Abstract
Precise control of cell division is essential for proper patterning and growth during the development of multicellular organisms. Coordination of formative divisions that generate new tissue patterns with proliferative divisions that promote growth is poorly understood. SHORTROOT (SHR) and SCARECROW (SCR) are transcription factors that are required for formative divisions in the stem cell niche of Arabidopsis roots1,2. Here we show that levels of SHR and SCR early in the cell cycle determine the orientation of the division plane, resulting in either formative or proliferative cell division. We used 4D quantitative, long-term and frequent (every 15 min for up to 48 h) light sheet and confocal microscopy to probe the dynamics of SHR and SCR in tandem within single cells of living roots. Directly controlling their dynamics with an SHR induction system enabled us to challenge an existing bistable model3 of the SHR–SCR gene-regulatory network and to identify key features that are essential for rescue of formative divisions in shr mutants. SHR and SCR kinetics do not align with the expected behaviour of a bistable system, and only low transient levels, present early in the cell cycle, are required for formative divisions. These results reveal an uncharacterized mechanism by which developmental regulators directly coordinate patterning and growth.
Similar content being viewed by others
Main
The final size, shape and function of tissues in multicellular organisms hinge upon the precise control of cell division4. Owing to intrinsic and extrinsic cell polarity, a 90° rotation of the division plane determines whether a cell will divide formatively (producing daughter cells with different fates) or proliferatively5,6 (producing daughter cells with similar fates). A wrong choice can lead to over-proliferation of cells, resulting in aberrant morphogenesis or tumorigenesis7,8. Developmental regulators that specify cell fate and interface directly with the cell cycle machinery9,10,11 are likely arbiters of this decision. However, we have limited knowledge about how these regulators dynamically control cell division in situ.
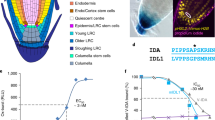
SHR and SCR control the formative division in the Arabidopsis root that gives rise to the endodermis and cortex cell types (ground tissue). SHR, a mobile intercellular signalling molecule, is produced in the central tissues of the root and moves outward into adjacent cells, including the endodermis and the cortex–endodermal initial daughter (CEID) cell, where it activates SCR expression12,13. SHR and SCR together activate the expression of the cell cycle regulator CYCLIND6;1 (CYCD6) only in the CEID, triggering formative division14. In shr and scr mutants, this division does not occur, resulting in a single ground tissue mutant layer, rather than distinct files of endodermis and cortex cells1,2,15 (Fig. 1a).
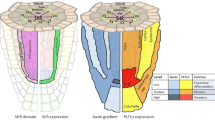
a, Diagram of Arabidopsis wild-type and SHR:GAL4–GR UAS:SHR–GFP shr2 mutant roots showing proliferative and formative division planes (adapted from ref. 52). SHR moves from the central tissues of the root into the adjacent cell layer. SHR expression and formative divisions occur in the inducible line upon treatment with dex. Yellow, QC (quiescent centre); orange, CEI (cortex–endodermal initial); red, CEID (cortex–endodermal initial daughter) and shr mutant layer; blue, cortex; purple, endodermis. b, Diagram of the SHR–SCR regulatory network controlling formative division based on Cruz–Ramirez et al.3. c, Confocal median longitudinal sections showing GFP-labelled SHR and H2B–RFP at timepoints after induction with 10 μM dex. Images are representative of independent timecourse experiments with eight roots. Numbers at the top left show the first five cell positions in the mutant ground tissue. Gamma is set to 0.75 to show signal in the mutant layer for the GFP-only images. Top and bottom show different roots. White arrows, formative divisions. Scale bars, 50 μm. d, Raw (grey) and smoothed (green) SHR trajectory (SHR–GFP/H2B–RFP fluorescence intensity) over time in the first five cells of a single cell file after full induction (10 μM dex). Plots are representative of 211 cells from independent time courses with 8 roots. Possible low and high steady states are indicated for cell 1. Black dashed line, proliferative division; orange dashed line, formative division. a.u., arbitrary units. e, SHR trajectory predicted by the Cruz–Ramirez model showing low and high steady states. f, SHR trajectories for cells that show a low peak of SHR accumulation hours prior to dividing formatively. Roots were treated with low dex (0.02 μM or 0.03 μM). Dark green, SHR trajectory corresponding to images in g. g, Median longitudinal sections through a root tip treated with low dex (0.02 μM) highlighting a cell with a low transient peak of SHR prior to dividing formatively. Plots and images in f and g are representative of 15 cells from 10 roots showing similar behaviour. Scale bars, 10 μm.
Cruz-Ramírez et al.3 proposed a bistable model to explain both how and where SHR and SCR trigger the decision to divide. According to the model, two positive feedback loops generate high stable steady states of SCR and nuclear SHR, triggering formative division (Fig. 1b). Bistability arising from positive feedback is at the heart of mathematical models of decision making in many systems16. However, positive feedback does not always lead to bistability17, and alternative decision-making mechanisms exist. For example, the simple presence of a factor at the right place and time can alter the cell cycle programme and lead to a different cell fate11.
Quantitative time-lapse imaging of transcription factor dynamics has provided key insights into gene-regulatory network function in single cell organisms and mammalian cell lines18,19,20. Assays of multiple transcription factors in tandem on a long timescale can enable examination of their regulatory relationships21. However, many technical challenges have made studies of network dynamics in vivo difficult22. Phototoxicity and photobleaching, in particular, restrict studies using confocal microscopy to short timescales or infrequent sampling and limit the number of fluorophores that can be imaged simultaneously. Owing to its lower phototoxicity, light sheet microscopy provides the means for longer-term multi-colour imaging of protein dynamics in vivo. This potential has been extolled for nearly two decades, but the technology has been used primarily for observation of cellular dynamics and morphology changes during development23,24,25.
Here, we use long-term 4D imaging of living roots and quantitative analysis to gain insight into the dynamics of the SHR–SCR gene-regulatory network that controls formative divisions in the root stem cell niche. Our measurements revealed a key aspect that was missing from the existing model: namely, that SHR and SCR levels are interpreted within the context of the cell cycle. We present evidence that low threshold levels of SHR and SCR are sufficient and act early in the cell cycle to change the orientation of the division plane.
SHR dynamics determine formative division
To investigate the mechanism by which SHR dynamically controls formative cell divisions, we generated a fluorescently labelled inducible SHR construct, SHR:GAL4–GR UAS:SHR–GFP, capable of rescuing the formative divisions absent in shr2 mutants1,2 (Fig. 1a). We induced transcription of SHR–GFP in its endogenous expression domain (the central tissues of the root; Fig. 1a) using diminishing concentrations of dexamethasone26 (dex) (10, 1, 0.05, 0.03, 0.02 and 0.01 μM), and imaged the roots in 3D every 15 min for up to 24 h using confocal microscopy (Fig. 1c and Supplementary Videos 1–4). We observed movement of SHR-GFP into the adjacent mutant ground tissue nuclei, as predicted from previous SHR localization studies27,28. We then quantified the fluorescence of SHR–GFP in the ground tissue nuclei (n = 935 cells from 29 roots) relative to a nuclei marker as a measure of SHR concentration (Extended Data Fig. 1a and Supplementary Methods), from the time of induction up to formative division or the end of the experiment if no division occurred (Fig. 1d and Supplementary Data 1).
The inducible SHR line enabled us to produce data from many cells in each root, and to produce a variety of SHR accumulation trajectories with different division outcomes (Fig. 1d and Extended Data Fig. 1b–f). We observed near complete rescue of meristematic formative divisions at 1 μM and 10 μM dex (96% and 99%, respectively) and no formative divisions at a concentration of 0.01 μM dex (Extended Data Fig. 1f). Occasionally, a cell divided anticlinally (proliferatively) before the periclinal formative division (Fig. 1d, cell 4). The rescued formative divisions are likely to be controlled by the SHR–SCR–CYCD6 pathway that controls CEID division in the stem cell niche of wild-type plants (Extended Data Fig. 2a–d and Supplementary Note 1), and levels of SHR–GFP in fully induced (10 μM dex) plants were comparable to levels of SHR:SHR–GFP in the relevant tissues (Extended Data Fig. 2e,f).
The bistable model3 postulates that SHR triggers formative division when nuclear SHR levels in the ground tissue flip to a high steady state (Fig. 1e). Consistent with this model, in many cases we observed a rapid increase in SHR levels followed by a period during which higher SHR levels were relatively constant prior to division (Fig. 1d, cell 2). However, in other cases, a transient low peak of SHR was able to trigger division many hours later (Fig. 1f,g, Extended Data Fig. 1b and Supplementary Video 5). These SHR kinetics are inconsistent with a bistable model in which high steady state levels of nuclear SHR are necessary to trigger division, where the scale of the model (predicted SHR levels) is comparable to the observed range of SHR protein levels (Fig. 1e–g). We sought next to directly examine SHR regulation of SCR levels.
Bistability is not required for SHR regulation of SCR
We measured SHR and SCR accumulation simultaneously in single nuclei, using a custom light sheet microscope built to image growing root tips under near physiological conditions29 (Extended Data Fig. 3a–c, Supplementary Video 6 and Supplementary Methods). We first introduced SCR:SCR–mKATE2 into the SHR:GAL4–GR UAS:SHR–GFP shr2 background. Next, we induced transcription of SHR–GFP and subsequent activation of SCR–mKATE2 expression using different concentrations of dex (40 μM, 20 μM and 0.4 μM; Supplementary Methods) to obtain a range of SHR and SCR accumulation profiles. We imaged and quantified the levels of SHR and SCR in the ground tissue nuclei every 15 min for up to 48 h after induction (Fig. 2a–d, Supplementary Videos 7 and 8 and Supplementary Data 2; n = 577 cells from 14 roots). In fully induced roots (40 μM dex), 89% of the meristematic cells had divided after 45 h.
a, 3D reconstruction of a z-stack showing induced SHR–GFP (green), SCR–mKATE2 (magenta) and H2B–CFP (blue) fluorescence in a SHR:GAL4-GR UAS:SHR–GFP SCR:SCR–mKATE2 UBQ10:H2B–CFP shr2 root. Scale bar, 50 μm. b, Endodermal nuclei detected in Imaris are selected for quantification. Colours specify different cell files. Scale bar, 50 μm. c, Median longitudinal sections of the root in a and b showing timepoints after induction with 40 μM dex. Images in a–c are representative of independent timecourse experiments with nine roots. White arrows, formative divisions. Scale bars, 50 μm. d, Quantification of SHR and SCR trajectories (SHR–GFP/H2B–CFP and SCR–mKATE2/H2B–CFP fluorescence intensity, respectively) for a single cell after full induction (40 μM dex). Measurement is representative of 274 cells from independent timecourse experiments with nine roots. SHR and SCR trajectory values are normalized to the 90th quantile (Supplementary Methods). Black dashed line, proliferative division; orange dashed line, formative division. e–h, Mean of all fully induced SHR (green) and SCR (magenta) normalized trajectories (Supplementary Methods) and predictions for SCR (grey lines) from the Cruz–Ramirez (e), Michaelis–Menten (f), Hill (g) and positive feedback (h) models. R2, adjusted R squared; n = 274 cells from 9 roots (treated with 40 μM dex; Supplementary Methods).
SHR and SCR levels appeared to follow simple dynamics (Fig. 2d). The average SHR and SCR curves closely follow a sigmoid pattern, with SCR having a slightly steeper and later rise (Fig. 2e–h). To further investigate the regulation of SCR by SHR, we fit the data to the bistable model3 and to three basic ODE models (Supplementary Methods). For each model, we used the averaged SHR dynamics as input to predict SCR expression. Using the published parameters3, the bistable model predicted a rapid jump in SCR levels that we did not observe in the data (Fig. 2e). By varying each parameter by two orders of magnitude, we were able to find parameter regimes that fit the data reasonably well (Extended Data Fig. 4a,b). However, in many of these cases the model no longer displays bistable properties (Extended Data Fig. 4c).
For the three basic ODE models, the fits were comparable (Fig. 2e–h; Michaelis–Menten: adjusted R2 = 0.987; Hill: adjusted R2 = 0.996; positive feedback: adjusted R2 = 0.994). The best fit Hill coefficient for the Hill model was larger than 1, which can occur due to the existence of positive feedback30 (Supplementary Methods). This model and the model explicitly incorporating positive feedback visually appeared to capture the rise of SCR better than the Michaelis–Menten model. This finding is consistent with previously described SCR autoregulation31.
It is not surprising that we were able to fit the data to the Cruz–Ramirez3 model given the larger number of parameters (which can lead to overfitting). However, three other more simple monostable alternatives with fewer parameters also fit the data well. We conclude that bistability is not required to explain the regulatory relationship between SHR and SCR.
SHR and SCR act early in the cell cycle
To further investigate the assumption that bistable steady-state levels of SHR and SCR determine whether a cell will divide formatively, we examined SHR and SCR accumulation just prior to division, when the trajectories of both factors have reached high levels. We found variability in SCR levels and did not observe a bimodal distribution corresponding with the fate of the cell (Extended Data Fig. 5a–d). Furthermore, SCR often did not appear to reach a stable point (Extended Data Fig. 5e,f).
Therefore, we hypothesized that a threshold amount of SHR and SCR triggers formative division at an earlier timepoint. To test this, we determined the accuracy of predicting formative division across a range of SHR and SCR thresholds (Fig. 3a and Supplementary Methods). Optimal thresholds were low relative to the full range of SHR and SCR levels and were able to accurately predict formative division 80% and 73% of the time, respectively. A similar analysis of the SHR confocal data found a maximum prediction accuracy of 87% (Fig. 3a).
a, Prediction accuracy of trajectory classification into formatively dividing and non-dividing cells for a range of SHR and SCR thresholds (Supplementary Methods). Light sheet (LS), n = 449 cells from 14 roots; confocal (conf), n = 743 cells from 29 roots. b, Maximum prediction accuracy of trajectory classification into proliferatively and formatively dividing cells for a given nuclear size window. ***P = 3.8 × 10−91, 8.9 × 10−61, 8.0 × 10−43 for conf - SHR, LS - SHR and LS - SCR, respectively; one-tailed binomial test. An approximate 50% accuracy is expected by chance. Light sheet, n = 500 cells from 14 roots; confocal, n = 633 cells from 29 roots. Top, example masks used to calculate the nuclear size trajectory for a single cell. NS, not significant. Data are mean ± s.e.m. c, Normalized nuclear size at the beginning of the time course for proliferatively (prolif.) and formatively (form.) dividing cells. Two-tailed Mann–Whitney test. Boxes encompass the IQR, centre lines show the median, and whiskers extend to the full range of the data. d, Quantified normalized CDT1a–CFP (G1 marker) fluorescence intensity and normalized nuclear size of a complete cell cycle from a light sheet PlaCCI time course. Top, CDT1a–CFP and H3.1–mCHERRY confocal images showing raw signal, and the Otsu threshold mask. Plot and images are representative of 45 cells from independent timecourse experiments with 2 roots. Scale bar, 5 μm. e–g, Frequency of dividing cells in unsynchronized roots (e), and roots synchronized with hydroxyurea at G1/S (f) or oryzalin at G2/M (g). Blue, formative; yellow, proliferative. h, Percentage of first divisions after dex induction that were formative for roots shown in e–g. Unpaired one-sided Student’s t-test P is shown. Data are mean ± s.e.m.
To improve these predictions, we considered the possibility that dynamic features of the SHR and SCR trajectories or position in the cell cycle may contribute to the decision to divide formatively. We took a simple machine learning approach (Supplementary Methods) to determine whether we could predict which cells divide using a set of features describing various aspects of the dynamics of the SHR, SCR and nuclear size trajectories (for example, maximum rate, mean SHR and area under the curve; Supplementary Table 1). We used nuclear size as a proxy for position in the cell cycle11 (Fig. 3b and Supplementary Methods). Our learning model was able to predict whether a cell divides formatively 89% and 92% of the time for the light sheet and confocal data, respectively.
To determine the most predictive features, we assessed the ability of each individual feature to discriminate between formatively dividing and non-dividing cells (Supplementary Methods). In addition to features associated with SHR levels, features relating to nuclear size were significant predictors of formative division (Supplementary Tables 2 and 3), suggesting that threshold levels of SHR might be required during a specific window of the cell cycle for formative division to occur.
To test this hypothesis, we determined the accuracy of predicting formative versus proliferative division for each individual cell cycle based on whether a threshold of SHR or SCR was reached during one of four quarters of the nuclear size range (Supplementary Methods). Requiring the threshold for SHR or SCR to be met in the first quarter of the nuclear size range resulted in higher accuracies than the predictions using threshold alone (90%, 94% and 89% for the confocal SHR, light sheet SHR and light sheet SCR, respectively) and higher accuracy than requiring the threshold to be met in any of the other three quarters of the nuclear size range (Fig. 3b). In addition, the feature most predictive of formative division over a single cell cycle was the maximum SHR level during the first quarter of the nuclear size range, which accurately predicted 94% and 89% of the confocal and light sheet datasets, respectively (Supplementary Tables 4 and 5). Cells that divided proliferatively had significantly larger nuclei at the beginning of the time course compared with formatively dividing cells (Fig. 3c), suggesting that these cells were already past a critical window of the cell cycle when SHR first reached threshold levels. Using the PlaCCI line32, which contains a marker for G1, we found that the 25th percentile of nuclear size falls within G1 or up to 1 h after G1 89% ± 1% of the time (mean ± s.e.m.; n = 36 cells from 2 roots) (Fig. 3d, Extended Data Fig. 6a and Supplementary Video 9). This suggests that SHR and SCR are required during G1 or early S phase to trigger a formative division.
To experimentally validate the hypothesis that position in the cell cycle determines sensitivity to SHR, we synchronized the cell cycle throughout the root meristem prior to induction with dex by treating roots containing the inducible SHR construct with hydroxyurea (see Supplementary Methods), which causes cell cycle arrest at the G1/S transition and early S phase33. We anticipated that this treatment would result in larger numbers of cells exposed to SHR during the critical early cell cycle window, leading to greater numbers of formatively dividing cells. Consistent with our hypothesis, 94% ± 3% of the first divisions after dex induction (n = 3 roots) in hydroxyurea-treated roots were formative compared to only 52% ± 4% of cells (n = 3 roots) treated with dex alone. Conversely, after synchronization with oryzalin at a later stage of the cell cycle (G2/M) followed by dex treatment, only 20% ± 6% of first divisions after dex treatment were formative (Fig. 3e–h, Extended Data Figs. 6b and 7a–c and Supplementary Video 10).
To understand how these findings inform division of the CEID in wild-type plants, we investigated the dynamics of SHR–GFP and SCR–mKATE2 driven by their native promoters in plants with a wild-type phenotype. We found that levels of SHR in the CEID fluctuated but never appeared entirely absent. SHR and SCR returned to pre-division levels quickly after division of the CEI (n = 6; Extended Data Fig. 8a–g, Supplementary Video 11 and Supplementary Data 3). Thus, it is likely that SHR and SCR are always present early in the cell cycle of the CEID, providing the conditions necessary for formative division.
Discussion
How developmental regulators control cell division is a central question in developmental biology, with potentially broad applications in understanding basic cell cycle control. Although we cannot exclude the possibility of bistability without a definitive test for hysteresis34,35, which would be nearly impossible to perform in our system, our data suggest that SHR and SCR are unlikely to regulate formative cell division through a bistable mechanism. Bistability was proposed to explain how and where the decision to divide formatively is made3. Our data suggest an alternative mechanism must exist to restrict SHR–SCR-regulated formative divisions to the CEID in wild-type plants. Levels of SHR and/or other coregulators are possible candidates36,37.
Our finding that low transient levels of SHR and SCR can alter the orientation of the division plane is consistent with previous reports12,31. A window of sensitivity to these transcription factors in G1 and early S is consistent with the known role of D-type cyclins, including CYCD6, in phosphorylating RBR during the G1/S transition3. RBR-associated kinase activity peaks during the G1/S transition and early S phase38. Previous studies have suggested that developmental cues specifying division plane orientation are perceived during G1, much earlier than the first visible signs of division plane formation in G25. SHR and SCR are transcription factors that need time to activate their targets in the regulation of division plane orientation. Thus, it may seem obvious they would need to act early in the cell cycle. However, transcription and translation occur on the order of minutes39, whereas the median cell cycle time in roots40 is approximately 12 h. Given these timescales, SHR and SCR could still function effectively much closer to the time of division plane formation. Understanding the early cellular events regulated by SHR and SCR that lead to the altered division plane is an intriguing avenue for future study.
D-type cyclins activate the RB–E2F bistable switch that commits the cell to DNA synthesis and irreversible progression through mitosis34. Notably, however, SHR induction cannot initiate formative division outside of the meristem, indicating that SHR is not sufficient to trigger cell division by itself. In addition, CYCD6 is not expressed prior to proliferative divisions in the meristem in our inducible system or in wild-type roots14 (Extended Data Fig. 2b). These findings suggest a non-canonical role for SHR, SCR and CYCD6 in determining the orientation of the division plane but not initiation and commitment to division. Thus, the presence or absence of SHR early in the cell cycle determines whether the cell will divide proliferatively or formatively, but other cyclins and other developmental cues must be present to initiate cell cycle progression (Fig. 4a,b). The RB–E2F bistable switch acts to integrate the many signals indicating a cell’s readiness to divide41. These findings suggest that both the timing and orientation of cell division are determined there.
a, Threshold levels of SHR and SCR specify formative division only when present during G1 or early S. b, The presence of SHR and SCR during G1 and early S activates CYCD6 to specify the orientation of the division plane, whereas other cyclins and developmental cues commit the cell to division. CYCD6 and other cyclins along with their associated kinases phosphorylate RBR, committing the cell to formative division. The two positive feedback loops (SCR autoregulatory loop and RBR release of SCR after phosphorylation by CYCD6) have a smaller role in the decision to divide formatively than previously predicted3.
The CYCLIN D–RBR–E2F pathway is highly conserved between plants and animals, including humans42,43. Coordination of axis determination and cell cycle progression by G1/S regulators is important for formative division in metazoans44,45,46, and D-type cyclins have been implicated in axis determination in metazoans such as Caenorhabditis elegans44. Thus, our findings may point to a shared mechanism used to coordinate axis and cell fate determination (patterning) with cell cycle progression (growth) across eukaryotic systems. Perturbation of the CYCLIN D–RBR–E2F pathway is estimated to occur during the development of nearly all cancers47,48,49,50,51, and defects in division plane orientation and formative division have recently been implicated in the genesis of breast and other cancers7,8. Most studies of cell cycle control have used single-cell organisms or cell lines. Future studies using an approach similar to the one described here could reveal mechanisms of cell cycle control that are important during the development of multicellular organisms and suggest opportunities for novel therapeutic interventions in cancer pathogenesis or prevention.
Methods
All methods are included in the Supplementary Information.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The pre-processed image files are available in the Duke Digital Research Data Repository (https://doi.org/10.7924/r46q2571m; datamanagement@duke.edu). Owing to their large size, original image files are available upon request (please contact researchdatasteward@duke.edu for the first 6 years from publication. To inquire about the availability of this dataset beyond 6 years, please contact caraw97@gmail.com). Complete trajectory data and all metadata needed to run the code are included in the Supplementary material. Source data for figures that were not generated by the code are provided in Excel files. Source data are provided with this paper.
Code availability
Custom code was central to the conclusions of the paper. The RootTracker (for microscope hardware control), image processing and quantification pipeline, and trajectory data analysis pipeline code are available at https://doi.org/10.5281/zenodo.10044880.
References
Benfey, P. N. et al. Root development in Arabidopsis: four mutants with dramatically altered root morphogenesis. Development 119, 57–70 (1993).
Scheres, B. et al. Mutations affecting the radial organisation of the Arabidopsis root display specific defects throughout the embryonic axis. Development 121, 53–62 (1995).
Cruz-Ramírez, A. et al. A bistable circuit involving SCARECROW–RETINOBLASTOMA integrates cues to inform asymmetric stem cell division. Cell 150, 1002–1015 (2012).
Baena-López, L. A., Baonza, A. & García-Bellido, A. The orientation of cell divisions determines the shape of Drosophila organs. Curr. Biol. 15, 1640–1644 (2005).
Facette, M. R., Rasmussen, C. G. & Van Norman, J. M. A plane choice: coordinating timing and orientation of cell division during plant development. Curr. Opin. Plant Biol. 47, 47–55 (2019).
Betschinger, J. & Knoblich, J. A. Dare to be different: asymmetric cell division in Drosophila, C. elegans and vertebrates. Curr. Biol. 14, R674–R685 (2004).
Chhabra, S. N. & Booth, B. W. Asymmetric cell division of mammary stem cells. Cell Div. 16, 5 (2021).
Powell, A. E. et al. Lessons from development: a role for asymmetric stem cell division in cancer. Stem Cell Res. 4, 3 (2010).
Kipreos, E. T. & van den Heuvel, S. Developmental control of the cell cycle: insights from Caenorhabditis elegans. Genetics 211, 797–829 (2019).
Komaki, S. & Sugimoto, K. Control of the plant cell cycle by developmental and environmental cues. Plant Cell Physiol. 53, 953–964 (2012).
Meyer, H. M. et al. Fluctuations of the transcription factor atml1 generate the pattern of giant cells in the Arabidopsis sepal. eLife 6, e19131 (2017).
Cui, H. et al. An evolutionarily conserved mechanism delimiting SHR movement defines a single layer of endodermis in plants. Science 316, 421–425 (2007).
Levesque, M. P. et al. Whole-genome analysis of the short-root developmental pathway in Arabidopsis. PLoS Biol. 4, 739–752 (2006).
Sozzani, R. et al. Spatiotemporal regulation of cell-cycle genes by SHORTROOT links patterning and growth. Nature 466, 128–132 (2010).
Di Laurenzio, L. et al. The SCARECROW gene regulates an asymmetric cell division that is essential for generating the radial organization of the Arabidopsis root. Cell 86, 423–433 (1996).
Tyson, J. J. & Novak, B. A dynamical paradigm for molecular cell biology. Trends Cell Biol. 30, 504–515 (2020).
Brandman, O. & Meyer, T. Feedback loops shape cellular signals in space and time. Science 322, 390–395 (2008).
Filipczyk, A. et al. Network plasticity of pluripotency transcription factors in embryonic stem cells. Nat. Cell Biol. 17, 1235–1246 (2015).
Hoppe, P. S. et al. Early myeloid lineage choice is not initiated by random PU.1 to GATA1 protein ratios. Nature 535, 299–302 (2016).
Park, B. O., Ahrends, R. & Teruel, M. N. Consecutive positive feedback loops create a bistable switch that controls preadipocyte-to-adipocyte conversion. Cell Rep. 2, 976–990 (2012).
Etzrodt, M. & Schroeder, T. Illuminating stem cell transcription factor dynamics: long-term single-cell imaging of fluorescent protein fusions. Curr. Opin. Cell Biol. 49, 77–83 (2017).
Skylaki, S., Hilsenbeck, O. & Schroeder, T. Challenges in long-term imaging and quantification of single-cell dynamics. Nat. Biotechnol. 34, 1137–1144 (2016).
Huisken, J., Swoger, J., Del Bene, F., Wittbrodt, J. & Stelzer, E. H. K. Optical sectioning deep inside live embryos by selective plane illumination microscopy. Science 305, 1007–1009 (2004).
Wan, Y., McDole, K. & Keller, P. J. Light-sheet microscopy and its potential for understanding developmental processes. Annu. Rev. Cell Dev. Biol. 35, 655–681 (2019).
Ovečka, M. et al. Multiscale imaging of plant development by light-sheet fluorescence microscopy. Nat. Plants 4, 639–650 (2018).
Aoyama, T. & Chua, N.-H. A glucocorticoid-mediated transcriptional induction system in transgenic plants. Plant J. 11, 605–612 (1997).
Helariutta, Y. et al. The SHORT-ROOT gene controls radial patterning of the Arabidopsis root through radial signaling. Cell 101, 555–567 (2000).
Nakajima, K., Sena, G., Nawy, T. & Benfey, P. N. Intercellular movement of the putative transcription factor SHR in root patterning. Nature 413, 307–311 (2001).
Trivedi, V. et al. Dynamic structure and protein expression of the live embryonic heart captured by 2-photon light sheet microscopy and retrospective registration. Biomed. Opt. Express 6, 2056–2066 (2015).
Zhang, Q., Bhattacharya, S. & Andersen, M. E. Ultrasensitive response motifs: basic amplifiers in molecular signalling networks. Open Biol. 3, 130031 (2013).
Heidstra, R., Welch, D. & Scheres, B. Mosaic analyses using marked activation and deletion clones dissect Arabidopsis SCARECROW action in asymmetric cell division. Genes Dev. 18, 1964–1969 (2004).
Desvoyes, B., Arana-Echarri, A., Barea, M. D. & Gutierrez, C. A comprehensive fluorescent sensor for spatiotemporal cell cycle analysis in Arabidopsis. Nat. Plants 6, 1330–1334 (2020). 2020 611.
Cools, T., Iantcheva, A., Maes, S., Van den Daele, H. & De Veylder, L. A replication stress-induced synchronization method for Arabidopsis thaliana root meristems. Plant J. Cell Mol. Biol. 64, 705–714 (2010).
Yao, G., Lee, T. J., Mori, S., Nevins, J. R. & You, L. A bistable Rb–E2F switch underlies the restriction point. Nat. Cell Biol. 10, 476–482 (2008).
Pomerening, J. R., Sontag, E. D. & Ferrell, J. E. Building a cell cycle oscillator: hysteresis and bistability in the activation of Cdc2. Nat. Cell Biol. 5, 346–351 (2003).
Koizumi, K., Hayashi, T., Wu, S. & Gallagher, K. L. The SHORT-ROOT protein acts as a mobile, dose-dependent signal in patterning the ground tissue. Proc. Natl Acad. Sci. USA 109, 13010–13015 (2012).
Crook, A. D. et al. BAM1/2 receptor kinase signaling drives CLE peptide-mediated formative cell divisions in Arabidopsis roots. Proc. Natl Acad. Sci. USA 117, 32750–32756 (2020).
Boniotti, M. B. & Gutierrez, C. A cell-cycle-regulated kinase activity phosphorylates plant retinoblastoma protein and contains, in Arabidopsis, a CDKA/cyclin D complex. Plant J. 28, 341–350 (2001).
Milo, R., Jorgensen, P., Moran, U., Weber, G. & Springer, M. BioNumbers-the database of key numbers in molecular and cell biology. Nucleic Acids Res. 38, D750–D753 (2010).
Rahni, R. & Birnbaum, K. D. Week-long imaging of cell divisions in the Arabidopsis root meristem. Plant Methods 15, 30 (2019).
Stallaert, W., Kedziora, K. M., Chao, H. X. & Purvis, J. E. Bistable switches as integrators and actuators during cell cycle progression. FEBS Lett. 593, 2805–2816 (2019).
Zluhan-Martínez, E. et al. Beyond what your retina can see: similarities of retinoblastoma function between plants and animals, from developmental processes to epigenetic regulation. Int. J. Mol. Sci. 21, 4925 (2020).
Shimotohno, A., Aki, S. S., Takahashi, N. & Umeda, M. Regulation of the plant cell cycle in response to hormones and the environment. Annu. Rev. Plant Biol. 72, 273–296 (2021).
Tilmann, C., Kimble, J. & Cyclin, D. Regulation of a sexually dimorphic asymmetric cell division. Dev. Cell 9, 489–499 (2005).
Costa, S. Are division plane determination and cell-cycle progression coordinated? New Phytol. 213, 16–21 (2017).
Noatynska, A., Tavernier, N., Gotta, M. & Pintard, L. Coordinating cell polarity and cell cycle progression: what can we learn from flies and worms? Open Biol. 3, 130083 (2013).
Li, X. Cyclin-dependent kinase 4 and 6 inhibitors as breast cancer therapy: research progress and prospects. Pharm. Chem. J. 56, 71–77 (2022).
Brighi, N. et al. The cyclin-dependent kinases pathway as a target for prostate cancer treatment: rationale and future perspectives. Crit. Rev. Oncol. Hematol. 157, 103199 (2021).
Kent, L. N. & Leone, G. The broken cycle: E2F dysfunction in cancer. Nat. Rev. Cancer 19, 326–338 (2019).
VanArsdale, T., Boshoff, C., Arndt, K. T. & Abraham, R. T. Molecular pathways: targeting the cyclin D–CDK4/6 axis for cancer treatment. Clin. Cancer Res. 21, 2905–2910 (2015).
Musgrove, E. A., Caldon, C. E., Barraclough, J., Stone, A. & Sutherland, R. L. Cyclin D as a therapeutic target in cancer. Nat. Rev. Cancer 11, 558–572 (2011).
Maizel, A. Arabidopsis root meristem. figshare https://doi.org/10.6084/m9.figshare.4786369.v1 (2017).
Acknowledgements
We express profound gratitude for the mentorship and unwavering support of Philip Benfey, who recently passed away. He was a guiding light whose visionary thinking shaped the lives and careers of many scientists. We are thankful that he was able to read the final version of this manuscript, and that he could share in the excitement of our findings. We also thank S. DiTalia, L. You, J. Socolar, R. Shahan, R. Sozzani, E. Pierre-Jerome, I. Taylor, T. Nolan, M. Zhu, S. Van Dierdonck, Q. Zhou and O. Szekely for critical reading and discussions of the manuscript; O. Szekely for help with graphics; D. Holland, F. Cutrale and J. Choi for contributing to the light sheet microscope design and construction; and L. Cameron and the Light Microscopy Core Facility at Duke for providing the workstations and support for Imaris image analysis. This work was funded by the US National Institutes of Health (NRSA 5F32GM106690-02 and MIRA 1R35GM131725) to C.M.W. and P.N.B., and by the Howard Hughes Medical Institute to P.N.B. as an Investigator. M.J., S.E.F. and T.V.T. were supported by the Translational Imaging Center, Bridge Institute, University of Southern California.
Author information
Authors and Affiliations
Contributions
C.M.W. and P.N.B. conceived the project and designed the experiments. C.M.W. and H.B. conducted experiments and generated transgenic plants. T.V.T. developed the light sheet imaging platform, with contributions from M.J. and S.E.F. V.P. developed the microscope control code and the image data extraction pipeline. C.M.W., H.B. and R.C. performed image analysis. P.S. developed computational tools and performed data analysis of the trajectories. C.M.W. and P.S. interpreted the results and wrote the paper with comments from all authors.
Corresponding authors
Ethics declarations
Competing interests
P.N.B. was the co-founder and Chair of the Scientific Advisory Board of Hi Fidelity Genetics, Inc, a company that works on crop root growth. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Krzysztof Wabnik and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer review reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Inducible SHR system produced a variety of protein accumulation trajectories and division outcomes.
a, Left, raw confocal images of SHR-GFP (green) and H2B-RFP (magenta), and nuclear mask (white; see Supplementary Methods) of a single cell over time. Every third timepoint is shown. Scale bar, 5 μm. Right, quantification of SHR-GFP (top) and H2B-RFP (middle) signal intensities, and the derived SHR trajectory (bottom). SHR-GFP and H2B-RFP signal intensities were extracted from the region demarcated by the nuclear mask at each timepoint. Images and plots are representative of 935 cells from independent timecourse experiments with 29 roots. b, Confocal median longitudinal sections through a root tip treated with low dex (0.02 μM) highlighting another cell that divides proliferatively hours after a transient low peak of SHR-GFP is detected. Quantified SHR trajectory is on the right. Scale bar, 10 μm. c, Confocal median longitudinal sections acquired 18 hrs after induction with 10, 1, 0.05, 0.03, 0.02, and 0.01 μM dex. Images are representative of 8, 2, 1, 8, 7 and 3 roots for 10, 1, 0.05, 0.03, 0.02 and 0.01 μM dex, respectively. Scale bar, 50 μm. d, SHR trajectories for all cells broken out by dex concentration and cell position. SHR trajectories show a quantitative response to different dex concentrations. Grey lines, raw data. Black lines, smoothed averages. 10 μM, n = 211 cells from 8 roots; 1 μM, n = 63 cells from 2 roots; 0.05 μM, n = 25 cells from 1 root; 0.03 μM, n = 291 cells from 8 roots; 0.02 μM, n = 221 cells from 7 roots; 0.01 μM, 124 cells from 3 roots. e, Boxplots of maximum SHR intensity (90th quantile) from all SHR trajectories from all roots treated with different concentrations of dex. Boxes, IQR; centre lines, median, whiskers, full range of the data. f, Percentage of cells that divided proliferatively by dex concentration. Data are mean ± s.d. For e,f, 10 μM, n = 158 cells from 8 roots; 1 μM, n = 46 cells from 2 roots; 0.05 μM, n = 19 cells from 1 root; 0.03 μM, n = 236 cells from 8 roots; 0.02 μM n = 180 cells from 7 roots; 0.01 μM, n = 104 cells from 3 roots.
Extended Data Fig. 2 Inducible SHR system controls the SHR-SCR-CYCD6 pathway active in the stem cell niche.
a, Confocal images of a SHR:SHR-GR CYCD6:GFP shr2 root 6 and 7 h after induction with 10 μM dex. White arrows, cell that divides proliferatively at 7 h. Scale bar, 10 μm. b, Confocal images showing SCR:SCR-mCHERRY and CYCD6:GFP-GUS expression after induction of SHR with 10 μM dex. Maximum pixel intensity for the magenta and green channels is adjusted to enhance visibility of the nucleus in the white box. Inset, image shown with a higher maximum pixel intensity to reduce saturation for that cell. Pink arrow, formatively divided cell; white arrow, proliferatively divided cell. Scale bar, 50 μm. Images in a,b are representative of independent time courses of 2 roots. c, Confocal images of inducible SHR-GFP and H2B-RFP in a shr2 scr3 (top) or shr2 (bottom) background after 18-hour 10 μM dex induction. Images are representative of four roots for each mutant line. Scale bar, 50 μm. d, Number of formative divisions present in the first five cells of 2 cell files in 6-day old inducible SHR-GFP roots in a shr2 (n = 4 roots) or shr2 scr3 (n = 4 roots) background after 18 h of dex. Unpaired two-sided Student’s t-test P is shown. Data are mean ± s.e.m. e, Confocal images (green channel only) of a SHR:SHR-GFP UBQ10:H2B-RFP shr2 root (left) and a fully induced (10 μM dex) SHR:GAL4-GR UAS:SHR-GFP EN7:H2B-RFP shr2 (right) root 12 h after dex treatment. Images are representative of 5 and 9 roots of the two respective genotypes. Scale bar, 10 μm. f, SHR-GFP fluorescence intensity in the stele (n = 10 from 5 roots), CEI (n = 9 from 5 roots) and CEID (n = 9 from 5 roots) of SHR:SHR-GFP UBQ10:H2B-RFP shr2 plants, and in the stele (n = 15 from 8 roots) and mutant cells (n = 66 from 9 roots) of SHR:GAL4-GR UAS:SHR-GFP EN7:H2B-RFP shr2 roots 10–15 h after induction with 10 μM dex. Mean SHR-GFP fluorescence in the shr2 mutant cells prior to formative division in the inducible SHR line is similar to mean levels of SHR-GFP in the CEI and CEID of shr2 roots. Mann-Whitney two-sided P is shown. N.S., not significant. Boxes, IQR; centre lines, mean, whiskers, full range of the data.
Extended Data Fig. 3 Custom light sheet microscope and analysis pipeline.
a, Imaging chamber. b, Capillary tube containing growing root mounted onto custom holder. The holder is lowered into the imaging chamber for imaging. c, Image acquisition and analysis pipeline to produce SHR and SCR trajectories for confocal and light sheet imaging.
Extended Data Fig. 4 Fitting the SCR data to the Cruz-Ramirez model.
a, Mean normalized trajectories of SHR-GFP (green) and SCR-mKATE2 (magenta) (n = 274 cells from 9 roots treated with 40 μM dex; Supplementary Methods) and predictions of SCR (grey lines) using the Cruz-Ramirez model3 (Supplementary Methods) starting from the published Cruz-Ramirez3 parameters. Each parameter (columns) was scaled separately by different values (rows). The resulting SCR curve was then scaled to be comparable with the measured SCR curve. b, The corresponding adjusted R2 for the plots in a. c, Example steady state plots for SHR/H2B and SCR/H2B using the Cruz-Ramirez model (Supplementary Methods) for six of the parameter sets from a. The plot on the left (where the Cruz-Ramirez parameter value for dR was multiplied by 10) shows bistability, while other parameter sets to the right are monostable.
Extended Data Fig. 5 SHR and SCR levels at the time of division are not a critical factor in the decision to divide formatively.
a, Median light sheet longitudinal sections of a SHR:GAL4-GR UAS:SHR-GFP SCR:SCR-mKATE2 UBQ10:H2B-CFP shr2 root treated with 40 μM dex showing two cells from a common progenitor with different levels of SCR just prior to formative division. Images are representative of 16 cell pairs from independent time courses of 10 roots. Scale bar, 50 μm. b,c, Quantification of transcription factor fluorescent protein (TF-FP) trajectories for the cells in a. d, Histograms of the average normalized (Supplementary Methods) SHR and SCR levels found during the last five timepoints of all light sheet SHR and SCR full trajectories (including all dex concentrations). Yellow, formatively dividing cells; grey, proliferatively dividing cells. n = 500 cells from independent time courses with 14 roots. e, Histograms of the slopes for SHR (blue) and SCR (orange) fully induced (40 μM) trajectories. n = 274 cells from independent time courses with 9 roots. f, Examples of SCR trajectories (blue) and their fitted slopes (orange). For e and f, slopes were calculated for trajectories between 1 and 5 h prior to division.
Extended Data Fig. 6 SHR levels are interpreted within the context of the cell cycle.
a, Left, histogram of normalized nuclear size one hour after the end of G1 (Supplementary Methods; n = 50 cells from independent time courses with 2 roots). Right, light sheet image of PlaCCI root showing CDT1a-CFP and H3.1-mCHERRY. Scale bar, 10 μm. b, Maximum projection confocal images of unsynchronized (top) and hydroxyurea- (middle) or oryzalin- (bottom) synchronized roots induced with 10 μM dex. Timepoints shown include the 8 h during which most cells first divide. Nuclei of cells are pseudo-colored according to the type of first division after dex treatment. Subsequent divisions maintain the pseudo-colour of the first. Yellow, proliferative division; blue, formative division; green, SHR-GFP; magenta, H2B-RFP expressed from the EN7 promoter. Each row shows a time course of a single root out of 3 roots per condition. Scale bar, 10 μm.
Extended Data Fig. 7 Division trajectories of synchronized cells.
Graphical representation showing the timing of proliferative (blue dots) and formative (red dots) divisions for each cell for roots pretreated for 17 h by transfer to plates containing no treatment (a), hydroxyurea (b), or oryzalin (c), followed by transfer to dex for imaging. Each row corresponds to a single root. Data are shown for two roots for each treatment. Each box contains dot plots for cells 2 (bottom) up to 9 (top) from a single cell file. Cells that have not divided proliferatively by the end of the time course have a cyan dot at the last timepoint.
Extended Data Fig. 8 SHR and SCR dynamics in CEI and CEID cells.
a, Light sheet images of a CEI cell that divides into a CEI and CEID from a time course of a SHR:SHR-GFP SCR:SCR-mKATE2 UBQ10:H2B-CFP shr2 root showing SHR-GFP and SCR-mKATE2 fluorescence just before and after CEI division. Images show a single cell representative of six cells from independent timecourse experiments with three roots. Scale bar, 10 μm. b, Normalized SHR-GFP and SCR-GFP accumulation in the CEID shown in a and its parent CEI during the time course. c–g, Additional examples of CEID and parent CEI SHR and SCR trajectories. Orange dashed line, formative division; black dashed line, proliferative division. Cells are from three roots.
Supplementary information
Supplementary Information
This file includes Supplementary Note 1, Supplementary Methods, Supplementary Figs 1 and 2 and Supplementary Tables 1–5.
Supplementary Data 1
Raw data extracted from time course images of the SHR:GAL4-GR UAS:SHR-GFP UBQ10:H2B-RFP shr2 line acquired with the confocal microscope (see Methods) and used to create the SHR confocal trajectories.
Supplementary Data 2
Raw data extracted from time course images of the SHR:GAL4-GR UAS:SHR-GFP SCR:SCR-mKATE2 UBQ10:H2B-CFP shr2 line acquired with the light sheet microscope (see Methods), used to create the SHR and SCR light sheet trajectories.
Supplementary Data 3
Raw data extracted from time course images of the SHR:SHR-GFP SCR:SCR-mKATE2 UBQ10:H2B-CFP shr2 line used to create the CEI/CEID trajectories.
Supplementary Video 1
Maximum projection of inducible SHR timecourse. Maximum intensity projection of a confocal time course of a growing SHR:GAL4-GR UAS:SHR-GFP 35S:H2B-RFP shr2 root after induction with 10 μM dex. Magenta, H2B-RFP used for normalization; Green, SHR-GFP.
Supplementary Video 2
Median slices of inducible SHR timecourse. Median longitudinal z-slices of a confocal time course of a growing SHR:GAL4-GR UAS:SHR-GFP 35S:H2B-RFP shr2 root after induction with 10 μM dex. Magenta, H2B-RFP; Green, SHR-GFP.
Supplementary Video 3
Registered median slices of inducible SHR timecourse. Registered median longitudinal z-slices from a confocal time course of a growing SHR:GAL4-GR UAS:SHR-GFP 35S:H2B-RFP shr2 root after induction with 10 μM dex. Magenta, H2B-RFP; Green, SHR-GFP. The white box highlights cell 3 in the left cell file up to an formative division.
Supplementary Video 4
Registered median slices of inducible SHR timecourse – full root. Registered median longitudinal z-slices of a confocal time course of a growing SHR:GAL4-GR UAS:SHR-GFP 35S:H2B-RFP shr2 root after induction with 10 μM dex showing only SHR-GFP along the full length of the root.
Supplementary Video 5
Registered median slices of low dex inducible SHR timecourse. Registered median longitudinal z-slice of a confocal time course of a growing SHR:GAL4-GR UAS:SHR-GFP 35S:H2B-RFP shr2 root after low dex (0.02uM) induction. The SHR levels peak at a low level, go back down, and after several hours the cell divides proliferatively. Corresponds to images shown in Figure 1e. Green, SHR-GFP; Magenta, H2B-RFP.
Supplementary Video 6
3D reconstruction of SCR expression from light sheet images. Reconstruction of a light sheet z-stack in Imaris showing 3D SCR:SCR-mKATE2 expression in endodermal cells of the root meristem.
Supplementary Video 7
Registered maximum projection of inducible SHR and SCR timecourse. Registered 3D reconstruction of a light sheet time course of a growing SHR:GAL4-GR UAS:SHR-GFP SCR:SCR-mKATE2 UBQ10:H2B-CFP shr2 root after induction with 10 μM dex. The spheres toward the end of the video show the nuclei detected in Imaris that were used for quantification of SHR, SCR, and H2B fluorescence intensity. Cyan, H2B-CFP used for normalization; Green, SHR-GFP; Magenta, SCR-mKate2.
Supplementary Video 8
Registered median slices of inducible SHR and SCR timecourse. Registered median longitudinal z-slices of a light sheet time course of a growing SHR:GAL4-GR UAS:SHR-GFP SCR:SCR-mKATE2 UBQ10:H2B-CFP shr2 root after induction with 10 μM dex. Cyan, H2B-CFP used for normalization; Green, SHR-GFP; Magenta, SCR-mKate2.
Supplementary Video 9
Registered maximum projection of PlaCCI timecourse. Maximum projection of a light sheet time course of a PlaCCI root used to correlate nuclear size with position in the cell cycle. Blue: CDT1a-CFP (G1 marker); Red: H3.1-mCHERRY; Green: CYCB1;1-GFP.
Supplementary Video 10
Registered median slices of inducible SHR timecourses in cell cycle synchronized roots. Registered median longitudinal z-slices of confocal time courses of growing SHR:GAL4-GR EN7:H2B-RFP shr2 roots after induction with 10 μM dex. Roots were pre-treated for 17 hours with 10 μM hydroxyurea (synchronizes cells at G1/S of the cell cycle), 2 μM oryzalin (synchronizes cells at G2/M of the cell cycle) treatment, or a control treatment (transfer to 1/2 MS only plates). The EN7 promoter is active only in the ground tissue, so H2B-RFP is expressed in the mutant ground tissue layer and in the endodermis and cortex after division. Green, SHR-GFP; Magenta, SCR-mKATE2.
Supplementary Video 11
Registered median slices of SHR and SCR timecourse. Registered median longitudinal z-slices of a light sheet time course of a growing SHR:SHR-GFP SCR:SCR-mKATE2 UBQ10:H2B-CFP shr2 root zoomed in to show a dividing CEI cell followed by formative division of the CEID. SHR and SCR levels return quickly to baseline levels after division of the CEI. Formative division of the CEID cell occurs in the last frame. Cyan: H2B-CFP; Green, SHR-GFP; Magenta, SCR-mKATE2.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Winter, C.M., Szekely, P., Popov, V. et al. SHR and SCR coordinate root patterning and growth early in the cell cycle. Nature 626, 611–616 (2024). https://doi.org/10.1038/s41586-023-06971-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06971-z
- Springer Nature Limited
This article is cited by
-
Unravelling how plant cells divide and differ
Nature (2024)
-
BIG coordinates auxin and SHORT ROOT to promote asymmetric stem cell divisions in Arabidopsis roots
Plant Cell Reports (2024)