses were performed using SPSS version 19.0 (SPSS Company, Chicago, IL).
References
Vasconcelos, I. et al. The St. Gallen surrogate classification for breast cancer subtypes successfully predicts tumor presenting features, nodal involvement, recurrence patterns and disease free survival. Breast 29, 181–185 (2016).
Stephens, P. J. et al. The landscape of cancer genes and mutational processes in breast cancer. Nature 486, 400–405 (2012).
Nik-Zainal, S. et al. Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature 534, 47–54 (2016).
Koboldt, D. C. et al. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Saunders, N. A. et al. Role of intratumoural heterogeneity in cancer drug resistance: molecular and clinical perspectives. EMBO Mol Med 4, 675–84 (2012).
Schmitt, M. W., Loeb, L. A. & Salk, J. J. The influence of subclonal resistance mutations on targeted cancer therapy. Nat Rev Clin Oncol 13, 335–47 (2016).
Crowley, E., Di Nicolantonio, F., Loupakis, F. & Bardelli, A. Liquid biopsy: monitoring cancer-genetics in the blood. Nat Rev Clin Oncol 10, 472–84 (2013).
Chabon, J. J. et al. Circulating tumour DNA profiling reveals heterogeneity of EGFR inhibitor resistance mechanisms in lung cancer patients. Nature Communications 7, 11815 (2016).
Yang, X. et al. Technical Validation of a Next-Generation Sequencing Assay for Detecting Clinically Relevant Levels of Breast Cancer-Related Single-Nucleotide Variants and Copy Number Variants Using Simulated Cell-Free DNA. J Mol Diagn. doi:10.1016/j.jmoldx.2017.04.007 (In Press).
Wang, E. et al. Cancer Systems Biology in the Genome Sequencing Era: Part 2, Evolutionary Dynamics of Tumor Clonal Networks and Drug Resistance. Semin Cancer Biol. 23, 286–292 (2013).
Wang, E. et al. Cancer Systems Biology in the Genome Sequencing Era: Part 1, Dissecting and Modeling of Tumor Clones and their Networks. Semin Cancer Biol. 23, 279–285 (2013).
Zaman, N. et al. Signaling Network Assessment of Mutations and Copy Number Variations Predict Breast Cancer Subtype-Specific Drug Targets. Cell Rep. 5, 216–223 (2013).
Zou, J. & Wang, E. ETumorType, an Algorithm of Discriminating Cancer Types for Circulating Tumor Cells or Cell-Free DNAs in Blood. Genomics, Proteomics & Bioinformatics. 15, 130–140 (2017).
Ma, C. X., Reinert, T., Chmielewska, I. & Ellis, M. J. Mechanisms of aromatase inhibitor resistance. Nature Reviews Cancer 15, 261–275 (2015).
Kim, J. et al. Association between Mutation and Expression of TP53 as a Potential Prognostic Marker of Triple-Negative Breast Cancer. Cancer Research and Treatment 48, 1338–1350 (2016).
Loibl, S. et al. PIK3CA mutations are associated with reduced pathological complete response rates in primary HER2-positive breast cancer: pooled analysis of 967 patients from five prospective trials investigating lapatinib and trastuzumab. Ann Oncol 27, 1519–25 (2016).
Arthur, L. M. et al. Changes in PIK3CA mutation status are not associated with recurrence, metastatic disease or progression in endocrine-treated breast cancer. Breast Cancer Research and Treatment 147, 211–219 (2014).
Oshiro, C. et al. PIK3CA mutations in serum DNA are predictive of recurrence in primary breast cancer patients. Breast Cancer Research and Treatment 150, 299–307 (2015).
Dirican, E., Akkiprik, M. & Özer, A. Mutation distributions and clinical correlations of PIK3CA gene mutations in breast cancer. Tumor Biology 37, 7033–7045 (2016).
Arsenic, R. et al. Analysis of PIK3CA mutations in breast cancer subtypes. Appl Immunohistochem Mol Morphol 22, 50–6 (2014).
Ahmad, F., Badwe, A., Verma, G., Bhatia, S. & Das, B.R. Molecular evaluation of PIK3CA gene mutation in breast cancer: determination of frequency, distribution pattern and its association with clinicopathological findings in Indian patients. Medical Oncology 33 (2016).
Kalinsky, K. et al. PIK3CA mutation associates with improved outcome in breast cancer. Clin Cancer Res 15, 5049–59 (2009).
Cizkova, M. et al. PIK3CA mutation impact on survival in breast cancer patients and in ERalpha, PR and ERBB2-based subgroups. Breast Cancer Res 14, R28 (2012).
Barbareschi, M. et al. Different prognostic roles of mutations in the helical and kinase domains of the PIK3CA gene in breast carcinomas. Clin Cancer Res 13, 6064–9 (2007).
Perez-Tenorio, G. et al. PIK3CA mutations and PTEN loss correlate with similar prognostic factors and are not mutually exclusive in breast cancer. Clin Cancer Res 13, 3577–84 (2007).
Ellis, M. J. et al. Phosphatidyl-inositol-3-kinase alpha catalytic subunit mutation and response to neoadjuvant endocrine therapy for estrogen receptor positive breast cancer. Breast Cancer Res Treat 119, 379–90 (2010).
Maruyama, N. et al. Clinicopathologic analysis of breast cancers with PIK3CA mutations in Japanese women. Clin Cancer Res 13, 408–14 (2007).
Schiavon, G. et al. Analysis of ESR1 mutation in circulating tumor DNA demonstrates evolution during therapy for metastatic breast cancer. Sci Transl Med 7, 313ra182 (2015).
Jeselsohn, R., Buchwalter, G., De Angelis, C., Brown, M. & Schiff, R. ESR1 mutations-a mechanism for acquired endocrine resistance in breast cancer. Nat Rev Clin Oncol 12, 573–83 (2015).
Niu, J. et al. Incidence and clinical significance of ESR1 mutations in heavily pretreated metastatic breast cancer patients. Onco Targets Ther 8, 3323–8 (2015).
Toy, W. et al. ESR1 ligand-binding domain mutations in hormone-resistant breast cancer. Nat Genet 45, 1439–45 (2013).
Andersson, E. R. & Lendahl, U. Therapeutic modulation of Notch signalling — are we there yet? Nature Reviews Drug Discovery 13, 357–378 (2014).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–74 (2011).
Yuan, X. et al. Expression of Notch1 Correlates with Breast Cancer Progression and Prognosis. PLOS ONE 10, e0131689 (2015).
Acknowledgements
The present study was supported by National Natural Science Foundation of China (81472453) and Major Project of Beijing Municipal Science and Technology Commission (D161100000816004).
Author information
Authors and Affiliations
Contributions
F.M. and Z.B.Y. conceived and designed the study. L.F.Y., X.Y.S., X.W.G., L.X.L., B.L.L. and B.H.X. Provided study materials or patients. Z.B.Y., C.X. L., R.R.C., Y.F.G., B.H.X and H.L.Q. performed the experiments. Z.B.Y. and F.M. analyzed the data and wrote the paper. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yi, Z., Ma, F., Li, C. et al. Landscape of somatic mutations in different subtypes of advanced breast cancer with circulating tumor DNA analysis. Sci Rep 7, 5995 (2017). https://doi.org/10.1038/s41598-017-06327-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-06327-4
- Springer Nature Limited
This article is cited by
-
Significant impact of circulating tumour DNA mutations on survival in metastatic breast cancer patients
Scientific Reports (2021)
-
Molecular landscape and efficacy of HER2-targeted therapy in patients with HER2-mutated metastatic breast cancer
npj Breast Cancer (2020)
-
Everolimus in hormone receptor-positive metastatic breast cancer: PIK3CA mutation H1047R was a potential efficacy biomarker in a retrospective study
BMC Cancer (2019)
-
Cell-free DNA analysis in healthy individuals by next-generation sequencing: a proof of concept and technical validation study
Cell Death & Disease (2019)