Abstract
The accumulation of deleterious mitochondrial DNA (∆mtDNA) causes inherited mitochondrial diseases and ageing-associated decline in mitochondrial functions such as oxidative phosphorylation. Following mitochondrial perturbations, the bZIP protein ATFS-1 induces a transcriptional programme to restore mitochondrial function. Paradoxically, ATFS-1 is also required to maintain ∆mtDNAs in heteroplasmic worms. The mechanism by which ATFS-1 promotes ∆mtDNA accumulation relative to wild-type mtDNAs is unclear. Here we show that ATFS-1 accumulates in dysfunctional mitochondria. ATFS-1 is absent in healthy mitochondria owing to degradation by the mtDNA-bound protease LONP-1, which results in the nearly exclusive association between ATFS-1 and ∆mtDNAs in heteroplasmic worms. Moreover, we demonstrate that mitochondrial ATFS-1 promotes the binding of the mtDNA replicative polymerase (POLG) to ∆mtDNAs. Interestingly, inhibition of the mtDNA-bound protease LONP-1 increased ATFS-1 and POLG binding to wild-type mtDNAs. LONP-1 inhibition in Caenorhabditis elegans and human cybrid cells improved the heteroplasmy ratio and restored oxidative phosphorylation. Our findings suggest that ATFS-1 promotes mtDNA replication in dysfunctional mitochondria by promoting POLG–mtDNA binding, which is antagonized by LONP-1.







Similar content being viewed by others
Data availability
The ChIP–seq data have been deposited to the Gene Expression Omnibus database under the BioProject accession code PRJNA590136. The next-generation sequencing data for mtDNA have been deposited in the NCBI Sequence Read Archive database under the BioProject accession code PRJNA780293. All other data supporting the findings of this study are available from the corresponding author on reasonable request. Source data are provided with this paper.
References
Baker, B. M. & Haynes, C. M. Mitochondrial protein quality control during biogenesis and aging. Trends Biochem. Sci 36, 254–261 (2011).
Wai, T., Teoli, D. & Shoubridge, E. A. The mitochondrial DNA genetic bottleneck results from replication of a subpopulation of genomes. Nat. Genet. 40, 1484–1488 (2008).
Gorman, G. S. et al. Prevalence of nuclear and mitochondrial DNA mutations related to adult mitochondrial disease. Ann. Neurol. 77, 753–759 (2015).
Srivastava, S. & Moraes, C. T. Manipulating mitochondrial DNA heteroplasmy by a mitochondrially targeted restriction endonuclease. Hum. Mol. Genet. 10, 3093–3099 (2001).
Tanaka, M. et al. Gene therapy for mitochondrial disease by delivering restriction endonuclease SmaI into mitochondria. J. Biomed. Sci. 9, 534–541 (2002).
Stewart, J.B. & Chinnery, P.F. Extreme heterogeneity of human mitochondrial DNA from organelles to populations. Nat. Rev. Genet. https://doi.org/10.1038/s41576-020-00284-x (2020).
Ahier, A. et al. Affinity purification of cell-specific mitochondria from whole animals resolves patterns of genetic mosaicism. Nat. Cell Biol. 20, 352–360 (2018).
Chiang, A. C. Y., McCartney, E., O’Farrell, P. H. & Ma, H. A genome-wide screen reveals that reducing mitochondrial DNA polymerase can promote elimination of deleterious mitochondrial mutations. Curr. Biol. 29, 4330–4336.e3 (2019).
Lin, Y.-F. et al. Maintenance and propagation of a deleterious mitochondrial genome by the mitochondrial unfolded protein response. Nature 533, 416–419 (2016).
Hahn, A. & Zuryn, S. The cellular mitochondrial genome landscape in disease. Trends Cell Biol. 29, 227–240 (2019).
Pereira, C. V. & Moraes, C. T. Current strategies towards therapeutic manipulation of mtDNA heteroplasmy. Front. Biosci. 22, 991–1010 (2017).
Larsson, N. G. & Clayton, D. A. Molecular genetic aspects of human mitochondrial disorders. Annu. Rev. Genet. 29, 151–178 (1995).
Schon, E. A., DiMauro, S., Hirano, M. & Gilkerson, R. W. Therapeutic prospects for mitochondrial disease. Trends Mol. Med. 16, 268–276 (2010).
Greaves, L. C. et al. Clonal expansion of early to mid-life mitochondrial DNA point mutations drives mitochondrial dysfunction during human ageing. PLoS Genet. 10, e1004620 (2014).
Bender, A. et al. High levels of mitochondrial DNA deletions in substantia nigra neurons in aging and Parkinson disease. Nat. Genet. 38, 515–517 (2006).
Gitschlag, B. L. et al. Homeostatic responses regulate selfish mitochondrial genome dynamics in C. elegans. Cell Metab. 24, 91–103 (2016).
Nargund, A. M., Fiorese, C. J., Pellegrino, M. W., Deng, P. & Haynes, C. M. Mitochondrial and nuclear accumulation of the transcription factor ATFS-1 promotes OXPHOS recovery during the UPRmt. Mol. Cell 58, 123–133 (2015).
Shpilka, T. et al. UPRmt scales mitochondrial network expansion with protein synthesis via mitochondrial import in Caenorhabditis elegans. Nat. Commun. 12, 479 (2021).
Nargund, A. M., Pellegrino, M. W., Fiorese, C. J., Baker, B. M. & Haynes, C. M. Mitochondrial import efficiency of ATFS-1 regulates mitochondrial UPR activation. Science 337, 587–590 (2012).
Bernstein, S. H. et al. The mitochondrial ATP-dependent Lon protease: a novel target in lymphoma death mediated by the synthetic triterpenoid CDDO and its derivatives. Blood 119, 3321–3329 (2012).
Tsang, W. Y. & Lemire, B. D. Stable heteroplasmy but differential inheritance of a large mitochondrial DNA deletion in nematodes. Biochem. Cell Biol. 80, 645–654 (2002).
Cristina, D., Cary, M., Lunceford, A., Clarke, C. & Kenyon, C. A regulated response to impaired respiration slows behavioral rates and increases lifespan in Caenorhabditis elegans. PLoS Genet. 5, e1000450 (2009).
Xu, C. et al. Genetic inhibition of an ATP synthase subunit extends lifespan in C. elegans. Sci. Rep. 8, 14836 (2018).
Deng, P. et al. Mitochondrial UPR repression during Pseudomonas aeruginosa infection requires the bZIP protein ZIP-3. Proc. Natl Acad. Sci. USA 116, 6146–6151 (2019).
Gustafsson, C. M., Falkenberg, M. & Larsson, N.-G. Maintenance and expression of mammalian mitochondrial DNA. Annu. Rev. Biochem. 85, 133–160 (2016).
Rauthan, M., Ranji, P., Aguilera Pradenas, N., Pitot, C. & Pilon, M. The mitochondrial unfolded protein response activator ATFS-1 protects cells from inhibition of the mevalonate pathway. Proc. Natl Aacd. Sci. USA 110, 5981–5986 (2013).
Bota, D. A. & Davies, K. J. A. Lon protease preferentially degrades oxidized mitochondrial aconitase by an ATP-stimulated mechanism. Nat. Cell Biol. 4, 674–680 (2002).
Liu, T. et al. DNA and RNA binding by the mitochondrial lon protease is regulated by nucleotide and protein substrate. J. Biol. Chem. 279, 13902–13910 (2004).
Chen, S. H., Suzuki, C. K. & Wu, S. H. Thermodynamic characterization of specific interactions between the human Lon protease and G-quartet DNA. Nucleic Acids Res. 36, 1273–1287 (2008).
Matsushima, Y., Goto, Y. I. & Kaguni, L. S. Mitochondrial Lon protease regulates mitochondrial DNA copy number and transcription by selective degradation of mitochondrial transcription factor A (TFAM). Proc. Natl Acad. Sci. USA 107, 18410–18415 (2010).
Göke, A. et al. Mrx6 regulates mitochondrial DNA copy number in S. cerevisiae by engaging the evolutionarily conserved Lon protease Pim1. Mol. Biol. Cell https://doi.org/10.1091/mbc.E19-08-0470 (2019).
Valenci, I., Yonai, L., Bar-Yaacov, D., Mishmar, D. & Ben-Zvi, A. Parkin modulates heteroplasmy of truncated mtDNA in Caenorhabditis elegans. Mitochondrion 20, 64–70 (2015).
Kukat, C. et al. Cross-strand binding of TFAM to a single mtDNA molecule forms the mitochondrial nucleoid. Proc. Natl Acad. Sci. USA 112, 11288–11293 (2015).
King, M. P. & Attardi, G. Human cells lacking mtDNA: repopulation with exogenous mitochondria by complementation. Science 246, 500–503 (1989).
Bruno, C. et al. A stop-codon mutation in the human mtDNA cytochrome c oxidase I gene disrupts the functional structure of complex IV. Am. J. Hum. Genet. 65, 611–620 (1999).
Moraes, C. T. et al. Mitochondrial DNA deletions in progressive external ophthalmoplegia and Kearns–Sayre syndrome. N. Engl. J. Med. 320, 1293–1299 (1989).
Moraes, C. T., Schon, E. A., DiMauro, S. & Miranda, A. F. Heteroplasmy of mitochondrial genomes in clonal cultures from patients with Kearns–Sayre syndrome. Biochem. Biophys. Res. Commun. 160, 765–771 (1989).
Yusoff, A. A. M., Abdullah, W. S. W., Khair, S. & Radzak, S. M. A. A comprehensive overview of mitochondrial DNA 4977-bp deletion in cancer studies. Oncol. Rev. 13, 409 (2019).
Lee, H. C., Pang, C. Y., Hsu, H. S. & Wei, Y. H. Differential accumulations of 4,977 bp deletion in mitochondrial DNA of various tissues in human ageing. Biochim. Biophys. Acta 1226, 37–43 (1994).
Munch, C. & Harper, J. W. Mitochondrial unfolded protein response controls matrix pre-RNA processing and translation. Nature 534, 710–713 (2016).
Gilkerson, R. W. et al. Mitochondrial autophagy in cells with mtDNA mutations results from synergistic loss of transmembrane potential and mTORC1 inhibition. Hum. Mol. Genet. 21, 978–990 (2012).
Paix, A., Folkmann, A., Rasoloson, D. & Seydoux, G. High efficiency, homology-directed genome editing in Caenorhabditis elegans using CRISPR–Cas9 ribonucleoprotein complexes. Genetics 201, 47–54 (2015).
Mukhopadhyay, A., Deplancke, B., Walhout, A. J. M. & Tissenbaum, H. A. Chromatin immunoprecipitation (ChIP) coupled to detection by quantitative real-time PCR to study transcription factor binding to DNA in Caenorhabditis elegans. Nat. Protoc. 3, 698–709 (2008).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Blankenberg, D. et al. Manipulation of FASTQ data with Galaxy. Bioinformatics 26, 1783–1785 (2010).
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30, 614–620 (2014).
Belmonte, F. R. et al. Digital PCR methods improve detection sensitivity and measurement precision of low abundance mtDNA deletions. Sci. Rep. 6, 25186 (2016).
Fiorese, C. J. et al. The transcription factor ATF5 mediates a mammalian mitochondrial UPR. Curr. Biol. 26, 2037–2043 (2016).
Pham, T.-N. D., Ma, W., Miller, D., Kazakova, L. & Benchimol, S. Erythropoietin inhibits chemotherapy-induced cell death and promotes a senescence-like state in leukemia cells. Cell Death Dis. 10, 22 (2019).
Haynes, C. M., Yang, Y., Blais, S. P., Neubert, T. A. & Ron, D. The matrix peptide exporter HAF-1 signals a mitochondrial UPR by activating the transcription factor ZC376.7 in C. elegans. Mol. Cell 37, 529–540 (2010).
Palikaras, K., Lionaki, E. & Tavernarakis, N. Coordination of mitophagy and mitochondrial biogenesis during ageing in C. elegans. Nature 521, 525–528 (2015).
Fang, E. F. et al. NAD+ augmentation restores mitophagy and limits accelerated aging in Werner syndrome. Nat. Commun. 10, 5284 (2019).
Weiss, K. R., Voigt, F. F., Shepherd, D. P. & Huisken, J. Tutorial: practical considerations for tissue clearing and imaging. Nat. Protoc. 16, 2732–2748 (2021).
Acknowledgements
We thank the Caenorhabditis Genetics Center for providing C. elegans strains (funded by the NIH Office of Research 362 Infrastructure Programs (P40 OD010440)). We thank C. Moraes for the KSS and G. Manfredi for the CoxI G6930A cybrid cell lines. This work was supported by the HHMI, the Mallinckrodt Foundation and National Institutes of Health grants (R01AG040061 and R01AG047182 to C.M.H., R01GM115911 and R01AI117839 to S.A.W., R01GM111706 and R35GM130320 to P.C., and F31HL147482 to K.L.).
Author information
Authors and Affiliations
Contributions
Q.Y. and C.M.H. planned the experiments. Q.Y., Y.D., T.S., N.U.N., J.L., R.D.Z. and P.C. generated the worm strains. R.L. and L.J.Z. analysed ATFS-1::GFP and TMRE quantification. Q.Y. performed the C. elegans and cybrid mtDNA analysis including ChIP and respiratory function. P.L., K.L. and S.A.W. performed and analysed mtDNA sequencing. Q.Y., N.S.A. and C.M.H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 OXPHOS dysfunction increases mtDNAs.
a, Quantification of total mtDNA in wildtype and nduf-7(et19) worms. n = 3, biologically independent samples (Each sample contains 40-60 animals; every dot stands for averaged value from 3 technical replicates; data shown represent mean ± S.E.M.). **P = 0.0015, Two-tailed Student’s t test. b, POLG immunoblot of wildtype worms following fractionation into total lysate (T), post-mitochondrial supernatant (S), and mitochondrial pellet (M). Tubulin (Tub) and the OXPHOS protein (NDUFS3) serve as loading controls. Representative immunoblots from four biological repeats. c, POLG immunoblot of lysates from wildtype worms raised on control(RNAi) or polg(RNAi). Tubulin (Tub) serves as a loading control. Representative immunoblots from seven biological repeats.
Extended Data Fig. 2 atfs-1-dependent transcription is impaired in atfs-1nuc(−) worms.
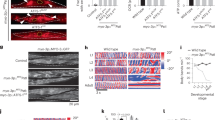
a, Schematic highlighting the R (Arginine) to A (Alanine) substitution to impair the nuclear localization sequence (NLS) within ATFS-1 yielding ATFS-1nuc(−) confirmed by Sanger sequencing. b, UPRmt signaling schematic highlighting the ATFS-1nuc(−) with an impaired NLS. c, Expression level of hsp-6 mRNA in wildtype and atfs-1nuc(−) worms raised on control(RNAi) or spg-7(RNAi) examined by qRT-PCR. n = 3, biologically independent samples. **P = 0.0028, One-way ANOVA. d–e, Photomicrographs of wildtype, atfs-1(et18) and atfs-1(et18)nuc(-);hsp-6pr::gfp worms (Scale bar 0.1 mm) (d); Quantification of fluorescence pixel intensity in wildtype (n = 117; Max: 12.51; Min: 1.87; Median: 5.12), atfs-1(et18) (n = 74; Max: 75.251; Min: 15.790; Median: 32.021) and atfs-1(et18)nuc(−) strains (n = 121; Max: 15.100; Min: 1.55; Median: 5.43). Box & whiskers plots Min to Max. ****P < 0.0001, One-way ANOVA. n means the number of sampling areas. Average pixel intensity signals were calculated from sampling areas at each condition in biological triplicates (e). f, hsp-6 mRNA expression in wildtype, atfs-1(et18) or atfs-1(et18)nuc(−) worms examined by qRT-PCR. n = 4 (wildtype), n = 5 (atfs-1(et18)), n = 3 (atfs-1(et18)nuc(−)), biologically independent samples. *P = 0.0114 (wildtype vs. atfs-1(et18)), *P = 0.0407 (atfs-1(et18) vs. atfs-1(et18)nuc(-)), one-way ANOVA. g, hsp-6 mRNA expression in wildtype and atfs-1nuc(−) worms raised on control(RNAi) or cco-1(RNAi) examined by qRT-PCR. n = 3, biologically independent samples. ***P = 0.0004, one-way ANOVA. h, polg mRNA expression in atfs-1nuc(−) worms raised on control(RNAi) or cco-1(RNAi) examined by qRT-PCR. n = 3, biologically independent samples. Two-tailed Student’s t test. i, POLG immunoblots of lysates from wildtype, atfs-1nuc(−) and atfs-1(null) worms raised on control or cco-1(RNAi). Representative immunoblots from four biological repeats. j, Immunoblots of lysates from wildtype and atfs-1nuc(−) worms raised on control or lonp-1(RNAi). ATFS-1 or ATFS-1nuc(−) are indicated with an arrowhead. Representative immunoblots from four biological repeats. In c, f–h, each dot represents the average from 3 technical replicates; data shown represent mean ± S.E.M.
Extended Data Fig. 3 LONP-1 inhibition promotes mtDNA content via ATFS-1.
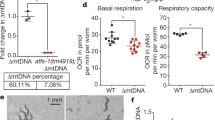
a, FLAG immunoblots of lysates from wildtype and LONP-1FLAG wildtype worms. Tubulin (Tub) serves as a loading control. Representative immunoblots from four biological repeats. b, Images of wildtype or LONP-1FLAG worms 48 hours after synchronization indicating worms expressing LONP-1FLAG at the endogenous locus develop normally (Scale bar 1 mm). Representative images from four biological repeats. c, Fluorescent photomicrographs of wildtype hsp-6pr::gfp or lonp-1FLAG;hsp-6pr::gfp worms 48 hours after synchronization indicating worms expressing LONP-1FLAG do not cause UPRmt activation (Scale bar 0.05 mm). Representative images from four biological repeats. d, Schematic of the putative ATFS-1 and LONP-1 binding sites within the mtDNA non-coding region (NCR) highlighting the proximity of both sites (~200 base pairs). e, POLG Immunoblots of lysates from wildtype worms raised on control or lonp-1(RNAi). Representative images from four biological repeats. f, Total mtDNA quantification in wildtype homoplasmic atfs-1nuc(−) worms raised on control(RNAi) or lonp-1(RNAi). n = 5, biologically independent samples. ***P = 0.0004, Two-tailed Student’s t test). g, Total mtDNA quantification in wildtype homoplasmic atfs-1mts(−);nuc(−) worms raised on control(RNAi) or lonp-1(RNAi). n = 3, biologically independent samples. Two-tailed Student’s t test. In f and g, each biologically independent sample contained 40-60 animals; every dot stands for averaged value from 3 technical replicates; data shown represent mean ± S.E.M. *p < 0.05, **p < 0.01, ****p < 0.0001.
Extended Data Fig. 4 Mitochondrial ATFS-1 is required to maintain ∆mtDNA in heteroplasmic worms.
a, Crossing strategy of atfs-1(null);pdr-1(tm598);uaDf5 strain. b. TMRE quantification of heteroplasmic (∆mtDNA) worms raised on control(RNAi) (n = 475; Max: 1.052; Min: 0.21; Median: 0.618), or wildtype worms raised on control (n = 232; Max: 1.318; Min: 0.725; Median: 0.995) or spg-7(RNAi) (n = 114; Max: 0.798; Min: 0.134; Median: 0.402). Box & whiskers plots Min to Max. n means the number of sampling areas. Average pixel intensity signals were calculated from sampling areas at each condition in biological triplicates. c,d, Photomicrographs of uaDf5 and atfs-1nuc(−);uaDf5;hsp-6pr::gfp worms (Scale bar 0.1 mm) (c); Quantification of fluorescence pixel intensity in uaDf5 (n = 199; Max: 30.89; Min: 3.430; Median:11.590) and atfs-1nuc(−);uaDf5;hsp-6pr::gfp (n = 234; Max: 15.640; Min: 2.540; Median: 6.915). Box & whiskers plots Min to Max. n means the number of sampling areas. Average pixel intensity signals were calculated from sampling areas at each condition in biological triplicates (d). e, ∆mtDNA quantification as determined by qPCR in heteroplasmic uaDf5 worms, atfs-1(null);uaDf5 worms and atfs-1nuc(−);uaDf5 worms. n = 3, biologically independent samples. f, ∆mtDNA quantification as determined by qPCR in heteroplasmic atfs-1nuc(−);uaDf5 worms and atfs-1mts(−);nuc(−);uaDf5. n = 3 (atfs-1nuc(−);uaDf5) and n = 4 (atfs-1mts(−);nuc(−);uaDf5), biologically independent samples. ***P = 0.0007. g, Quantification of total mtDNA following POLG ChIP-mtDNA in homoplasmic wildtype or uaDf5 worms. n = 4 (wildtype) and n = 3 (uaDf5), biologically independent samples. *P = 0.0229. In e and f, each biologically independent sample contained 40-60 animals; in g, each biologically independent sample contained about 150,000 animals; each dot stands for averaged value from 3 technical replicates in f,g; Two-tailed Student’s t test was used in d, f and g, One-way ANOVA was used in b; data shown represent mean ± S.E.M. *p < 0.05, **p < 0.01, ****p < 0.0001.
Extended Data Fig. 5 ATFS-1 and POLG primarily interact with ∆mtDNAs in heteroplasmic worms.
a, Overview of the qPCR strategy to quantify the ∆mtDNA percentage in heteroplasmic worms or heteroplasmic cells. Plasmids containing a sequence specific to the ∆mtDNA or wildtype mtDNA were created7. Standard curves were generated using the indicated concentration of each plasmid harboring sequences specific to either wildtype or ∆mtDNAs. Both PCR reactions were carried out simultaneously in the same qPCR machine. b,c, Scatter plots (b) and results (c) of 3D digital PCR quantification of wildtype mtDNA and ∆mtDNA following ATFS-1 ChIP-mtDNA in heteroplasmic uaDf5 worms. n = 4, biologically independent samples. d-e, Scatter plots (d) and results (e) of 3D digital PCR quantification of wildtype mtDNA and ∆mtDNA following POLG ChIP-mtDNA in heteroplasmic uaDf5 worms. n = 4, biologically independent samples. f, HMG-5/TFAM immunoblot of wildtype worms following fractionation into total lysate (T), post-mitochondrial supernatant (S), and mitochondrial pellet (M). Tubulin (Tub) and the OXPHOS component (NDUFS3) serve as loading controls. Representative immunoblots from two biological repeats. g, HMG-5/TFAM immunoblots of lysates from wildtype worms raised on control or hmg-5/tfam(RNAi). Tubulin (Tub) serves as a loading control. Representative immunoblots from three biological repeats. Each biologically independent sample contained 150,000 animals in c,e; data shown represent mean ± S.E.M.
Extended Data Fig. 6 Inhibition of LONP-1 improves the deleterious heteroplasmy ratio.
a, LONP-1 immunoblots of lysates from wildtype worms raised on control(RNAi) or lonp-1(RNAi). Tubulin (Tub) serves as a loading control. Representative immunoblots from four biological repeats. b, ChIP-mtDNA using ATFS-1 or LONP-1 antibodies in wildtype worms followed by quantification of total mtDNA. n = 3, biologically independent samples. **P = 0.0042. c, ChIP-mtDNA using LONP-1 antibodies in wildtype or heteroplasmic worms followed by quantification of total mtDNA (both wildtype and ∆mtDNA). n = 3, biologically independent samples. d, ∆mtDNA quantification in atfs-1nuc(−);uaDf5 worms raised on control(RNAi) or lonp-1(RNAi). n = 3, biologically independent samples. *P = 0.0168. e, The brood size of heteroplasmic worms raised on control or lonp-1(RNAi). n = 9 worms. f, ∆mtDNA and wildtype mtDNA quantification following HMG-5/TFAM ChIP-mtDNA in uaDf5 heteroplasmic worms raised on lonp-1(RNAi) indicating that the binding of HMG-5/TFAM to wildtype mtDNAs or ∆mtDNAs is similar the input ratio. n = 4, biologically independent samples. g, wildtype mtDNA quantification in uaDf5 heteroplasmic worms raised on control(RNAi) or cco-1(RNAi). n = 3, biologically independent samples. **P = 0.0075. h, wildtype mtDNA quantification in uaDf5 or clk-1(qm30);uaDf5 heteroplasmic worms. n = 3, biologically independent samples. **P = 0.0029. In b,c and f, Each biologically independent sample contained 150,000 animals; in d,g,h each biologically independent sample contained 40-60 animals; every dot stands for averaged value from 3 technical replicates in b-d and f-h; Two-tailed Student’s t test was used; data shown represent mean ± S.E.M.
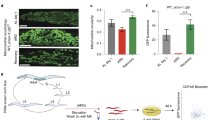
Extended Data Fig. 7 Pharmacological inhibition of LONP1 improves heteroplasmy and OXPHOS function in heteroplasmic cybrid cells.
a, Mutant (G6930A) mtDNA ratio confirmation by sanger sequencing in CoxI G6930A cells treated by CDDO. b, Oxygen consumption rates (OCR) of 143B (wildtype) cells treated with DMSO (ctrl), 0.1 μM or 0.25 μM CDDO for 3 days. n = 22 (ctrl) and n = 24 (0.1 μM and 0.25 μM CDDO), biologically independent samples. c, Cell viability of 143b (WT) and KSS ∆mtDNA cells exposed to various concentrations of CDDO for 72 hours. n = 3, biologically independent samples. d, Basal respiration of KSS heteroplasmic cells treated with DMSO (ctrl), 0.1 μM or 0.25 μM CDDO for 4 or 13 weeks. n = 14 (ctrl) and n = 16 (0.1 μM and 0.25 μM CDDO), biologically independent samples. ****P < 0.0001, Two-tailed Student’s t test. Data shown represent mean ± S.E.M.
Supplementary information
Supplementary Tables
Supplementary Table 1: primers for qPCR, qRT–PCR, mtDNA quantification and guide RNAs for gene editing. Supplementary Table 2: antibodies used in this study.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 2
Unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 6
Unprocessed western blots
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 6
Statistical source data,
Source Data Extended Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
About this article
Cite this article
Yang, Q., Liu, P., Anderson, N.S. et al. LONP-1 and ATFS-1 sustain deleterious heteroplasmy by promoting mtDNA replication in dysfunctional mitochondria. Nat Cell Biol 24, 181–193 (2022). https://doi.org/10.1038/s41556-021-00840-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-021-00840-5
- Springer Nature Limited
This article is cited by
-
Mitochondrial disorders as a mechanism for the development of obese Sarcopenia
Diabetology & Metabolic Syndrome (2023)
-
ATFS-1 counteracts mitochondrial DNA damage by promoting repair over transcription
Nature Cell Biology (2023)
-
Hypoxia-induced mitochondrial stress granules
Cell Death & Disease (2023)
-
Mitochondrial stress induces hepatic stellate cell activation in response to the ATF4/TRIB3 pathway stimulation
Journal of Gastroenterology (2023)
-
Pleiotropic effects of mitochondria in aging
Nature Aging (2022)