Abstract
Cyclic nucleotide–gated (CNG) channels convert cyclic nucleotide (CN) binding and unbinding into electrical signals in sensory receptors and neurons. The molecular conformational changes underpinning ligand activation are largely undefined. We report both closed- and open-state atomic cryo-EM structures of a full-length Caenorhabditis elegans cyclic GMP−activated channel TAX-4, reconstituted in lipid nanodiscs. These structures, together with computational and functional analyses and a mutant channel structure, reveal a double-barrier hydrophobic gate formed by two S6 amino acids in the central cavity. cGMP binding produces global conformational changes that open the cavity gate located ~52 Å away but do not alter the structure of the selectivity filter—the commonly presumed activation gate. Our work provides mechanistic insights into the allosteric gating and regulation of CN-gated and nucleotide-modulated channels and CNG channel−related channelopathies.







Similar content being viewed by others
Data availability
The cryo-EM density maps have been deposited in the Electron Microscopy Data Bank under accession numbers EMD-21649 for cGMP-unbound WT TAX-4, EMD-21650 for cGMP-bound WT TAX-4 and EMD-21651 for cGMP-unbound F403V V407A mutant TAX-4. The coordinates of the atomic models have been deposited in the Protein Data Bank under accession numbers PDB 6WEJ for cGMP-unbound WT TAX-4, PDB 6WEK for cGMP-bound WT TAX-4 and PDB 6WEL for cGMP-unbound F403V V407A mutant TAX-4. Source data for Fig. 4a,b and Extended Data Figs. 1b, 3b and 4b are available with the paper online.
References
Kaupp, U. B. & Seifert, R. Cyclic nucleotide-gated ion channels. Physiol. Rev. 82, 769–824 (2002).
Zagotta, W. N. & Siegelbaum, S. A. Structure and function of cyclic nucleotide–gated channels. Annu. Rev. Neurosci. 19, 235–263 (1996).
Varnum, M. D. & Dai, G. in The Hankbook of Ion Channels (eds J. Zheng & M.C. Trudeau) 361−382 (CRC Press, 2015).
Michalakis, S., Becirovic, E. & Biel, M. Retinal cyclic nucleotide-gated channels: from pathophysiology to therapy. Int. J. Mol. Sci. 19, 749 (2018).
Fesenko, E. E., Kolesnikov, S. S. & Lyubarsky, A. L. Induction by cyclic GMP of cationic conductance in plasma membrane of retinal rod outer segment. Nature 313, 310–313 (1985).
Nakamura, T. & Gold, G. H. A cyclic nucleotide-gated conductance in olfactory receptor cilia. Nature 325, 442–444 (1987).
Kaupp, U. B. et al. Primary structure and functional expression from complementary DNA of the rod photoreceptor cyclic GMP-gated channel. Nature 342, 762–766 (1989).
Dhallan, R. S., Yau, K. W., Schrader, K. A. & Reed, R. R. Primary structure and functional expression of a cyclic nucleotide-activated channel from olfactory neurons. Nature 347, 184−187 (1990).
Goulding, E. H., Tibbs, G. R., Liu, D. & Siegelbaum, S. A. Role of H5 domain in determining pore diameter and ion permeation through cyclic nucleotide-gated channels. Nature 364, 61−64 (1993).
Li, M. et al. Structure of a eukaryotic cyclic-nucleotide-gated channel. Nature 542, 60–65 (2017).
Gordon, S. E. & Zagotta, W. N. A histidine residue associated with the gate of the cyclic nucleotide-activated channels in rod photoreceptors. Neuron 14, 177–183 (1995).
Gordon, S. E. & Zagotta, W. N. Localization of regions affecting an allosteric transition in cyclic nucleotide-activated channels. Neuron 14, 857–864 (1995).
Brown, R. L., Snow, S. D. & Haley, T. L. Movement of gating machinery during the activation of rod cyclic nucleotide-gated channels. Biophys. J. 75, 825–833 (1998).
Zong, X., Zucker, H., Hofmann, F. & Biel, M. Three amino acids in the C-linker are major determinants of gating in cyclic nucleotide-gated channels. EMBO J. 17, 353−362 (1998).
Zhou, L., Olivier, N. B., Yao, H., Young, E. C. & Siegelbaum, S. A. A conserved tripeptide in CNG and HCN channels regulates ligand gating by controlling C-terminal oligomerization. Neuron 44, 823−834 (2004).
Paoletti, P., Young, E. C. & Siegelbaum, S. A. C-linker of cyclic nucleotide–gated channels controls coupling of ligand binding to channel gating. J. Gen. Physiol. 113, 17–34 (1999).
Flynn, G. E. & Zagotta, W. N. Conformational changes in S6 coupled to the opening of cyclic nucleotide-gated channels. Neuron 30, 689–698 (2001).
Contreras, J. E. & Holmgren, M. Access of quaternary ammonium blockers to the internal pore of cyclic nucleotide-gated channels: implications for the location of the gate. J. Gen. Physiol. 127, 481−494 (2006).
Contreras, J. E., Srikumar, D. & Holmgren, M. Gating at the selectivity filter in cyclic nucleotide-gated channels. Proc. Natl Acad. Sci. USA 105, 3310−3314 (2008).
Sun, Z. P., Akabas, M. H., Goulding, E. H., Karlin, A. & Siegelbaum, S. A. Exposure of residues in the cyclic nucleotide–gated channel pore: P region structure and function in gating. Neuron 16, 141–149 (1996).
Becchetti, A. & Roncaglia, P. Cyclic nucleotide-gated channels: intra- and extracellular accessibility to Cd2+ of substituted cysteine residues within the P-loop. Pflug. Arch. 440, 556–565 (2000).
James, Z. M. & Zagotta, W. N. Structural insights into the mechanisms of CNBD channel function. J. Gen. Physiol. 150, 225−244 (2018).
Flynn, G. E., Johnson, J. P., Jr. & Zagotta, W. N. Cyclic nucleotide-gated channels: shedding light on the opening of a channel pore. Nat Rev Neurosci. 2, 643−651 (2001).
Cukkemane, A., Seifert, R. & Kaupp, U. B. Cooperative and uncooperative cyclic-nucleotide-gated ion channels. Trends Biochem. Sci. 36, 55−64 (2011).
Clayton, G. M., Altieri, S., Heginbotham, L., Unger, V. M. & Morais-Cabral, J. H. Structure of the transmembrane regions of a bacterial cyclic nucleotide-regulated channel. Proc. Natl Acad. Sci. USA 105, 1511−1515 (2008).
James, Z. M. et al. CryoEM structure of a prokaryotic cyclic nucleotide-gated ion channel. Proc. Natl Acad. Sci. USA 114, 4430–4435 (2017).
Rheinberger, J., Gao, X., Schmidpeter, P. A. & Nimigean, C. M. Ligand discrimination and gating in cyclic nucleotide-gated ion channels from apo and partial agonist-bound cryo-EM structures. Elife 7, e39775 (2018).
Komatsu, H., Mori, I., Rhee, J. S., Akaike, N. & Ohshima, Y. Mutations in a cyclic nucleotide–gated channel lead to abnormal thermosensation and chemosensation in C. elegans. Neuron 17, 707–718 (1996).
Komatsu, H. et al. Functional reconstitution of a heteromeric cyclic nucleotide-gated channel of Caenorhabditis elegans in cultured cells. Brain Res. 821, 160–168 (1999).
Klesse, G., Rao, S., Sansom, M. S. P. & Tucker, S. J. CHAP: a versatile tool for the structural and functional annotation of ion channel pores. J. Mol. Biol. 431, 3353–3365 (2019).
Rao, S., Klesse, G., Stansfeld, P. J., Tucker, S. J. & Sansom, M. S. P. A heuristic derived from analysis of the ion channel structural proteome permits the rapid identification of hydrophobic gates. Proc. Natl Acad. Sci. USA 116, 13989−13995 (2019).
Trick, J. L. et al. Functional annotation of ion channel structures by molecular simulation. Structure 24, 2207–2216 (2016).
Doyle, D. A. et al. The structure of the potassium channel: molecular basis of K+ conduction and selectivity. Science 280, 69–77 (1998).
Cao, E., Liao, M., Cheng, Y. & Julius, D. TRPV1 structures in distinct conformations reveal activation mechanisms. Nature 504, 113−118 (2013).
Lee, C. H. & MacKinnon, R. Structures of the human HCN1 hyperpolarization-activated channel. Cell 168, 111–120.e11 (2017).
Flynn, G. E. & Zagotta, W. N. A cysteine scan of the inner vestibule of cyclic nucleotide-gated channels reveals architecture and rearrangement of the pore. J. Gen. Physiol. 121, 563−582 (2003).
Zagotta, W. N. et al. Structural basis for modulation and agonist specificity of HCN pacemaker channels. Nature 425, 200–205 (2003).
Saponaro, A. et al. Structural basis for the mutual antagonism of cAMP and TRIP8b in regulating HCN channel function. Proc. Natl Acad. Sci. USA 111, 14577–14582 (2014).
Xu, X., Vysotskaya, Z. V., Liu, Q. & Zhou, L. Structural basis for the cAMP-dependent gating in the human HCN4 channel. J. Biol. Chem. 285, 37082−37091 (2010).
Lolicato, M. et al. Tetramerization dynamics of C-terminal domain underlies isoform-specific cAMP gating in hyperpolarization-activated cyclic nucleotide-gated channels. J. Biol. Chem. 286, 44811–44820 (2011).
Taraska, J. W., Puljung, M. C., Olivier, N. B., Flynn, G. E. & Zagotta, W. N. Mapping the structure and conformational movements of proteins with transition metal ion FRET. Nat. Methods 6, 532−537 (2009).
Altieri, S. L. et al. Structural and energetic analysis of activation by a cyclic nucleotide binding domain. J. Mol. Biol. 381, 655–669 (2008).
Goldschen-Ohm, M. P. et al. Structure and dynamics underlying elementary ligand binding events in human pacemaking channels. Elife 5, e20797 (2016).
Kowal, J. et al. Ligand-induced structural changes in the cyclic nucleotide-modulated potassium channel MloK1. Nat. Commun. 5, 3106 (2014).
Marchesi, A. et al. An iris diaphragm mechanism to gate a cyclic nucleotide-gated ion channel. Nat. Commun. 9, 3978 (2018).
Kim, D. M. & Nimigean, C. M. Voltage-gated potassium channels: a structural examination of selectivity and gating. Cold Spring Harb. Perspect. Biol. 8, a029231 (2016).
Zhou, X. et al. Cryo-EM structures of the human endolysosomal TRPML3 channel in three distinct states. Nat. Struct. Mol. Biol. 24, 1146–1154 (2017).
Fodor, A. A., Black, K. D. & Zagotta, W. N. Tetracaine reports a conformational change in the pore of cyclic nucleotide-gated channels. J. Gen. Physiol. 110, 591–600 (1997).
Fodor, A. A., Gordon, S. E. & Zagotta, W. N. Mechanism of tetracaine block of cyclic nucleotide-gated channels. J. Gen. Physiol. 109, 3–14 (1997).
Craven, K. B., Olivier, N. B. & Zagotta, W. N. C-terminal movement during gating in cyclic nucleotide-modulated channels. J. Biol. Chem. 283, 14728–14738 (2008).
Craven, K. B. & Zagotta, W. N. Salt bridges and gating in the COOH-terminal region of HCN2 and CNGA1 channels. J. Gen. Physiol. 124, 663–677 (2004).
Mazzolini, M. et al. The gating mechanism in cyclic nucleotide-gated ion channels. Sci. Reports 8, 45 (2018).
Martinez-Francois, J. R., Xu, Y. & Lu, Z. Mutations reveal voltage gating of CNGA1 channels in saturating cGMP. J. Gen. Physiol. 134, 151−164 (2009).
Mazzolini, M., Anselmi, C. & Torre, V. The analysis of desensitizing CNGA1 channels reveals molecular interactions essential for normal gating. J. Gen. Physiol. 133, 375−386 (2009).
Crary, J. I., Dean, D. M., Nguitragool, W., Kurshan, P. T. & Zimmerman, A. L. Mechanism of inhibition of cyclic nucleotide–gated ion channels by diacylglycerol. J. Gen. Physiol. 116, 755–768 (2000).
Womack, K. B. et al. Do phosphatidylinositides modulate vertebrate phototransduction? J. Neurosci. 20, 2792–2799 (2000).
Bright, S. R., Rich, E. D. & Varnum, M. D. Regulation of human cone cyclic nucleotide-gated channels by endogenous phospholipids and exogenously applied phosphatidylinositol 3,4,5-trisphosphate. Mol. Pharmacol. 71, 176−183 (2007).
Spehr, M., Wetzel, C. H., Hatt, H. & Ache, B. W. 3-phosphoinositides modulate cyclic nucleotide signaling in olfactory receptor neurons. Neuron 33, 731–739 (2002).
Zhainazarov, A. B., Spehr, M., Wetzel, C. H., Hatt, H. & Ache, B. W. Modulation of the olfactory CNG channel by Ptdlns(3,4,5)P3. J. Membr. Biol. 201, 51–57 (2004).
Gordon, S. E., Downing-Park, J., Tam, B. & Zimmerman, A. L. Diacylglycerol analogs inhibit the rod cGMP-gated channel by a phosphorylation-independent mechanism. Biophys. J. 69, 409−417 (1995).
Dai, G., Peng, C., Liu, C. & Varnum, M. D. Two structural components in CNGA3 support regulation of cone CNG channels by phosphoinositides. J. Gen. Physiol. 141, 413−430 (2013).
Ritchie, T. K. et al. Chapter 11 — Reconstitution of membrane proteins in phospholipid bilayer nanodiscs. Methods Enzymol. 464, 211–231 (2009).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Feng, X. et al. A fast and effective microfluidic spraying-plunging method for high-resolution single-particle cryo-EM. Structure 25, 663–670.e3 (2017).
Rice, W. J. et al. Routine determination of ice thickness for cryo-EM grids. J. Struct. Biol. 204, 38–44 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, e42166 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta. Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta. Crystallogr. D Struct. Biol. 74, 531–544 (2018).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta. Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Amunts, A. et al. Structure of the yeast mitochondrial large ribosomal subunit. Science 343, 1485–1489 (2014).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta. Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Quick, M. & Javitch, J. A. Monitoring the function of membrane transport proteins in detergent-solubilized form. Proc. Natl Acad. Sci. USA 104, 3603–3608 (2007).
Acknowledgements
We thank S. Tucker and G. Klesse (University of Oxford) for assisting with the installation and use of CHAP and developing a new CHAP module for this work, K. Zhang (Yale University) and Z. Li (Purdue University) for discussion on structure validation, B. Carragher and C. Potter for access to and L. Yen and E. Eng for help with the use of the electron microscopy facility at the New York Structural Biology Center (NYSBC). This work was supported by grants to J.Y. from the National Institutes of Health (NIH) (RO1EY027800 and RO1GM085234), to J.F. from the NIH (RO1GM55440), to G.L. from the National Natural Science Foundation of China (NSFC) (21625302 and 21933010) and to Y.Z. from the NSFC (31700647) and the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB17000000). Some of the cryo-EM work was performed at the Simons Electron Microscopy Center and National Resource for Automated Molecular Microscopy located at NYSBC, supported by grants from the Simons Foundation (SF349247), NYSTAR and the NIH National Institute of General Medical Sciences (GM103310), with additional support from Agouron Institute (F00316) and NIH (OD019994). Some cryo-EM work was performed at the National Center for CryoEM Access and Training (NCCAT) and the Simons Electron Microscopy Center located at NYSBC, supported by the NIH Common Fund Transformative High Resolution Cryo-Electron Microscopy program (U24 GM129539) and by grants from the Simons Foundation (SF349247) and NY State Assembly Majority.
Author information
Authors and Affiliations
Contributions
M.L. and J.Y. conceived and initiated the structural project, and X.Z., Z.F., D.S., Y.Z., M.L., X.L., Y.P., M.Z., G.L., J.F. and J. Y. designed experiments, analyzed results and wrote the manuscript. X.Z., D.S. and M.L. performed molecular biological and biochemical experiments. Z.F. and X.Z. performed cryo-EM data acquisition and processing, assisted by R.A.G. and S.L. X.Z. built the atomic models. G.L. designed the molecular dynamics simulation (MDS) strategy, and G.L. and Y.Z. performed MDS. D.S. carried out HEK 293T cell recordings, assisted by H.L. M.Z. supervised and Y.P. performed liposome recordings and analysis, and Z.H. performed supplemental single-channel analysis. H.L. performed confocal imaging, and Z.R. performed liposome flux assays (data not shown). All authors contributed to manuscript discussion, preparation and editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Peer reviewer reports are available. Katarzyna Marcinkiewicz was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cryo-EM single-particle analysis of cGMP-unbound TAX-4 reconstituted in lipid nanodiscs.
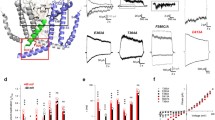
a, Gel filtration profile of cGMP-unbound TAX-4 nanodisc sample. b, SDS-PAGE of cGMP-unbound TAX-4 nanodisc sample used for cryo-EM. c, A representative motion-corrected micrograph. d, Fourier power spectrum of the micrograph shown in (c). e, Gallery of typical averages from 2D classification. f, Flow chart of cryo-EM image processing. g, Euler angle distribution of particles used in the final 3D reconstruction with C4 symmetry. h, Local resolution of the final density map reconstructed with C4 symmetry. i, Densities of the SF generated from the two half maps, contoured at 5σ and overlaid with the model. j, Gold-standard FSC curves of the final 3D reconstructions with C4 and C1 symmetry, respectively. k, FSC curves for cross-validation between maps and model. Black, model versus the summed map. Blue, model versus the half map that was used for model refinement (called ‘work’). Red, model versus another half map that was not used for model refinement (called ‘free’). Uncropped image for panel b is available as source data.
Extended Data Fig. 2 Cryo-EM density maps and atomic models of selected key regions in cGMP-unbound and -bound structures.
a, cGMP-unbound. The map was low-pass filtered to 2.6 Å, sharpened with a temperature factor of -117 Å2 and contoured at 5σ. b, cGMP-bound. The map was low-pass filtered to 2.7 Å, sharpened with a temperature factor of -113 Å2 and contoured at 5σ.
Extended Data Fig. 3 Cryo-EM single-particle analysis of cGMP-bound TAX-4 reconstituted in lipid nanodiscs.
a, Gel filtration profile of cGMP-bound TAX-4 nanodisc sample. b, SDS-PAGE of cGMP-bound TAX-4 nanodisc sample used for cryo-EM. c, A representative motion-corrected micrograph. d, Fourier power spectrum of the micrograph shown in (c). e, Gallery of typical averages from 2D classification. f, Flow chart of cryo-EM image processing. g, Euler angle distribution of particles used in the final 3D reconstruction with C4 symmetry. h, Local resolution of the final density map reconstructed with C4 symmetry. i, Densities of the SF generated from the two half maps, contoured at 5σ and overlaid with the model. j, Gold-standard FSC curves of the final 3D reconstructions with C4 and C1 symmetry, respectively. k, FSC curves for cross-validation between maps and model. Black, model versus the summed map. Blue, model versus the half map that was used for model refinement (called ‘work’). Red, model versus another half map that was not used for model refinement (called ‘free’). Uncropped image for panel b is available as source data.
Extended Data Fig. 4 Cryo-EM single-particle analysis of cGMP-unbound F403V V407A (VA) mutant TAX-4 reconstituted in lipid nanodiscs.
a, Gel filtration profile of cGMP-bound VA TAX-4 nanodisc sample. b, SDS-PAGE of cGMP-bound VA TAX-4 nanodisc sample used for cryo-EM. c, A representative motion-corrected micrograph. d, Fourier power spectrum of the micrograph shown in (c). e, Gallery of typical averages from 2D classification. f, Flow chart of cryo-EM image processing. g, Euler angle distribution of particles used in the final 3D reconstruction with C4 symmetry. h, Local resolution of the final density map reconstructed with C4 symmetry. i, Densities of the SF generated from the two half maps, contoured at 5σ and overlaid with the model. j, Gold-standard FSC curves of the final 3D reconstructions with C4 and C1 symmetry, respectively. k, FSC curves for cross-validation between maps and model. Black, model versus the summed map. Blue, model versus the half map that was used for model refinement (called ‘work’). Red, model versus another half map that was not used for model refinement (called ‘free’). Uncropped image for panel b is available as source data.
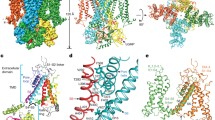
Extended Data Fig. 5 Comparison of the CNBD and C-linker structures of TAX-4 and HCN1.
a,b, Superposition of the CNBD and C-linker structures in unliganded states in stereo view. The α-carbon r.m.s.d is 9.2 Å. c,d, Superposition of the CNBD and C-linker structures in liganded states in stereo view. The α-carbon r.m.s.d is 9.4 Å.
Extended Data Fig. 6 Superposition of S6 of TAX-4 orthologs.
a,b, Superposition of S6 (viewed from the extracellular side) of cGMP-bound (open state) TAX-4, apo (closed state) TAX-4, apo SthK (PDB ID: 6CJQ) and apo LliK (PDB ID: 5V4S), showing the presence of a hydrophobic cavity gate in all the apo structures.
Extended Data Fig. 7 TAX-4 and lipid interactions.
a, Cryo-EM density map of cGMP-unbound TAX-4 reconstituted in lipid nanodiscs. Densities colored in blue are not modeled into TAX-4 and likely represent lipids. b,c, Possible lipid binding in several different regions in the closed state. The extra densities were fit with phosphatidylcholine (lipid 1 and 2) or phosphatidic acid (lipid 3 and 4). d, Cryo-EM density map of cGMP-bound TAX-4 reconstituted in lipid nanodiscs. Densities colored in blue are not modeled into TAX-4 and likely represent lipids. e, Possible lipid binding in the open state. The extra densities were fit with phosphatidylcholine (lipid 1 and 2) or phosphatidic acid (lipid 3). Density maps were contoured at 5σ (b,e) or 3σ (c).
Supplementary information
Supplementary Information
Supplementary Figure 1.
Supplementary Video 1
Conformational changes produced by cGMP binding and unbinding. Morph of closed- and open-state structures of full-length TAX-4, highlighting the movement of the C-helix of the CNBD, the E′F′ helices of the C-linker, the A′B′ helices of the gating ring, and S4, S5 and S6, viewed parallel to the membrane (side view) and then from the intracellular side (bottom-up view).
Supplementary Video 2
Movement of the S6 hydrophobic cavity gate upon cGMP binding and unbinding. Morph of closed- and open-state structures of S6, highlighting the rotation of F403 and V407 side chains, viewed from the intracellular side (bottom-up view).
Source data
Source Data Fig. 4
Statistical Source Data
Source Data Extended Data Fig. 1
Unprocessed gels
Source Data Extended Data Fig. 3
Unprocessed gels
Source Data Extended Data Fig. 4
Unprocessed gels
Rights and permissions
About this article
Cite this article
Zheng, X., Fu, Z., Su, D. et al. Mechanism of ligand activation of a eukaryotic cyclic nucleotide−gated channel. Nat Struct Mol Biol 27, 625–634 (2020). https://doi.org/10.1038/s41594-020-0433-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-020-0433-5
- Springer Nature America, Inc.
This article is cited by
-
The open gate of the AMPA receptor forms a Ca2+ binding site critical in regulating ion transport
Nature Structural & Molecular Biology (2024)
-
Conformational trajectory of allosteric gating of the human cone photoreceptor cyclic nucleotide-gated channel
Nature Communications (2023)
-
Structural insights into the mechanism of pancreatic KATP channel regulation by nucleotides
Nature Communications (2022)
-
Redefining the role of Ca2+-permeable channels in photoreceptor degeneration using diltiazem
Cell Death & Disease (2022)
-
Gating intermediates reveal inhibitory role of the voltage sensor in a cyclic nucleotide-modulated ion channel
Nature Communications (2022)