Abstract
Novel therapeutics designed to target the polymeric matrix of biofilms requires innovative techniques to accurately assess their efficacy. Here, multiple particle tracking (MPT) was developed to characterize the physical and mechanical properties of antimicrobial resistant (AMR) bacterial biofilms and to quantify the effects of antibiotic treatment. Studies employed nanoparticles (NPs) of varying charge and size (40–500 nm) in Pseudomonas aeruginosa PAO1 and methicillin-resistant Staphylococcus aureus (MRSA) biofilms and also in polymyxin B (PMB) treated Escherichia coli biofilms of PMB-sensitive (PMBSens) IR57 and PMB-resistant (PMBR) PN47 strains. NP size-dependent and strain-related differences in the diffusion coefficient values of biofilms were evident between PAO1 and MRSA. Dose-dependent treatment effects induced by PMB in PMBSens E. coli biofilms included increases in diffusion and creep compliance (P < 0.05), not evident in PMB treatment of PMBR E. coli biofilms. Our results highlight the ability of MPT to quantify the diffusion and mechanical effects of antibiotic therapies within the AMR biofilm matrix, offering a valuable tool for the pre-clinical screening of anti-biofilm therapies.
Similar content being viewed by others
Introduction
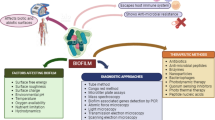
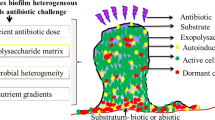
The important role of bacterial biofilms in chronic human diseases, such as cystic fibrosis, otitis media, chronic skin wounds and implant- and catheter-associated infections, has been increasingly recognized1. Within these biofilms, the bacteria are embedded in a complex, charged, self-produced extracellular polymeric matrix (EPS). The EPS matrix is an entangled polymer network2 predominately composed of polysaccharides, extracellular DNA (eDNA), proteins and lipids, which facilitates biofilm formation and maturation3. The matrix confers considerable fitness advantages over planktonic bacteria in hydration, protection from environmental tensile or shear forces, increased cell–cell communication and enhanced horizontal gene transfer4. The EPS matrix also offers protection from antimicrobials, as bacteria within biofilm structures can resist conventional antibiotic therapies up to 103-fold5,6,7. This inherent ability of biofilms to resist antibiotics occurs through reduced metabolic activity, development of persister cells8 and reduced antibiotic/small molecular diffusion through the EPS polymeric network via charge-interactions9,10,11.
The global rise of antibiotic resistance is an increasing problem, threatening the ability of healthcare providers to treat common bacterial infections, such as hospital-acquired methicillin-resistant Staphylococcus aureus (MRSA)12. Multidrug resistance in bacteria is also of increasing concern due to their ability to acquire multiple resistance mechanisms through horizontal gene transfer. More worrying still is the emergence of resistance to the so called ‘antibiotics of last resort;’ i.e. the polypeptide polymyxin antibiotics13,14, particularly those carried on mobile genetic elements such as plasmids. The colistin-resistant mcr-1 gene is most often isolated in E. coli15,16, although recently it has also been found to have spread to Klebsiella sp. carried on a broad host range, self-transferable IncP plasmid17 suggesting the high likelihood of imminent further spread to other Gram-negative species.
Modification of biofilm assembly and targeted disruption of the cross-linked network of “entangled polymers” within the biofilm EPS matrix would be advantageous in both clinical and industrial applications, especially with the global rise of antibiotic resistance. Most strategies fall into one of two categories; biofilm disruption or biofilm prevention. To date, anti-biofilm strategies have amongst others, included surface/substrate modification (to modify initial bacterial adhesion)18, disruption of the biofilm matrix using conventional antibiotics19, antimicrobial peptides20, dispersal agents21, detergents22, chelators (e.g., EDTA)23, EPS synthesis inhibitors24, and dysregulation of quorum sensing within biofilms25,26. In attempting to design and deliver new antimicrobial and anti-biofilm therapies, the ability to accurately measure the effects of such potential therapies upon the biofilm and biofilm matrix, and accurately quantify the complexity and variability of these biofilms is currently challenging. Conventional biofilm characterization techniques include scanning electron microscopy (SEM) and confocal laser scanning microscopy (CLSM), often combined with image analysis. These techniques, however, fail to accurately characterize the EPS structure of treated biofilms, due to extensive sample preparation and/or sample dehydration, or the lack of universal dyes for EPS staining27. Moreover, the EPS biofilm matrix varies depending on species, strain and growth environment, for example, P. aeruginosa produces different types of polysaccharides with varying charge namely, cationic Pel and neutral Psl, as well as anionic alginate28,29,30 which further complicates characterization.
To understand the key ‘fitness’ advantages that biofilms possess against antimicrobial treatment, workers have sought to characterize the diffusion and material properties of the whole biofilm matrix using mesoscale and nanoscale technologies, without the need to visualize the actual EPS matrix itself. Such technologies include shear and extensional rheology31,32, fluorescence correlation spectroscopy (FCS)33,34 and fluorescence recovery after photobleaching (FRAP)35. Both FCS and FRAP employ fluorescently-labelled particles which can be traced within the biofilm structures. In FRAP, an area of biofilm is photo-bleached and fluorescence recovery (into the area) is modelled to determine diffusion parameters, whereas FCS is based on diffusion measurements from single-molecule fluorescence intensity fluctuations within a discrete region of the biofilm34,36.
Multiple particle tracking (MPT) is a recently described technique, allowing simultaneous tracking of nano-sized particles using fluorescent microscopy, from which the diffusion-based parameters of embedded particles within the EPS of bacterial biofilms can be determined37. MPT also facilitates measurement of the micro-rheological properties of biofilms38. MPT, when used in conjunction with nanoparticles (NPs) of discrete size and charge, represents a non-invasive technique which can be readily employed in situ within biofilms. MPT has subsequently been employed to characterize the diffusion properties of NPs within biofilms of a range of bacterial species including P. aeruginosa, E. coli, P. fluorescens and S. aureus and also to determine time-dependent changes in the biofilm matrix following adhesion36,37,39,40.
In this study, we sought to develop the MPT biofilm model to detect variations in the biofilm structure of Gram-negative P. aeruginosa and Gram-positive S. aureus biofilms and examine its sensitivity and correlation to CLSM imaging. Using polymyxin B-sensitive (PMBSens) and resistant (PMBR) E. coli strains, we examined the sensitivity of the MPT biofilm model to detect variations in the biofilm structure after dose-dependent polymyxin B antibiotic therapy in comparison to traditional confocal microscopy. By using commercially available NPs, this study highlights the usefulness and sensitivity of the MPT technique in the development of novel anti-biofilm therapeutics for AMR infections.
Results
CLSM imaging and MPT measurements identify distinct variations in S. aureus and P. aeruginosa biofilms
CLSM imaging of S. aureus 1004A (MRSA) and P. aeruginosa PAO1 revealed variation in the structural properties of the biofilms produced by the two strains. While MRSA formed a thin, but bacterially-dense biofilm structure, PAO1 biofilms possessed greater height, but appeared less bacterially-dense (Supplementary Fig. 1). These CLSM images correlated well with diffusion of the negatively charged NPs (40–500 nm) through the biofilm, with NP diffusion being lower in the MRSA biofilms when compared to those of PAO1 (40 and 200 nm NPs; P < 0.01). In contrast, diffusion of the positively charged NPs (200 nm) in both biofilms were similar (P > 0.05; Table 1).
MPT revealed that as the size of the negatively charged particles increased, the diffusion coefficient of the particles within PAO1 biofilms decreased significantly (40–200 nm; P < 0.05). This significant trend was also evident in the MRSA biofilms for particle sizes of 40–200 nm. Also, while diffusion of the positively charged 200 nm NPs in PAO1 biofilms was significantly reduced when compared with the diffusion of the negatively charged NPs of the same size (P < 0.05), in MRSA biofilms, the diffusion of the positively charged NPs was not significantly different when compared to the negatively charged particles (P > 0.05; Table 1).
The ratio of the biofilm diffusion coefficient to the diffusion coefficient in water (<Deff>/D°) allows measurement of NP diffusion through the biofilm structure in relation to the intrinsic free Brownian motion of the NPs in water, thereby taking into account the impact of NP size on its unrestricted diffusion in liquid. The percentage ratio of <Deff>/D° revealed that the diffusion of 200 and 500 nm sized NPs were significantly lower than the diffusion of 40 and 100 nm NPs in both MRSA and PAO1 biofilms (P < 0.05). However, negatively charged NP diffusion was still greater (by at least 2.5 times) in PAO1 biofilms when compared to their diffusion in MRSA biofilms (Table 1).
PAO1 biofilms displayed greater heterogeneity in negatively charged NP movement with resultant 90th/10th percentile ratios of 100–987, when compared to MRSA biofilms which displayed much lower ratios of 3–85 (Fig. 1). The heterogeneity data again confirmed the observed variations in the structural properties between MRSA and PAO1 biofilms seen in the CLSM images. The data also revealed that there was increasing heterogeneity in NP diffusion with increasing particle size until the NP size reached 100 nm for S. aureus and 200 nm for P. aeruginosa biofilm systems, after which heterogeneity in NP diffusion then decreased.
Heterogeneity of 40, 100, 200 and 500 nm negatively charged carboxylate-modified FluoSphere® NP movement through P. aeruginosa and S. aureus biofilms. For each particle type, the effective diffusion coefficient ratio of <Deff> was calculated for 360 individual particles (n = 3 for biofilm experiments, each comprised of 120 particles) over a time interval of 20 s and then data was ranked into percentiles from the 90th through to 10th percentile. a, e Negatively charged 40 nm carboxylate-modified FluoSphere® in P. aeruginosa and S. aureus biofilms, respectively; b, f negatively charged 100 nm carboxylate-modified FluoSphere® in P. aeruginosa and S. aureus biofilms, respectively; c, g negatively charged 200 nm carboxylate-modified FluoSphere® in P. aeruginosa and S. aureus biofilms, respectively; d, h negatively charged 500 nm carboxylate-modified FluoSphere® in P. aeruginosa and S. aureus biofilms, respectively. Fold difference in the figure indicates the fold difference between the 90th and 10th percentiles.
The exponential anomalous values of both PAO1 and MRSA biofilms calculated using 40–500 nm NPs appeared to demonstrate a viscous response of these biofilms (α > 0.5; Fig. 2).
Exponential anomalous values <α> of a P. aeruginosa and b S. aureus biofilms using 40, 100, 200 and 500 nm negatively charged carboxylate-modified FluoSpheres®. α is measured based on the relation between the ensemble mean square displacement <MSD> versus time scale of the traced FluoSphere® particles and reflects the micro-rheological degree of resistance of the biofilm towards traced particles where; α > 0.5 indicates viscous resistance; α < 0.5 indicates elastic resistance. <MSD> represents the geometric mean of MSDs of 360 particles (n = 3, each 120 particles).
CLSM and MPT measurements describe dose-dependent disruption of E. coli IR57 biofilms treated with polymyxin B
MIC assays of PMBSens E. coli IR57 and PMBR E. coli PN47 were performed to confirm strain sensitivity to the antibiotic, giving MIC values against polymyxin B of 0.06 and 2 µg/ml, respectively.
The CLSM assay revealed cellular aggregation and disruption of the PMBSens E. coli IR57 biofilm matrix in a dose-dependent manner following polymyxin B treatment (Fig. 3a). This finding was confirmed by MPT through dose-dependent increases in the effective diffusion coefficients (<Deff>) within the treated biofilms for all three NP sizes tested (100, 200 and 500 nm; Fig. 3), with the 100 nm NPs revealing the greatest dose-dependent sensitivity in diffusion coefficients to antibiotic treatment. The dose-dependent changes in NP diffusion coefficients, starting from 2 µg/ml polymyxin B treatment, were all significantly different from the control (P < 0.05).
Polymyxin B (PMB)-treated (2, 8, 16, 32, 64 µg/ml) and untreated PMBSens E. coli IR57 biofilms showing: a CLSM 3D and side-on imaging of biofilms grown for 48 h followed by polymyxin B treatment for a further 24 h at 37 °C, visualized using Syto9® staining (scale bar, 40 µm; n = 3). b Diffusion coefficient <Deff> of 100, 200 and 500 nm negatively charged carboxylate-modified FluoSphere® particles in non-treated versus polymyxin B treated biofilms (n = 3, ± SEM). *indicates FluoSphere® particle sizes of 100, 200 and 500 nm.
Polymyxin B treatment (2–8 µg/ml) revealed a greater increase in the mean square displacement <MSD> versus time measurements for the 200 and 500 nm particles when compared to the 100 nm particles, indicative of increasing pore size within the biofilm structure following polymyxin B treatment (Fig. 4). The exponential anomalous values of the E. coli biofilms demonstrated that polymyxin B treatment had little effect on the viscoelastic response of the biofilms (Fig. 4). However, the dose-dependent disruption of the E. coli IR57 biofilms was reflected in increasing biofilm creep compliance (J(t)) with increasing polymyxin B dose, observed with all 3 NP sizes tested (Fig. 5).
Exponential anomalous values <α> of E. coli IR57 biofilms in response to polymyxin B treatment using 100, 200 and 500 nm negatively charged carboxylate-modified FluoSpheres® a untreated control; b 2; c, 8; d, 16; e, 32; and f 64 µg/ml polymyxin B treatment. α is measured based on the relation between the ensemble mean square displacement (MSD) versus time scale of the traced FluoSphere® particles and reflects the micro-rheological degree of resistance of the biofilm towards traced particles where; α > 0.5 indicates viscous resistance; α < 0.5 indicates elastic resistance. <MSD> represents the geometric mean of MSDs of 360 particles (n = 3, each 120 particles).
Creep compliance (J(t)) of E. coli IR57 (PMBSens) biofilms (measured using the ensemble mean square displacement <MSD> versus lag time of 100, 200 and 500 nm negatively charged carboxylate-modified FluoSphere® particles) representing biofilm deformation in response to polymyxin B treatment (2, 8, 16, 32 and 64 µg/ml) versus untreated biofilm. a Untreated E. coli biofilms for 100, 200, and 500 nm FluoSphere® particles, b 100 nm FluoSphere® particles, c 200 nm FluoSphere® particles, and d 500 nm FluoSphere® particles, in response to polymyxin B treatment.
MPT measurements describe distinct variations in the response of PMBSens E. coli IR57 and PMBR E. coli PN47 biofilms to polymyxin B treatment
To assess the impact of resistance to polymyxin B on treatment of E. coli biofilms, two strains of E. coli (PMBR PN47 and PMBSens IR57) were selected based on their susceptibility to polymyxin B. The MPT assay revealed that the diffusion coefficient <Deff> of 200 nm NPs was greatly increased in the PMBSens IR57 biofilms (0.0643 vs. 0.34 cm2 s−1 × 10−9) following treatment, while NP diffusion within the PMBR PN47 biofilms remained largely unchanged (Table 2). This trend was reflected in the heterogeneity of NP diffusion measurements (Fig. 6), where the PMBSens IR57 biofilm demonstrated decreased 90th/10th percentile ratio of 2577 to 198 following treatment, revealing more homogenous NP diffusion, indicative of increasing biofilm pore size with antibiotic treatment. As before, the heterogeneity of the NP diffusion measurements within the PMBR PN47 biofilms demonstrated little response to treatment. This result was also confirmed by the creep compliance data showing increasing values following polymyxin B treatment in the PMBSens strain, but not in the PMBR strain (Fig. 7). Interestingly, NP diffusion (<Deff>) within the PMBSens IR57 biofilm was more than three times greater than the <Deff> for PMBR PN47 biofilm (Table 2). NP diffusion within the PMBSens strain appeared to be vastly more heterogeneous than for the PMBR strain, indicating that pore size was more heterogeneous in the biofilms of the PMBSens strain (Fig. 6).
Heterogeneity of 200 nm negatively charged carboxylate-modified FluoSpheres® movement within E. coli biofilms. Untreated controls a PMBSens E. coli IR57, b PMBR E. coli PN47. Polymyxin B (PMB; 8 µg/ml) treated biofilms, c PMBsens E. coli IR57, d PMBR E. coli PN47. For each particle type, the effective diffusion coefficient <Deff> was calculated for 360 individual particles (n = 3 for biofilm experiments, each comprised of 120 particles) over a time interval of 20 s and the data ranked into percentiles (90th to 10th). Fold difference in the figure indicates the fold difference between the 90th and 10th percentiles.
Discussion
NPs are being increasingly applied in a variety of medical applications, such as in the diagnosis and treatment of human disease (e.g. drug- and gene delivery)41 and as novel research tools (both in vitro and in vivo) to improve our understanding of biological systems and human illness42,43,44. This study has demonstrated the use of MPT with traceable NPs as a robust, non-invasive, in situ technique to inform our understanding of biofilms and further our insight into the potential effects of antimicrobial and anti-biofilm therapies on the biofilm matrix, and on biofilm-related changes induced with the acquisition of antibiotic resistance.
The EPS matrix of biofilms varies not only between species and strains, but also with differences in environmental growth conditions, such as surface composition/roughness, nutrient availability, temperature, and hydrodynamic shear38,45. As a result, the net charge and functional groups present within the matrix, as well as biofilm pore size, exhibit considerable variations. To assess variations in biofilm structure between bacterial species, the diffusion coefficients of fluorescently-labelled NPs were initially measured in both Gram-positive S. aureus 1004A (MRSA) and Gram-negative P. aeruginosa PAO1, revealing the influence of both NP size and surface-charge on diffusion through the respective biofilm structures. The influence of NP charge was clearly evident in the diffusion coefficient measurements of 200 nm NPs within P. aeruginosa PAO1 biofilms, with reduced diffusion of amine-modified (positively charged) particles compared to the carboxylate-modified (negatively charged) NPs. In P. aeruginosa PAO1, the exopolysaccharides form a major part of the EPS matrix and are composed of negatively charged alginate, neutrally-charged Psl and positively charged Pel, while eDNA in biofilm matrixes has been shown to be negatively charged46. Our results indicate a net negative charge of the P. aeruginosa EPS matrix, where the existence of increased negative charges within the EPS matrix reduced the diffusion coefficient of the positively charged amine-modified NPs. In S. aureus MRSA biofilms, while the diffusion coefficient of the amine-modified NPs was twice that of the carboxylated-modified NPs, these values were not significantly different. In S. aureus, the two major components of EPS matrix are the positively charged poly-N-acetylgluosamine (PNAG) polysaccharides and negatively charged eDNA47,48. These results potentially indicate a net positive charge of the MRSA EPS matrix; increased positive charges within the matrix reducing the diffusion coefficient of the negatively charged carboxylated NPs. However, as the diffusion coefficient values achieved with 200 nm carboxylate-modified NPs were similar to those observed with 500 nm NPs in MRSA (0.0031 vs 0.0038 cm2 S−1 × 10−9, respectively), this indicated the inability of the 200 nm NPs to freely move within the dense MRSA biofilm matrix. The restricted movement of 200 nm NPs within the MRSA biofilms resulted in small, distinct differences in diffusion coefficient between the amine-modified NPs and carboxylate-modified NPs, which may have been more pronounced with the use of smaller sized amine-modified NPs.
In this model, NP diffusion into the biofilm EPS matrix was not only clearly influenced by charge, but also by their size49,50,51. Large particles may not be able to permeate into the biofilm EPS structure due to steric hindrance or biofilm pore size, while hydrophobic/electrostatic interactions may influence the diffusion of smaller particles38. Previous researchers have used a variety of techniques to characterise the internal structure and pore size of biofilms. Zhang et al.52 originally employed sectioning by microtome and dye adsorption to describe the heterogeneity in pore size within wastewater biofilms, where biofilm pore sizes ranged between 0.3 and 2.7 µm. More recently, Rosenthal et al.53 used optical coherence tomography (OCT) to characterize the heterogeneous network of a thick, multi-species biofilm, where pore diameters as large as 110 µm were measured, while other researchers have demonstrated pore-size ranges of 500–1000 nm54 and >100 nm55 using single-particle tracking techniques.
In the current study, the effect of NP size on diffusion was clearly evident in PAO1 and MRSA biofilms, where a reduction in NP size (<200 nm), induced a dramatic rise in particle diffusion; a finding possibly indicative of a mean biofilm pore size <200 nm. Similarly, Peulen and Wilkinson56 demonstrated increasing diffusion coefficients with decreasing size and negative charge of silver NPs in Pseudomonas fluorescens biofilms, where the optimum particle size for diffusion in these dense biofilms was <50 nm. Increased NP diffusion coefficients in PAO1 biofilms when compared to the MRSA biofilms (40–500 nm carboxylate-modified NPs) were evident in this study and reflected the biofilm architecture; CLSM imaging demonstrating the more dense structure of the MRSA biofilms.
Profiling of the diffusion coefficients of 360 individual particles through ranking <Deff> data into percentiles (highest 90th to lowest 10th percentiles) provides insight into the heterogeneity of particle movement, which may be indicative of heterogeneous pore sizes within the biofilm itself57,58. This study revealed that NP diffusion coefficients became more heterogeneous with increasing particle size until a NP size of 100 nm for S. aureus and 200 nm for P. aeruginosa biofilms was reached, at which point NP diffusion became more homogeneous (the particle size becoming too large to facilitate free movement within the biofilm structure) with NP movement occurring only through larger pore sizes. This data demonstrates contrasting inter-species biofilm architecture; P. aeruginosa forming a pore structure that was larger in size and more heterogeneous compared to that of S. aureus biofilms.
MPT may, in the future, offer valuable insight in characterizing the EPS matrix of heterogeneous polymicrobial biofilms which are commonly found in human disease states59,60. While these biofilms are heterogeneous, they are often composed of distinct, individual ‘pockets’ of homogenous single-species growth and MPT coupled with GFP-labelled bacterial populations or fluorescence in situ hybridization (FISH) labelling could be employed to visualize and analyse the biofilm matrix of these bacterial populations.
Structurally, biofilms possess a complex architecture, being composed of cell clusters surrounded by voids and water channels61. Biofilm structures possess viscoelastic properties (exhibiting both elastic and viscous properties) which can aid biofilm survival under mechanical and chemical loads62,63. Cao et al.37 employed single-particle tracking with CLSM imaging to characterize the viscoelastic properties of cell clusters and voids within P. fluorescens biofilms, demonstrating that the viscoelastic properties (creep compliance) of the ‘biofilm void’ zones was the primary contributor to the viscoelastic properties of the biofilm. Importantly, while the larger (500 nm) NPs employed in this study were unable to diffuse into the bacterial cell clusters, their diffusion into the biofilm voids could still provide a sensitive and reproducible model to monitor changes in the viscoelastic properties of biofilms. In this study, calculation of the exponential anomalous values revealed that PAO1 and MRSA biofilms displayed viscous behaviour when assessed using 40–500 nm NPs, however, no micro-rheological differences could be determined between the well-established 72 h grown PAO1 and MRSA biofilms.
The resistance of bacterial biofilms to antibiotic therapy has been widely reported in the literature, with high levels of antibiotic tolerance arising due to a number of different factors64. Penetration of antibiotics through the biofilm EPS matrix can vary considerably depending on how they interact with the charged components of the EPS10,11. Previous research has shown that cationic antibiotics (e.g. tobramycin) exhibit charge-mediated binding to polyanions within the biofilm EPS matrix, resulting in reduced penetration of antibiotics into the biofilm structure9,65. Sankaran et al.34, however, demonstrated that both labelled aminoglycoside tobramycin (positively-charged) and the fluoroquinolone ciprofloxacin (neutrally-charged) were able to diffuse into, and remained mobile within the interior of P. aeruginosa biofilms, demonstrating that reduced antibiotic diffusion in the EPS matrix is not solely responsible for the antibiotic tolerance of biofilms66. Moreover, the development of a sub-population of persister cells (with low or dormant metabolic activity) may ensure that a small cohort of cells within the biofilm can withstand multiple doses of antimicrobial therapy67. Expression of biofilm-specific genetic mechanisms may also occur11, making the biofilm less susceptible to antimicrobial attack.
Zrelli et al.68 and Reighard et al.39 demonstrated the failure of antibiotic treatment (afloxacin, ticarcillin, tobramycin) to induce changes in the NP diffusion and mechanical properties of E. coli and P. aeruginosa biofilms using particle tracking methodologies. In contrast, here we demonstrate the dose-dependent disruption of the PMBsens E. coli biofilm matrix induced by exposure to polymyxin B, with significant increases in NP diffusion and creep compliance in the treated biofilms. Klinger-Strobel et al.69 demonstrated the ability of colistin (polymyxin E) to disrupt 48 h E. coli biofilms at concentrations of 4–16 µg/ml (with MICs ranging from 1 to 0.0625 µg/ml for laboratory strains and clinical isolates, respectively). The authors suggested that colistin destabilized the biofilm matrix (even in strains with intrinsic polymyxin resistance) leading to the dispersal of planktonic cells that were then more susceptible to antibiotics. The published susceptibility breakpoints for polymyxins are S ≤ 2 µg/ml > R for E. coli (EuCAST 2018)70. The concentrations of polymyxin B used in our study were relatively high (2–64 µg/ml), equal to, or above, the MIC values (0.06–2 µg/ml), to induce effective disruption of the biofilm structure. However, differences in NP diffusion were detected at lower concentrations (2 µg/ml) than that employed by Klinger-Strobel et al.69.
Witten and Ribbeck38 proposed that biofilm permeability may be a biomarker for antimicrobial resistance, and the data here demonstrates the sensitivity of the MPT biofilm assay in this respect. In the PMBsens E. coli IR57 (MIC 0.06 µg/ml) biofilms, MPT was able to detect structural, pore-size related changes induced within the biofilm matrix with antibiotic dosing as low as 2 µg/ml (a clinically-relevant concentration) where changes in the treated biofilm structure were indiscernible at this concentration using conventional CLSM imaging. The disruptive effect of polymyxin B on the E. coli biofilm structure was, however, evident in CLSM imaging at higher concentrations (>32 µg/ml). We also sought to model biofilm disruption by polymyxin B in PMBR E. coli as we have previously demonstrated that mcr-1 and -3 expression in E. coli is associated with alterations in bacterial viability within the biofilm matrix16,71, hypothesizing that the acquisition of polymyxin resistant mcr-1 was an evolutionary ‘trade-off’; resistance to polymyxin being associated with decreased biofilm biomass and growth in 24 h biofilm formation assays71. Here, we clearly demonstrate the resistance to biofilm disruption of PMBR PN47 following treatment with polymyxin B (8 µg/ml) in well-established 72 h grown biofilms.
The observed variations in particle diffusion and creep compliance with antibiotic/antimicrobial treatment in this study could also be crucial in understanding the interaction of the innate immune system with biofilm infections. Increased pore size and decreased mechanical properties of the biofilm structure may facilitate increased inflammatory cell penetration into the biofilm matrix, thereby improving bacterial clearance72. Moreover, increased porosity of biofilm structures may allow greater penetration of antimicrobial agents and may be of clinical benefit, especially in treating implant biofilm-related infections73.
This study demonstrated not only the ability of MPT to detect disruption in biofilm structures, but also revealed the ability of the technique to dissect the effects of antimicrobial and anti-biofilm therapies, not always discernible using conventional technologies. Here, the ability of MPT to inform understanding of, and test therapies against, emergent antimicrobial-resistant pathogens is clear. As anti-biofilm therapies play a clear role in increasing susceptibility of resistant bacteria to existing therapies, the utility of MPT in assessing novel therapies to disrupt the biofilm matrix, e.g. chelating agents23 and G-block alginate oligomers31,74 may be invaluable in the future. MPT may further our insight into the potential effects of such antimicrobial therapies in vivo and provide increased understanding into the biofilm matrix and biofilm-related changes induced with the acquisition of antibiotic resistance.
Methods
Bacterial strains, growth media and culture conditions
The following strains were used in this study; Pseudomonas aeruginosa PAO1, methicillin-resistant Staphylococcus aureus (MRSA 1004A)75, polymyxin B-resistant (PMBR) Escherichia coli PN47 (carrying the colistin resistance plasmids mcr-1 and mcr-3)76 and polymyxin B-sensitive (PMBSens) E. coli IR57. Bacterial colonies were sub-cultured on LB agar plates supplemented with/without 2 µg/ml polymyxin B (Sigma-Aldrich). Overnight bacterial cultures were grown in Tryptone Soy Broth (TSB; Oxoid) for 37 °C at 120 rpm. Biofilms were grown in cation-adjusted Mueller Hinton Broth (MHB; LabM) with/without supplementation with the antibiotic polymyxin B.
Minimum inhibitory concentration (MIC) measurements
Overnight bacterial cultures were adjusted to a standardized cell suspension of ~108 colony forming units (CFU)/ml (equivalent to 0.5 McFarland standard). Two-fold serial dilutions in polymyxin B were prepared in MHB within flat-bottom 96-well microtiter plates (100 µl per well). The adjusted O/N bacterial cultures were then diluted 10-fold in MHB and 5 µl added to the microtiter plate containing the antibiotic serial dilutions to give a final concentration of 5 × 105 CFU/ml. The plates were incubated for 16–20 h (37 °C) and MICs determined as the lowest concentration at which there was no visible growth.
Biofilm growth
Overnight bacterial cultures were adjusted to a standardized cell suspension of 1 × 107 CFU/ml in MHB. First, 0.2 ml of adjusted O/N culture was placed into the centre of each glass well within a 12-well dish (glass thickness 1.5 and 14 mm diameter; MatTek) and incubated statically for 1 h. Then, 1.8 ml of fresh MHB was placed into each well and incubated statically for 24 h at 37 °C. A further 1 ml of MHB was then placed into each well and plates incubated for a further 24 h (37 °C). Following 48 h growth, biofilms were treated with a further 1 ml of MHB which was added into each well with a further incubation of 24 h (37 °C), resulting in a total biofilm growth time of 72 h.
Polymyxin B treatment of E. coli biofilms
Following 48 h growth, E. coli biofilms were treated with either 1 ml of MHB (control) or 1 ml polymyxin B (2, 8, 16, 32, 64 µg/ml; treatment) which was added into each well with a further incubation of 24 h (37 °C), resulting in a total biofilm growth time of 72 h.
SYTO 9 staining and CLSM imaging of biofilms
After 72 h growth, the bacterial supernatant was carefully removed and the bacterial cells within the biofilms stained with 0.5% Syto9® dye (Invitrogen; 400 µl) for 1 h. After staining, the biofilms were washed with phosphate-buffered saline (PBS; x2) prior either to CLSM Z-stack imaging using a Leica TCS SP5 CLSM or NP addition.
MPT measurement of bacterial biofilms
NPs used in this study were either negatively charged carboxylate-modified Fluospheres® (40, 100, 200 and 500 nm) or positively charged amino-modified Fluospheres® (200 nm) ([Ex/Em]: [580/605 nm]; ThermoFisher Scientific). The independent assessment of size and zeta potential values of the NPs were characterized in PBS buffer using a Malvern Zetasizer Nano ZS prior to MPT studies. For MPT experiments, the Fluospheres® suspension was vortexed for 1 min then diluted in sterilized PBS buffer (0.0025%), before addition of 10 µl diluted Fluospheres® suspension onto the biofilms followed by a 2 h incubation. Biofilms were stained with SYTO 9® before addition of the Fluospheres® to visualize the lower layers of the biofilm matrix using a Leica DM IRB wide-field Epifluorescence microscope (×63 oil immersion lens). Videos of particle movement within the biofilms were captured at a frame rate of 33 ms (600 frames, 20 s) using a high-speed camera (Allied Vision Technologies, UK) and then particle trajectories were tracked using ImageJ software (Mosaic) over 2 s to convert NP movements into metric displacements in both the X and Y directions43,77. Ensemble mean square displacement <MSD>, effective diffusion coefficient <Deff>, and heterogeneity of particle diffusion were measured as described in supplementary materials57,78. Here, 120 particle movements were captured in each biofilm well, and each bacterial strain was tested in triplicate (i.e. 360 particles in total for each biofilm species).
The viscoelastic properties of the biofilms were assessed by determining the anomalous diffusion exponent (α). This was calculated by fitting the power law to log(<MSD>) versus log(Δt) and calculating the slope of this data79, where α = 1 for a completely viscous system (liquid), α = 0 for a completely elastic system (solid) and 1 > α > 0 for a viscoelastic system39,40. In addition, the micro-rheological properties of the biofilms were further defined by calculation of the creep compliance (J(t)), where the MSD represents deformation of the biofilm within time (Δt) under constant pressure/shear force represented by the temperature of the atmosphere. Creep compliance (J(t)) was calculated by the following equation:
where kB is the Boltzmann constant, T is absolute temperature and d is the diameter of the particle80.
Statistical analysis
Statistical software (Minitab, State College, PA) was used to calculate significant differences with ANOVA testing with post hoc Tukey multi-comparison tests for the statistical analyses presented.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Data generated and analysed during this study are included in this published article and its Supplemental information file. Additional details are available upon reasonable request.
References
Bjarnsholt, T. The role of bacterial biofilms in chronic infections. APMIS Suppl. 136, 1–51 (2013).
Mazza, M. G. The physics of biofilms—an introduction. J. Phys. D. Appl. Phys. 49, 203001 (2016).
Flemming, H.-C. et al. Biofilms: an emergent form of bacterial life. Nat. Rev. Microbiol. 14, 563–575 (2016).
Boudarel, H., Mathias, J.-D., Blaysat, B. & Grediac, M. Towards standardized mechanical characterization of microbial biofilms: analysis and critical review. NPJ Biofilms Microbiomes 4, 17 (2018).
Mah, T. F. Biofilm-specific antibiotic resistance. Future Microbiol. 7, 1061–1072 (2012).
Ceri, H. et al. The Calgary biofilm device: new technology for rapid determination of antibiotic susceptibilities of bacterial biofilms. J. Clin. Microbiol. 37, 1771–1776 (1999).
Moskowitz, S. M., Foster, J. M., Emerson, J. & Burns, J. L. Clinically feasible biofilm susceptibility assay for isolates of Pseudomonas aeruginosa from patients with cystic fibrosis. J. Clin. Microbiol. 42, 1915–1922 (2004).
Høiby, N., Bjarnsholt, T., Givskov, M., Molin, S. & Ciofu, O. Antibiotic resistance of bacterial biofilms. Int. J. Antimicrob. Agents 35, 322–332 (2010).
Tseng, B. S. et al. The extracellular matrix protects Pseudomonas aeruginosa biofilms by limiting the penetration of tobramycin. Environ. Microbiol. 15, 2865–2878 (2013).
Hunt, B. E., Weber, A., Berger, A., Ramsey, B. & Smith, A. L. Macromolecular mechanisms of sputum inhibition of tobramycin activity. Antimicrob. Agents Chemother. 39, 34–39 (1995).
Mah, T. F. et al. A genetic basis for Pseudomonas aeruginosa biofilm antibiotic resistance. Nature 426, 306–310 (2003).
Lindsay, J. A. Hospital-associated MRSA and antibiotic resistance—what have we learned from genomics? Int J. Med. Microbiol. 303, 318–323 (2013).
Corona, A. & Cattaneo, D. Dosing colistin properly: let’s save ‘Our Last Resort Old Drug. Clin. Infect. Dis. 65, 870 (2017).
Bulman, Z. P. et al. Polymyxin combinations combat Escherichia coli harboring mcr-1 and blaNDM-5: preparation for a post-antibiotic era. MBio 8, e00540-17 (2017).
Poirel, L. et al. Antimicrobial resistance in Escherichia coli. Microbiol. Spectr. 6, 4 (2018).
Yang, Q. et al. Balancing mcr-1 expression and bacterial survival is a delicate equilibrium between essential cellular defense mechanisms. Nat. Commun. 8, 2054 (2017).
Zhao, F., Feng, Y., Lü, X., Mcnally, A. & Zong, Z. IncP plasmid carrying colistin resistance gene mcr-1 in Klebsiella pneumoniae from hospital sewage. Antimicrob. Agents Chemother. 61, e02229-16 (2017).
Lemire, J. A., Harrison, J. J. & Turner, R. J. Antimicrobial activity of metals: mechanisms, molecular targets and applications. Nat. Rev. Micro 11, 371–384 (2013).
Bayramov, D. F. & Neff, J. A. Beyond conventional antibiotics—new directions for combination products to combat biofilm. Adv. Drug Deliv. Rev. 112, 48–60 (2016).
Pletzer, D., Coleman, S. R. & Hancock, R. E. Anti-biofilm peptides as a new weapon in antimicrobial warfare. Curr. Opin. Microbiol. 33, 35–40 (2016).
Barraud, N., Kelso, M. J., Rice, S. A. & Kjelleberg, S. Nitric oxide: a key mediator of biofilm dispersal with applications in infectious diseases. Curr. Pharm. Des. 21, 31–42 (2015).
Otzen, D. E. Biosurfactants and surfactants interacting with membranes and proteins: same but different. BBA: Biomembranes 1859, 639–649 (2017).
Finnegan, S. & Percival, S. L. EDTA: an antimicrobial and antibiofilm agent for use in wound care. Adv. Wound Care 4, 415–421 (2015).
Pandit, S. et al. Low concentrations of Vitamin C reduce the synthesis of extracellular polymers and destabilize bacterial biofilms. Front. Microbiol. 8, 2599 (2017).
Jack, A. A. et al. Alginate oligosaccharide-induced modification of the lasI-lasR and rhlI-rhlR quorum sensing systems in Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 62, e02318–17 (2018).
Brackman, G. & Coenye, T. Quorum sensing inhibitors as anti-biofilm agents. Curr. Pharm. Des. 21, 5–11 (2015).
Pan, M., Zhu, L., Chen, L., Qiu, Y. & Wang, J. Detection techniques for extracellular polymeric substances in biofilms: a review. BioRes 11, 8092–8115 (2016).
Jennings, L. K. et al. Pel is a cationic exopolysaccharide that cross-links extracellular DNA in the Pseudomonas aeruginosa biofilm matrix. Proc. Natl Acad. Sci. USA. 112, 11353–11358 (2015).
Billings, N. et al. The extracellular matrix component Psl provides fast-acting antibiotic defense in Pseudomonas aeruginosa biofilms. PLoS Pathog. 9, e1003526 (2013).
Franklin, M. J., Nivens, D. E., Weadge, J. T. & Howell, P. L. Biosynthesis of the Pseudomonas aeruginosa extracellular polysaccharides, alginate, Pel, and Psl. Front. Microbiol. 167, 1–16 (2011).
Powell, L. C. et al. The effect of alginate oligosaccharides on the mechanical properties of Gram-negative biofilms. Biofouling 29, 413–421 (2013).
Pritchard, M. F. et al. A low-molecular-weight alginate oligosaccharide disrupts pseudomonal microcolony formation and enhances antibiotic effectiveness. Antimicrob. Agents Chemother. 61, e00762-17 (2017).
Gulot, E. et al. Heterogeneity of diffusion inside microbial biofilms determined by fluorescence correlation spectroscopy under two-photon excitation. Photochem Photobio. 75, 570–578 (2002).
Sankaran, J. et al. Single microcolony diffusion analysis in Pseudomonas aeruginosa biofilms. NPJ Biofilms Microbiomes 5, 35 (2019).
Waharte, F., Steenkeste, K., Briandet, R. & Fontaine-Aupart, M. P. Diffusion measurements inside biofilms by image-based fluorescence recovery after photobleaching (FRAP) analysis with a commercial confocal laser scanning microscope. Appl. Environ. Microbiol. 76, 5860–5869 (2010).
Billings, N., Birjiniuk, A., Samad, T. S., Doyle, P. S. & Ribbeck, K. Materials properties of biofilms—a review of methods for understanding permeability and mechanics. Rep. Prog. Phys. 78, 036601 (2015).
Cao, H. et al. Revealing region-specific biofilm viscoelastic properties by means of a micro-rheological approach. NPJ Biofilms Microbiomes 2, 5 (2016).
Witten, J. & Ribbeck, K. The particle in the spider’s web: transport through biological hydrogels. Nanoscale 9, 8080–8095 (2017).
Reighard, K. P., Hill, D. B., Dixon, G. A., Worley, B. V. & Schoenfisch, M. H. Disruption and eradication of P. aeruginosa biofilms using nitric oxide-releasing chitosan oligosaccharides. Biofouling 31, 775–787 (2015).
Chew, S. C., Rice, S. A., Kjelleberg, S. & Yang, L. In situ mapping of the mechanical properties of biofilms by particle-tracking microrheology. J. Vis. Exp. 4, e53093 (2015).
Zazo, H., Colino, C. I. & Lanao, J. M. Current applications of nanoparticles in infectious diseases. J. Control Release 224, 86–102 (2016).
Natan, M. & Banin, E. From nano to micro: using nanotechnology to combat microorganisms and their multidrug resistance. FEMS Microbiol. Rev. 41, 302–322 (2017).
Inchaurraga, L. et al. Modulation of the fate of zein nanoparticles by their coating with a Gantrez® AN-thiamine polymer conjugate. Int J. Pharm. X 1, 100006 (2019).
Brotons-Canto, A. et al. Evaluation of nanoparticles as oral vehicles for immunotherapy against experimental peanut allergy. Int. J. Biol. Macromol. 110, 328–335 (2018).
Fulaz, S., Vitale, S., Quinn, L. & Casy, E. Nanoparticle-biofilm interactions: the role of the EPS matrix. Trends Microbiol. 27, 915–926 (2019).
Jennings, L. K. et al. Pel is a cationic exopolysaccharide that cross-links extracellular DNA in the Pseudomonas aeruginosa biofilm matrix. PNAS 112, 11353–11358 (2015).
Dengler, V., Foulston, L., DeFrancesco, A. & Losick, R. An electrostatic net model for the role of extracellular DNA in biofilm formation by Staphylococcus aureus. J. Bacteriol. 197, 3779–3787 (2015).
Hiltunen, A. K. et al. Structural and functional dynamics of Staphylococcus aureus biofilms and biofilm matrix proteins on different clinical materials. Microorganisms 7, 584 (2019).
Campoccia, D., Montanaro, L. & Arciola, C. R. A review of the biomaterials technologies for infection-resistant surfaces. Biomaterials 34, 8533–8554 (2013).
Habimana, O., Steenkeste, K., Fontaine-Aupart, M. P., Bellon-Fontaine, M.-N. & Briandet, R. Diffusion of nanoparticles in biofilms is altered by bacterial cell wall hydrophobicity. Appl. Environ. Microbiol. 77, 367–368 (2011).
Ikuma, K., Decho, A. W. & Lau, B. L. T. When nanoparticles meet biofilms—interactions guiding the environmental fate and accumulation of nanoparticles. Front. Microbiol. 6, 591 (2015).
Zhang, T. C. & Bishop, P. L. Density, porosity and pore structure of biofilms. Water Res. 28, 2267–2277 (1994).
Rosenthal, A. F. et al. Morphological analysis of pore size and connectivity in a thick mixed culture biofilm. Biotechnol. Bioeng. 115, 2268–2279 (2018).
Chew, S. C. et al. Dynamic remodeling of microbial biofilms by functionally distinct exopolysaccharides. MBio 5, e01536-14 (2014).
Forier, K. et al. Transport of nanoparticles in cystic fibrosis sputum and bacterial biofilms by single-particle microscopy. Nanomedicine 8, 935–949 (2013).
Peulen, T. O. & Wilkinson, K. J. Diffusion of nanoparticles in a biofilm. Environ. Sci. Technol. 45, 3367–3373 (2011).
Abdulkarim, M. et al. Nanoparticle diffusion within intestinal mucus: three-dimensional response analysis dissecting the impact of particle surface charge, size and heterogeneity across polyelectrolyte, pegylated and viral particles. Eur. J. Pharm. Biopharm. 97, 230–238 (2015).
Sahle-Demessie, E. & Tadesse, H. Kinetics and equilibrium adsorption of nano-TiO2 particles on synthetic biofilm. Surf. Sci. 60, 1177–1184 (2011).
Zijnge, V. et al. Oral biofilm architecture on natural teeth. PLoS ONE 5, e9321 (2010).
Bjarnsholt, T. et al. Why chronic wounds will not heal: a novel hypothesis. Wound Repair Regen. 16, 2–10 (2008).
Donlan, R. M. & Costerton, J. W. Biofilms: survival mechanisms of clinically relevant microorganisms. Clin. Microb. Rev. 15, 167–193 (2002).
Peterson, B. W. et al. Viscoelasticity of biofilms and their recalcitrance to mechanical and chemical challenges. FEMS Microbiol. Rev. 39, 234–245 (2015).
Rogers, S. S., van der Walle, C. & Waigh, T. A. Microrheology of bacterial biofilms in vitro: Staphylococcus aureus and Pseudomonas aeruginosa. Langmuir 24, 13549–13555 (2008).
Lebeaux, D., Ghigo, J.-M. & Beloin, C. Biofilm-related infections: bridging the gap between clinical management and fundamental aspects of recalcitrance toward antibiotics. Microbiol. Mol. Biol. Rev. 78, 510–543 (2014).
Huang, J. X. et al. Mucin binding reduces colistin antimicrobial activity. Antimicrob. Agents Chemother. 59, 5925–5931 (2015).
Walters, M. C. III, Roe, F., Bugnicourt, A., Franklin, M. J. & Stewart, P. S. Contributions of antibiotic penetration, oxygen limitation, and low metabolic activity to tolerance of Pseudomonas aeruginosa biofilms to ciprofloxacin and tobramycin. Antimicrob. Agents Chemother. 47, 317–323 (2003).
Percival, S. L., Hill, K. E., Malic, S., Thomas, D. W. & Williams, D. W. Antimicrobial tolerance and the significance of persister cells in recalcitrant chronic wound biofilms. Wound Repair Regen. 19, 1–9 (2011).
Zrelli, K. et al. Bacterial biofilm mechanical properties persist upon antibiotic treatment and survive cell death. N. J. Phys. 15, 125026 (2013).
Klinger-Strobel, M., Stein, C., Forstner, C., Makarewicz, O. & Pletz, M. W. Effects of colistin on biofilm matrices of Escherichia coli and Staphylococcus aureus. Int J. Antimicrob. Agents 49, 471–479 (2017).
EUCAST. Breakpoint tables for interpretation of MICs and zone diameters. http://www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/Breakpoint_tables/v_8.1_Breakpoint_Tables.pdf (2018).
Yang, Q. E. et al. Compensatory mutations modulate the competitiveness and dynamics of plasmid-mediated resistance in Escherichia coli clones. ISME J. 14, 861–865 (2020).
Geddes-McAlister, J., Kugadas, A. & Gadjeva, M. Tasked with a challenging objective: Why do neutrophils fail to battle Pseudomonas aeruginosa biofilms. Pathogens 8, 283 (2019).
Connaughton, A., Childs, A., Dylewski, S. & Sabesan, V. J. Biofilm disrupting technology for orthopedic implants: what’s on the horizon? Front. Med. 1, 22 (2014).
Powell, L. C. et al. Targeted disruption of the extracellular polymeric network of Pseudomonas aeruginosa biofilms by alginate oligosaccharides. NPJ Biofilms Microbiomes 4, 13 (2018).
Howell-Jones, R. S. Antibiotic use in the Treatment of Chronic Wounds. PhD Thesis, Cardiff University (2007).
Yang, Q. E. et al. Environmental dissemination of mcr-1 positive Enterobacteriaceae by Chrysomya spp. (common blowfly): an increasing public health risk. Environ. Int 122, 281–290 (2018).
Bonengel, S. et al. Impact of different hydrophobic ion pairs of octreotide on its oral bioavailability in pigs. J. Control Release 273, 21–29 (2018).
Rohrer, J. et al. Mucus permeating thiolated self-emulsifying drug delivery systems. Eur. J. Pharm. Biopharm. 98, 90–97 (2016).
Schuster, B. S., Suk, J. S., Woodworth, G. F. & Hanes, J. Nanoparticle diffusion in respiratory mucus from humans without lung disease. Biomaterials 34, 3439–3446 (2013).
Birjiniuk, A. et al. Single particle tracking reveals spatial and dynamic organization of the Escherichia coli biofilm matrix. N. J. Phys. 16, 085014 (2014).
Acknowledgements
We thank Prof. Timothy Walsh and Dr Brad Spiller for the E. coli PN47 strain. We thank the National Research Network for Life Sciences and Health (NRN) and MRC-Proximity to Discovery Scheme (MC-PC_17186) for funding. The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication.
Author information
Authors and Affiliations
Contributions
L.C.P. and M.A. are co-first authors. Funding acquisition: D.W.T., M.G., L.C.P., M.A. and K.E.H. Conceived and designed experiments: D.W.T., M.G., L.C.P. and M.A. Performed the experiments: L.C.P., M.A., J.S. and K.E.H. Analysed the data: M.A. and L.C.P. Contributed reagents/materials/analysis tools: Q.E.Y. and T.R.W. Wrote and edited the paper: L.C.P., M.A., K.E.H., M.G. and D.W.T.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Powell, L.C., Abdulkarim, M., Stokniene, J. et al. Quantifying the effects of antibiotic treatment on the extracellular polymer network of antimicrobial resistant and sensitive biofilms using multiple particle tracking. npj Biofilms Microbiomes 7, 13 (2021). https://doi.org/10.1038/s41522-020-00172-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41522-020-00172-6
- Springer Nature Limited
This article is cited by
-
The biofilm matrix: multitasking in a shared space
Nature Reviews Microbiology (2023)
-
Alginate oligosaccharides enhance diffusion and activity of colistin in a mucin-rich environment
Scientific Reports (2022)
-
Supporting the strategic pillars of translational research in biofilms
npj Biofilms and Microbiomes (2022)