Abstract
Kaposi’s sarcoma-associated herpesvirus (KSHV) is the etiological agent of Kaposi’s sarcoma, primary effusion lymphoma and multicentric Castleman’s disease, etc. In this study, we firstly systematically constructed the KSHV-encoded miRNA-regulated co-expressed protein-protein interaction network (CePPIN), which display the biological knowledge regarding the mechanism of miRNA-regulated KSHV pathogenesis. Then, we investigated the topological parameters for the proteins in CePPIN, especially for those miRNA targets and we found that cellular target genes of KSHV-encoded miRNAs tend to be hubs and bottlenecks in the network. Then the GO and KEGG pathway analysis suggests that miRNA targets are involved in various cellular processes mostly related to immune regulate and cell cycle. Enrichment analysis was also performed to identify the six important functional modules which are proven to be highly related to KSHV pathogenesis. Finally, difference analysis of common targets and specific targets shows that two kinds of targets are different in terms of both topological properties and enriched functions, thus we can extrapolate that the functions of KSHV-encoded miRNAs can be also classified into two generic groups, one can act as functional mimics of some oncogenic human miRNAs which contribute to tumorigenesis and the other can contribute to maintaining viral survival.
Similar content being viewed by others
Introduction
microRNAs (miRNAs) are a set of small non-coding RNA molecules with a length of 21~24 nucleotides and function as major gene expression regulators at the posttranscriptional level in eukaryotic cells1. However, some viruses also express miRNAs in host cells and 5 virally-encoded miRNAs were firstly discovered by Pfeffer et al. in human B cells after infection with the herpesvirus Epstein–Barr virus (EBV)2. Since then, more than 200 mature miRNAs have been identified in all herpesviruses, except for Varicella Zoster virus3, 4. miRNAs are complementary to specific sequence motifs in the 3′ untranslated regions (UTRs) of their target mRNAs and negatively regulate gene expression at the post-transcriptional level by inhibiting translation or promoting mRNA degradation, based on the degree of complementary base pairing between the miRNA and mRNA. More and more evidence have revealed that miRNAs are involved in diverse physiological processes, such as development5, cell proliferation and differentiation6, 7, apoptosis8 and a variety of pathological conditions9, 10.
The human tumor virus Kaposi’s sarcoma-associated herpesvirus (KSHV), also called human herpesvirus 8 (HHV8), is the etiological agent of Kaposi’s sarcoma (KS), the most common malignance in AIDS patients, primary effusion lymphoma (PEL) and some types of multicentric Castleman’s disease (MCD)11,12,13. KSHV has been confirmed to encode at least 25 mature miRNAs14,15,16,17. Currently 16 experimentally confirmed KSHV miRNAs and their targets are available, but knowledge about their functions remains scarce since only a small number of mRNA targets have been experimentally validated. Study on the function of viral miRNA targets would contribute to further dissecting the role of viral miRNAs in viral-host interactions, and then might contribute to an elucidation of virus infection mechanism and potential identification of antiviral targets. Characterizing the direct targets of miRNAs has been widely employed to map miRNAs to biological pathways and diseases18,19,20. However, the regulatory effects of a miRNA are not only due to the direct RNA-induced silencing complex (RISC)-dependent targeting. Both direct and indirect targets are integral components of gene regulatory networks and should be together considered in functional analysis. In addition, proteins that physically associate with direct targets to function together in a complex may also be affected. Hsu et al.21 named the direct targets of miRNAs “L0 proteins” and the interacting partners of these targets “L1 proteins” in their protein-protein interaction network (PPIN). By comparing four topological parameters, namely degree, betweenness centrality, clustering coefficient and closeness centrality of L0 proteins and L1 proteins jointly with randomly selected proteins, it reveals that miRNA-regulated targets and their interacting partners jointly show significantly higher connectivity and modularity than randomly selected proteins, even than direct targets alone. In their following work22, they demonstrate that L0 proteins and L1 proteins together also show a stronger functional relationship than L0 proteins alone. They further named the network formed by L0 proteins and L1 proteins the co-expressed protein-protein interaction network (CePPIN). Recent studies have shown that integrating miRNA targets and PPIN makes it more efficient to dissect the regulation rules of miRNAs23.
In this study, to further understand the regulation rules of KSHV-encoded miRNAs, we constructed the miRNA-regulated CePPIN for 16 KSHV miRNAs and 109 experimentally validated human target genes. We firstly investigated the topological properties of the proteins in CePPIN, especially for those miRNA targets. Subsequently we analyzed the function of the proteins in CePPIN from multi-perspectives including the GO and KEGG pathway analysis and functional module extraction. Finally, to further explore difference between common targets and specific targets, we investigated the topological properties and functional annotation for genes targeted by both KSHV-encoded miRNAs and human miRNAs and genes only targeted by KSHV-encoded miRNAs, respectively. Consequently, these results provide new clues for elucidating the regulation rules of KSHV-encoded miRNAs in viral-host interactions.
Result and Discussion
Statistical analysis between KSHV-encoded miRNAs and their human targets
For 16 KSHV miRNAs and 109 human targets, we counted the number of human targets that each KSHV miRNA regulates. As shown in Fig. 1A, almost three quarters (80) of the 109 targets were regulated by miR-K12-11. miR-K12-11 is the ortholog of host hsa-miR-155 and these miRNAs have identical seed sequence. Has-miR-155 is a multifunctional miRNA that is important in immunity, hematopoiesis, inflammation, and oncogenesis. In vitro studies have shown that both miRNAs can regulate a common set of cellular target genes, suggesting that KSHV miR-K12-11 may mimic hsa-miR-155 function36, 37. Moreover, the KSHV-encoded miRNAs that regulate relatively more human targets, such as miR-K12-10a and miR-K12-3, are also the ortholog of host hsa-miR-142-3p and hsa-miR-2338, respectively. The results of sequence alignment between KSHV miRNAs and human miRNA36,37,38 also suggests that some KSHV miRNAs might act as functional mimics of some oncogenic human miRNAs which contribute to tumorigenesis.
On the other hand, we counted the number of targets regulated by different numbers of KSHV miRNAs. As can be seen from Fig. 1B, only 6 human targets are regulated by 3 or 4 kinds of KSHV miRNAs. They are BCLAF1 (4 miRNAs), MAF (4 miRNAs), THBS1 (4 miRNAs), WEE1 (3 miRNAs), BACH1 (3 miRNAs) and CDKN1A (3 miRNAs) respectively and they are all protein coding genes. BCLAF1 (Bcl-2-associated factor), a pro-apoptotic protein, was identified as a target of miR-K12-5, -9, -3 and -10b. The repression of BCLAF1 by KSHV miRNAs enabled the cells to overcome etoposide-induced caspase activation39. THBS1 (thrombospondin 1) was identified as a target of miR-K12-1, -3, -6-3p and -11 and it is a protein of the TGF-β pathway. The TGF-β pathway confers a strong anti-proliferative phenotype to many epithelial and endothelial cells and hence, it is presumable that KSHV has developed mechanisms to escape from this inhibition40. BACH1 (BTB and CNC homology 1), a transcription repressor involved in the regulation of genes regulating cell cycle and oxidative stress41. Down-regulation of the LEC-specific transcription repressor, MAF, by KSHV miRNAsmiR-K12-6 and miR-K12-11, was shown to induce the expression of several BEC-specific markers like CXCR4, thereby contributing to transcriptional reprogramming42. KSHV miR-K12-1 down-regulated the levels of the cyclin-dependent kinase inhibitor, CDKN1A (p21), thereby allowing the infected cells to overcome p21-mediated cell cycle arrest43. WEE1 is a key regulator of cell cycle progression. Down-regulated of WEE1 might decrease the efficiency of DNA safeguard mechanisms thus enhance the rate of spontaneous mutations44. These six targets are all important to KSHV infection. It is possible that only a relatively small subset of KSHV-miRNAs targets provides a strong selective advantage to KSHV, while many other miRNAs-targets interactions could be largely inconsequential to KSHV biology. Such particularly important targets should include those with multiple KSHV miRNA binding sites, for example CDKN1A (p21) or Wee1.
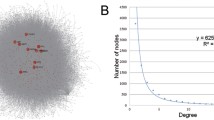
Construction of CePPIN for human targets of KSHV miRNAs
We used the 109 human targets as source to compile the human protein-protein interaction (PPI) information from InnateDB26. Overall, we obtained 3281 proteins (100 targets and 3181 co-expressed proteins) and 5730 PPIs as the original dataset of CePPIN. 9 targets were discarded for lacking the interactome information. The visualization of CePPIN is shown in Fig. 2A. A scale-free network is a network whose degree distribution follows a power law. As we can see from Fig. 2B, the degree distribution of CePPIN follows a power law. Small world networks are those that have a relative small mean path length but high clustering coefficient. Since the average shortest path length of all the nodes in CePPIN is about 3.5124 and the average clustering coefficient is 0.0823, so we can see that CePPIN presents the properties of small-world and scale-free network, which indicates that a small number of proteins called hubs directly regulate most of other members in the network. This network is more robust against random variation but more fragile against some aberration of the hubs compared with random network. In Fig. 2A, the bigger red nodes with labels are the top sixteen-ranked proteins by their degrees in CePPIN. Their degrees are all higher than 100 and they are all miRNA targets. For example, NFKB Inhibitor Alpha (NFKBIA) is down-regulated by kshv-mir-K12-1, leading to NFKB activation and maintenance of latency. The cyclin-dependent kinase inhibitor p21(CDKN1A) down-regulated by KSHV miR-K12-1, allowing the infected cells to overcome p21-mediated cell cycle arrest43. More information of the sixteen proteins is also shown in Table 1. It can also be seen explicitly that the overwhelming majority of miRNA targets have more interactions than other nodes and may play an important role in CePPIN.
Topological analysis for KSHV miRNA-regulated CePPIN
Four important topological parameters were computed for all nodes in CePPIN, namely degree, betweenness centrality, clustering coefficient and closeness centrality, respectively. Table 2 shows the topological parameters of all nodes and miRNA targets respectively in CePPIN. Since the range of the four topological parameters are very wide, the deviation values are very high. For example, among all proteins, the minimum degree is 1 and the maximum is 644, so the deviation is as high as 19.0992. In fact, the four topological parameters are all nonnegative number. We can see that the average degree and betweenness of miRNA targets are much higher than those of all nodes and the p-values are much lower than 0.05 by t-test. Degree and betweenness centrality are considered to be the best predictor of the essentiality of a node for network robustness, cooperation and communication and closeness centrality is a measure of centrality in a network. Thus, we can conclude that miRNA targets are essential in the CePPIN. At the same time, clustering coefficient of miRNA targets which quantifies the cohesiveness of the neighborhood of a node are much lower, indicating that miRNA targets are dispersed in the network. We further counted the percentage of targets in top 50, 100, 200, 500 and 1000-ranked nodes by their degrees and betweenness centralities respectively in the whole CePPIN. As we can see from Fig. 3, most of the targets have the highest degree and betweenness centrality in CePPIN, indicating these targets tend to be hub or bottleneck proteins in the network. Among the 100 miRNA targets, there are 88 ones with degrees higher than the average value of all proteins and 81 with betweenness centrality higher than the average value of all proteins. Owing to the existence of these miRNA targets, the CePPIN remains robust against random attack, but it is more sensitive to even slight changes of miRNA expression and more efficient to transfer the changes to other proteins, which can make a global influence on the whole PPIN.
Functional analysis of human targets of KSHV-miRNAs
We performed gene ontology (GO) and KEGG pathway analysis on all 109 miRNA targets. Totally, 83 GO: biological process (GO: bp), 27 GO: molecular function (GO: mf) and 19 KEGG pathway terms (p-value < 0.05) were found for these targets. Figure 4 shows the selected top 10 enriched GO terms and KEGG pathway terms for these targets. We can see that most of these enriched biological process terms are associated with oncogenesis, including cell death, apoptosis, transcriptional regulation and so on. For example, apoptosis is significantly enriched by targets of KSHV miRNAs, and inhibiting apoptosis contributes to maintaining KSHV survival. Several KSHV miRNAs can inhibit apoptosis such as miR-K5, -K9, and -K10a/b by targeting Bcl-2 associated factor (BCLAF1), a pro-apoptotic protein. On the other hand, all the molecular function terms are about the kinase activity and transcription factor binding, which are all molecular mechanisms related to cell cycle. KSHV genomes replicate once per cell cycle in latent cells and depends fully on cellular replication machinery for replication and genome maintenance, so cell cycle control regulated by KSHV miRNAs is essential for KSHV genomes replication. As for KEGG pathway, most of the enriched pathways are cell cycle and some cancer-related pathways such as pathways in cancer, TGF-β and so on. There are also some immune-related pathways such as B cell receptor signaling pathway included. These enriched GO terms and KEGG pathways are consistence with the conclusion that KSHV-encoded microRNAs tend to target immune modulatory genes and cell cycle control genes45.
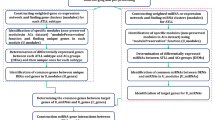
Furthermore, we identified and extracted functional modules from the KSHV miRNA-regulated CePPIN by using enrichment analysis in Reactome46 database with Network Analyst. As a result, six network modules of highly-interacting groups of genes were identified and extracted, namely innate immune system, cell cycle, signaling by the B Cell Receptor (BCR), herpes simplex infection, pathways in cancer and B cell receptor signaling pathway, respectively. The sub-networks of these six extracted functional modules were shown in Fig. 5. All these six important functional modules are proven to be highly related to KSHV pathogenesis. For example, I-kappa-B kinase epsilon (IKKɛ, gene symbol IKBKE) is a target of KSHV-miR-K12-11 in functional module innate immune system. KSHV-miR-K12-11 is critical for the modulation of Type I interferon (IFN) signaling, the principal response mediating antiviral innate immunity and acts through targeting IKKɛ. Importantly, expression of miR-K12-11 attenuated IFN signaling by decreasing IKKɛ-mediated IRF3/IRF7 phosphorylation and further inhibiting the activation IKKɛ-dependent IFN stimulating genes (ISGs), allowing miR-K12-11 suppression of antiviral immunity47. Therefore, it can be asserted that miR-K12-11 can contribute to maintenance of KSHV latency by targeting IKKɛ.
Comparative analysis on common targets and specific targets
As shown in Fig. 6B, through mapping the 109 human targets of KSHV-encoded miRNAs to 1727 targets of human miRNAs, we found an overlap of 40 genes, which indicating that a common set of targets are co-regulated by both some KSHV miRNAs and human miRNAs. We called the genes targeted by both KSHV miRNAs and human miRNAs as the common targets and the genes only specifically targeted by KSHV as the specific targets. To further investigate the difference between these two kinds of targets, we constructed miRNA-regulated CePPIN for the common targets (Fig. 7A) and specific targets (Fig. 7B), respectively. After that, we also performed gene ontology (GO) and KEGG pathway analysis on these two kinds of targets respectively. Figure 6A displays the number of enriched GO terms and KEGG pathway terms for common targets and specific targets and the number of overlap terms between them. It demonstrates that few common enriched GO terms and KEGG pathways can be found between the common targets and specific ones. In addition, we also performed gene ontology (GO) and KEGG pathway analysis on the remaining human targets and the KSHV targets. Compared to 113 GO: biological process, 32 GO: molecular function and 60 KEGG pathway terms of the remaining 1687 targets, only a minor portion of them are overlapped with those of the 109 KSHV targets, including 17 GO: biological process, 10 GO: molecular function and 14 terms KEGG terms (p-value < 0.05). It also shows that a very few common enriched GO terms and KEGG pathways can be found between the remaining human targets and the KSHV targets.
Difference analysis of common targets and specific targets. (A) The number and overlaps of GO: biological process, GO: molecular function and KEGG pathway terms (p-value < 0.05) for common targets and specific targets. (B) Overlaps of targets between KSHV miRNAs and human miRNAs. Blue represents KSHV miRNAs and orange represents human miRNAs. (C) Difference analysis of common targets and specific targets in terms of degree and betweenness centrality (t-test, *p < 0.05).
Moreover, we computed degree and betweenness centrality for common targets and specific targets in the whole miRNA-regulated CePPIN. Figure 6C shows the degree and betweenness centrality of common targets and specific targets. We can see that common targets and specific targets have significant difference in both degree and betweenness centrality by t-test (p-value < 0.05). These findings also suggest that KSHV miRNAs can be also classified into two generic groups, in which one can act as functional mimics of some oncogenic human miRNAs that contribute to tumorigenesis and the other could contribute to its own latent viral replication.
Conclusion
This work focuses on dissecting the regulation role of KSHV-encoded miRNAs through topological and functional analysis of human targets in miRNA-regulated CePPIN. We found that miRNA targets are essential in CePPIN with high degrees, betweenness centralities, closeness centralities and low clustering coefficients, thus they can make a global influence on the whole PPIN. Moreover, these targets are involved in various cellular processes especially related to immune regulate and cell cycle control, which contributes to KSHV immune escape and maintaining KSHV survival. In addition, the comparisons between the two sub-CePPINs for common targets and specific targets on the topological and function analysis indicate that significant difference are found in terms of both topological properties and enriched functions. So we can speculate that KSHV miRNAs mainly have two roles in viral-host interactions. One role is acting as functional mimics of some oncogenic human miRNAs which contribute to tumorigenesis and the other can facilitate to maintaining KSHV survival. Overall, the functional dissection of human targets for KSHV-encoded miRNAs in our work provides helpful information for further understanding the role of KSHV-encoded miRNAs in viral-host interaction from the perspective of the co-expressed protein-protein interaction network and hopefully may facilitate the researches on the current prevention and treatment of KSHV-induced cancers.
Material and Methods
Dataset of miRNAs and targets
The data of KSHV miRNAs and the corresponding human targets were downloaded from VmiReg database24. This database collects experimentally validated and predicted target genes of viral miRNAs. Here, we focused on experimentally validated human targets only and totally, we collected 16 KSHV miRNAs and the corresponding 109 human targets. In addition, an experimentally validated dataset of 1727 human targets of human miRNAs was also collected from the work of Shao et al.24.
PPIN construction and visualization
In our study, the construction and visualization of PPIN was generated by Network Analyst25 which contains the initial human PPI data from InnateDB26. In addition, NetworkAnalyst not only uses a comprehensive high-quality protein-protein interaction (PPI) database based on InnateDB but also contains manually curated protein interaction data from published literature as well as experimental data from several PPI databases including IntAct, MINT, DIP, BIND, and BioGRID. We mapped 109 human targets of 16 KSHV miRNAs to the PPI data to find the co-expressed proteins of these targets. 100 human targets and their co-expressed proteins were extracted from the original data, since 9 targets were discarded for lack of interactome information. Then, we used selected PPIs to construct the virally-encoded miRNA-regulated CePPIN for the human targets of the KSHV miRNAs. The topological parameters of CePPIN were computed using the Cytoscape (version 3.2.0)27 plugin Network Analyzer.
Topological parameters of PPIN
Topological analysis of biological networks is a topic of great interest in the field of current bioinformatics and systems biology. It provides quantitative insight into biological systems, especially for PPIN. It is a more effective strategy to identify important molecules, analyze functional modules or pathways and further uncover the mechanism of molecular interaction28, even to predict new molecular interaction29, 30.
In this study, four important topological parameters were used to describe the characterization of the nodes in CePPIN, including degree, betweenness centrality, clustering coefficient and closeness centrality respectively. Degree denotes the number of edges linked to the specified node in the network. In scale-free network, the nodes with degrees much higher than the average degree of the whole network are usually defined as hubs31. Betweenness centrality measures the number of non-redundant shortest paths going through a node. Studies have reported that nodes with high betweenness centralities called bottlenecks might act as an important link between two groups of nodes32. In most cases, nodes with high degrees tend to have high betweenness centralities and the two parameters are considered to be the best predictor of the essentiality of a node for network robustness, cooperation and communication. Clustering coefficient is defined as the ratio between the number of edges linking adjacent nodes and the total number of possible edges among them. Closeness centrality can be calculated as the reciprocal of the average shortest path length between a node and any other node in network. Nodes with higher closeness centralities are closer to other nodes in location and can also be more efficient to transmit information.
Functional analysis
To understand the underlying biological processes and pathways of downstream PPIs, we used the DAVID (The Database for Annotation, Visualization and Integrated Discovery)33 online tool (http://david.abcc.ncifcrf.gov/) to conduct PPI enrichment analysis. Investigations were implemented on GO (Gene Ontology)34 biological process, GO molecular function and KEGG (Kyoto Encyclopedia of Genes and Genomes)35 pathways at the significant level (p-value < 0.05). Furthermore, we also identified and extracted functional modules from the KSHV miRNA-regulated CePPIN by using enrichment analysis with Network Analyst25.
References
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Pfeffer, S. et al. Identification of virus-encoded microRNAs. Science 304, 734–736 (2004).
Boss, I. W., Plaisance, K. B. & Renne, R. Role of virus-encoded microRNAs in herpesvirus biology. Trends Microbiol 17, 544–553 (2009).
Skalsky, R. L. & Cullen, B. R. Viruses, microRNAs, and host interactions. Annu. Rev. Microbiol 64, 123–141 (2010).
Olena, A. F., Rao, M. B., Thatcher, E. J., Wu, S. Y. & Patton, J. G. miR-216a regulates snx5, a novel notch signaling pathway component, during zebrafish retinal development. Dev. Biol 400, 72–81 (2015).
Gioia, U. et al. mir-23a and mir-125b regulate neural stem/progenitor cell proliferation by targeting Musashi1. RNA Biol 11, 1105–1112 (2014).
Möhnle, P. et al. MicroRNA-146a controls Th1-cell differentiation of human CD4+ T lymphocytes by targeting PRKCε. Eur. J. Immunol 45, 260–272 (2015).
Zhu, H. Q. et al. MicroRNA-29b promotes high-fat diet-stimulated endothelial permeability and apoptosis in apoE knock-out mice by down-regulating MT1 expression. Int. J. Cardiol 176, 764–770 (2014).
Sandhu, R. et al. Overexpression of miR-146a in basal-like breast cancer cells confers enhanced tumorigenic potential in association with altered p53 status. Carcinogenesis 35, 2567–2575 (2014).
Berry, G. J., Budgeon, L. R., Cooper, T. K., Christensen, N. D. & Waldner, H. The type 1 diabetes resistance locus B10 Idd9.3 mediates impaired B-cell lymphopoiesis and implicates microRNA-34a in diabetes protection. Eur. J Immunol 44, 1716–1727 (2014).
Chang, Y. et al. Identification of herpesvirus-like DNA sequences in AIDS-associated Kaposi’s sarcoma. Science 266, 1865–1869 (1994).
Cesarman, E., Chang, Y., Moore, P. S., Said, J. W. & Knowles, D. M. Kaposi’s sarcoma-associated herpesvirus-like DNA sequences in AIDS-related body-cavity-based lymphomas. N. Engl. J. Med 332, 1186–1191 (1995).
Soulier, J. et al. Kaposi’s sarcoma-associated herpesvirus-like DNA sequences in multicentric Castleman’s disease. Blood 86, 1276–1280 (1995).
Cai, X. et al. Kaposi’s sarcoma-associated herpesvirus expresses an array of viral microRNAs in latently infected cells. Proc. Natl. Acad. Sci. USA 102, 5570–5575 (2005).
Grundhoff, A., Sullivan, C. S. & Ganem, D. A combined computational and microarray-based approach identifies novel microRNAs encoded by human gamma-herpesviruses. RNA 12, 733–750 (2006).
Pfeffer, S. et al. Identification of microRNAs of the herpesvirus family. Nat. Meth 2, 269–276 (2005).
Samols, M. A., Hu, J., Skalsky, R. L. & Renne, R. Cloning and identification of a microRNA cluster within the latency-associated region of Kaposi’s sarcoma-associated herpesvirus. J. Virol 79, 9301–9305 (2005).
Ambros, V. The function of animal miRNAs. Nature 431, 350–355 (2004).
Lee, Y. & Dutta, A. MicroRNAs in cancer. Annu. Rev. Pathol 4, 199–227 (2009).
Garzon, R., Calin, G. & Croce, C. MicroRNAs in cancer. Annu. Rev. Med 60, 167–179 (2009).
Hsu, C. W., Juan, H. F. & Huang, H. C. Characterization of microRNA-regulated protein-protein interaction network. Proteomics 8, 1975–1979 (2008).
Tseng, C. W., Lin, C. C., Chen, C. N., Huang, H. C. & Juan, H. F. Integrative network analysis reveals active microRNAs and their functions in gastric cancer. BMC Syst. Biol 5, 99 (2011).
Liu, Z., Guo, Y., Pu, X. & Li, M. Dissecting the regulation rules of cancer-related miRNAs based on network analysis. Sci. Rep 6, 34172 (2016).
Shao, T. et al. Functional dissection of virus-human crosstalk mediated by miRNAs based on the VmiReg database. Mol. Biosyst 11, 1319–1328 (2015).
Xia, J., Gill, E. E. & Hancock, R. E. Network Analyst–integrative approaches for protein-protein interaction network analysis and visual exploration. Nucleic Acids Res 42, W167–174 (2014).
Lynn, D. J. et al. InnateDB: facilitating systems-level analyses of the mammalian innate immune response. Mol. Syst. Biol 4, 218 (2008).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13, 2498–2504 (2003).
Aittokallio, T. & Schwikowski, B. Graph-based methods for analysing networks in cell biology. Brief Bioinform 7, 243–55 (2006).
Paladugu, S. R., Zhao, S., Ray, A. & Raval, A. Mining protein networks for synthetic genetic interactions. BMC Bioinformatics 9, 426–439 (2008).
You, Z. H., Yin, Z., Han, K., Huang, D. S. & Zhou, X. A semi-supervised learning approach to predict synthetic genetic interactions by combining functional and topological properties of functional gene network. BMC Bioinformatics 11, 343–355 (2010).
Albert, R. Scale-free networks in cell biology. J. Cell Sci 118, 4947–4957 (2005).
Joy, M. P., Brock, A., Ingber, D. E. & Huang, S. High-betweenness proteins in the yeast protein interaction network. J. Biomed. Biotechnol 2005, 96–103 (2005).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc 4, 44–57 (2009).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet 25, 25–29 (2000).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28, 27–30 (2000).
Gottwein, E. et al. A viral microRNA functions as an orthologue of cellular miR-155. Nature 450, 1096–1099 (2007).
Skalsky, R. L. et al. Kaposi’s sarcoma-associated herpesvirus encodes an ortholog of miR-155. J. Virol 81, 12836–12845 (2007).
Qin, Z. et al. Role of Host MicroRNAs in Kaposi’s Sarcoma-Associated Herpesvirus Pathogenesis. Viruses 6, 4571–4580 (2014).
Ziegelbauer, J. M., Sullivan, C. S. & Ganem, D. Tandem array-based expression screens identify host mRNA targets of virus-encoded microRNAs. Nat. Genet 41, 130–134 (2009).
Ramalingam, D., Kieffer-Kwon, P. & Ziegelbauer, J. M. Emerging themes from EBV and KSHV microRNA targets. Viruses 4, 1687–710 (2012).
Warnatz, H. J. et al. The BTB and CNC homology 1 (BACH1) target genes are involved in the oxidative stress response and in control of the cell cycle. J. Biol. Chem 286, 23521–23532 (2011).
Hansen, A. et al. KSHV-encoded miRNAs target MAF to induce endothelial cell reprogramming. Genes Dev 24, 195–205 (2010).
Gottwein, E. & Cullen, B. R. A human herpesvirus microRNA inhibits p21 expression and attenuates p21-mediated cell cycle arrest. J. Virol 84, 5229–5237 (2010).
Tili, E. et al. Mutator activity induced by microRNA-155 (miR-155) links inflammation and cancer. Proc. Natl. Acad. Sci. USA 108, 4908–4913 (2011).
Zhu, Y. et al. γ-Herpesvirus-encoded miRNAs and their roles in viral biology and pathogenesis. Curr Opin Virol 3, 266–275 (2013).
Joshi-Tope, G. et al. Reactome: a knowledgebase of biological pathways. Nucleic Acids Res 33, D428–432 (2005).
Liang, D. et al. A human herpesvirus miRNA attenuates interferon signaling and contributes to maintenance of viral latency by targeting IKKε. Cell Res 21, 793–806 (2011).
Acknowledgements
This work was funded by the National Natural Science Foundation of China (No. 21675114, 21573151).
Author information
Authors and Affiliations
Contributions
Y.W. and Y.L. performed research, analyzed data and wrote the paper; Y.G. designed research and wrote the paper; X.P. and M.L. analyzed data. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, Y., Lin, Y., Guo, Y. et al. Functional dissection of human targets for KSHV-encoded miRNAs using network analysis. Sci Rep 7, 3159 (2017). https://doi.org/10.1038/s41598-017-03462-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-03462-w
- Springer Nature Limited
This article is cited by
-
Individually double minimum-distance definition of protein–RNA binding residues and application to structure-based prediction
Journal of Computer-Aided Molecular Design (2018)