Abstract
Paspalum notatum Flüggé is an economically important subtropical fodder grass that is widely used in the Americas. Here, we report a new chromosome-scale genome assembly and annotation of a diploid biotype collected in the center of origin of the species. Using Oxford Nanopore long reads, we generated a 557.81 Mb genome assembly (N50 = 56.1 Mb) with high gene completeness (BUSCO = 98.73%). Genome annotation identified 320 Mb (57.86%) of repetitive elements and 45,074 gene models, of which 36,079 have a high level of confidence. Further characterisation included the identification of 59 miRNA precursors together with their putative targets. The present work provides a comprehensive genomic resource for P. notatum improvement and a reference frame for functional and evolutionary research within the genus.
Similar content being viewed by others
Background & Summary
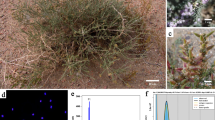
Paspalum notatum Flüggé (bahiagrass) is a subtropical grass native to South America that is widespread on lightly textured soils in warm, humid regions of the Western Hemisphere and extensively used as a pasture and ground cover1,2. The species forms a multiploid complex in which the diploid (2n = 2x = 20) plants are self-sterile and sexual, while the polyploids (3x = 30, 4x = 40, 5x = 50) are pseudogamous aposporous apomicts, i.e. they form seeds containing maternal embryos3,4. The diploid form, var. saurae, also known as Pensacola bahiagrass, occurs naturally in a restricted geographical area of Argentina stretching between the western and eastern banks of the Uruguay and Paraná rivers, respectively2. It owes its name to the fact that it was inadvertently introduced in the Pensacola area of Florida before 1926 and subsequently naturalized as a warm-season perennial pasture throughout the coastal plain and Gulf Coast regions of the United States5. Today, it is one of the most important grasses for pastures and lawns in the southeastern United States6. The search for the origin of Pensacola bahiagrass led the agricultural scientist Glenn W. Burton to travel through Brazil, Uruguay, and Argentina, where he eventually found highly diverse populations in a small area of the province of Santa Fe, on the banks of the Paraná River and the island of Berduc, near the city of Cayastá5 (Fig. 1a,b). Since cytogenetic studies indicate that polyploid P. notatum races (var. notatum) are autotetraploid and share homologous chromosomes with the saurae plants7,8, this region was then considered to be the center of origin of the species2,5.
General view of the proposed center of diversity of P. notatum. (a) Map of the area of natural distribution of the diploid cytotype in the provinces of Santa Fe and Entre Ríos, Argentina. The arrow indicates the location of the city of Cayastá (31° 12′ 0″ S; 60° 10′ 0″ W), close to the sampling site of the #R1 plant. (b) Photograph of the herbarium voucher of diploid bahiagrass collected by Prof. Camilo Quarin in 1992 kept at the Carmen L. Cristobal herbarium at the Instituto de Botánica del Nordeste (IBONE), CONICET-UNNE, Corrientes, Argentina. (c) Photograph of the banks of the Paraná River in the Cayastá area (top), where natural bahiagrass populations were found (bottom left), and a close-up of a bahiagrass inflorescence at anthesis (bottom right). (d) Duplicate of the P. notatum var. saurae #R1 plant used for genome sequencing.
Because P. notatum establishes well in poor-quality sandy soils and tolerates drought, sporadic flooding, and continuous grazing, the species has been selected and improved by classical and molecular methods for almost 80 years, with about 20 cultivars released to date9. While the diploid sexual races could be crossed to generate improved hybrids, tetraploid cultivars were traditionally obtained through ecotype selection due to their apomictic mode of reproduction9. However, the experimental production of tetraploid sexual individuals by doubling the chromosomes of diploids and the creation of synthetic sexual tetraploid populations have increased the variability for breeding programs through crosses with natural apomictic pollen donors9,10,11.
P. notatum ecotypes have relatively small genomes, with 1 C values ranging from 0.55 to 0.60 pg12. Recent studies have provided a wealth of information on the species’ genetic, transcriptomic, and genomic data11 and have set up strategies for the functional characterization of agronomically important genes using genetic transformation and gene editing13,14,15. Available resources include leaf and flower transcriptomes of sexual and apomictic genotypes16,17,18, a catalog of small RNAs present during the sexual and apomictic reproductive development19, and a chromosome-scale de novo genome assembly (514 Mb) of the species20. However, information on gene content annotation and miRNA genes is not yet available.
Long-read sequencing technologies have proven to be extremely effective in improving the quality of assembly in complex genomes, with high levels of heterozygosity, polyploidy, and repetitive elements21,22,23, particularly for non-model species and orphan crops24,25,26,27,28. Here, we report a chromosome-level genome assembly and annotation of a natural diploid P. notatum biotype (#R1) collected at the species center of diversity using Oxford Nanopore Technology (ONT). The plant #R1 reproduces sexually but occasionally produces aposporous embryo sacs, which is the first step of apomictic reproduction29. Further extensive genomic characterization using Illumina short reads, together with the existing and newly generated transcriptomes, makes the #R1 genome assembly and annotation a valuable resource for providing new insights into the gene content and genome evolution, and for elucidating the developmental genetics of agronomically valuable traits.
Methods
Sample collection
The #R1 plant is a diploid individual collected in a natural population established near the city of Cayastá, Santa Fe Province, Argentina29 (Fig. 1a,c), which belongs to the living germplasm collection of Paspalum spp. of the Instituto de Botánica del Nordeste (IBONE), CONICET-UNNE, Corrientes, Argentina (voucher CTES0553130; Herbarium Carmen L. Cristobal) (Fig. 1b). Several duplicates generated by vegetative propagation through rhizomes are also maintained at the Instituto de Investigaciones en Ciencias Agrarias de Rosario (IICAR), CONICET-UNR, Rosario, Argentina, and at the French National Research Institute for Sustainable development (IRD), Montpellier, France (Fig. 1d). For ONT sequencing, we used ~5 gr of fresh leaf tissue to extract high molecular weight genomic DNA (HMW gDNA) from nuclei isolation and performed quality control, both according to Mariac et al.30. We also extracted total RNA for cDNA synthesis and ONT sequencing from flowers of #R1 immature inflorescences collected before anthesis using a method adapted from Azevedo et al.31. Briefly, the plant material was ground in liquid nitrogen, mixed with the extraction buffer, incubated for 15 min at room temperature, and finally extracted using chloroform-isoamyl alcohol. We preserved RNA integrity by avoiding vortexing and keeping samples on ice throughout the extraction process. The genomic DNA used for preparing Illumina sequencing libraries was extracted from ~3 gr of fresh leaf tissue using a CTAB (cetyltrimethylammonium bromide) method32 and qualified for concentration and purity using a NanoDrop 2000 (Thermo Scientific, USA).
DNA sequencing
Nanopore sequencing
DNA libraries of the #R1 genotype were prepared from non-fragmented HMW gDNA using the ligation sequencing Kit 1D SQK-LSK109 (Oxford Nanopore Technology). ONT sequencing was carried out using either a MinION MK1b (Oxford Nanopore Technology) at IRD or a PromethION (Oxford Nanopore Technology, UK) at Novogene (Cambridge, UK) employing R9.4.1 Spot-On Flow Cells (Oxford Nanopore Technology). ONT sequencing FAST5 files were base-called using GUPPY v6.0.6 software and the dna_r9.4.1_450bps_hac_prom.cfg model. The quality control of raw reads in FASTQ format was conducted using NanoPlot v1.31.0 software33.
Illumina sequencing
Illumina sequencing was carried out at the Instituto de Agrobiotecnología de Rosario (INDEAR; Rosario, Argentina). Sequencing libraries were prepared from 50 ng of genomic DNA using the Nextera DNA Library Prep Kit (Illumina, Inc., San Diego, CA, USA) according to the manufacturer’s instructions and sequenced using a 2 × 250 paired-end Illumina HiSeq. 1500 platform.
Assessing the heterozygosity level of the #R1 genome
Illumina reads were trimmed to remove adaptors and filtered by quality using Trimmomatic v0.3334. Approximately, 277 million high-quality Illumina reads (Q > 37) (Supplementary Table 1) were used as input to count 21k-mers using Jellyfish v2.3.035, followed by a genome scan using GenomeScope36.
cDNA sequencing
cDNA from flowers of immature #R1 inflorescences was synthesized from 50 ng of total RNA using the SMART-Seq V4 low-input RNA kit (Takara Bio Europe, France). Of the 10 µl reverse transcription reaction, 1 µl was used for quality control and the remaining 9 µl were amplified using Seq Amp DNA Polymerase with seqAmp CB PCR Buffer for long fragment amplification (Takara Bio Europe, France). A sequencing cDNA library with an estimated concentration of 80 fmol (2000 bp average library size) was prepared using the SQK-LSK 109 ligation sequencing kit (Oxford Nanopore Technologies, UK). Preparation included RNA and cDNA purification steps using dAMPure XP Beads (Beckman Coulter, France). RNA quality was assessed using the Agilent High Sensitivity DNA Reagent Kit (Agilent Technologies, France). ONT sequencing and base calling were performed at IRD, as described above. Raw reads were filtered for quality (Q > 10) and length (>300 bp) and trimmed (85 bp at both ends) using Nanofilt v1.036.
Genome survey and assembly
Preliminary k-mer analysis carried out with the Illumina reads predicted a total genome size of 513 Mb, an abundance of repetitive elements of approximately 50.0% and a heterozygosity rate of 1.73%, as indicated by the bimodal k-mer profile (Fig. 2). This high level of heterozygosity was expected for the #R1 genotype based on previous genetic analysis of the natural population from which the plant was collected29 and is similar to that reported for other self-incompatible grasses37,38. To achieve genome assembly, we first generated 72.13 Gb of ONT long reads (Q > 7) (19.98 Gb from MinION and 52.15 Gb from PromethION) with a N50 = 19.71 kb of read length and a GC content of 45.56% (Supplementary Table 139). The reads were then filtered for quality (Q > 10) and length (>5 kb) using NanoFilt v1.033 resulting in of 68 Gb of data with a %GC of 45.60 and an N50 of 20.41 kb (Supplementary Table 1), which were assembled using Flye v2.940. The de novo assembled contigs were polished using Racon v1.4.1041 and scaffolded by RagTag v2.1.042 using the available P. notatum genome reference20 (NCBI Genome assembly ASM2253091v1), excluding the unassigned contigs. The new assembly was polished with the 70 × coverage of Illumina pair-end sequences. Illumina short reads mapping was performed using BWA-MEM v0.7.1743, and error correction was performed with Pilon v1.2344 in two successive iterations. This procedure resulted in a 557.8 Mb #R1 genome (GenBank GCA_036689595.1), including the ten expected chromosome-length scaffolds (N50 = 56.10 Mb) and a GC content of 45.80% (Table 1). Of the total ONT reads used as input, 99.14% were mapped within the assembly, indicating a high degree of raw data inclusiveness. #R1 pseudomolecules were named based on their sequence similarity to the reference chromosomes20. Chromosome size varied between 46.63 and 85.72 Mb, with a mean of 55.78 ± 10.93 Mb (Supplementary Fig. 1). Some of the unassigned contigs reported by Yan et al.20 showed similarity with sequences within the #R1 chromosomes. These additions probably contribute to the increase in the genome length from 541 Mb of the reference20 to 557.8 Mb of the new assembly.
Flowers and leaves transcriptome assembly
The #R1 genome was used for a reference-guided transcriptome assembly of flowers and leaves. From a total of 11.9 Gb of ONT cDNA reads from flower transcriptome, ~10 Gb of filtered reads (Q > 10) were assembled using Stringtie v 2.1.445. The resulting flower transcriptome assembly consisted of 36,317 transcripts with a GC content of 51.68% and an N50 of 2,382 bp (Table 2; Supplementary Table 2) (GenBank GKQU01000000.1). Furthermore, the Illumina cDNA paired-end reads (QC > 30) from leaves of diploid genotypes available from NCBI database SRR7347364, SRR7347365, SRR7347366, SRR7347367, SRR7347368, SRR734736917 were reference-based assembled using Trinity v2.0.246 and produced 76,682 transcripts with a %GC content of 46.69% and N50 of 1,545 bp (Table 2, Supplementary Table 2). The features of both transcriptomes were consistent with previous reports for the species16,17,18 and were subsequently used as biological evidence for the #R1 genome annotation (see below).
Genome annotation
Repetitive sequences
Repetitive sequences in the #R1 genome assembly were assessed using the filtered Illumina paired-end reads and the RepeatExplorer2 pipeline integrated into the Galaxy platform (https://repeatexplorer-elixir.cerit-sc.cz/) following the protocol described by Novak et al.47. Briefly, a clustering analysis was performed using RepeatExplorer2 and the TAREAN tandem repeat analyzer module. The DANTE tool was used to extract the consensus sequences of transposable elements (TEs) and classify them based on the REXdb database Viridiplantae 3.0 release48, using ‘BLOSUM80’ as scoring matrix and no iterative search. RepeatModeler v4.1.249 (RM2) was used to generate a custom library of P. notatum TEs, and RepeatMasker v4.1.2-p150 was used to determine the frequency of repeat DNA families. The RM2 output was then parsed (modified ParseRM.pl script51) to identify and quantify TE families. The putative centromeric regions of #R1 chromosomes were localized using the centromere-specific satellite sequences of eight grass species (Oryza sativa, Setaria viridis, Setaria italica, Panicum hallii, Panicum capillare, Panicum virgatum, Zea mays and Zea luxurians) described by Melters et al.52. Chromosomal positions were determined by BLASTN analysis53 using the satellite sequences as query and considering only the alignments longer than 100 bp and identities >80%54. Telomeric regions were identified using the quarTeT tool55.
Analysis of the Illumina reads with RepeaExplorer2 identified a total of 320 Mb of repetitive sequences (57.36% of the #R1 assembly), predominantly consisting of retrotransposons (82.12%) and DNA transposons (7.17%) and including a significantly large proportion of unclassified elements (Fig. 3a, Table 3). When mapped onto the #R1 genome, repetitive sequences occupied a minimum of 44.96% (chr. 02) and a maximum of 71.21% (chr. 08) of the chromosome length (Table 4). As expected, the density distribution of the different repeat elements varied along the chromosomes. LTRs were most abundant in putative centromeric regions, whereas retroelements (LINE and SINE), DNA transposons, and rolling circles were prevalent in chromosome arms (Fig. 3b,c). Simple repeats and satellite repeats appeared regularly distributed along all ten chromosomes (Fig. 3c). The putative centromeric regions could be assigned to eight of the ten chromosomes. For chromosomes 2 and 10, these regions could not be properly defined, probably due to a low assembly resolution in these areas and therefore, the proposed locations are hypothetical (Table 4, Supplementary Fig. 1). Similarly, the putative locations of the telomeric regions of chromosomes 2, 3, 4, 9 and 10 were recognized. However, for chromosomes 1, 5, 6, 7 and 8 the positions given are provisional due to the short alignments obtained (Table 4). The average length of the putative telomeres was 6,255 bp, ranging from 70 bp (Chr. 09) to 26,929 bp (Chr. 03) (Table 4).
#R1 genomic sequences annotation. (a) Percentages of the repetitive and non-repetitive sequences (RepeatMasker results) in the #R1 genome, and proportions of the different types of TEs (estimated over the total of repetitive sequences). (b, c) Density distribution over the #R1 genome of (b) retroelements (from outside to inside LTR, LINE and SINE) and (c) DNA transposons, simple repeats, satellite repeats and rolling circles (from outside to inside).
Gene annotation
Gene prediction and annotation were performed using the MAKER v2.31.9 pipeline56 by integrating ab initio gene model predictions with biological transcriptomic and proteomic data through multiple BLAST steps using Exonerate v2.4.057. The soft-repeat-masked version of the #R1 genome together with flower and leaf transcriptomes (this work) merged and filtered for redundancy (similarity threshold of 90%) using CD-HIT58 were used as input. In addition, the transcriptome of Sorghum bicolor NCBIv3 (GeneBank GCA_000003195.3) and the proteome of Oryza sativa Japonica Group cv. Nipponbare (Genebank GCA_001433935.1) were included as expressed sequence evidence of related species. Two MAKER iterations were performed to obtain the final annotation. In the first one, ab initio gene predictions were carried out using AUGUSTUS v3.2.259 with the EST trust-blindly option enabled and Oryza sativa as the model species. The resulting gene models were filtered to retain only those with an annotation error distance (AED) <0.556. The outcome of this first annotation was then used to train new species models for AUGUSTUS and SNAP60 for the second run of MAKER. Gene models with an AED score > 0.5 and transcripts <50 nt were filtered out. The predicted coding sequences (CDS) obtained with MAKER were then translated to protein sequences using the program GffReadv0.12.761 with parameter “-y”. Predicted protein sequences were checked for CDS features (presence of start and stop codons) and for homology with known domains using InterProScan v5.53.87.062 (consulting the databases TIGRFAM, SFLD, SUPERFAMILY, PANTHER, SMART, CDD, PIRSR, Pfam, on April 2023). Gene models that fitted with both criteria were considered as “high confidence”.
Using this strategy, a total of 51,249 transcripts with an AED < 0.5 (85.18% of the total predicted) (Fig. 4a), which defined 45,074 gene models with approximately 1.14 transcripts per gene, were obtained (Supplementary Table 3). The average lengths of mRNA and CDS were 3,679 nt and 1,258 nt, respectively. Each predicted gene contained an average of 4.4 exons, and the exons’ mean length was 346 nt. Of the total predicted gene models, 36,079 (80.04%) were classified as high-confidence (HC) genes. The complete list of genes, their genomic coordinates and corresponding A. thaliana and rice homologs, together with their functional annotation, are summarized from the GFF file in the Supplementary Table 3. As expected, over 99% of the flower and leaf transcripts mapped in the #R1 genome showing a high density towards the ends of the chromosome arms and a low density in most of the putative centromeric regions (Fig. 4b). The number and density of genes per #R1 chromosome are shown in Table 5.
Transcripts prediction and distribution: (a) Histogram of the Annotation Edit Distances (AED) scores of the predicted transcripts. Green and blue bars show the number of transcripts with and without hits in the InterProScan database, respectively. (b) Circle plot showing (from outside to inside) the density distribution of predicted, leaf and flower transcripts in the #R1 genome.
Identification of rRNA and tRNAs
rRNA genes were identified using Barrnap v0.963 software with an e-value cut-off for similarity of 1e−10 and a minimum length threshold of 0.9. In addition, tRNA genes were identified using tRNAscan-SE V1.3.164 with the ‘-infernal’ mode. These analyses resulted in the annotation of 354 rRNA genes and 544 tRNA genes in the #R1 genome (Table 1), which localization is presented in the GFF annotation file deposited in the NCBI database accession number (GCA_036689595.1).
Prediction of microRNA (miRNA) genes and targets
MicroRNA (miRNA) gene precursors present in the #R1 genome were searched using the small RNA (sRNA) sequence database of the reproductive development of sexual and apomictic P. notatum genotypes19 available at the NCBI BioProject Accession: PRJNA373857 and the software ShortStack 3.8.465. miRNA precursors, miRNA mature sequences and putative targets in the #R1 genome were detected as described in Ortiz et al.19 using the #R1 assembly as a reference. The putative miRNA’s target regions were analyzed using the #R1 GFF annotation file to determine the location of the mature miRNA alignment (5′ UTR, exon, intron, or 3′ UTR regions) within the genes. Following these procedures, a total of 59 clusters distributed across the 10 chromosomes containing sRNAs were detected (Supplementary Table 4, sheet 1), most of them producing mature miRNAs of 21 nt (47 clusters) and 22 nt (9 clusters). A total of 52 unique mature miRNAs were predicted, corresponding to 21 known families and including all miRNAs previously described in the species, with the exception of the miR39019 (Supplementary Table 4, sheet 2). Moreover, two new miRNAs (miR827 and miR3979) were identified in the species (Supplementary Table 4, sheet 2). Fourteen precursors generate putative mature miRNAs with no significant match in MirBase and, therefore, may represent novel Paspalum-specific miRNAs. A search for target regions in the #R1 genome performed with TargetFinder66 identified 1,456 unique genomic regions (TF score < 4), of which 1,324 have homology with known proteins (Supplementary Table 4, sheet 3).
Data Records
The raw reads derived from the #R1 genome sequencing using Oxford Nanopore (ONT) technology were deposited in the NCBI Sequence Read Archive (SRA) database under accession Nos. SRS1997548067 and SRS1997548268. The sequencing Illumina raw data were deposited in the NCBI SRA database SRS1997548369 and SRS1997548470. The #R1 genome assembly and annotation were deposited in the NCBI database under accession No GCA_036689595.171. The reads of the #R1 flower cDNA ONT sequencing were deposited in SRA database SRS1997548172, and the #R1 flower transcriptome assembly were deposited in the NCBI database under accession No. GKQU00000000.173. The raw reads from leaves were downloaded from the NCBI Sequence Read Archive (SRA) database accession Nos. SRR734736474, SRR734736575, SRR734736676, SRR734736777, SRR734736878, SRR734736979. The leaf transcriptome assembly was deposited in the NCBI under the accession number DAWXED00000000080. The precursor and mature miRNA sequence data recovered from the #R1 genome has been incorporated in the Supplementary Table 4, sheets 1 and 2.
Technical Validation
Assessing the quality of HMW genomic DNA for ONT sequencing
The quality and integrity of the #R1 genomic DNA for ONT sequencing was evaluated using a NanoDrop One/One Spectrophotometer and a Pulsed-Field Gel Electrophoresis system (PFGE BioRad) according to Mariac et al.30 (https://www.protocols.io/view/high-molecular-weight-dna-extraction-from-plant-nu-83shyne). DNA preparations consistently showed spectrophotometric ratios 260/280 nm 1.8–2.0 and 260/230 2.0-2.2, confirming the purity of the extraction. On the other hand, the high molecular weight of the DNA preparation was checked out by loading 1.5–5.5 µg of genomic DNA in 1% agarose gel (0.5 × TAE) with 5 µl of 6 × loading buffer and electrophoresed using the following parameters: pulse time: initial = 5 s, final = 117 s, running time = 20.5 h, V/cm = 5, Angle = 120, Temp = 14° and mA end of run = 255. The molecular weight of the genomic DNA preparation obtained ranged from 48 to 200 kb (Supplementary Fig. 2).
Assessment of genome and transcriptome assembly and annotation quality
The NCBI-FCS-GX scan tool81 was used to find contaminants in the assembly, setting the taxon in Viridiplantae. In addition, the presence of organellar DNA was assessed by BLASTn analysis (query coverage >30% and % of identity >60%) using the Oryza sativa IRGSP-1.0 organellar data set as query. No contaminants or organellar DNA were detected in the #R1 assembly. The software Merqury82 was used to estimate the base-level accuracy and k-mer completeness of the #R1 genome. This analysis showed an assembly consensus quality value (QV) of 30.2, which correspond to an accuracy of 99.9%, and a k-mer completeness value of 84.3%. Nevertheless, we cannot discard that some regions may include both haplotypes (Supplementary Fig. 3). In addition, the #R1 assembly quality was evaluated using BUSCO v583 using the Liliopsida gene set as a reference, and by mapping the Illumina paired-end reads over the genome. The BUSCO score showed the presence of 94.7% of complete genes,(with 91.8% of them corresponding to single genes), 4% of fragmented genes and 1.3% of missing genes (Supplementary Fig. 4a). Furthermore, the percentage of the total core genes with more than one ortholog was only 3.1%. Moreover, 97.7% of the paired-end Illumina reads were properly mapped by BWA-mem v0.7.17 to the #R1 genome, with an estimated average coverage depth of 93.2×. Using the same procedure for assessing MAKER gene annotation, the BUSCO score showed that 94.4% of the 3,236 Lilliopsida single-copy genes were properly annotated, with an average of 1.19 orthologs for each gene (Supplementary Fig. 4b). In this case, the percentage of duplicate transcripts increase up to 10.1%, probably due to the inclusion of splicing variants. On the other hand, BUSCO analysis performed to evaluate both the flower and leaf transcriptome assemblies revealed 87.9% and 82.2% of complete, 3.7% and 8.6% of fragmented, and 8.4% and 9.2% of missing genes, respectively (Supplementary Fig. 4c,d). Overall, these results indicate that both transcriptomes have a high level of completeness, and therefore represent comprehensive evidence of the expressed sequences of the #R1 genome.
Code availability
All software packages used in this study were run according to their user manuals. The version and parameters used are listed in the Table 6. No specific custom codes were used in this study.
References
Chase, A. The North American species of Paspalum. In Systematic plant studies. 1–310 (1929).
Gates, R. N., Quarin, C. L. & Pedreira, C. G. S. Bahiagrass. In: Warm‐season (C4) grasses 45, 651-680 (2004).
Burton, G. W. The method of reproduction in common bahia grass, Paspalum notatum. Agron. J. 40(5), 443–452 (1948).
Burton, G. W. Breeding Pensacola Bahiagrass, Paspalum notatum: Method of reproduction. Agron. J. 47(7), 311–314 (1955).
Burton, G. W. A search for the origin of Pensacola Bahia grass. Econ. Bot. 21(4), 379–382 (1967).
Acuña, C. A. et al. Bahiagrass tetraploid germplasm: reproductive and agronomic characterization of segregating progeny. Crop Sci. 49, 581–588 (2009).
Forbes, I. Jr & Burton, G. W. Cytology of diploids, natural and Induced tetraploids, and intra‐species hybrids of Bahiagrass, Paspalum Notatum Flügge. Crop Sci. 1(6), 402–406 (1961).
Quarin, C. L., Burson, B. L. & Burton, G. W. Cytology of intra-and interspecific hybrids between two cytotypes of Paspalum notatum and P. cromyorrhizon. Bot. Gaz. 145(3), 420–426 (1984).
Acuña, C. A. et al. Reproductive systems in Paspalum: Relevance for germplasm collection and conservation, breeding techniques, and adoption of released cultivars. Front. Plant Sci. 10, 1377 (2019).
Zilli, A. L. et al. Widening the gene pool of sexual tetraploid bahiagrass: generation and reproductive characterization of a sexual synthetic tetraploid population. Crop Sci. 58(2), 762–772 (2018).
Ortiz, J. P. A., Pupilli, F., Acuña, C. A., Leblanc, O. & Pessino, S. C. How to become an apomixis model: the multifaceted case of Paspalum. Genes 11(9), 974 (2020).
Galdeano, F. et al. Relative DNA content in diploid, polyploid, and multiploid species of Paspalum (Poaceae) with relation to reproductive mode and taxonomy. J. Plant Res. 129(4), 697–710 (2016).
Mancini, M. et al. The MAP3K-coding QUI-GON JINN (QGJ) gene is essential to the formation of unreduced embryo sacs in Paspalum. Front. Plant Sci. 9, 1547 (2018).
Colono, C. et al. A plant-specific TGS1 homolog influences gametophyte development in sexual tetraploid Paspalum notatum ovules. Front. Plant Sci. 10, 1566 (2019).
May, D., Sanchez, S., Gilby, J. & Altpeter, F. Multi-allelic gene editing in an apomictic, tetraploid turf and forage grass (Paspalum notatum Flüggé) using CRISPR/Cas9. Front. Plant Sci. 14 (2023).
Ortiz, J. P. A. et al. A reference floral transcriptome of sexual and apomictic Paspalum notatum. BMC Genom. 18, 1–14 (2017).
de Oliveira, F. A. et al. Coexpression and transcriptome analyses identify active apomixis-related genes in Paspalum notatum leaves. BMC Genom. 21(1), 1–15 (2020).
Podio, M., Colono, C., Siena, L., Ortiz, J. P. A. & Pessino, S. C. A study of the heterochronic sense/antisense RNA representation in florets of sexual and apomictic Paspalum notatum. BMC Genom. 22, 1–19 (2021). (2021).
Ortiz, J. P. A. et al. Small RNA-seq reveals novel regulatory components for apomixis in Paspalum notatum. BMC Genom. 20(1), 1–17 (2019).
Yan, Z. et al. High-quality chromosome-scale de novo assembly of the Paspalum notatum ‘Flugge’ genome. BMC Genom. 23(1), 293 (2022).
Pucker, B., Irisarri, I., de Vries, J. & Xu, B. Plant genome sequence assembly in the era of long reads: Progress, challenges and future directions. Quant. Plant. Biol. 3, e5 (2022).
Sahu, S. K. & Liu, H. Long-read sequencing (method of the year 2022): the way forward for plant omics research. Mol. Plant 16(5), 791–793 (2023).
Warburton, P. E. & Sebra, R. P. Long-Read DNA Sequencing: Recent Advances and Remaining Challenges. Annu Rev Genomics Hum Genet. 24 (2023).
Siadjeu, C., Pucker, B., Viehöver, P., Albach, D. C. & Weisshaar, B. High contiguity de novo genome sequence assembly of trifoliate yam (Dioscorea dumetorum) using long read sequencing. Genes 11(3), 274 (2020).
Hunt, S. P. et al. A chromosome-scale assembly of the garden orach (Atriplex hortensis L.) genome using Oxford Nanopore sequencing. Front. Plant Sci. 11, 624 (2020).
Carballo, J. et al. A high-quality genome of Eragrostis curvula grass provides insights into Poaceae evolution and supports new strategies to enhance forage quality. Sci. Rep. 9(1), 10250 (2019).
Sun, G. et al. Genome of Paspalum vaginatum and the role of trehalose mediated autophagy in increasing maize biomass. Nat. Commun. 13(1), 7731 (2022).
Wu, D. et al. Genomic insights into the evolution of Echinochloa species as weed and orphan crop. Nat. Commun. 13(1), 689 (2022).
Quarin, C. L., Espinoza, F., Martinez, E. J., Pessino, S. C. & Bovo, O. A. A rise of ploidy level induces the expression of apomixis in Paspalum notatum. Sex. Plant Reprod. 13, 243–249 (2001).
Mariac, C., Zekraoui, L. & Leblanc, O. High molecular weight DNA extraction from plant nuclei isolation. Protocols.io. https://doi.org/10.17504/protocols.io.83shyne (2019).
Azevedo, H., Lino-Neto, T. & Tavares, R. M. An improved method for high-quality RNA isolation from needles of adult maritime pine trees. Plant Mol. Biol. Rep. 21, 333–338 (2003).
Clarke, J. D. Cetyltrimethyl ammonium bromide (CTAB) DNA miniprep for plant DNA isolation. Cold Spring Harb. Protoc. 3, pdb–prot5177 (2009).
De Coster, W. & Rademakers, R. NanoPack2: population-scale evaluation of long-read sequencing data. Bioinform. 39(5), btad311 (2023).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinform. 30, 2114–20 (2014).
Marçais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinform. 27(6), 764–770 (2011).
Vurture, G. W. et al. GenomeScope: fast reference-free genome profiling from short reads. Bioinform. 33(14), 2202–2204 (2017).
Yan, Q. et al. The elephant grass (Cenchrus purpureus) genome provides insights into anthocyanidin accumulation and fast growth. Mol Ecol Resour. 21(2), 526–542 (2021).
Zhang, B. et al. A high-quality haplotype-resolved genome of common bermudagrass (Cynodon dactylon L.) provides insights into polyploid genome stability and prostrate growth. Front. Plant Sci. 13, 890980 (2022).
Doležel, J., Greilhuber, J. & Suda, J. Estimation of nuclear DNA content in plants using flow cytometry. Nat. Protoc. 2(9), 2233–2244 (2007).
Kolmogorov, M., Yuan, J., Lin, Y. & Pevzner, P. A. Assembly of long, error-prone reads using repeat graphs. Nat Biotechnol. 37, 540–546 (2019).
Vaser, R., Sović, I., Nagarajan, N. & Šikić, M. Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res. 27(5), 737–746 (2017).
Alonge, M. et al. Automated assembly scaffolding using RagTag elevates a new tomato system for high-throughput genome editing. Genome Biol. 23(1), 1–19 (2022).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinform. 26(5), 589–595 (2010).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PloS One 9(11), e112963 (2014).
Kovaka, S. et al. Transcriptome assembly from long-read RNA-seq alignments with StringTie2. Genome Biol. 20, 1–13 (2019).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29(7), 644–652 (2011).
Novák, P., Neumann, P. & Macas, J. Global analysis of repetitive DNA from unassembled sequence reads using RepeatExplorer2. Nat. Protoc. 15(11), 3745–3776 (2020).
Neumann, P., Novák, P., Hoštáková, N. & Macas, J. Systematic survey of plant LTR-retrotransposons elucidates phylogenetic relationships of their polyprotein domains and provides a reference for element classification. Mob. DNA 10, 1–17 (2019).
Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl. Acad. Sci. USA 117(17), 9451–9457 (2020).
Smith, A., Hubley, R., & Green, P. RepeatMasker Open-4.0. RepeatMasker Open-4.0 (2013).
Kapusta, A., Suh, A. & Feschotte, C. Dynamics of genome size evolution in birds and mammals. Proc. Natl. Acad. Sci. USA 114(8), E1460–E1469 (2017).
Melters, D. P. et al. Comparative analysis of tandem repeats from hundreds of species reveals unique insights into centromere evolution. Genome Biol. 14(1), 1–20 (2013).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215(3), 403–410 (1990).
Salson, M. et al. An improved assembly of the pearl millet reference genome using Oxford Nanopore long reads and optical mapping. G3-Genes, Genom. Genet. 13(5), jkad051 (2023).
Lin, Y. et al. quarTeT: a telomere-to-telomere toolkit for gap-free genome assembly and centromeric repeat identification. Hortic. Res 10(8), uhad127 (2023).
Holt, C. & Yandell, M. MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinform. 12(1), 1–14 (2011).
Slater, G. S. C. & Birney, E. Automated generation of heuristics for biological sequence comparison. BMC Bioinform. 6, 31, https://doi.org/10.1186/1471-2105-6-31 (2005).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinform. 28(23), 3150–3152 (2012).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. 34(suppl_2), W435–W439 (2006).
Korf, I. Gene finding in novel genomes. BMC Bioinform. 5(1), 1–9 (2004).
Pertea, G. & Pertea, M. GFF utilities: GffRead and GffCompare. F1000Research 9 (2020).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinform. 30(9), 1236–1240 (2014).
Seemann T. barrnap 0.9: rapid ribosomal RNA prediction. v0.9 (2018).
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25(5), 955–964 (1997).
Axtell, M. J. ShortStack: comprehensive annotation and quantification of small RNA genes. RNA 19(6), 740–751 (2013).
Fahlgren, N. & Carrington, J. C. miRNA target prediction in plants. Plant MicroRNAs: Methods and Protocols. Springer; New York, NY, USA. pp. 51–57 (2010).
NCBI Sequence Read Archive. https://identifiers.org/ncbi/insdc.sra:SRS19975480 (2024).
NCBI Sequence Read Archive. https://identifiers.org/ncbi/insdc.sra:SRS19975482 (2024).
NCBI Sequence Read Archive. https://identifiers.org/ncbi/insdc.sra:SRS19975483 (2024).
NCBI Sequence Read Archive. https://identifiers.org/ncbi/insdc.sra:SRS19975484 (2024).
NCBI GenBank. https://identifiers.org/ncbi/insdc.gca:GCA_036689595.1 (2024).
NCBI Sequence Read Archive. https://identifiers.org/ncbi/insdc.sra:SRS19975481 (2024).
NCBI Transcriptome Shotgun Assembly. https://identifiers.org/ncbi/insdc:GKQU00000000.1 (2024).
NCBI Sequence Read Archive. https://identifiers.org/ncbi/insdc.sra:SRR7347364 (2019).
NCBI Sequence Read Archive. https://identifiers.org/ncbi/insdc.sra:SRR7347365 (2019).
NCBI Sequence Read Archive. https://identifiers.org/ncbi/insdc.sra:SRR7347366 (2019).
NCBI Sequence Read Archive. https://identifiers.org/ncbi/insdc.sra:SRR7347367 (2019).
NCBI Sequence Read Archive. https://identifiers.org/ncbi/insdc.sra:SRR7347368 (2019).
NCBI Sequence Read Archive. https://identifiers.org/ncbi/insdc.sra:SRR7347369 (2019).
NCBI Transcriptome Shotgun Assembly. https://identifiers.org/ncbi/insdc:DAWXED000000000 (2024).
Astashyn, A. et al. Rapid and sensitive detection of genome contamination at scale with FCS-GX. Genome Biol. 25(1), 60 (2024).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biol. 21, 1–27 (2020).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinform. 31(19), 3210–3212 (2015).
Acknowledgements
This research was funded by the European Union’s Horizon 2020 Research and Innovation Program under the Marie Skłodowska-Curie Grant Agreements 872417 (MAD) and 101007438 (POLYPLOID). J.P.A.O., L.A.S., M.P. and S.C.P. were supported by the National Agency for Scientific and Technological Promotion (ANPCyT), Argentina (PICT-2017-1956 and PICT 2019-03414, PICT 2019-02153); CONICET, Argentina Projects PUE 22920160100043CO and PIP 11220200101680CO, and National University of Rosario, Argentina Project 80020190300021UR. E.A. and F.P. were funded by the European Union-Next Generation EU through the Ministero dell’ Università e della Ricerca (MUR), Italy with the projects PRIN 2022 (2022Z4HLLJ) and PRIN 2022 PNRR (P2022KFJB5), respectively. This work was carried out using the facilities of the CCT-Rosario Computational Center, a member of the High-Performance Computing National System (SNCAD, MincyT-Argentina), and the ISO 9001 certified IRD i-Trop HPC (South Green Platform) at IRD Montpellier, France. L.A.S., M.P., S.C.P. and J.P.A.O. are research staff members of CONICET, Argentina, and J.M.V. was supported by a PhD grant from CONICET.
Author information
Authors and Affiliations
Contributions
Conceptualization, J.O., J.M.V., J.P.A.O., S.C.P. and O.L.; Methodology, C.M., M.C.C. and L.A.S.; Software, J.O., J.M.V., M.P. and O.L.; Formal Analysis, J.O., J.M.V. and M.P.; Investigation, C.M., M.C.C., J.O., J.M.V., M.P., S.C.P. and L.A.S.; Data Curation, J.O., M.P., J.P.A.O. and O.L.; Writing – Original Draft, J.P.A.O. and O.L.; Writing – Review & Editing, C.M., E.A., F.P., M.C.C., M.P., J.M.V., J.O. and S.P.; Visualization, J.M.V., M.P., J.P.A.O.; Supervision, J.P.A.O. and O.L.; Funding Acquisition, E.A., F.P., S.C.P., M.P., J.P.A.O. and O.L. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vega, J.M., Podio, M., Orjuela, J. et al. Chromosome-scale genome assembly and annotation of Paspalum notatum Flüggé var. saurae. Sci Data 11, 891 (2024). https://doi.org/10.1038/s41597-024-03731-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-024-03731-0
- Springer Nature Limited