Abstract
Biobased furans have emerged as chemical building blocks for the development of materials because of their diverse scaffolds and as they can be directly prepared from sugars. However, selective, efficient, and cost-effective scalable conversion of biobased furans remains elusive. Here, we report a robust transaminase (TA) from Shimia marina (SMTA) that enables the scalable amination of biobased furanaldehydes with high activity and broad substrate specificity. Crystallographic and mutagenesis analyses provide mechanistic insights and a structural basis for understanding SMTA, which enables a higher substrate conversion. The enzymatic cascade process established in this study allows one-pot synthesis of 2,5-bis(aminomethyl)furan (BAMF) and 5-(aminomethyl)furan-2-carboxylic acid from 5-hydroxymethylfurfural. The biosynthesis of various furfurylamines, including a one-pot cascade reaction for BAMF generation using whole cells, demonstrates their practical application in the pharmaceutical and polymer industries.
Similar content being viewed by others
Introduction
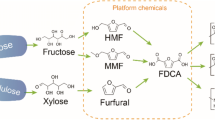
The “greening” of chemical manufacturing and the transition to a carbon-neutral, sustainable economy based on renewable biomass as a raw material represents the major challenges of the 21st century1,2,3. Biomass is one of the best organic carbon sources and is equivalent to petroleum for fuels, biopolymers, and fine commodity chemicals, with net zero carbon emissions4,5. In particular, the biomass-derived 5-hydroxymethylfurfural (HMF) has been called a “sleeping giant” of sustainable chemistry owing to its enormous synthetic potential and high reactivity and can be further transformed into numerous high-valued derivatives. For example, 2,5-furan dicarboxylic acid, an oxidation product of HMF, has been utilized to produce polyethylene furandicarboxylate, which is a 100% renewable alternative to poly(ethylene terephthalate)5.
Furfurylamines are promising precursors of bio-renewable polymers, such as polyamides, polyimides, and polyureas, and have potential applications as intermediates in the manufacturing of pharmaceuticals, such as antiseptic and antihypertensive agents6,7,8. Polymeric Schiff bases made of furans are potential biobased polymers, exhibiting intriguing properties due to their dynamic nature linked to covalent bonds and an improved oxygen, carbon dioxide, and water vapor barrier, conferring better mechanical properties and enhanced thermal stability. These properties confer novel qualities to the corresponding biomaterials9,10,11,12. 2,5-bis(aminomethyl)furan (BAMF), a diamine derivative of furan, is an outstanding monomer for the synthesis of amine-containing polymers, such as polyurethanes and polyamides12,13. Owing to the presence of aldehyde and carboxylic acid groups, 5-(aminomethyl)furan-2-carbaldehyde (AMFC) and 5-(aminomethyl)furan-2-carboxylic acid (MAFCA) can also serve as building blocks for polymer synthesis. Moreover, MAFCA, an amino acid, can be used to produce unnatural peptides, such as cyclopeptides14. The synthesis of furfurylamines via chemocatalytic pathways frequently requires protective groups, harsh reductive agents, and multiple steps, thereby being less economical15,16. Therefore, developing a methodology to produce furfurylamines in large quantities would help broaden their use for polymer and pharmaceutic syntheses. To achieve this, biocatalysts are an efficient option because they tend to direct reactions toward specific products more selectively under mild conditions17,18,19,20. Recent advances in ω-transaminase- (ω-TA)-mediated biocatalytic pathways have attracted widespread attention for their application in furfurylamine biosynthesis5,8,21,22,23. However, their industrial application is hindered by several factors, including equilibrium thermodynamics, product inhibition, poor substrate tolerance, low product recovery, and the excessive use of alanine amino donors, even when using alanine regeneration systems21,22,23,24. Moreover, no methods for the biocatalytic synthesis of BAMF and MAFCA from HMF have yet been developed owing to the lack of an efficient and selective oxidation method to obtain the corresponding aldehydes/acids without the formation of undesired by-products. Therefore, the development of highly efficient and selective biocatalytic systems for these conversions remains challenging.
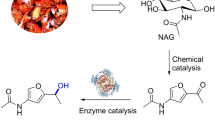
Herein, we report a robust transaminase (TA) derived from Shimia marina (SMTA) that efficiently aminates biobased furan aldehydes with high catalytic activity (Fig. 1a). We also demonstrate a one-pot multi-enzymatic reaction for generating BAMF and MAFCA from HMF, which includes an in situ cofactor-recycling system containing an aldehyde reductase from Synechocystis sp. PCC 6906 (SAHR). We further performed a whole-cell-based biosynthesis of various furan-based amines, including the efficient conversion of HMF to BAMF. This approach, comprising productive biocatalysts and methodologies, constitutes an effective and high-yield system for the production of diverse furfurylamines from biobased furans.
a Generalized synthetic route toward the synthesis of oxidized/reduced/aminated intermediate products from biobased furans. b Whole-cell-based colorimetric TA screening using 2-(4-Nitrophenyl)ethan-1-amine and HMF (R = -CH2OH), FFCA (R = -COOH), and DFF (R = -CHO) as an amino acceptor; screening conditions: described in the supplementary methods section. c Specific activities of SMTA toward HMF and HMF-based derivatives and substituted furfurals. Reaction conditions: 10 mM of amino acceptors, 100 mM of IPA, 0.1 mM of PLP, and 100 mM of Tris-HCl (pH 8.0), 37 °C. Source data is provided as a Source Data file. Data in c are mean values of triplicate experiments with error bars indicating the s. d. (n = 3).
Results
Screening TAs for furan-based aldehyde conversion
To study the conversion of furan-based aldehydes into their corresponding amines, the substrate specificity of an in-house library of whole cells individually expressing 30 TAs was screened using HMF, DFF, and 5-formyl-2-furancarboxylic acid (FFCA); 96-well plates; and 2-(4-nitrophenyl)ethan-1-amine (NPD)25 as the amine donor (Supplementary Fig. 1). Five (S)-amine TAs CVTA, ATTA, VPTA, SMTA, and VFTA (TAs from Chromobacterium violaceum, Agrobacterium tumefaciens, Variovorax paradoxus, Shimia marina, and Vibrio fluvialis respectively) exhibited strong coloration (Fig. 1b). In contrast, the control reaction without an amine acceptor resulted in a faint orange color due to the small amount of remaining intracellular pyruvate, which was different from that of the reaction containing furan-based aldehydes (Supplementary Fig. 1). In particular, CVTA is an efficient catalyst for furfurylamine synthesis8,26. Then, a new set of experiments was designed using selected purified TAs containing various amino donors and higher substrate concentrations (HMF, DFF, and FFCA; 50 and 100 mM) (Supplementary Figs. 2–5). Among the tested TAs, SMTA was the most efficient biocatalyst when isopropylamine (IPA) was used as an amino donor, completely converting the selected furan aldehydes into the corresponding amines (100 mM). IPA is an ideal amino donor because both IPA and its co-product acetone are cost-effective and volatile, enabling an easier scale-up and product isolation27.
The kinetic experiments using SMTA revealed higher catalytic efficiencies, with catalytic constant (kcat) for HMF, FFCA, and furfural of 2.5, 33.3, and 28.1 s−1 respectively, and very high substrate affinities (the KM of HMF, FFCA, and furfural was 0.74, 0.59, and 0.75 mM, respectively). Determining the amount of monoaminated product in the case of DFF was not possible because it was unstable during the synthetic investigations, forming complex polymerization mixtures via imine formation8.
The effect of pH on SMTA activity was determined in different buffer solutions (100 mM) (Supplementary Fig. 6). SMTA was active at pH 6.5–9.5, with optimum activity at pH 8.0 in Tris-HCl buffer. SMTA retained 52% activity when the pH was increased to 9.5. To examine the applicability of SMTA at high substrate loadings, transamination using IPA as the amino donor was investigated (Supplementary Tables 3–5). The reactions yielded an efficient amination of up to 500 mM for HMF and FFCA, and 400 mM for furfural using purified SMTA (Table 1). Although the selectivity towards DFF was far less satisfactory because of the high propensity of diamine products to polymerize with dialdehydes (Supplementary Fig. 10), the selected SMTA led to a BAMF formation of 90% from 50 mM DFF, whereas the reaction with 100 mM DFF resulted in the formation of 65 mM BAMF using 0.5 mg/mL SMTA. Notably, DFF transamination with SMTA resulted in the complete consumption of the substrate (Supplementary Fig. 8). To further enhance the BAMF formation from DFF, we examined the effects of various water-miscible cosolvents, as well as enzyme concentrations (Supplementary Figs. 10–13). A BAMF formation of 87%, a promising result, was observed when using 10% DMSO, 2.0 mg/mL SMTA and 100 mM DFF substrate. The residual activity of SMTA was investigated in the presence of an amine acceptor (FFCA, 100/200 mM) with an excess supply of amine donor (IPA, 1 M) to assess the operational stability. The residual activity of SMTA remained above 90% for 1 h but the activity decreased to 74% and 54% after 6 h in 100 mM and 200 mM of FFCA, respectively (Supplementary Fig. 14).
Then, the substrate scope of SMTA was examined, and DFF, FFCA, and furfural showed higher activity than HMF (706 ± 20 mU/mg) (Fig. 1c). This activity value was higher than that obtained for CVTA (Supplementary Table 6). While the -Br substitution at the 5-position of furfural led to a 1.7-fold higher activity than that of HMF, the substitution of -Br at the 4-position reduced the activity by 0.80-fold. The activity resulting from the -Cl substitution at the 5-position of furfural was 0.84-fold lower than that of HMF. In addition, the methyl, ethyl, and propyl side chain substitutions at the 5-position of furan resulted in activities that were 2.5-, 1.4-, and 2-fold higher than those obtained from HMF.
Crystal structure of SMTA
To elucidate the structural basis for the remarkable activity and stability of SMTA, we determined the crystal structure of this enzyme in the presence of pyridoxal-5′-phosphate (PLP). X-ray diffraction data were collected from SMTA crystals at a resolution of 2.14 Å, and the crystal structure was determined via molecular replacement and refined to a Rwork/Rfree of 0.176/0.216 (Supplementary Table 7). Based on the size exclusion chromatography results, SMTA was assumed to function as a homodimer in solution. One copy of dimeric SMTA was present in an asymmetric unit of the crystal and no notable packing interactions with symmetry-related neighboring molecules were observed. The overall structure of SMTA and its interaction with PLP were highly similar to those of other dimeric TAs28,29 (Fig. 2a). The apparent electron density indicated a Schiff base between PLP and the catalytic lysine K291 (Fig. 2c).
a Overall structure of dimeric SMTA. Protomers and bound PLP molecules are represented as a partially transparent cartoon model (pale yellow and cyan) and as a stick model (green), respectively. The N-terminal residues (aa 2–10) involved in the dimeric interaction are presented in purple and orange for each protomer. b The unique interaction of the N-terminal residues (purple and pale cyan) with the residues of the other protomer (yellow) in the SMTA structure. The hydrogen bond is indicated by a dashed line. c The 2fofc omit map (pink, 1.5 σ contour level) is shown on the internal aldimine of PLP that is covalently linked to Lys291 and hydrogen bond-mediating water molecules, which were omitted during the calculation made for map generation. d Superposition of the PLP binding site for SMTA (yellow) and CVTA (gray, PDB entry 4A6T) shows the details of the interaction of the aspartate residue with a salt bridge with the pyridinium nitrogen of PLP. The hydrogen bonds and an ionic interaction are depicted by dashed lines.
Notably, all residues, except for the N-terminal methionine (Met1) in the refined structure of both protomers of the SMTA dimer, displayed interpretable electron density, considering that the N-terminal regions of dimeric TAs are not conserved and are not visible in other structures, probably because of their conformational flexibility (Supplementary Fig. 15). Notably, the N-terminal region of each SMTA protomer was expected to play a role in the dimer interactions through additional contact with the other protomer (Fig. 2b). As the dimeric form is the functional unit, TA dissociation into monomers can lead to protein unfolding. Mutations in the N-terminal residues reduced the thermal stability of SMTA, determined using differential scanning fluorimetry (Supplementary Fig. 16). The double mutant, I4G and N6D, which can disrupt the hydrophobic interaction and hydrogen bond with the other protomer, lowered the melting temperature (Tm) by 19 °C, probably due to the loss of the interactions (Fig. 2b). Replacing L13 with tryptophan reduced the thermal stability of SMTA by lowering the Tm by 19 °C, probably by weakening the dimeric interaction. In the CVTA structure, the N-terminal region is invisible, implying the absence of any interaction involving this region. The N-terminal region of CVTA could not bend toward the dimeric interface owing to the steric hindrance by the large size of the corresponding residue W10 (Supplementary Fig. 17). Since the N-terminal region, including L13 of SMTA, is not conserved among transaminases, it is likely related to the different stability and activity of each transaminase, albeit in a minor way (Supplementary Fig. 15). The temperature dependency of the SMTA activity in the wild-type and double mutant, I4G and N6D, was investigated after incubation at different temperatures (25–70 °C) for 1 hr (Supplementary Fig. 18a). SMTA maintained >97% residual activity till 37 °C for 1 hr. Notably, incubation of SMTA with 0.1 mM PLP resulted in the retainment of >98% residual activity until 50 °C for 1 hr. However, the N-terminal double mutant I4G and N6D resulted in a significant loss of residual activity (54% at 37 °C). Even in the presence of 0.1 mM PLP, the I4G and N6D double mutant exhibited maximal residual activity of 61% at 37 °C and 12% at 50 °C. We investigated the residual activity by prolonging the incubation of enzymes at 50 °C (Supplementary Fig. 18b). SMTA retained substantial residual activity (51%) after 3 hr, while the I4G and N6D double mutant showed a complete loss of residual activity after 3 hr of incubation at 50 °C.
In SMTA, the pyridinium nitrogen in PLP forms an ionic interaction with the side chain of D262, which is conserved in other TAs. The conformation of D262 in SMTA is supported by a hydrogen bond network involving N127 and N130 (Fig. 2d). However, the CVTA residues at the equivalent positions, V124 and M127, cannot form such an interaction network. The cofactor release assay showed that the N127V mutant significantly decreased the binding affinity of both PLP and PMP compared to that of wild-type SMTA (Supplementary Fig. 19). The N130M mutant could not be evaluated because it was expressed in a completely insoluble form in E. coli. This mutation may hinder protein folding through a putative clash of the mutated methionine with surrounding residues of SMTA, which are not conserved in CVTA. The hydrogen-bond network mediated by N127 and N130 in SMTA may facilitate the ionic interaction between D262 and its cofactor by optimally positioning D262, and thereby preventing the release of PLP and PMP during the catalytic cycle. Other transaminases, including VFTA (Vibrio fluvialis TA), HETA (Halomonas elongate TA), and PPTA (Pseudomonas putida TA), contain an asparagine residue at N127 in SMTA, whereas N130 of SMTA shows little conservation among transaminases (Supplementary Fig. 15). Thus, this hydrogen-bond network mediated by N130, N127, and D262 could be a unique characteristic of SMTA, which provides a preformed structure for optimal interaction with the cofactors. Cofactor loss in dimeric TAs induces a significant structural rearrangement within the dimer interface, thereby promoting dimer dissociation and the unfolding of the enzyme into an irreversibly inactivated state30,31,32. Thus, the high activity and robustness of SMTA, even in the presence of high substrate concentration, can be attributed in part to its unique structural features, including the role of the N-terminal region in reinforcing the dimer interaction and the ability of the hydrogen-bond network to prevent cofactor leakage from the enzyme.
Molecular basis for the catalytic activity of SMTA
The docking structure of SMTA with the external aldimine intermediate of PLP-HMF was modeled using its crystal structure to provide molecular insights into the high activity and broad substrate specificity of SMTA. In the hypothetical structure, the furan ring moiety of PLP-HMF is surrounded by hydrophobic residues from the two protomers of the SMTA dimer via van der Waals interactions (Fig. 3). Furthermore, the oxygen atom in the furan ring forms a hydrogen bond with the side chain of W63. The high activity of SMTA toward furfural may be attributed to these interactions, which enable an efficient binding to the furan ring of the substrates.
a Hypothetical model of the docking interaction of the external aldimine intermediate of PLP-HMF (green) with the residues (yellow and blue from each protomer) in SMTA. The hydrogen bonds and ionic interaction are indicated by dashed lines. Water molecules involved in the interaction are excluded from the figure for clarity. b Detailed interactions around the furan ring moiety. The van der Waals interaction by the residues of SMTA (yellow and cyan for each protomer) surrounding the furan ring moiety is represented as a partially transparent surface (black) and the hydrogen bond is represented as a dashed line. c Sliced surfaces through the active site of SMTA (left) and CVTA (right). PLP-HMF (green) docked to SMTA is overlaid onto the active site of CVTA for comparison. The arrow indicates putative steric collision of the furan ring moiety in the CVTA active site.
The active site of SMTA contains sufficient space to accommodate various furan derivatives, particularly those substituted at the 4 and 5 positions (Fig. 3c). When PLP-HMF was overlaid on the active site of CVTA while maintaining the interactions with the PLP moiety, the furan ring sterically collided with the CVTA surface. This collision was primarily due to the switch-in conformation of R416 in CVTA. TAs often adopt an arginine switch strategy for dual specificity, accepting both negatively charged and neutral substrates at the same site33,34. Carboxylated substrates form bidentate salt bridges with the arginine residue in a canonical switch-in conformation, whereas large aromatic substrates can be accommodated by moving the arginine residue from the active site. In the structure of CVTA in complex with PLP, R416 assumed a switch-in conformation, which was stabilized by a water-mediated hydrogen bond to Y168 (Supplementary Fig. 20). In the SMTA structure, the switch-out conformation of R418 in both protomers was stabilized by stacking interactions with R407 and F92. The optimal conformation of R407 for the stacking interaction is supported by its ionic interaction with E408, whereas the equivalent residue in CVTA, D406, tilts the R405 conformation. Moreover, the I425 residue may sterically interfere with the putative switch-in conformation of R418 in SMTA, since I425 has an extra δ-carbon compared to the V423 residue of CVTA. This switch-in conformation of CVTA would not be readily compatible with the binding of the PLP-HMF intermediate owing to this steric occlusion (Fig. 3c; Supplementary Figs. 20, 21). A significant catalytic activity reduction was observed in the SMTA mutants whose residues were altered to CVTA, which destabilized and stabilized the switch-out and switch-in conformation of R418, respectively. This suggests that the preference of SMTA for the switch-out arginine conformation may contribute to the high activity and broad specificity of the enzyme for various furfural types as amino acceptors (Supplementary Fig. 22). The switch-out conformation of SMTA creates a wide substrate binding site, allowing ready accessibility to both amino donors, alanine and IPA. However, the switch-in conformation in CVTA may facilitate the ideal positioning of alanine’s carboxylic group, thereby establishing alanine as a more favorable amino donor than IPA (Supplementary Fig. 4). As expected, the experiment to assess the initial reaction rate demonstrated that the relative preference for alanine over IPA is more significant in CVTA (3.4-fold higher) than in SMTA (1.5-fold higher) (Supplementary Fig. 23).
Investigation of CVTA structures revealed that R416 of CVTA adopted diverse conformations depending on the enzyme’s state, such as the apo, PLP-bound, PMP-bound, and mutant forms (Supplementary Fig. 24). Most CVTA structures feature R416 in switch-in conformations, except for the W60C mutant. In the W60C mutant, one protomer of the dimeric enzyme allows Arg416 to adopt a complete switch-out conformation similar to SMTA, while the other protomer maintains a switch-in conformation. The mutation, W60C, boosts CVTA’s specific activity for some substrates, suggesting its effect on electrochemical properties in addition to making more space in the active site35,36. W60C may contribute to the higher activity of the enzyme by increasing the population of the enzyme with a switch-out conformation of R416. However, the mechanism through which this mutation induces the switch-out conformation in the enzyme remains unclear. Subtle variations in transamidases’ active sites might affect the conformational equilibrium of their switchable arginine residue, which in turn could influence their catalytic properties.
Cascade catalysis for selective BAMF and MAFCA generation from HMF
In principle, BAMF and MAFCA can be enzymatically synthesized by the amination of HMF-oxidized DFF and FFCA, the chemical or biocatalytic synthesis of which is challenging37,38. Recently, Turner et al. engineered a galactose oxidase enzyme (Variant GOM7-2A) that can be used in organic solvent/buffer biphasic systems at a high substrate concentration for the oxidation of HMF to DFF37. To catalyze the initial step of this oxidation, the reaction was tested at different HMF concentrations up to 500 mM using DMSO as a cosolvent and a biocatalytic system comprising GOM7-2A, horseradish peroxidase (HRP), and catalase (Supplementary Table 8). The complete conversion of HMF to DFF was achieved up to a substrate concentration of 200 mM but decreased to 90%, 76%, and 70% as the substrate concentration of 300, 400, and 500 mM, respectively. To catalyze the conversion of HMF to BAMF, we performed a one-pot reaction employing SMTA using a developed biocatalytic system (GOM7-2A, HRP, and catalase). However, an equimolar conversion of the HMF substrate (50 mM) to 5-hydroxymethylfurfurylamine (HMFA) (48%) and BAMF (52%) was observed after 18 h, and the BAMF concentration did not increase further until 48 h (Supplementary Fig. 26). This equimolar conversion can be attributed to the multifaceted substrate HMF, which serves as a common substrate for both SMTA and GOM7-2A. Furthermore, the accumulated HMFA could not be oxidized by GOM7-2A, indicating the lack of specificity of GOM7-2A toward HMFA. Therefore, the selective conversion of HMF into BAMF was achieved in two sequential one-pot reaction steps. In the first step, the HMF substrate was completely oxidized to DFF using a GOM7-2A, HRP, and catalase system, followed by the SMTA-mediated transamination of the generated DFF into BAMF. Notably, the complete conversion of HMF to DFF is important for the selective synthesis of BAMF since separating the remnant amination byproduct of HMF (i.e., HMFA) from BAMF is challenging. After the completion of the first step, the reaction was diluted to achieve a final substrate (DFF) concentration of 50 mM. This one-pot, two-step reaction using 100 to 200 mM of HMF led to a production of the intermediate DFF > 99%, which was subsequently aminated to produce 87% of BAMF in the second step (Fig. 4a; Table 2). However, increasing the initial substrate concentration of HMF to 300 mM resulted in a 76% conversion to BAMF with a 10% formation of HMFA as a byproduct.
a, Time course measuring the one-pot enzyme cascade reaction for the conversion of HMF to BAMF. S1 oxidation conditions: 100 mM HMF, 0.05 mM CuSO4, 0.625 mg/mL purified GOM7-2A, 0.0128 mg/mL HRP, 880 U/mL catalase, 100 mM NaPi (pH 7.4), 10% DMSO, 20 °C. S2 amination conditions: the S1 oxidation reaction volume was increased to a final substrate and IPA concentration of 50 and 500 mM, respectively; 0.5 mg/mL purified SMTA, 0.5 mM PLP, and 100 mM of NaPi (pH 7.4), 37 °C. b, Time course measuring the one-pot enzyme cascade reaction for the conversion of HMF to MAFCA. S1 oxidation conditions: 100 mM HMF, 0.05 mM CuSO4, 0.625 mg/mL purified GOM7-2A, 0.0128 mg/mL HRP, 880 U/mL catalase, 0.5 mg/mL purified SAHR, 0.1 mM NAD(P)+, 200 mM NaPi (pH 7.4), 25 °C. S2 amination conditions: the S1 oxidation reaction volume was increased to a final substrate and IPA concentration of 50 and 500 mM, respectively; 0.5 mg/mL purified SMTA; 0.5 mM PLP; 100 mM NaPi (pH 7.4), 37 °C. c, Bar chart comparing the study of SMTA-overexpressed in BL-21 and RARE cells for the transamination of an increasing concentration of HMF. Reaction conditions: 100–500 mM HMF, 1 M of IPA for 100–300 mM of HMF, 1.5 M of IPA for 400–500 mM of HMF, 9 mgCDW/mL SMTA-overexpressed in BL-21 and RARE cells, 100 mM Tris-HCl (pH 8.0), 37 °C, 18 h. d, Time course measuring the transamination of HMF to HMFA by SMTA-overexpressed in BL-21 and RARE cells. Reaction conditions: 200 mM HMF, 1 M IPA, 9 mgCDW/mL SMTA overexpressed in BL-21 and RARE cells, 100 mM Tris-HCl (pH 8.0), 37 °C, 6 h. e, One-pot enzyme cascade reaction for the conversion of HMF to BAMF. S1 oxidation conditions: 100 mM HMF, GOM7-2A, HRP co-overexpressed RARE cells, 18 mgCDW/mL, 0.2 mM CuSO4, 10% DMSO, 100 mM NaPi (pH 7.4), 20 °C; S2 amination conditions: the S1 oxidation reaction was diluted to a final substrate and IPA concentration of 50 and 500 mM, respectively; 9 mgCDW/mL SMTA, 10% DMSO, and 100 mM NaPi (pH 7.4), 37 °C. Source data are provided as a Source Data file. Data in a–e are mean values of triplicate experiments with error bars indicating the s. d. (n = 3).
The synthesis of MAFCA from HMF was achieved by generating FFCA, a partially oxidized derivative of HMF containing a carboxylic acid group and a formyl group5 (Fig. 1a). The synthesis of FFCA is challenging because of its highly oxidized yet incomplete state. However, biocatalysts capable of producing FFCA are scarce. Therefore, numerous biocatalysts were screened for the conversion of the HMF substrate to the desired FFCA (Supplementary Table 9). The screened enzymes exhibited a substrate scope toward aliphatic and aromatic alcohols. Most enzymes showed activity toward HMF to produce 5-hydroxymethyl-2-furoic acid (HMFCA); however, none could directly produce FFCA from HMF. Alternatively, FFCA could be obtained by oxidizing the HMFCA intermediate. The GOM7-2A variant showed excellent activity in generating FFCA, leading to a conversion of >99% using 50 and 100 mM of the HMFCA substrate (Supplementary Table 10). However, the multi-enzymatic FFCA generation from HMF is technically challenging by utilizing two oxidases, especially at high substrate concentrations due to the lower availability of O2 in the reaction solutions, especially at high temperatures (0.26 mM at 25 °C)38,39. Unlike oxidases, DHs typically use nicotinamide cofactors instead of O2 to oxidize substrates. Hemoproteins/oxidases can regenerate NAD(P)+ for DH-catalyzed oxidations38,40,41. The combination of the DH from Synechocystis sp. (SADH) with an oxidase can be used to produce FFCA from HMF38. To identify efficient DHs, we tested the aldehyde reductase SAHR (Supplementary Figs. 28 and 29). The SADH-combined GOM7-2A-HRP-catalase system produced 43 mM of the desired FFCA and 7 mM of the DFF intermediate from 50 mM of HMF within 24 h. However, SAHR combined with GOM7-2A-HRP-catalase resulted in the complete formation of the desired FFCA within 6 h. The applicability of this cascade reaction was further evaluated at a larger scale, using 100 mM of HMF, for which 58 mM of FFCA was obtained and 9 and 31 mM of the HMFCA and DFF intermediates, respectively, were formed. The selectivity of SAHR toward the HMF and DFF substrates was tested (Supplementary Tables 11 and 12). The SAHR-based reaction with DFF yielded 20% FFCA, 10% HMFCA, and 19% HMF. Notably, no overoxidation of FFCA was observed, indicating that the GOM7-2A/SAHR system-based oxidation of HMF could be controlled to obtain the desired amount of FFCA. To drive the reaction equilibrium by efficiently regenerating NAD(P)+, the HMF to FFCA reaction was performed by combining the NADPH oxidase from Lactobacillus reuteri (LreNOX)42,43,44 with the SAHR-GOM7-2A-HRP-catalase system. The reaction using 100 mM of HMF led to a 99% conversion of HMF: mostly into the desired FFCA product (63%), but also into the DFF intermediate (39%) (Supplementary Table 13). Because the formation of FFCA resulted in a drop in the pH of the reaction mixture down to a pH of 6.5, the effect of increasing the buffering capacity of NaPi from 50 to 500 mM was evaluated (Supplementary Table 14). Increasing the buffer capacity to 200 mM drastically improved the conversion toward the desired product, resulting in the complete conversion of HMF to FFCA. Notably, the reaction without NOX proceeded well and resulted in the complete conversion of HMF to FFCA within 6 h (Table 2). Even at an HMF concentration of 300 mM, the tandem oxidation yielded up to 73% of FFCA. To catalyze the conversion of HMF to MAFCA, we performed a one-pot reaction by combining SMTA with a newly developed oxidation system (GOM7-2A, HRP, catalase, and SAHR). HMF (50 mM) yielded 40% of HMFA and 60% of BAMF (Supplementary Table 15), demonstrating that the GOM7-2A and SAHR variants lack specificity toward HMFA oxidation. A sequential one-pot, two-step reaction was performed (Fig. 4b). In the first step, the oxidation of HMF to FFCA was completed before the second transamination step. This stepwise reaction produced MAFCA from 200 mM of HMF, yielding a conversion >99% (Table 2).
Whole-cell-based generation of furfurylamines
Whole-cell biotransformations are advantageous because of their low downstream requirements and better enzyme stability45. To minimize the rapid endogenous conversion of aldehydes into their corresponding alcohols, we tested an engineered E. coli strain with reduced aromatic aldehyde reduction (RARE)46. The whole-cell-based transamination reactions of HMF and furfural (at a 100–500 mM concentration) using the BL-21 and RARE strains were compared. The amination conversion capacity was significantly higher in SMTA-overexpressed RARE than in BL-21 cells. HMF (400 mM) (Fig. 4c) and furfural (300 mM) (Supplementary Fig. 33) were successfully aminated using RARE cells (9 mgCDW/mL) expressing SMTA and IPA amino donors. The enhanced biotransamination obtained in RARE cells (Fig. 4d) was likely associated with the minimized futile conversion of aldehyde to alcohol47. The RARE strain overexpressing SMTA was selected to transform FFCA and DFF. Using FFCA as a substrate (400 mM) led to a conversion into MAFCA > 99%, whereas using 100 mM of DFF led to a 95% conversion into BAMF (Table 16; Supplementary Figs. 35 and 36). The reduced mass transfer of AMFC/BAMF within whole cells led to a reduced interaction with DFF, thereby leading to a higher formation of BAMF than of the purified enzyme.
We then performed further experiments on the oxidation of HMF to DFF in whole cells. Initially, GOM7-2A was overexpressed in RARE cells used to oxidize HMF at 50 and 100 mM without the addition of catalase or HRP (Supplementary Table 17). The reaction using 50 mM of HMF led to the conversion into DFF without the formation of overoxidation/reduction side products. Whereas the reaction with 100 mM of HMF led to the formation of 64 mM of DFF with 34 mM of remaining HMF and >1 mM of overoxidized FFCA. This satisfactory conversion may also be attributed to the presence of inherent catalases in E. coli48. HRP reactivates GO, likely by utilizing the H2O2 formed during the reaction and enhancing the GO activity to shorten the conversion time in a biocatalytic reaction49. Unfortunately, the active expression of HRP in microbial cells is complex50. Sushma et al. effectively produced active HRP fused with E. coli phosphoglycerate kinase (PGK) and co-expressed it with ferrochelatase (FC) in E. coli cells51. To establish RARE cells expressing functional GO, the pET24ma plasmid encoding GOM7-2A and the pETduet-1 plasmid encoding PGK-HRP in MCS-1 cells and FC in MCS-2 cells were constructed, transformed, and overexpressed (Supplementary Fig. 37). RARE whole cells co-expressing functional GOM7-2A with HRP converted 100 mM of HMF into 74 mM of DFF. The use of DMSO 10% (v/v) increased the conversion to 82% (Supplementary Fig. 38). Because the complete saturation of GOM7-2A with Cu2+ is critical for the activity of GO49, the addition of 0.2 mM of Cu2+ improved the DFF formation by up to 93% (Supplementary Table 19; Supplementary Fig. 39). The sequestration of Cu2+ by cell proteins and the NaPi buffer is a more likely explanation for the enhanced product formation regarding the optimal CuSO4 concentration. Nevertheless, increasing the HMF concentration to 200 mM led to the formation of 60% DFF and <1% overoxidized FFCA as a side product.
Subsequently, the reaction was combined with transamination to produce the desired BAMF. This one-pot two-step reaction using 50 and 100 mM of HMF produced 87% and 85% (Fig. 4e) of the desired BAMF, respectively. Finally, to demonstrate the practical utility of the whole-cell-based multi-enzymatic process for generating BAMF, we carried out a one-pot preparative-scale reaction (100 mL) using 100 mM (1.26 gm) of HMF, which led to the formation of 92% of DFF. The SMTA-catalyzed transamination of the DFF intermediate resulted in the formation of 85% of the final BAMF product.
Discussion
Furan-based amines are potential biobased monomers that are utilized in the production of several promising biopolymers and intermediates in the pharmaceutical and food industries6,7,8. Owing to the challenges associated with the available biocatalysts and the complexity of controlling the oxidation status, efficient and scalable methods for the production of diverse furfurylamines remain rare. To the best of our knowledge, the finding of SMTA is useful for the scalable amination of various biobased furan aldehydes at higher concentrations using IPA, which is an affordable amino donor with excellent catalytic activity and broad substrate scope toward substituted furfurals.
The crystal structure and molecular basis for the higher catalytic activity of SMTA offer an important precedent that can be applied for further engineering of TAs whose practical application is challenging. The elucidated 3D structure of SMTA reveals unique characteristics, such as the role of the N-terminal region in dimerization and the ability of the hydrogen-bond network to prevent cofactor leaks, which leads to better SMTA operational stability. In addition, the switch-out conformation of R418 at the active site contributes to the high activity and broad specificity of SMTA toward various furfurals; these characteristics can be harnessed to devise new TAs with superior functionality.
In addition, we developed a one-pot enzymatic cascade reaction for generating BAMF and MAFCA from HMF. The developed oxidative multi-enzymatic cascade reaction with a novel SAHR combined with GOM7-2A, HRP, and catalase, based on an in situ NAD(P)+ recycling system, provides complete and controlled access for FFCA formation at higher HMF concentrations. The demonstrated whole-cell method for creating diverse furfural amines using a one-pot oxidative cascade reaction by co-expressing active HRP and GOM7-2A in the RARE strain efficiently synthesized DFF from HMF and subsequently produced BAMF via amination, which further demonstrates its practical applications. Collectively, these results highlight the potential to tackle challenging problems associated with the broader utilization of biobased furans in polymer and pharmaceutical syntheses.
Methods
One hundred milliliter scale cascade biotransformation
GOM7-2A and PGK-HRP with FC were co-expressed in the RARE strain and harvested using NaPi buffer (pH 7.4, 100 mM). The reaction components included co-overexpressed GOM7-2A and PGK-HRP with FC (18 mgCDW/mL) RARE cells, HMF (100 mM), CuSO4 (0.2 mM), DMSO (10%), and NaPi (pH 7.4, 100 mM). The total volume of the reaction was 100 mL. The reaction was performed in a reactor with sufficient headspace. After adding HMF, the reaction was initiated at 20 °C and 350 rpm. Oxygen in the headspace of the reaction vessel was refreshed by opening it every 4 hr. After the maximum conversion of the reaction was reached (monitored using HPLC, as described in the SI), the reaction was diluted by half by adding SMTA- overexpressed RARE cells (9 mgCDW/mL), IPA (1 M), and NaPi buffer (pH 7.4, 100 mM). After completing the reaction (monitored using HPLC), the reaction was quenched using an amount of methanol that was twofold in excess, and the denatured protein was removed via centrifugation (4416 × g at 4 °C for 30 min). The supernatant, which contained the organic layer, was removed under reduced pressure. The isolation process is described in the Supplementary Methods.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The atomic coordinates and structural factors of the crystal structure of SMTA in complex with PLP were deposited in the Protein Data Bank (www.rcsb.org) under the accession number 8WQJ. The data generated in this study are provided within the paper and its Supplementary Information. Source data are provided in this paper.
References
Bozell, J. J. & Petersen, G. R. Technology development for the production of biobased products from biorefinery carbohydrates - The US Department of Energy’s “Top 10” revisited. Green. Chem. 12, 539–554 (2010).
Sheldon, R. A. Biocatalysis and biomass conversion: enabling a circular economy. Philos. Trans. R. Soc. A. 378, 20190274 (2020).
Ragauskas, A. J. et al. The path forward for biofuels and biomaterials. Science 311, 484–489 (2006).
Isikgor, F. H. & Becer, C. R. Lignocellulosic biomass: a sustainable platform for the production of bio-based chemicals and polymers. Polym. Chem. 6, 4497–4559 (2015).
Li, N. & Zong, M. H. (Chemo) biocatalytic upgrading of biobased furanic platforms to chemicals, fuels, and materials: a comprehensive review. ACS Catal. 12, 10080–10114 (2022).
Froidevaux, V., Negrell, C., Caillol, S., Pascault, J. P. & Boutevin, B. Biobased amines: from synthesis to polymers; present and future. Chem. Rev. 116, 14181–14224 (2016).
Chakraborty, T. K., Tapadar, S. & Kumar, S. K. Cyclic trimer of 5-(aminomethyl)-2-furancarboxylic acid as a novel synthetic receptor for carboxylate recognition. Tetrahedron Lett. 43, 1317–1320 (2002).
Dunbabin, A., Subrizi, F., Ward, J. M., Sheppard, T. D. & Hailes, H. C. Furfurylamines from biomass: transaminase catalysed upgrading of furfurals. Green. Chem. 19, 397–404 (2017).
Xiang, T. et al. Schiff base polymers derived from 2, 5‐diformylfuran. Polym. Int. 62, 1517–1523 (2013).
Tachibana, Y., Hayashi, S. & Kasuya, K. I. Biobased poly (schiff-base) composed of bifurfural. ACS Omega 3, 5336–5345 (2018).
Dhers, S., Vantomme, G. & Avérous, L. A fully bio-based polyimine vitrimer derived from fructose. Green. Chem. 21, 1596–1601 (2019).
Wei, Z. et al. One‐step reductive amination of 5‐hydroxymethylfurfural into 2, 5‐bis (aminomethyl) furan over Raney Ni. ChemSusChem 14, 2308–2312 (2021).
Delidovich, I. et al. Alternative monomers based on lignocellulose and their use for polymer production. Chem. Rev. 116, 1540–1599 (2016).
Lancien, A. et al. Hybrid conversion of 5‐hydroxymethylfurfural to 5‐aminomethyl‐2‐furancarboxylic acid: toward new bio‐sourced polymers. ChemCatChem 13, 247–259 (2021).
Chatterjee, M., Ishizaka, T. & Kawanami, H. Reductive amination of furfural to furfurylamine using aqueous ammonia solution and molecular hydrogen: an environmentally friendly approach. Green. Chem. 18, 487–496 (2016).
Chieffi, G., Braun, M. & Esposito, D. Continuous reductive amination of biomass‐derived molecules over carbonized filter paper‐supported FeNi alloy. ChemSusChem 8, 3590–3594 (2015).
Arnold, F. H. Directed evolution: bringing new chemistry to life. Angew. Chem. Int. Ed. 57, 4143–4148 (2018).
Winkler, C. K., Schrittwieser, J. H. & Kroutil, W. Power of biocatalysis for organic synthesis. ACS Cent. Sci. 7, 55–71 (2021).
Benítez-Mateos, A. I., Roura Padrosa, D. & Paradisi, F. Multistep enzyme cascades as a route towards green and sustainable pharmaceutical syntheses. Nat. Chem. 14, 489–499 (2022).
Wu, S., Snajdrova, R., Moore, J. C., Baldenius, K. & Bornscheuer, U. T. Biocatalysis: enzymatic synthesis for industrial applications. Angew. Chem. Int. Ed. 60, 88–119 (2021).
Koszelewski, D., Tauber, K., Faber, K. & Kroutil, W. ω-Transaminases for the synthesis of non-racemic α-chiral primary amines. Trends Biotechnol. 28, 324–332 (2010).
Slabu, I., Galman, J. L., Lloyd, R. C. & Turner, N. J. Discovery, engineering, and synthetic application of transaminase biocatalysts. ACS Catal. 7, 8263–8284 (2017).
Patil, M. D., Grogan, G., Bommarius, A. & Yun, H. Oxidoreductase-catalyzed synthesis of chiral amines. ACS Catal. 8, 10985–11015 (2018).
Khobragade, T. P. et al. Promoter engineering‐mediated Tuning of esterase and transaminase expression for the chemoenzymatic synthesis of sitagliptin phosphate at the kilogram‐scale. Biotechnol. Bioeng. 118, 3263–3268 (2021).
Baud, D., Ladkau, N., Moody, T. S., Ward, J. M. & Hailes, H. C. A rapid, sensitive colorimetric assay for the high-throughput screening of transaminases in liquid or solid-phase. Chem. Commun. 51, 17225–17228 (2015).
Kaulmann, U., Smithies, K., Smith, M. E., Hailes, H. C. & Ward, J. M. Substrate spectrum of ω-transaminase from Chromobacterium violaceum DSM30191 and its potential for biocatalysis. Enzym. Microb. Technol. 41, 628–637 (2007).
Kelefiotis-Stratidakis, P., Tyrikos-Ergas, T. & Pavlidis, I. V. The challenge of using isopropylamine as an amine donor in transaminase catalysed reactions. Org. Biomol. Chem. 17, 1634–1642 (2019).
Humble, M. S. et al. Crystal structures of the Chromobacterium violaceum ω‐transaminase reveal major structural rearrangements upon binding of coenzyme PLP. FEBS J. 279, 779–792 (2012).
Planchestainer, M., Hegarty, E., Heckmann, C. M., Gourlay, L. J. & Paradisi, F. Widely applicable background depletion step enables transaminase evolution through solid-phase screening. Chem. Sci. 10, 5952–5958 (2019).
Roura Padrosa, D. et al. Enhancing PLP-binding capacity of class-III ω-transaminase by single residue substitution. Front. Bioeng. Biotechnol. 7, 282 (2019).
Börner, T. et al. Explaining operational instability of amine transaminases: substrate-induced inactivation mechanism and influence of quaternary structure on enzyme–cofactor intermediate stability. ACS Catal. 7, 1259–1269 (2017).
Ruggieri, F. et al. Insight into the dimer dissociation process of the Chromobacterium violaceum (S)-selective amine transaminase. Sci. Rep. 9, 16946 (2019).
Eliot, A. C. & Kirsch, J. F. Pyridoxal phosphate enzymes: mechanistic, structural, and evolutionary considerations. Annu. Rev. Biochem. 73, 383–415 (2004).
Malashkevich, V. N., Onuffer, J. J., Kirsch, J. F. & Jansonius, J. N. Alternating arginine-modulated substrate specificity in an engineered tyrosine aminotransferase. Nat. Struct. Mol. Biol. 2, 548–553 (1995).
Cassimjee, K. E., Humble, M. S., Land, H., Abedi, V. & Berglund, P. Chromobacterium violaceum ω-transaminase variant Trp60Cys shows increased specificity for (S)-1-phenylethylamine and 4′-substituted acetophenones, and follows Swain–Lupton parameterisation. Org. Biomol. Chem. 10, 5466–5470 (2012).
Humble, M. S., Cassimjee, K. E., Abedi, V., Federsel, H. J. & Berglund, P. Key amino acid residues for reversed or improved enantiospecificity of an ω‐transaminase. ChemCatChem 4, 1167–1172 (2012).
Birmingham, W. R. et al. Toward scalable biocatalytic conversion of 5-hydroxymethylfurfural by galactose oxidase using coordinated reaction and enzyme engineering. Nat. Commun. 12, 4946 (2021).
Jia, H. Y., Zong, M. H., Zheng, G. W. & Li, N. One‐pot enzyme cascade for controlled synthesis of furancarboxylic acids from 5‐hydroxymethylfurfural by H2O2 internal recycling. ChemSusChem 12, 4764–4768 (2019).
Al-Shameri, A., Siebert, D. L., Sutiono, S., Lauterbach, L. & Sieber, V. Hydrogenase-based oxidative biocatalysis without oxygen. Nat. Commun. 14, 2693 (2023).
Puetz, H., Puchľová, E., Vranková, K. & Hollmann, F. Biocatalytic oxidation of alcohols. Catalysts 10, 952 (2020).
Golub, E., Freeman, R. & Willner, I. A hemin/G‐quadruplex acts as an NADH oxidase and NADH peroxidase mimicking DNAzyme. Angew. Chem. Int. Ed. 123, 11914–11918 (2011).
Rehn, G., Pedersen, A. T. & Woodley, J. M. Application of NAD (P) H oxidase for cofactor regeneration in dehydrogenase catalyzed oxidations. J. Mol. Catal. B Enzym. 134, 331–339 (2016).
Aalbers, F. S. & Fraaije, M. W. Design of artificial alcohol oxidases: alcohol dehydrogenase–NADPH oxidase fusions for continuous oxidations. ChemBioChem 20, 51–56 (2019).
Lim, S., Yoo, H. W., Sarak, S., Kim, B. G. & Yun, H. A multi-enzyme cascade reaction for the production of α, ω-dicarboxylic acids from free fatty acids. J. Ind. Eng. Chem. 98, 358–365 (2021).
de Carvalho, C. C. Whole cell biocatalysts: essential workers from nature to the industry. Microb. Biotechnol. 10, 250–263 (2017).
Kunjapur, A. M., Tarasova, Y. & Prather, K. L. Synthesis and accumulation of aromatic aldehydes in an engineered strain of Escherichia Coli. J. Am. Chem. Soc. 136, 11644–11654 (2014).
Kunjapur, A. M. & Prather, K. L. Microbial engineering for aldehyde synthesis. Appl. Environ. Microbiol. 81, 1892–1901 (2015).
Tan, H., Zhou, F., Liao, D., Ouyang, J. & Zheng, Z. Improved biosynthesis of 2, 5-Furandicarboxylic acid through coupling of heterologous pathways in Escherichia coli and native pathways in Pseudomonas putida. Biochem. Eng. J. 161, 107657 (2020).
Toftgaard Pedersen, A. et al. Process requirements of galactose oxidase catalyzed oxidation of alcohols. Org. Process Res. Dev. 19, 1580–1589 (2015).
Morawski, B. et al. Functional expression of horseradish peroxidase in Saccharomyces cerevisiae and Pichia pastoris. Protein Eng. 13, 377–384 (2000).
Chauhan, S. & Kang, T. J. Soluble expression of horseradish peroxidase in Escherichia coli and its facile activation. J. Biosci. Bioeng. 126, 431–435 (2018).
Acknowledgements
This study was supported by the Basic Science Research Program (RS-2023-00208582, NRF-2023R1A2C1006874), Bio & Medical Technology Development Program (NRF-2022M3A9B6082669 and NRF-2022M3A9I3082366), and the Enzyme Engineering for Next-Generation Biorefineries (NRF-2022M3J5A1056169 and NRF-2022M3J5A1085235) through the National Research Foundation of Korea (NRF) funded by the Korean government. We appreciate the assistance of the staff of Beamline 5 C at the Pohang Accelerator Laboratory with the X-ray diffraction experiments. We are thankful to Professor Kristina L. Jones Prather at MIT for providing the RARE strain for whole-cell studies.
Author information
Authors and Affiliations
Contributions
P.G. designed and performed experiments, data analysis and characterization, and data collection, and wrote the manuscript. S.L. designed and performed experiments, data analysis, and collection, and edited the draft. T.P.K., A.D.P., M.D.P., S.S., and H.J. analyzed the data and edited the draft. S. Joo, Y.G., and S. Jung contributed to the purification of the biocatalysts. Y.J.J. and S.B.C. collected data from crystallization and X-ray diffraction studies. Y.C.K. performed differential scanning fluorimetry experiments. T.J.K. contributed to the cloning and optimization of the HRP gene. Y.-S.H. determined the crystal structure, performed docking modeling, and wrote the manuscript. H.Y. supervised the project, conceptualization, investigation, designed experiments, and visualization, writing, review, and finalized the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ioannis Pavlidis, Andy-Mark Thunnissen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Giri, P., Lim, S., Khobragade, T.P. et al. Biocatalysis enables the scalable conversion of biobased furans into various furfurylamines. Nat Commun 15, 6371 (2024). https://doi.org/10.1038/s41467-024-50637-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-50637-x
- Springer Nature Limited