Abstract
During bacterial cell division, filaments of tubulin-like FtsZ form the Z-ring, which is the cytoplasmic scaffold for divisome assembly. In Escherichia coli, the actin homologue FtsA anchors the Z-ring to the membrane and recruits divisome components, including bitopic FtsN. FtsN regulates the periplasmic peptidoglycan synthase FtsWI. To characterize how FtsA regulates FtsN, we applied electron microscopy to show that E. coli FtsA forms antiparallel double filaments on lipid monolayers when bound to the cytoplasmic tail of FtsN. Using X-ray crystallography, we demonstrate that Vibrio maritimus FtsA crystallizes as an equivalent double filament. We identified an FtsA–FtsN interaction site in the IA–IC interdomain cleft of FtsA using X-ray crystallography and confirmed that FtsA forms double filaments in vivo by site-specific cysteine cross-linking. FtsA–FtsN double filaments reconstituted in or on liposomes prefer negative Gaussian curvature, like those of MreB, the actin-like protein of the elongasome. We propose that curved antiparallel FtsA double filaments together with treadmilling FtsZ filaments organize septal peptidoglycan synthesis in the division plane.






Similar content being viewed by others
Data availability
Atomic coordinates have been deposited in the PDB with accession codes 7Q6D (E. coli FtsA1–405), 7Q6G (X. poinarii FtsA1–396), 7Q6F (V. maritimus FtsA1–396, antiparallel double filament) and 7Q6I (V. maritimus FtsA1–396 and FtsN1–29, bent tetramers in antiparallel double filament arrangement). The next-generation sequencing data associated with this study are available from the Sequence Read Archive at BioProject PRJNA852398. PDB entries 1E4F, 1E4G, 2YCH, 3WQT, 3WQU, 3WT0, 4A2A, 4A2B and 4CZJ were used for structural superpositions and analyses. Source data are provided with this paper.
References
Typas, A., Banzhaf, M., Gross, C. A. & Vollmer, W. From the regulation of peptidoglycan synthesis to bacterial growth and morphology. Nat. Rev. Microbiol. 10, 123–136 (2011).
Szwedziak, P. & Löwe, J. Do the divisome and elongasome share a common evolutionary past. Curr. Opin. Microbiol. 16, 745–751 (2013).
Pichoff, S. & Lutkenhaus, J. Tethering the Z ring to the membrane through a conserved membrane targeting sequence in FtsA. Mol. Microbiol. 55, 1722–1734 (2005).
Salje, J., van den Ent, F., de Boer, P. & Löwe, J. Direct membrane binding by bacterial actin MreB. Mol. Cell 43, 478–487 (2011).
Pichoff, S. & Lutkenhaus, J. Unique and overlapping roles for ZipA and FtsA in septal ring assembly in Escherichia coli. EMBO J. 21, 685–693 (2002).
Nogales, E., Downing, K. H., Amos, L. A. & Löwe, J. Tubulin and FtsZ form a distinct family of GTPases. Nat. Struct. Biol. 5, 451–458 (1998).
Bisson-Filho, A. W. et al. Treadmilling by FtsZ filaments drives peptidoglycan synthesis and bacterial cell division. Science 355, 739–743 (2017).
Yang, X. et al. GTPase activity-coupled treadmilling of the bacterial tubulin FtsZ organizes septal cell wall synthesis. Science 355, 744–747 (2017).
Monteiro, J. M. et al. Peptidoglycan synthesis drives an FtsZ-treadmilling-independent step of cytokinesis. Nature 554, 528–532 (2018).
Squyres, G. R. et al. Single-molecule imaging reveals that Z-ring condensation is essential for cell division in Bacillus subtilis. Nat. Microbiol. 6, 553–562 (2021).
Whitley, K. D. et al. FtsZ treadmilling is essential for Z-ring condensation and septal constriction initiation in Bacillus subtilis cell division. Nat. Commun. 12, 2448 (2021).
Karimova, G., Dautin, N. & Ladant, D. Interaction network among Escherichia coli membrane proteins involved in cell division as revealed by bacterial two-hybrid analysis. J. Bacteriol. 187, 2233–2243 (2005).
Du, S. & Lutkenhaus, J. Assembly and activation of the Escherichia coli divisome. Mol. Microbiol. 105, 177–187 (2017).
Pichoff, S., Du, S. & Lutkenhaus, J. Disruption of divisome assembly rescued by FtsN–FtsA interaction in Escherichia coli. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.1806450115 (2018).
Gerding, M. A. et al. Self-enhanced accumulation of FtsN at division sites and roles for other proteins with a SPOR domain (DamX, DedD, and RlpA) in Escherichia coli cell constriction. J. Bacteriol. 191, 7383–7401 (2009).
Liu, B., Persons, L., Lee, L. & de Boer, P. A. Roles for both FtsA and the FtsBLQ subcomplex in FtsN-stimulated cell constriction in Escherichia coli. Mol. Microbiol. 95, 945–970 (2015).
Tsang, M. J. & Bernhardt, T. G. A role for the FtsQLB complex in cytokinetic ring activation revealed by an ftsL allele that accelerates division. Mol. Microbiol. 95, 925–944 (2015).
Marmont, L. S. & Bernhardt, T. G. A conserved subcomplex within the bacterial cytokinetic ring activates cell wall synthesis by the FtsW–FtsI synthase. Proc. Natl Acad. Sci. USA 117, 23879–23885 (2020).
Li, Y. et al. Genetic analysis of the septal peptidoglycan synthase FtsWI complex supports a conserved activation mechanism for SEDS–bPBP complexes. PLoS Genet. 17, e1009366 (2021).
Busiek, K. K. & Margolin, W. A role for FtsA in SPOR-independent localization of the essential Escherichia coli cell division protein FtsN. Mol. Microbiol. 92, 1212–1226 (2014).
van den Ent, F. & Löwe, J. Crystal structure of the cell division protein FtsA from Thermotoga maritima. EMBO J. 19, 5300–5307 (2000).
Busiek, K. K., Eraso, J. M., Wang, Y. & Margolin, W. The early divisome protein FtsA interacts directly through its 1c subdomain with the cytoplasmic domain of the late divisome protein FtsN. J. Bacteriol. 194, 1989–2000 (2012).
Pichoff, S., Du, S. & Lutkenhaus, J. The bypass of ZipA by overexpression of FtsN requires a previously unknown conserved FtsN motif essential for FtsA–FtsN interaction supporting a model in which FtsA monomers recruit late cell division proteins to the Z ring. Mol. Microbiol. 95, 971–987 (2015).
Baranova, N. et al. Diffusion and capture permits dynamic coupling between treadmilling FtsZ filaments and cell division proteins. Nat. Microbiol. https://doi.org/10.1038/s41564-019-0657-5 (2020).
Corbin, B. D., Geissler, B., Sadasivam, M. & Margolin, W. Z-ring-independent interaction between a subdomain of FtsA and late septation proteins as revealed by a polar recruitment assay. J. Bacteriol. 186, 7736–7744 (2004).
Rico, A. I., Garcia-Ovalle, M., Mingorance, J. & Vicente, M. Role of two essential domains of Escherichia coli FtsA in localization and progression of the division ring. Mol. Microbiol. 53, 1359–1371 (2004).
Bernard, C. S., Sadasivam, M., Shiomi, D. & Margolin, W. An altered FtsA can compensate for the loss of essential cell division protein FtsN in Escherichia coli. Mol. Microbiol. 64, 1289–1305 (2007).
Pichoff, S., Shen, B., Sullivan, B. & Lutkenhaus, J. FtsA mutants impaired for self-interaction bypass ZipA suggesting a model in which FtsA’s self-interaction competes with its ability to recruit downstream division proteins. Mol. Microbiol. 83, 151–167 (2012).
Radler, P. et al. In vitro reconstitution of Escherichia coli divisome activation. Nat. Commun. 13, 2635 (2022).
Szwedziak, P., Wang, Q., Freund, S. M. & Löwe, J. FtsA forms actin-like protofilaments. EMBO J. 31, 2249–2260 (2012).
Wagstaff, J. & Löwe, J. Prokaryotic cytoskeletons: protein filaments organizing small cells. Nat. Rev. Microbiol. 16, 187–201 (2018).
Schoenemann, K. M. et al. Gain-of-function variants of FtsA form diverse oligomeric structures on lipids and enhance FtsZ protofilament bundling. Mol. Microbiol. https://doi.org/10.1111/mmi.14069 (2018).
van den Ent, F., Izore, T., Bharat, T. A., Johnson, C. M. & Löwe, J. Bacterial actin MreB forms antiparallel double filaments. eLife 3, e02634 (2014).
Hussain, S. et al. MreB filaments align along greatest principal membrane curvature to orient cell wall synthesis. eLife https://doi.org/10.7554/eLife.32471 (2018).
Wong, F., Garner, E. C. & Amir, A. Mechanics and dynamics of translocating MreB filaments on curved membranes. eLife https://doi.org/10.7554/eLife.40472 (2019).
Dion, M. F. et al. Bacillus subtilis cell diameter is determined by the opposing actions of two distinct cell wall synthetic systems. Nat. Microbiol. 4, 1294–1305 (2019).
Krupka, M. et al. Escherichia coli FtsA forms lipid-bound minirings that antagonize lateral interactions between FtsZ protofilaments. Nat. Commun. 8, 15957 (2017).
Hayward, S. & Lee, R. A. Improvements in the analysis of domain motions in proteins from conformational change: DynDom version 1.50. J. Mol. Graph. Model. 21, 181–183 (2002).
Karuppiah, V. & Derrick, J. P. Structure of the PilM–PilN inner membrane type IV pilus biogenesis complex from Thermus thermophilus. J. Biol. Chem. 286, 24434–24442 (2011).
Szwedziak, P., Wang, Q., Bharat, T. A., Tsim, M. & Löwe, J. Architecture of the ring formed by the tubulin homologue FtsZ in bacterial cell division. eLife 3, e04601 (2014).
Bendezu, F. O., Hale, C. A., Bernhardt, T. G. & de Boer, P. A. RodZ (YfgA) is required for proper assembly of the MreB actin cytoskeleton and cell shape in E. coli. EMBO J. 28, 193–204 (2009).
McQuillen, R. & Xiao, J. Insights into the structure, function, and dynamics of the bacterial cytokinetic FtsZ-ring. Annu. Rev. Biophys. https://doi.org/10.1146/annurev-biophys-121219-081703 (2020).
Du, S. & Lutkenhaus, J. At the heart of bacterial cytokinesis: the Z ring. Trends Microbiol. 27, 781–791 (2019).
Park, K. T., Pichoff, S., Du, S. & Lutkenhaus, J. FtsA acts through FtsW to promote cell wall synthesis during cell division in Escherichia coli. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.2107210118 (2021).
Taguchi, A. et al. FtsW is a peptidoglycan polymerase that is functional only in complex with its cognate penicillin-binding protein. Nat. Microbiol. 4, 587–594 (2019).
Cho, H. et al. Bacterial cell wall biogenesis is mediated by SEDS and PBP polymerase families functioning semi-autonomously. Nat. Microbiol. 1, 16172 (2016).
Sjodt, M. et al. Structural coordination of polymerization and crosslinking by a SEDS–bPBP peptidoglycan synthase complex. Nat. Microbiol. htttps://doi.org/10.1038/s41564-020-0687-z (2020).
Jacquier, N., Viollier, P. H. & Greub, G. The role of peptidoglycan in chlamydial cell division: towards resolving the chlamydial anomaly. FEMS Microbiol. Rev. 39, 262–275 (2015).
Gorrec, F. & Löwe, J. Automated protocols for macromolecular crystallization at the MRC laboratory of molecular biology. J. Vis. Exp. https://doi.org/10.3791/55790 (2018).
Gorrec, F. An anticipated optimization approach to macromolecular crystallization. Preprint at bioRxiv https://doi.org/10.1101/620328 (2019).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Turk, D. MAIN software for density averaging, model building, structure refinement and validation. Acta Crystallogr. D 69, 1342–1357 (2013).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Schuck, P. On the analysis of protein self-association by sedimentation velocity analytical ultracentrifugation. Anal. Biochem. 320, 104–124 (2003).
Chaturvedi, S. K., Ma, J., Zhao, H. & Schuck, P. Use of fluorescence-detected sedimentation velocity to study high-affinity protein interactions. Nat. Protoc. 12, 1777–1791 (2017).
Brautigam, C. A. Calculations and publication-quality illustrations for analytical ultracentrifugation data. Methods Enzymol. 562, 109–133 (2015).
Abràmoff, M. D., Magalhães, P. J. & Ram, S. J. Image processing with ImageJ. Biophotonics Int. 11, 36–42 (2004).
Levy, D. et al. Two-dimensional crystallization on lipid layer: a successful approach for membrane proteins. J. Struct. Biol. 127, 44–52 (1999).
Russo, C. J., Scotcher, S. & Kyte, M. A precision cryostat design for manual and semi-automated cryo-plunge instruments. Rev. Sci. Instrum. 87, 114302 (2016).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife https://doi.org/10.7554/eLife.42166 (2018).
Kimanius, D., Dong, L., Sharov, G., Nakane, T. & Scheres, S. H. W. New tools for automated cryo-EM single-particle analysis in RELION-4.0. Biochem. J. 478, 4169–4185 (2021).
Silva, J. C. et al. Quantitative proteomic analysis by accurate mass retention time pairs. Anal. Chem. 77, 2187–2200 (2005).
Jaravine, V. A., Zhuravleva, A. V., Permi, P., Ibraghimov, I. & Orekhov, V. Y. Hyperdimensional NMR spectroscopy with nonlinear sampling. J. Am. Chem. Soc. 130, 3927–3936 (2008).
Jung, Y. S. & Zweckstetter, M. Mars—robust automatic backbone assignment of proteins. J. Biomol. NMR 30, 11–23 (2004).
Lee, W., Tonelli, M. & Markley, J. L. NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31, 1325–1327 (2015).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Wang, K. et al. Defining synonymous codon compression schemes by genome recoding. Nature 539, 59–64 (2016).
Yu, D. et al. An efficient recombination system for chromosome engineering in Escherichia coli. Proc. Natl Acad. Sci. USA 97, 5978–5983 (2000).
Miyazaki, K. Molecular engineering of a PheS counterselection marker for improved operating efficiency in Escherichia coli. Biotechniques 58, 86–88 (2015).
Fredens, J. et al. Total synthesis of Escherichia coli with a recoded genome. Nature 569, 514–518 (2019).
Inoue, H., Nojima, H. & Okayama, H. High efficiency transformation of Escherichia coli with plasmids. Gene 96, 23–28 (1990).
Engler, C., Kandzia, R. & Marillonnet, S. A one pot, one step, precision cloning method with high throughput capability. PLoS ONE 3, e3647 (2008).
Deatherage, D. E. & Barrick, J. E. Identification of mutations in laboratory-evolved microbes from next-generation sequencing data using breseq. Methods Mol. Biol. 1151, 165–188 (2014).
Acknowledgements
We thank D. Bellini, F. Gorrec and M. Yu (MRC LMB) for many helpful discussions on crystallization and help with synchrotron data collection; the staff at Diamond Light Source beamlines I03 and I04 for excellent technical support; G. Cannone, C. Savva and all of the members of the LMB electron microscopy facility for training and support; G. Cannone and Y. Liu for help with electron microscopy data collection; T. Darling and J. Grimmett for help with computing infrastructure; K. Liu for next-generation-sequencing library preparation; and all members of the Löwe group for discussions and technical help (all MRC LMB). T.N. was supported by a Boehringer Ingelheim Fonds PhD fellowship and a Vice-Chancellor’s Award by the Cambridge Commonwealth, European and International Trust. F.B. was supported by an EMBO Advanced fellowship (grant no. ALTF 605-2019). This work was funded by the Medical Research Council (grant nos U105181009 and UPA0241008 to J.W.C, and U105184326 to J.L.) and Wellcome (grant no. 202754/Z/16/Z to J.L.).
Author information
Authors and Affiliations
Contributions
T.N. performed protein purifications, electron microscopy, co-pelleting, in vivo cysteine cross-linking and strain characterization. S.H.M. performed the SPR, fluorescence polarization and FDS–AUC experiments. T.N., D.K.-C. and J.L. performed crystallization and crystallography. S.L.M. and J.M.S. performed the HDX-MS experiments. C.W.H.Y. and S.M.V.F. performed the NMR experiments and assignments. L.F.H.F. designed the genome mutagenesis strategy under the supervision of J.W.C., which was adapted by F.B. for combinatorial mutagenesis. T.N. and F.B. performed genome engineering. T.N. and L.F.H.F. performed next-generation sequencing. J.L. supervised the study. T.N. and J.L. wrote the manuscript, with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Rut Carballido-Lopez, William Margolin and Felipe Merino for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The position of the IC domain within the FtsA monomer varies.
a, Comparison of longitudinal filament contacts in FtsA crystal structures from E. coli (PDB 7Q6D), X. poinarii (PDB 7Q6G), V. maritimus (PDB 7Q6F) and T. maritima (PDB 4A2B). E. coli and X. poinarii FtsA form ‘loose’ protofilaments with detached IIA and IIB domains (arrowhead). IIA and IIB domains are in close contact in the VmFtsA and TmFtsA structures, which form continuous filaments in the crystals. b, The IC domain of FtsA is flexible. Left: an arrow along the first principal axis of inertia of the IC domain (purple) can be used to indicate IC domain orientation. Right: FtsA structures in the PDB aligned on their IA, IIA and IIB domains, with arrows indicating the position of the IC domains, showing that the IC domain orientation is variable within the FtsA monomer. There is no correlation between IC domain position and species (different colours) or formation of continuous filaments in the crystals (‡). Principal axes of inertia were calculated using main chain atoms (N, CA, C) of IC domains. c, Comparison between the FtsA and MreB double filaments. Because the lateral interface is formed by the IC domain in the FtsA double filament, it is wider than the MreB double filament. The membrane-proximal side of both double filaments is flat.
Extended Data Fig. 2 VmFtsA forms antiparallel double filaments upon binding VmFtsN1–29.
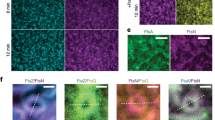
a, Schematic overview of EcFtsN. b, SPR equilibrium response titration of VmFtsN1-29 binding to immobilized VmFtsA1-396. Binding affinity is about threefold lower than for the EcFtsA1-405-EcFtsN1-32 interaction (Fig. 2b). c, VmFtsA1-396 titration into VmFtsN1-29-C-Atto 495. Data were fitted with a two-step model, with transitions being indicative of FtsN binding and polymerization (d). A representative quadruplicate is shown. Kds are given as mean ± SEM from five independent experiments. d, Weight-averaged sedimentation coefficients of a VmFtsA1–396 titration into VmFtsN1–29-C-Atto 495 by FDS–AUC show that VmFtsN1-29 is part of higher order FtsA polymers. Data were fitted to a two-step model, recapitulating the FP data in c. e, Co-pelleting assay of VmFtsN1–29 titrated into VmFtsA1–396 indicates that VmFtsN1–29 induces FtsA polymerization. A representative SDS–PAGE gel is shown. Given are mean ± sd (black lines) of technical duplicates (white dots). P: pellet, S: supernatant. f, Negative-stain electron micrographs of VmFtsA with and without VmFtsN1–29 on supported lipid monolayers. VmFtsA forms ‘mini-rings’ in the absence of FtsN and double filaments at tenfold molar excess of VmFtsN1–29. Two independent grids were examined per condition. Scale bar, 50 nm, 20 nm (inset). g, Multiple sequence alignment of 245 FtsN sequences comprising cytoplasmic and transmembrane domains. EcFtsN1-32 and VmFtsN1–29 sequences are highlighted in bold. h, Mapping of the FtsA-interacting region in EcFtsN1–32 using the lipid monolayer assay. EcFtsN1–32 mutants were in tenfold molar excess of FtsA. In contrast to EcFtsN1-32, EcFtsN1-32, D5N and a scrambled version of the EcFtsN1-32 peptide24 did not induce FtsA double filaments. EcFtsN4-26 and EcFtsN1-32, ΔRK1 led to formation of fewer double filaments. At least two independent grids were examined per condition. Scale bar, 20 nm. i, Summary of EcFtsN1–32 peptides. Mutations are highlighted in bold. Equilibrium dissociation constants (Keqd) are given as mean ± SEM (n ≥ 2 for each construct). For weak binders the maximum response was fixed during fitting, hence these are only approximate values as indicated by asterisks. ND: not determinable. The predominant higher order polymer observed in the monolayer assay is given in the ‘EM’ column. Note that EcFtsN4–26 and EcFtsN1-32, ΔRK1 still lead to formation of a few FtsA double filaments.
Extended Data Fig. 3 Modelling of VmFtsN1-29 binding to VmFtsA1-396.
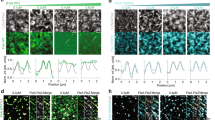
a, Comparison between the peptide-bound closed (coloured) and peptide-free open conformer (black) of FtsA in the VmFtsA1–396-VmFtsN1–29 co-crystal structure (PDB 7Q6I). The IC domain of the open conformer is rotated 13.8° downwards compared to the closed conformer, as determined by analysis with DynDom38. Consequently, the open conformation is likely incompatible with VmFtsN1–29 binding. The inset shows the position of open and closed conformers within the tetramer. b, Left: EcFtsA forms ‘mini-rings’ on lipid monolayers as determined by cryo-EM. Two independent grids were examined. Data was collected on one grid. Right: a computed 2D projection after expansion of a longitudinal dimer from the VmFtsA1-396-VmFtsN1–29 co-crystal structure (PDB 7Q6I) is shown for comparison. The expanded longitudinal dimer does not form a closed ring but a helix. The comparison illustrates that FtsA’s IA domains are facing outwards. Scale bar, 20 nm. c, Comparison between the V. maritimus FtsA–FtsN and Thermus thermophilus PilM-PilN (PDB 2YCH) interaction sites39. Both binding sites are in the IA–IC interdomain cleft of FtsA and PilM but occupy distinct subspaces. FtsN predominantly contacts the IC domain of FtsA, whereas PilN binds closer to the IA domain of PilM. d, Stereo images of the FtsA–FtsN interaction site in the VmFtsA1-396-VmFtsN1–29 co-crystal structure (PDB 7Q6I). Top: our preferred interpretation of the electron density corresponding to VmFtsN1–29, with residues M1-R8 modelled (purple). Side chains of FtsA residues in the interaction site are shown as sticks and polar contacts are marked with black, dashed lines. Bottom: electron density interpretation guided by the NMR data instead (e and f), with residues Y6-K11 of VmFtsN1-29 modelled (purple). Electron density maps (grey) are shown at 1.2 sigma. e, 1H, 15N 2D-HSQC NMR spectrum of free GG-VmFtsN2–29 (blue) and with equimolar amounts of FtsA added (orange). To follow VmFtsN numbering, the first glycine of GG-VmFtsN2–29 is assigned as G0. f, Changes in relative peak intensity expressed as Ibound/Ifree with intensities normalized to IR29, which is assumed not to be involved in the VmFtsA-FtsN interaction.
Extended Data Fig. 4 FtsA cysteine mutant strains: generation and absence of phenotype changes.
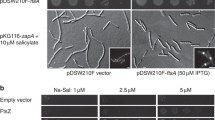
a, Workflow for REXER72-based strain construction. PCR products containing the 3×HA tag or cysteine point mutations were inserted into a shuttle vector by Golden Gate assembly77. Assembled shuttle vectors were transformed into the donor strain, conjugated, and excised in vivo using Cas9. Targeting constructs contain homology regions for 𝜆-Red mediated recombination into the target locus. Recombinants were selected for neoR and tetR markers and against the pheS* marker. Strains were cured of the helper plasmid pKW20 by growth in the absence of selection. b, Growth of strains containing single or double cysteine point mutations and a 3×HA tag in the endogenous ftsA gene, and a kanamycin resistance cassette inserted after the lpxC gene. Parent strains and the original MG1655 strain are also shown. SW: sandwich fusion. c, Growth curves of the same strains in liquid LB medium. Plotted are traces of technical octuplicates (coloured) with mean (black). d, DIC images of the same strains in the exponential phase (OD600 = 0.2-0.3) demonstrating the absence of elongated cells. Similar results were achieved in biological triplicate, of which one was imaged using DIC and two were imaged using phase contrast.
Extended Data Fig. 5 Distances between endogenous and mutated cysteines in the FtsA double filament.
a, Positions of endogenous cysteines in EcFtsA (grey spheres) and of all cysteine mutations used for in vivo cysteine cross-linking (black sticks) (Fig. 5a) are highlighted on the VmFtsA double filament structure (PDB 7Q6F). Dotted lines indicate intermolecular Cβ-Cβ (putative cross-link) distances between selected cysteine mutations and the closest endogenous cysteine, the shortest distance being 15.9 Å (FtsAi C163-FtsAi* D123C). The inset highlights the interfaces in the VmFtsA double filament. SW: sandwich fusion. b, Single cysteine point mutations serve as controls for distance-independent intermolecular cross-linking because of the symmetry of the FtsA double filament, as illustrated on the example of P98C. The P98C mutation used for in vivo cysteine cross-linking is highlighted on the VmFtsA double filament structure (PDB 7Q6F). Cβ-Cβ distances between intermolecular P98C mutations are indicated by dotted lines. The inset provides a comparison between experimentally sampled intermolecular Cβ-Cβ distances by single cysteine point mutations (orange) and all intermolecular Cβ-Cβ distances between amino acids P98, S118, E199 and S252 (blue). Single cysteine point mutations sample intermolecular distances similar to those between double cysteine mutations. Calculated intermolecular Cβ-Cβ distances were rounded to one digit and duplicate values removed prior to plotting.
Supplementary information
Supplementary Information
Supplementary Tables 1–6 and supplementary references.
Supplementary Video 1
Overview of the VmFtsA1–396–VmFtsN1–29 co-crystal structure (PDB 7Q6I). The 16 FtsA monomers are organized in short, antiparallel and curved tetramers. Each protofilament contains one FtsA monomer in the open and closed conformation. Density for VmFtsN1–29 is only observed in the IA–IC interdomain cleft of closed conformers. The IC domain of open conformers is rotated downwards and is probably incompatible with VmFtsN1–29 binding.
Supplementary Data 1
Vector maps.
Supplementary Data 2
Crystallographic models.
Supplementary Data 3
Annotated genomic loci of E. coli strains.
Supplementary Data 4
Summary of the next-generation-sequencing analysis.
Source data
Source Data Fig. 2
Numerical data for Fig. 2b–e.
Source Data Fig. 2
Unprocessed SDS–PAGE gel for Fig. 2e.
Source Data Fig. 5
Unprocessed western blots for Fig. 5c,e.
Source Data Extended Data Fig. 2
Numerical data for Extended Data Fig. 2b–e.
Source Data Extended Data Fig. 2
Unprocessed SDS–PAGE gel for Extended Data Fig. 2e.
Source Data Extended Data Fig. 3
Numerical data for Extended Data Fig. 3f.
Source Data Extended Data Fig. 4
Numerical data for Extended Data Fig. 4c.
Source Data Extended Data Fig. 5
Numerical data for Extended Data Fig. 5b.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Nierhaus, T., McLaughlin, S.H., Bürmann, F. et al. Bacterial divisome protein FtsA forms curved antiparallel double filaments when binding to FtsN. Nat Microbiol 7, 1686–1701 (2022). https://doi.org/10.1038/s41564-022-01206-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-022-01206-9
- Springer Nature Limited
This article is cited by
-
Insights into the assembly and regulation of the bacterial divisome
Nature Reviews Microbiology (2024)
-
Chiral and nematic phases of flexible active filaments
Nature Physics (2023)
-
Conformational changes in the essential E. coli septal cell wall synthesis complex suggest an activation mechanism
Nature Communications (2023)