Abstract
Macrobenthic traits, for example feeding mode, life history, morphology, are increasingly used for determining responses of macrobenthic fauna to environmental change and influences on ecosystem functioning. Yet, trait information is scarce or non-existent in several parts of the world, such as New Zealand. This deficit makes collecting trait data a difficult and time-consuming task, limiting its potential use in trait-based assessments. Here, we present the New Zealand Trait Database (NZTD) for marine benthic invertebrates, the first comprehensive assessment of macrobenthic traits in New Zealand. The NZTD provides trait information for more than 700 macrobenthic taxa, categorised by 18 traits and 77 trait modalities. The NZTD includes five freely downloadable datasets, (1) the macrobenthic trait dataset, with outcomes from a fuzzy coding procedure, (2) the trait source information, (3) the references by taxa, (4) the full references list, and (5) the full taxa list used in the NZTD. Establishing the NZTD closes the trait knowledge gap in New Zealand and facilitates future research applying trait-based approaches to New Zealand’s coastal macrofauna.
Similar content being viewed by others
Background & Summary
Traits are properties of organisms that can be measured, usually at the individual organism level and used comparatively across species1,2,3. Traits are generally assigned based on feeding mode, life history, morphology, physiology, and behavioural characteristics of species4,5,6. In recent decades, the use of trait-based analyses has expanded, advancing our understanding of marine ecosystem functioning3,7,8,9 and how organisms are responding to environmental change (e.g. environmental gradients, anthropogenic disturbances, and climate change)3,6,10,11.
One challenge in applying trait-based approaches is the lack of knowledge and availability of species trait information. Macrobenthic fauna, defined as organisms retained on 0.5 mm mesh size, usually include large numbers of invertebrate taxa (e.g., polychaetes, crustaceans and molluscs), many of which are rare, small, cryptic, and inadequately studied. Therefore, gathering trait information from the literature or biological collections can be difficult and time-consuming3,6,12,13. Worldwide, several efforts have been made to alleviate the lack of macrobenthic trait information, for example establishing trait databases for the Arctic14, Southern Australia13, and Northwest Europe12, in addition to well-established online databases (e.g. MArLIN15, WoRMS16), and taxa-specific trait databases (e.g. Polytraits17). Despite these efforts, information about traits of macrobenthic fauna in several regions around the World is scarce or non-existent3, limiting the use of trait-based assessments that could assist conservation and management actions and increase understanding of the functioning of benthic ecosystems in those regions.
This article presents the New Zealand Trait Database (NZTD), aiming to (i) close knowledge gaps on macrobenthic trait information, and (ii) advance trait-based approaches for New Zealand. The NZTD is an open access database that followed the structure and traits presented in previous research7,13,14 for easy comparability and sharing among researchers. The NZTD provides trait information for more than 700 taxa. This is the first comprehensive assessment focusing on traits of marine macrobenthic fauna of New Zealand. Our aim was to provide a reliable source of macrobenthic trait information and facilitate further research using trait-based perspectives in New Zealand marine waters.
Methods
Data acquisition
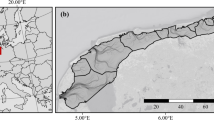
Macrobenthic data (see full macrobenthic taxa list18) were compiled from previous published research on macrobenthic fauna19,20,21,22,23,24,25,26,27,28,29,30,31,32,33 carried out by the authors in 20 different localities within New Zealand (Fig. 1, see also technical validation section). Briefly, sediment samples for assessing macrobenthic fauna were collected using a hand-held PVC corer (70 mm diameter) pushed into the sediment up to 10 cm depth. Macrobenthic fauna samples were then sieved through a 0.5 mm mesh size and preserved in 70% isopropanol. Benthic macrofauna were then sorted, identified to the lowest possible taxonomic level, and counted. The dataset encompasses a mix of records from soft sediments of New Zealand, including muddy-to-sandy substrates, vegetated (seagrass, mangrove) and unvegetated (microphytobenthos only) coastal ecosystems in estuaries and on the exposed coast.
Localities sampled across New Zealand from where information about macrobenthic fauna was retrieved for the assessment of macrobenthic traits used for the NZTD. Blue dots show the main regions sampled over time, while the green solid line shows the overall extent of macrobenthic distribution covered (ranging from 0 to 20 m water depth) in the database.
Selection of traits
The traits presented in the NZTD were selected based on a published global review3 describing their relevance for the assessment of ecosystem functioning and identified as the most commonly used traits for assessing macrobenthic fauna3. Our selection also considered traits that could be compared across studies and geographical areas, i.e., are applicable to most benthic assemblages6,7,13. In total, 18 traits and 77 trait modalities allocated across four different subject areas (e.g. biology, habitat, life-history and larval type)13 were assessed (Tables 1–4).
Trait allocation
Trait information was retrieved from different published primary literature (e.g., peer reviewed journal articles, books, thesis), secondary literature (e.g. online resources, reports), and expert knowledge (see full trait source database34), depending on the availability of information for each taxon. We categorised the sources of the trait information into four categories based on origin: (1) New Zealand literature, (2) Australian literature, (3) Overseas literature, and (4) Expert knowledge/Online resources. We followed the approach used in the South Australian Trait Database13 and allocated the trait information based on the available information for each taxon. When trait information of a particular taxon was missing, trait information from the closest phylogenetic taxon was used. For example, if no trait information was available at Species level, trait information was used from another species within the same Genus; if information was unavailable at Genus level, we considered information at Family level. Additional considerations such as taxa distribution, resemblance, and expert judgment were also applied.
Trait expression
A fuzzy coding procedure was applied to describe trait expression, scoring each of the taxa analysed depending on the affinity that a taxon displayed with a trait-modality13,35,36,37. Our fuzzy coding used a scoring range from 0–1, with 0 being no affinity and 1 being high affinity to a trait. For example, coding the trait ‘Feeding mode’ for Amalda novaezelandiae (Gastropoda), considered that A. novaezelandiae is a predatory species, however it also exhibits scavenger feeding, giving a fuzzy coding of 0.5 as predator, and 0.5 as scavenger, completing the full allocation of 1 for the feeding mode trait.
Data Records
The New Zealand Trait Database (NZTD) and related datasets are available in xlsx-formatted files and can be freely viewed and downloaded from the repository Figshare38 and National Institute of Water and Atmospheric Research (NIWA) website39. Five csv files are available for download: (1) The New Zealand Trait Database of marine benthic invertebrates38, (2) the trait information sources34, (3) the references by taxa dataset40, (4) the full references list41, and (5) the full taxa list18 used in the NZTD.
Taxa included
In total, we provide trait information for 702 taxa18. Different levels of taxonomic identification were assessed, 225 at Species level, 254 at Genus level, 139 at Family level, and the remaining 28 taxa at higher levels (Order, Class, or Phyla; Fig. 2a). The phylum with most records was Mollusca (244 records, 35% of all taxa), followed by Annelida (224 records, 32% of all taxa) and Arthropoda (178 records, 25% of all taxa), with the remaining 8% of taxa belonging to 15 different phyla (Fig. 2b).
Trait sources
Trait information was retrieved from several sources and a database containing this information was created for easy interpretation and useability34. Including all the traits assessed, 90% of the information was retrieved from primary and secondary sources, which included 45% from New Zealand literature, 39% from Australian literature and 7% from overseas literature. The remaining 9% of information was obtained from reputable resources online and expert knowledge34. Yet, we found that the source of trait information differed between types of traits (Fig. 3a). Across taxonomic levels, most of the trait information retrieved was available at the Family (41%), Genus (30%), and Species (21%) levels, with proportionally less at the Order/Class levels (6%; Fig. 3b). It was also evident that the traits larval type, life span, reproductive frequency and technique are less studied for the New Zealand macrobenthic fauna.
Technical Validation
The New Zealand Trait Database (NZTD) is based on macrobenthic fauna datasets previously validated, used, and published in different articles (e.g.19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,) and official reports, in which macrobenthic fauna were used to understand ecosystem functioning and to evaluate responses to environmental change.
Trait information was compiled using the same approach as for the SAMT database13, i.e., the NZTD retrieved information from the most reliable sources available with an accompanying dataset presenting all the references used for each taxon (trait references by taxa40; full references list41). In addition, the NZTD delivered a detailed dataset showing the taxonomic resolution of the trait information used for each taxa34.
Usage Notes
The New Zealand Trait Database (NZTD) can be freely viewed and downloaded from the repository Figshare38 and NIWA website39. The information allocated on the Figshare repository is a static version of the data last reviewed on July 2023, further updates will be released in the same Figshare project, but the DOI could be different. A report record will be also included to control further updates. Yet, the datasets available on the NIWA website are dynamically updated.
The NZTD is an ongoing project, with continuous updates and refinements as additional taxa and trait information becomes available. Future updated releases will be published in the same host repositories38,39, seeking to keep the structure of NZTD as simple as possible, avoiding complexity, redundancy, and duplication between traits as it expands to include more taxa and traits. As the NZTD evolves, we strongly suggest users to approach the database with awareness of its limitations of available taxonomic and trait-based information, ongoing changes to taxonomic nomenclature, trait information, and trait classification. Future developments may also include an expanded trait database encompassing New Zealand and Australian macrobenthic fauna.
Code availability
This research did not use or generate any coding to present the data described in the manuscript.
References
Petchey, O. L. & Gaston, K. J. Functional diversity: back to basics and looking forward. Ecology Letters 9, 741–758, https://doi.org/10.1111/j.1461-0248.2006.00924.x (2006).
Weiss, K. C. B. & Ray, C. A. Unifying functional trait approaches to understand the assemblage of ecological communities: synthesizing taxonomic divides. Ecography 42, 2012–2020, https://doi.org/10.1111/ecog.04387 (2019).
Lam-Gordillo, O., Baring, R. & Dittmann, S. Ecosystem functioning and functional approaches on marine macrobenthic fauna: A research synthesis towards a global consensus. Ecological Indicators 115, https://doi.org/10.1016/j.ecolind.2020.106379 (2020).
Díaz, S. & Cabido, M. Vive la difference: plant functional diversity matters to ecosystem. Trends in Ecology and Evolution 16, 646–655, https://doi.org/10.1016/S0169-5347(01)02283-2 (2001).
Reiss, J., Bridle, J. R., Montoya, J. M. & Woodward, G. Emerging horizons in biodiversity and ecosystem functioning research. Trends in Ecology and Evolution 24, 505–514, https://doi.org/10.1016/j.tree.2009.03.018 (2009).
Degen, R. et al. Trait-based approaches in rapidly changing ecosystems: A roadmap to the future polar oceans. Ecological Indicators 91, 722–736, https://doi.org/10.1016/j.ecolind.2018.04.050 (2018).
Costello, M. J. et al. Biological and ecological traits of marine species. PeerJ 3, e1201, https://doi.org/10.7717/peerj.1201 (2015).
Cano‐Barbacil, C., Radinger, J. & García‐Berthou, E. Reliability analysis of fish traits reveals discrepancies among databases. Freshwater Biology 65, 863–877, https://doi.org/10.1111/fwb.13469 (2020).
Tavares, D. C., Moura, J. F., Acevedo-Trejos, E. & Merico, A. Traits Shared by Marine Megafauna and Their Relationships With Ecosystem Functions and Services. Frontiers in Marine Science 6, https://doi.org/10.3389/fmars.2019.00262 (2019).
Beauchard, O., Veríssimo, H., Queirós, A. M. & Herman, P. M. J. The use of multiple biological traits in marine community ecology and its potential in ecological indicator development. Ecological Indicators 76, 81–96, https://doi.org/10.1016/j.ecolind.2017.01.011 (2017).
Dissanayake, N. G., Frid, C. L. J., Drylie, T. P. & Caswell, B. A. Ecological functioning of mudflats: global analysis reveals both regional differences and widespread conservation of functioning. Marine Ecology Progress Series 604, 1–20, https://doi.org/10.3354/meps12728 (2018).
Clare, D. S. et al. Biological traits of marine benthic invertebrates in Northwest Europe. Scientific Data 9, 339, https://doi.org/10.1038/s41597-022-01442-y (2022).
Lam-Gordillo, O., Baring, R. & Dittmann, S. Establishing the South Australian Macrobenthic Traits (SAMT) database: A trait classification for functional assessments. Ecol Evol 10, 14372–14387, https://doi.org/10.1002/ece3.7040 (2020).
Degen, R. & Faulwetter, S. The Arctic Traits Database – a repository of Arctic benthic invertebrate traits. Earth System Science Data 11, 301–322, https://doi.org/10.5194/essd-11-301-2019 (2019).
MArLIN. The Marine Life Information Network https://www.marlin.ac.uk (2023).
WoRMS. World Register of Marine Species https://www.marinespecies.org/traits/index.php (2023).
Faulwetter, S. et al. Polytraits: A database on biological traits of marine polychaetes. Biodivers Data J, e1024, https://doi.org/10.3897/BDJ.2.e1024 (2014).
Lam-Gordillo, O., Lohrer, A. M., Hewitt, J. E. & Dittmann, S. Full Macrobenthic taxa list. Figshare https://doi.org/10.6084/m9.figshare.21939620.v2 (2023).
Hewitt, J., Gammal, J. & Ellis, J. Assessing ecological health in areas with limited data by using biological traits. Mar Pollut Bull 181, 113900, https://doi.org/10.1016/j.marpolbul.2022.113900 (2022).
de Juan, S., Thrush, S. F., Hewitt, J. E., Halliday, J. & Lohrer, A. M. Cumulative degradation in estuaries: contribution of individual species to community recovery. Marine Ecology Progress Series 510, 25–38, https://doi.org/10.3354/meps10904 (2014).
Townsend, M., Lohrer, A. M., Rodil, I. F. & Chiaroni, L. D. The targeting of large-sized benthic macrofauna by an invasive portunid predator: evidence from a caging study. Biological Invasions 17, 231–244, https://doi.org/10.1007/s10530-014-0722-1 (2014).
Thrush, S. F. et al. Cumulative stressors reduce the self-regulating capacity of coastal ecosystems. Ecological Applications 31, e02223, https://doi.org/10.1002/eap.2223 (2021).
Thrush, S. F., Hewitt, J. E., Gibbs, M., Lundquist, C. & Norkko, A. Functional Role of Large Organisms in Intertidal Communities: Community Effects and Ecosystem Function. Ecosystems 9, 1029–1040, https://doi.org/10.1007/s10021-005-0068-8 (2006).
Pratt, D. R. et al. Detecting Subtle Shifts in Ecosystem Functioning in a Dynamic Estuarine Environment. PLoS One 10, e0133914, https://doi.org/10.1371/journal.pone.0133914 (2015).
O’Meara, T. A., Hewitt, J. E., Thrush, S. F., Douglas, E. J. & Lohrer, A. M. Denitrification and the Role of Macrofauna Across Estuarine Gradients in Nutrient and Sediment Loading. Estuaries and Coasts 43, 1394–1405, https://doi.org/10.1007/s12237-020-00728-x (2020).
Mangan, S. et al. Shady business: the darkening of estuaries constrains benthic ecosystem function. Marine Ecology Progress Series 647, 33–48, https://doi.org/10.3354/meps13410 (2020).
Mangan, S. et al. Resilience and Species Accumulation across Seafloor Habitat Transitions in a Northern New Zealand Harbour. Diversity 14, https://doi.org/10.3390/d14110998 (2022).
Gammal, J. et al. Stressors Increase the Impacts of Coastal Macrofauna Biodiversity Loss on Ecosystem Multifunctionality. Ecosystems https://doi.org/10.1007/s10021-022-00775-4 (2022).
Gammal, J., Järnström, M., Bernard, G., Norkko, J. & Norkko, A. Environmental Context Mediates Biodiversity–Ecosystem Functioning Relationships in Coastal Soft-sediment Habitats. Ecosystems 22, 137–151, https://doi.org/10.1007/s10021-018-0258-9 (2018).
Douglas, E. J., Lohrer, A. M. & Pilditch, C. A. Biodiversity breakpoints along stress gradients in estuaries and associated shifts in ecosystem interactions. Scientific Reports 9, 17567, https://doi.org/10.1038/s41598-019-54192-0 (2019).
Douglas, E. J. et al. Macrofaunal Functional Diversity Provides Resilience to Nutrient Enrichment in Coastal Sediments. Ecosystems 20, 1324–1336, https://doi.org/10.1007/s10021-017-0113-4 (2017).
Douglas, E. J., Hewitt, J., Lohrer, A. M. & Stephenson, F. Changing intra- and interspecific interactions across sedimentary and environmental stress gradients. Ecosphere 14, e4373, https://doi.org/10.1002/ecs2.4373 (2023).
Gammal, J. et al. Stressors Increase the Impacts of Coastal Macrofauna Biodiversity Loss on Ecosystem Multifunctionality. Ecosystems 26, 539–552, https://doi.org/10.1007/s10021-022-00775-4 (2023).
Lam-Gordillo, O., Lohrer, A. M., JE, H. & Dittmann, S. Trait Source dataset. Figshare https://doi.org/10.6084/m9.figshare.21939632 (2023).
Bremner, J. Species’ traits and ecological functioning in marine conservation and management. Journal of Experimental Marine Biology and Ecology 366, 37–47, https://doi.org/10.1016/j.jembe.2008.07.007 (2008).
Bremner, J., Rogers, S. & Frid, C. Methods for describing ecological functioning of marine benthic assemblages using biological traits analysis (BTA). Ecological Indicators 6, 609–622, https://doi.org/10.1016/j.ecolind.2005.08.026 (2006).
Chevenet, F., Doledec, S. & Chessel, D. A fuzzy coding approach for the analysis of long-term ecological data. Freshwater Biology 3, 295–309, https://doi.org/10.1111/j.1365-2427.1994.tb01742.x (1994).
Lam-Gordillo, O., Lohrer, A. M., JE, H. & Dittmann, S. The New Zealand Trait Database (NZTD). Figshare https://doi.org/10.6084/m9.figshare.21939647.v2 (2023).
NIWA. The New Zealand Trait Database (NZTD) https://niwa.co.nz/coasts-and-estuaries/research-projects/NZTD (2023).
Lam-Gordillo, O., Lohrer, A. M., JE, H. & Dittmann, S. Trait references by taxa list. Figshare https://doi.org/10.6084/m9.figshare.21939659.v2 (2023).
Lam-Gordillo, O., Lohrer, A. M., JE, H. & Dittmann, S. Full references list. Figshare https://doi.org/10.6084/m9.figshare.21939671.v3 (2023).
Kristensen, E. et al. What is bioturbation? The need for a precise definition for fauna in aquatic sciences. Marine Ecology Progress Series 446, 285–302, https://doi.org/10.3354/meps09506 (2012).
Queiros, A. M. et al. A bioturbation classification of European marine infaunal invertebrates. Ecology and Evolution 3, 3958–3985, https://doi.org/10.1002/ece3.769 (2013).
Liu, K. et al. Functional trait composition and diversity patterns of marine macrobenthos across the Arctic Bering Sea. Ecological Indicators 102, 673–685, https://doi.org/10.1016/j.ecolind.2019.03.029 (2019).
van der Linden, P. et al. A biological trait approach to assess the functional composition of subtidal benthic communities in an estuarine ecosystem. Ecological Indicators 20, 121–133, https://doi.org/10.1016/j.ecolind.2012.02.004 (2012).
van der Linden, P. et al. Functional changes in polychaete and mollusc communities in two tropical estuaries. Estuarine, Coastal and Shelf Science 187, 62–73, https://doi.org/10.1016/j.ecss.2016.12.019 (2017).
Acknowledgements
We gratefully acknowledge for the assistance in collating trait information to Simon Thrush (University of Auckland), Tom Ysebaert (NIOZ), John Gray, Greig Funnell (Department of Conservation), Carolyn Lundquist (NIWA), Sarah Hailes (NIWA), Samantha Parkes (NIWA), Barry Greenfield (NIWA), Emily Douglas (NIWA), Casper Kraan, Ivan Rodil (Universidad de Cadiz), Silvia de Juan (Institut de Ciencies del Mar), Megan Carbines (Auckland Council), Hannah Jones (Ministry for Environment), and others.
Author information
Authors and Affiliations
Contributions
O.L.-G. collected and analysed the data, developed the outline for the manuscript, and wrote the manuscript. D.L. contributed to the collection of data and writing of the manuscript. J.H. contributed to the collection of data and provided substantial comments on the manuscript. S.D. reviewed, edited, and provided substantial comments on the manuscript. All authors contributed to the article and approved the submitted version.
Corresponding author
Ethics declarations
Competing interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lam-Gordillo, O., Lohrer, A.M., Hewitt, J. et al. NZTD - The New Zealand Trait Database for shallow-water marine benthic invertebrates. Sci Data 10, 502 (2023). https://doi.org/10.1038/s41597-023-02414-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02414-6
- Springer Nature Limited