Abstract
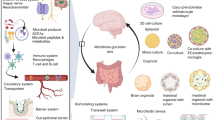
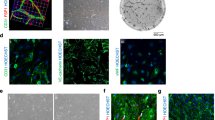
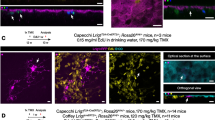
Angiogenesis and neurogenesis are functionally interconnected during brain development. However, the study of the vasculature has trailed other brain cell types because they are delicate and of low abundance. Here we describe a protocol extension to purify prenatal human brain endothelial and mural cells with FACS and utilize them in downstream applications, including transcriptomics, culture and organoid transplantation. This approach is simple, efficient and generates high yields from small amounts of tissue. When the experiment is completed within a 24 h postmortem interval, these healthy cells produce high-quality data in single-cell transcriptomics experiments. These vascular cells can be cultured, passaged and expanded for many in vitro assays, including Matrigel vascular tube formation, microfluidic chambers and metabolic measurements. Under these culture conditions, primary vascular cells maintain expression of cell-type markers for at least 3 weeks. Finally, we describe how to use primary vascular cells for transplantation into cortical organoids, which captures key features of neurovascular interactions in prenatal human brain development. In terms of timing, tissue processing and staining requires ~3 h, followed by an additional 3 h of FACS. The transplant procedure of primary, FACS-purified vascular cells into cortical organoids requires an additional 2 h. The time required for different transcriptomic and epigenomic protocols can vary based on the specific application, and we offer strategies to mitigate batch effects and optimize data quality. In sum, this vasculo-centric approach offers an integrated platform to interrogate neurovascular interactions and human brain vascular development.
Key points
-
This protocol extension describes the purification of prenatal human brain endothelial and mural cells with FACS and their utilization in downstream applications, including cell culture, organoid transplantation and single-cell transcriptomics.
-
This simple, efficient protocol has relatively few steps compared with other methods and uses inexpensive reagents. Robust yields of healthy vascular and perivascular cells can be obtained in 6 h.






Similar content being viewed by others
Data availability
ScRNA-seq data have been deposited at GEO and are publicly available with code GEO: PRJNA803255. In addition, the data can be downloaded at https://cells.ucsc.edu/?ds=vascular-dev. No original code was generated using this data. The flow cytometry data can be accessed at Mendeley Data (https://doi.org/10.17632/9j6chfbvxm.1). Other data produced during and/or analyzed in this study are available from the corresponding authors upon request.
References
Paredes, I., Himmels, P. & Ruiz de Almodóvar, C. Neurovascular communication during CNS development. Dev. Cell 45, 10–32 (2018).
Silva-Vargas, V., Crouch, E. E. & Doetsch, F. Adult neural stem cells and their niche: a dynamic duo during homeostasis, regeneration, and aging. Curr. Opin. Neurobiol. 23, 935–942 (2013).
Gould, D. B. et al. Mutations in Col4a1 cause perinatal cerebral hemorrhage and porencephaly. Science 308, 1167–1171 (2005).
Xu, H. et al. Maturational changes in laminin, fibronectin, collagen IV, and perlecan in germinal matrix, cortex, and white matter and effect of betamethasone. J. Neurosci. Res. 86, 1482–1500 (2008).
Garcia, F. J. et al. Single-cell dissection of the human brain vasculature. Nature 603, 893–899 (2022).
Winkler, E. A. et al. A single-cell atlas of the normal and malformed human brain vasculature. Science 375, eabi7377 (2022).
Ricci-Vitiani, L. et al. Tumour vascularization via endothelial differentiation of glioblastoma stem-like cells. Nature 468, 824–828 (2010).
Wang, R. et al. Glioblastoma stem-like cells give rise to tumour endothelium. Nature 468, 829–833 (2010).
Yang, A. C. et al. A human brain vascular atlas reveals diverse mediators of Alzheimer’s risk. Nature 603, 885–892 (2022).
Daneman, R. & Prat, A. The blood–brain barrier. Cold Spring Harb. Perspect. Biol. 7, a020412 (2015).
Armulik, A. et al. Pericytes regulate the blood–brain barrier. Nature 468, 557–561 (2010).
Daneman, R., Zhou, L., Kebede, A. A. & Barres, B. A. Pericytes are required for blood–brain barrier integrity during embryogenesis. Nature 468, 562–566 (2010).
Ayloo, S. et al. Pericyte-to-endothelial cell signaling via vitronectin–integrin regulates blood–CNS barrier. Neuron 110, 1641–1655.e6 (2022).
Crouch, E. E. et al. Ensembles of endothelial and mural cells promote angiogenesis in prenatal human brain. Cell 185, 3753–3769.e18 (2022).
Kisler, K. et al. Pericyte degeneration leads to neurovascular uncoupling and limits oxygen supply to brain. Nat. Neurosci. 20, 406–416 (2017).
Hill, R. A. et al. Regional blood flow in the normal and ischemic brain is controlled by arteriolar smooth muscle cell contractility and not by capillary pericytes. Neuron 87, 95–110 (2015).
Hall, C. N. et al. Capillary pericytes regulate cerebral blood flow in health and disease. Nature 508, 55–60 (2014).
Parker, K. R. et al. Single-cell analyses identify brain mural cells expressing CD19 as potential off-tumor targets for CAR-T immunotherapies. Cell 183, 126–142.e17 (2020).
Duan, L. et al. PDGFRβ cells rapidly relay inflammatory signal from the circulatory system to neurons via chemokine CCL2. Neuron 100, 183–200.e8 (2018).
Crouch, E. E., Liu, C., Silva-Vargas, V. & Doetsch, F. Regional and stage-specific effects of prospectively purified vascular cells on the adult V-SVZ neural stem cell lineage. J. Neurosci. 35, 4528–4539 (2015).
Crouch, E. E. & Doetsch, F. FACS isolation of endothelial cells and pericytes from mouse brain microregions. Nat. Protoc. 13, 738–751 (2018).
Nowakowski, T. J. et al. Spatiotemporal gene expression trajectories reveal developmental hierarchies of the human cortex. Science 358, 1318–1323 (2017).
Crisan, M. et al. A perivascular origin for mesenchymal stem cells in multiple human organs. Cell Stem Cell 3, 301–313 (2008).
Lui, J. H. et al. Radial glia require PDGFD–PDGFRβ signalling in human but not mouse neocortex. Nature 515, 264–268 (2014).
Seandel, M. et al. Generation of a functional and durable vascular niche by the adenoviral E4ORF1 gene. Proc. Natl Acad. Sci. USA 105, 19288–19293 (2008).
Watson, C. A., Camera-Benson, L., Palmer-Crocker, R. & Pober, J. S. Variability among human umbilical vein endothelial cultures. Science 268, 447–448 (1995).
Sarkanen, J.-R. et al. Intra-laboratory pre-validation of a human cell based in vitro angiogenesis assay for testing angiogenesis modulators. Front. Pharmacol. 1, 147 (2010).
Paredes, M. F. et al. Nests of dividing neuroblasts sustain interneuron production for the developing human brain. Science 375, eabk2346 (2022).
Sun, X.-Y. et al. Generation of vascularized brain organoids to study neurovascular interactions. eLife 11, e76707 (2022).
Mansour, A. A. et al. An in vivo model of functional and vascularized human brain organoids. Nat. Biotechnol. 36, 432–441 (2018).
Cakir, B. et al. Engineering of human brain organoids with a functional vascular-like system. Nat. Methods 16, 1169–1175 (2019).
Bhaduri, A. et al. Cell stress in cortical organoids impairs molecular subtype specification. Nature 578, 142–148 (2020).
Chen, J. et al. Proinflammatory milieu disrupts homeostatic microglia-vascular interactions to promote germinal matrix hemorrhage. Cell Sneak Peak https://doi.org/10.2139/ssrn.4260494 (2022).
Polacheck, W. J., Kutys, M. L., Tefft, J. B. & Chen, C. S. Microfabricated blood vessels for modeling the vascular transport barrier. Nat. Protoc. 14, 1425–1454 (2019).
Nzou, G. et al. Human cortex spheroid with a functional blood brain barrier for high-throughput neurotoxicity screening and disease modeling. Sci. Rep. 8, 7413 (2018).
Cho, C.-F. et al. Blood–brain-barrier spheroids as an in vitro screening platform for brain-penetrating agents. Nat. Commun. 8, 15623 (2017).
Wang, L. et al. A human three-dimensional neural-perivascular ‘assembloid’ promotes astrocytic development and enables modeling of SARS-CoV-2 neuropathology. Nat. Med. 27, 1600–1606 (2021).
Sheikh, B. N. et al. Systematic identification of cell-cell communication networks in the developing brain. iScience 21, 273–287 (2019).
Spitzer, D. et al. A flow cytometry-based protocol for syngenic isolation of neurovascular unit cells from mouse and human tissues. Nat. Protoc. https://doi.org/10.1038/s41596-023-00805-y (2023).
Pebworth, M.-P., Ross, J., Andrews, M., Bhaduri, A. & Kriegstein, A. R. Human intermediate progenitor diversity during cortical development. Proc. Natl Acad. Sci. Usa. 118, e2019415118 (2021).
Eze, U. C., Bhaduri, A., Haeussler, M., Nowakowski, T. J. & Kriegstein, A. R. Single-cell atlas of early human brain development highlights heterogeneity of human neuroepithelial cells and early radial glia. Nat. Neurosci. 24, 584–594 (2021).
Paşca, A. M. et al. Functional cortical neurons and astrocytes from human pluripotent stem cells in 3D culture. Nat. Methods 12, 671–678 (2015).
Kimble, A. L. et al. A method for rapid flow-cytometric isolation of endothelial nuclei and RNA from archived frozen brain tissue. Lab. Invest. 102, 204–211 (2022).
Kadoshima, T. et al. Self-organization of axial polarity, inside-out layer pattern, and species-specific progenitor dynamics in human ES cell-derived neocortex. Proc. Natl Acad. Sci. USA 110, 20284–20289 (2013).
Lancaster, M. A. et al. Cerebral organoids model human brain development and microcephaly. Nature 501, 373–379 (2013).
Lancaster, M. A. & Knoblich, J. A. Generation of cerebral organoids from human pluripotent stem cells. Nat. Protoc. 9, 2329–2340 (2014).
Spitzer, D. et al. Profiling the neurovascular unit unveils detrimental effects of osteopontin on the blood-brain barrier in acute ischemic stroke. Acta Neuropathol. 144, 305–337 (2022).
Foster, B., Prussin, C., Liu, F., Whitmire, J. K. & Whitton, J. L. Detection of intracellular cytokines by flow cytometry. Curr. Protoc. Immunol. 6, 6.24.1–6.24.21 (2007).
Walls, P. L. L. et al. Quantifying the potential for bursting bubbles to damage suspended cells. Sci. Rep. 7, 15102 (2017).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
Acknowledgements
The authors thank E. Winkler and A. Yang for their discussion on the strengths and challenges of different methods to purify human brain vascular cells (Table 1). This study has been supported by the Eli and Edythe Broad Regeneration Medicine and Stem Cell Fellowship, Pediatric Scientist Development Program (5K12HD000850-34), UCSF Physician Scientist Scholars Program (PSSP), American Heart Association Career Development Grant 857876, CIRM Alpha Stem Cell Clinic Fellowship and National Institutes of Health (NIH) grants K08 NS116161 to E.E.C., NIH K99/R00 NS111731 to A.B., NIH K99/R00 MH125329 to M.G.A., and California Institute of Regenerative Medicine (CIRM) Bridges Science Master’s Fellowship (SFSU EDUC2-12693) to E.J.V., P01 NS083513 and U01 MH105989 to E.J.H. The UCSF Flow Cytometry Core is supported by DRC Center Grant NIH P30 DK063720.
Author information
Authors and Affiliations
Contributions
E.E.C. and E.J.H. conceived the project and designed the experiments. E.E.C. performed FACS experiments, 2D culture experiments and organoid transplants. L.D. performed FACS experiments with passaged cells and organoid transplants. E.J.V. performed FACS experiments. J.O., K.W.-P., J.C., T.J. and A.B. performed scRNA-seq bioinformatics. L.D., A.B. and M.G.A. performed and designed organoid experiments. E.E.C. and E.J.H. wrote the manuscript with inputs from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Related links
Key reference using this protocol
Crouch, E. E. et al. Cell 185, 3753–3769.e18 (2022): https://doi.org/10.1016/j.cell.2022.09.004
This protocol is an extension to: Nat. Protoc. 13, 738–751 (2018): https://doi.org/10.1038/nprot.2017.158
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Crouch, E.E., Diafos, L.N., Valenzuela, E.J. et al. Profiling human brain vascular cells using single-cell transcriptomics and organoids. Nat Protoc 19, 603–628 (2024). https://doi.org/10.1038/s41596-023-00929-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00929-1
- Springer Nature Limited