Abstract
Cryptococcus spp. are environmental fungi that first must adapt to the host environment before they can cause life-threatening meningitis in immunocompromised patients. Host CO2 concentrations are 100-fold higher than the external environment and strains unable to grow at host CO2 concentrations are not pathogenic. Using a genetic screening and transcriptional profiling approach, we report that the TOR pathway is critical for C. neoformans adaptation to host CO2 partly through Ypk1-dependent remodeling of phosphatidylserine asymmetry at the plasma membrane. We also describe a C. neoformans ABC/PDR transporter (PDR9) that is highly expressed in CO2-sensitive environmental strains, suppresses CO2-induced phosphatidylserine/phospholipid remodeling, and increases susceptibility to host concentrations of CO2. Interestingly, regulation of plasma membrane lipid asymmetry by the TOR-Ypk1 axis is distinct in C. neoformans compared to S. cerevisiae. Finally, host CO2 concentrations suppress the C. neoformans pathways that respond to host temperature (Mpk1) and pH (Rim101), indicating that host adaptation requires a stringent balance among distinct stress responses.
Similar content being viewed by others
Introduction
Cryptococcus spp. cause over one hundred thousand deaths per year across the globe with the majority occurring in those living with HIV1. Cryptococcus spp. are environmental basidiomycetes that occupy a variety of niches and have broad genetic diversity2. However, only a minority of these species cause human disease3. Accordingly, the mechanisms and traits that distinguish pathogenic and non-pathogenic Cryptococcus spp. have been of intense interest4. Three key virulence traits have emerged from these studies: (1) the ability to grow at host body temperature (37 °C); (2) the production of a polysaccharide capsule; and (3) the generation of melanin pigment1. Strains or species that lack these traits are hypo-virulent and, conversely, expression of the three traits correlates with patient outcome among strains that infect humans5,6.
The big three virulence traits do not, however, completely explain the variations in virulence among Cryptococcus spp. and strains7. For example, Litvintseva and Mitchell showed that the virulence of C. neoformans strains isolated from the environment was greatly reduced relative to clinical strains despite no difference in the expression of the big three traits3. In addition, a systematic comparison of virulence traits and outcomes of genetically similar C. neoformans strains isolated from patients demonstrated wide variation in patient outcomes that could not be explained by differences in high temperature growth, capsule formation, or melaninization8,9. These observations clearly indicate that additional virulence traits must distinguish Cryptococcus strains that are successful mammalian pathogens from those that are not.
Our group recently found that C. neoformans strains isolated from the environment and unable to infect humans or mammals grow poorly in an atmosphere that contained carbon dioxide (CO2) concentrations (5%) corresponding to the host environment10. In contrast, strains isolated from patients were much more tolerant to an atmosphere of 5% CO2. Indeed, the few environmental isolates that grew well in 5% CO2 were also virulent in a mouse model of cryptococcosis. Importantly, the effects of CO2 on the growth of environmental and clinical C. neoformans strains were independent of medium pH. The environmental niche of C. neoformans contains approximately 0.04% CO2 while most tissues in the human host have CO2 concentrations that correspond to 5%. Therefore, the transition from an environmental niche to the mammalian host requires the fungus to tolerate ~100-fold increase in CO2 concentrations and our previous work indicates that this is a significant, independent host-related stress that the fungus must overcome10.
To identify the mechanisms that underlie CO2 tolerance in C. neoformans, we screened systematic libraries of deletion mutants covering protein kinases and transcription factors in the CO2-tolerant reference strain H9911,12. Coupling these screens with transcriptional profiling revealed that CO2 tolerance is dependent upon the target of rapamycin (TOR) pathway and membrane phospholipid asymmetry. Importantly, we have also found evidence that the TOR pathway’s role in phospholipid asymmetry has been significantly rewired relative to the model yeast, Saccharomyces cerevisiae, and we identified a putative ABC/PDR family transporter, PDR9, that plays a key role C. neoformans CO2 tolerance. Finally, the TOR-mediated response to host CO2 suppresses the activation of cell pathways that respond to host temperature and pH. As a result, C. neoformans cells must strike an exquisite balance between potentially contradicting stress responses to successfully transition from the environment to the host.
Results
The TOR and cell wall integrity MAPK pathways play opposite roles during CO2 stress
Protein kinases (PK) and transcription factors (TF) are key regulators of the cellular response to extracellular stresses. To identify PKs and TFs that affect CO2 tolerance, we screened systematic deletion mutant libraries covering the majority of non-essential PKs and TFs in the CO2 tolerant C. neoformans strain background H9911,12. Host levels of CO2 are fungistatic to intolerant strains and mutants rather than fungicidal10. To generate quantitative data on the relative fitness of these mutants, we developed a competitive fitness assay (Fig. 1a). H99 expressing mNeonGreen was used as a reference strain and inoculated into a 96-well microtiter plate with an unlabeled PK or TF mutant at a 1:1 ratio in host-relevant RPMI 1640 medium buffered to pH 7 at 30 °C under either ambient or 5% CO2 for 24 h. The screen was performed at 30 °C rather than at the host temperature of 37 °C so that the CO2 phenotype of temperature sensitive mutants could be assessed. The relative ratio of mutant to reference was determined by flow cytometry and the ratio at 5% CO2 was normalized to growth under ambient air to generate a CO2 fitness score. Control experiments comparing H99 to CO2-intolerant strains demonstrated that the assay identified strains with fitness defects at 5% CO2 (Supplementary Fig. 1a).
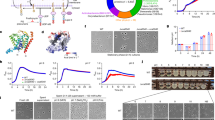
a Schematic representation of competition assay, created with BioRender.com. Overnight cultures of mNeon-Green labeled H99 and unlabeled mutant cells were combined in a 1:1 ratio in RPMI 1640 medium with 165 mM MOPS, pH 7 and incubated at 30 °C for 24 h in ambient air or with 5% CO2. Cell populations were characterized by flow cytometry and the percentage of mNeonGreen negative cells in 5% CO2 normalized to those in ambient conditions to determine a competitive fitness score for each mutant strain, plotted in (b) for the kinase deletion library and (c) for the transcription factor deletion library. Colored data points indicate values that were statistically significant by CHI square test (P < 0.05). Source data and specific p-values reported in Supplementary Data 1 and 2. Pathways of interest with multiple significant hits are depicted above each graph with data points involved coordinately colored. Panels (d) and (e) represent the competitive index of independently generated mutant isolates of strains of interest. Bars represent the average and SEM of three biological replicates. Source data are provided as a Source Data file.
We screened a total of 129 PK and 155 TF mutants in duplicate11,12. Scatter plots summarizing the screens are shown in Fig. 1b & c. A total of 21 PK mutants showed statistically significant alterations in competitive fitness relative to H99 with 14 mutants hyper-susceptible to CO2 while 7 mutants displayed increased fitness. Importantly, the cbk1∆ mutant showed reduced fitness which corroborates Chadwick et al. who showed that the RAM pathway is important for CO2 tolerance13. In contrast, all but one of the TF mutants with altered CO2 fitness were more resistant than H99. The full results of the screen are provided in Supplementary Data 1 and 2. We re-constructed 7 PK and 4 TF deletion mutants with altered CO2 fitness and determined their competitive fitness in independent experiments (Fig. 1d, e). The majority of mutants tested on agar plates showed reduced growth, although some mutants (e.g., sch9∆ bwc2∆) only showed a phenotype in liquid media; this is likely due to the increased sensitivity of direct competition assay (Supplementary Fig. 1b).
Two major protein kinase signaling pathways emerged from this data set. First, deletion mutants of YPK1, SCH9, and GSK3 were hypersusceptible to elevated CO2 concentrations and all are part of the TOR pathway11,14, strongly suggesting that TOR is required for CO2 tolerance (Fig. 1b, d). Although Ark1 has not been characterized in C. neoformans, its S. cerevisiae homolog negatively regulates Tor-dependent endocytosis, tying it to the TOR pathway as well15. Second, deletion of the MAPKKK, MAPK, and MAPK kinases (MKK1, BCK1, and MPK1) of the cell wall integrity (CWI) pathway increased CO2 fitness (Fig. 1b, d), suggesting that this pathway may be maladaptive for CO2 stress. Importantly, the TOR and CWI pathways have been shown to negatively regulate the other in both S. cerevisiae and C. neoformans14,16.
The only TF mutant that showed reduced fitness (albeit modest) in 5% CO2 was BWC2, a component of the blue-white light sensing system in C. neoformans17. In contrast, multiple TF deletion mutants showed increased fitness in 5% CO2 relative to H99. Among these, a previously defined pathway of Yap1, Gat201 and Gat20418 emerged (Fig. 1c). Gat201 and Gat204 are GATA TFs that regulate multiple virulence traits in C. neoformans19. Additionally, Gat201 binds the GAT204 promoter19 and regulates its expression of Gat204 (Fig. 1c). Furthermore, Jang et al. found that Yap1 functions upstream of Gat201 during capsule induction18. The consistent phenotypes observed across components of previously described regulatory pathways provide confidence that these genes are important for CO2 responses.
Finally, the deletion mutant of Rim101, a TF that is critical for adaptation to host pH20, is also CO2 resistant (Fig. 1c, e). The cAMP-PKA pathway functions through Rim101 under some conditions21 and the pka1∆ mutant is modestly resistant to CO2 as well (Fig. 1b). From these genetic data, it appears that signaling pathways that are required for adaptation to other host-related environmental stresses such as elevated temperature (CWI pathway) and alkaline pH (Rim101 pathway) may reduce adaptation to host CO2 concentrations. It is also important to note that 9 PKs not related to the TOR pathway with diverse functions are also required for CO2 tolerance (Fig. 1b), further supporting the notion that it represents a significant cellular stress for C. neoformans.
The TOR pathway is required for C. neoformans tolerance of elevated CO2 concentrations
A single, essential Tor kinase is present in C. neoformans and, consequently, the construction of TOR1 deletion mutants is not possible14. To test the hypothesis that CO2 tolerance is dependent on TOR, we compared the antifungal activity of the TOR inhibitor rapamycin (RAP) under ambient and 5% CO2 conditions. RAP was more potent in 5% CO2 compared to ambient conditions (Fig. 2a). RAP was also more active against an environmental strain, A7-35-23, at elevated CO2 (Fig. 2b). Because deletion mutants of YPK1 and SCH9 are both CO2 sensitive and hypersensitive to RAP14, we asked if resistance to CO2 of TF mutants correlated with RAP resistance. Interestingly, rim101∆ mutants were relatively resistant to RAP at elevated CO2 compared to H99 but not under ambient conditions (Fig. 2c). Thus, it is possible that these TF mutants have elevated TOR pathway activity at baseline as a compensatory response to the defects induced by the mutations and this leads to increased fitness in elevated CO2.
Ten-fold serial dilutions from overnight cultures of (a) H99, (b) environmental strain A7-35-23 and (c) transcription factor mutant rim101Δ were spotted on solid RPMI medium with 165 mM MOPS, pH 7 alone or with increasing concentrations of rapamycin as indicated at 30 °C in ambient air or at 5% CO2 for 48 hours before images were acquired. Images are representative of three biological replicates for each condition.
The TOR pathway is a conserved regulator of many cellular processes including ribosome biosynthesis, protein translation, amino acid transport, autophagy, actin polarization, and membrane homeostasis among others22. Previous studies of the C. neoformans TOR pathway confirmed these conserved functions14. Specifically, inhibition of TOR with RAP down-regulated genes involved in rRNA processing, ribosome biogenesis, and actin cytoskeleton while transmembrane transporters and carbohydrate metabolic genes were up-regulated14. To further test the hypothesis that CO2 triggers a TOR-mediated cellular response, we characterized the transcriptional profile of H99 cells grown in buffered RPMI medium at 37 °C under ambient or 5% CO2 concentrations (Supplementary Data 3; (Wald test p values and Benjamini-Hochberg adjustment for multiple comparisons)). Somewhat surprisingly, CO2 induced a relatively slow transcriptional response with only 9 genes differentially expressed (adjusted P value < 0.05, log2 ± 1) relative to ambient conditions after a 4-hour exposure to 5% CO2 (Fig. 3a).
a Volcano plot of genes identified by RNA-Seq in H99 cultured in RPMI 1640 medium with 165 mM MOPS, pH 7 at 4-, 8- or 24-hours incubation at 37 °C in 5% CO2 compared to ambient air. Source data and specific p-values (Wald test p values with Benjamini-Hochberg) are reported in Supplementary Data 3. GO terms for the differentially expressed genes (±log2 1 and FDR < 0.05, Benjamini–Hochberg) at 24 h are represented in semantic similarity scatterplots for genes with (b) elevated expression or (c) decreased expression in 5% CO2 compared to ambient air. Source data and specific p-values reported in Supplementary Data 4. d Venn diagram of differentially expressed genes at 24 h in 5% CO2 compared to ambient air and differentially expressed genes in H99 with 3 ng/mL rapamycin or YPD alone at 3 hours (reported in ref. 14). Source data are provided as a Source Data file.
Eight hours after exposure to 5% CO2, 199 genes were differentially expressed with 192 of those genes downregulated. This set of downregulated genes was enriched for membrane (FDR .012, Benjamini-Hochberg) and integral membrane proteins (FDR 0.024); indeed, 60 of the 199 differentially expressed genes were membrane-associated proteins. By 24 h; however, a total of 1204 genes were differentially expressed (log2 ± 1, FDR < 0.05) with the expression of 528 genes downregulated and 677 upregulated (Fig. 3a). GO term analysis (FDR < 0.05, Benjamini-Hochberg) indicates that the upregulated genes are enriched for rRNA processing, ribosome biogenesis, DNA replication and aromatic compound synthesis (Fig. 3b and Supplementary Data 4) while the down-regulated genes were enriched for transmembrane transport, carbohydrate metabolism, redox process, and cellular response to heat (Fig. 3c and Supplementary Data 4). As indicated by the Venn diagram shown in Fig. 3d, 22% of the genes upregulated in CO2 (151/677) are downregulated in cells treated with RAP while 33% (176/528) of genes downregulated in CO2 are upregulated in the presence of RAP. Overall, the transcriptional response to CO2 occurs over multiple hours and appears to have two phases: an early phase that is highly enriched for membrane-associated genes and a late phase that is consistent with the activation of the TOR pathway. The delayed nature of this response would suggest that gene expression may be compensating for the physiological and biophysical effects of elevated CO2 rather than through a direct sensing of the elevated CO2 concentrations.
Elevated CO2 suppresses CWI MAPK signaling in CO2-tolerant but not intolerant strains
Genes involved in the cellular response to heat are downregulated when cells are shifted from ambient to host-like concentrations of CO2 (Fig. 3c). The CWI pathway is a key positive regulator of the C. neoformans response to elevated temperature23,24,25. Because deletion mutants of PKs in the CWI pathway (BCK1, MKK2, MPK1) in H99 background are resistant to CO2 (Fig. 1b, d), it appears that the expression of temperature stress-related genes may be maladaptive during CO2 stress. Indeed, we have previously reported that elevated temperature exacerbates CO2 stress13. In S. cerevisiae and C. neoformans, activation of the TOR pathway negatively regulates the CWI pathway14,16, suggesting the hypothesis that 5% CO2 may blunt CWI pathway activation. To test this hypothesis, we used a phospho-specific antibody to monitor the phosphorylation of Mpk1, the terminal MAPK of the CWI pathway, following temperature shift from 30 °C to 37 °C in ambient or 5% CO2 (Fig. 4a). Consistent with previous literature23,24, Mpk1 phosphorylation is increased in CO2-tolerant H99 at 37 °C in ambient air conditions and this level of phosphorylation is maintained for the 4-hour time course. In 5% CO2, temperature-induced Mpk1 phosphorylation was delayed and was consistently reduced relative to ambient CO2 conditions over the time course (Fig. 4a, left panel). These data clearly demonstrate that the activation of the CWI pathway is blunted by host levels of CO2.
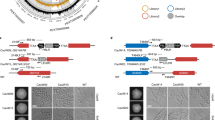
a Ten μg of total cell lysate from H99 or A7-35-23 cells harvested at 0-, 1-, 2-, 3-, and 4-hours post-shift from 30 °C to 37 °C were analyzed by western blot for phosphorylated-Mpk1 in ambient or 5% CO2 conditions. Cells were grown in YPD, shaking at 200 rpm. Total loading was visualized by Ponceau. Band intensity relative to the 0-hour time point for each strain is indicated. Results are representative of three biological replicates. b Schematic representation of Rim101 processing pathway. Cleavage of Rim101 by the Rim13 protease leads to activation and translocation to the nucleus. c Overnight cultures of mNeon-Green labeled H99 and unlabeled mutant cells were combined in a 1:1 ratio in RPMI 1640 medium with 165 mM MOPS, pH 7 and incubated at 30 °C for 24 h in ambient air or with 5% CO2. Cell populations were characterized by flow cytometry and the percentage of mNeonGreen negative cells in 5% CO2 normalized to those in ambient conditions to determine a competitive fitness score for each mutant strain as indicated. Bars represent the average and SD from three technical replicates. An ordinary one-way ANOVA with Tukey’s multiple comparisons test was performed using GraphPad Prism. Significance relative to H99 is represented; ****P < 0.0001; Prim13 = 2.172−7; Prim20 = 5.014−8; Prim101 = 5.507−5. Results are representative of three biological replicates. d Overnight cultures of H99 expressing GFP-tagged Rim101 were diluted to an OD600:0.2 and grown to an OD600:1 in RPMI 1640 with 165 mM MOPS, pH 7 shaking at 200 rpm in ambient air or 5% CO2. Rim101-GFP was pulled down from the whole cell lysate of an equivalent number of cells with GFP-trap beads. An anti-GFP antibody was used to probe the separated proteins on a nitrocellulose membrane. Full-length (FL) and processed (P) Rim101-GFP are indicated. Results are representative of three biological replicates. e Venn diagram of genes downregulated by rapamycin treatment in H99 compared to those that are upregulated under CO2 in the rim101Δ mutant compared to CO2-exposed H99. RNA-Seq source data and specific p-values (Wald test p values and Benjamini-Hochberg) are reported in Supplementary Data 5. Source data are provided as a Source Data file.
We next asked if temperature-induced Mpk1 phosphorylation was also blunted by 5% CO2 in a CO2-sensitive strain. To do so, we performed the same temperature-shift experiment with A7-35-233,10, a CO2 sensitive environmental isolate (Fig. 4a, right panel). Mpk1 phosphorylation increased at 37 °C in A7-35-23 but the extent of Mpk1 phosphorylation was unaffected by growth in 5% CO2. These data, combined with the genetic and transcriptional results discussed above, support a model in which CO2 suppresses temperature-induced activation of the CWI pathway and that this suppression may improve tolerance to host CO2 concentrations.
Reduced Rim101 pathway activity leads to CO2 and rapamycin resistance
The Rim101 pathway regulates cellular responses to alkaline pH, capsule formation, and cell wall biosynthesis20,21,26; our data indicate that it, too, is maladaptive during CO2 stress (Fig. 1c, e). Rim101 is activated by proteolysis which in turn is mediated by a well-defined protein complex in C. neoformans (Fig. 4b). Deletion mutants of two additional components of the Rim101 pathway, the calpain protease Rim13 and the scaffolding protein Rim20, were also resistant to CO2 relative to H99 (Fig. 4c). Next, we characterized the effect of CO2 on the proteolytic processing of Rim101 using western blot analysis of cells containing a Rim101-GFP allele as previously reported20,21. Under ambient CO2 in buffered RPMI medium, both unprocessed (~150 kD band; FL) and processed (~100 kD band; P) Rim101-GFP are detectable (Fig. 4d); additional degradation products are also present in the blot. In 5% CO2, neither the processed nor the unprocessed forms of Rim101 are detectable and increased amounts of a band corresponding to free GFP are present. These data suggest that Rim101 is degraded in 5% CO2 and are consistent with the idea that Rim101 activity is reduced to compensate for CO2 stress.
We next characterized the effect of the Rim101 pathway on the transcriptional response to growth at 5% CO2. The rim101∆ mutant was incubated in buffered RPMI medium in ambient or 5% CO2 for 24 h. The Rim101 TF has a broad effect on gene expression with 680 genes upregulated and 859 genes downregulated in the rim101∆ mutant compared to H99 in 5% CO2 (log2 ± 1, FDR < 0.05, Supplementary Data 5; (Wald test p values and Benjamini-Hochberg adjustment for multiple comparisons)). Because the rim101∆ mutant is resistant to RAP, we hypothesized that genes upregulated in the mutant may correspond to genes downregulated by RAP. Indeed, 150 of the 360 genes (41%) downregulated by inhibition of TOR14 are upregulated in the rim101∆ mutant (Fig. 4e). GO term analysis of this set demonstrates that ribosome biogenesis (40 genes; p = 8.3 × 10−37), ribonucleoprotein complex processing (40 genes, p = 1.8 ×10−33), and RNA processing (38 genes; p = 5.8 × 10−22) are strongly enriched and represent 25% of the upregulated genes in rim101∆. These genes are key effectors of the TOR pathway and are upregulated beyond the level of expression induced by CO2 in H99. These transcriptional data support the hypothesis that deletion of Rim101 leads to compensatory activation of the TOR pathway and, consequently, increased CO2 tolerance.
Elevated expression of putative ABC transporter CNAG_07799/PDR9 reduces CO2 fitness
Next, we sought to identify non-regulatory genes that modulate CO2 fitness. To do so, we examined the expression of CO2-responsive genes in CO2-tolerant and -intolerant strains using Nanostring nCounter technology. A focused set of 118 genes covering a diverse set of functions that showed differential regulation in CO2 ( ± 1 log2, FDR < 0.05) based on the RNA seq data at the 24-hour time point above (see Supplementary Data 6 for full set of genes, raw and processed data). The transcriptional profiles of CO2-tolerant and intolerant strains were distinct as shown in Fig. 5a. The sensitive strains show a more consistent profile across different strains than the CO2-tolerant strains. For example, the profile for CO2-tolerant strain C23 is quite distinct from the other two CO2-tolerant strains. In contrast, the CO2-sensitive strains have sets of genes that are consistently highly expressed or lowly expressed. No genes were consistently highly expressed in all tolerant strains relative to intolerant genes. However, a set of 13 genes was expressed higher in the CO2-sensitive stains relative to the tolerant strains (Fig. 5a inset). Five out of these 13 genes have predicted functions related to membrane and lipid homeostasis: ECM2201, CNAG_03227, HAPX, CNAG_07799/PDR9, IPC1, and SRE1. ECM2201, HAPX, and SRE1 are TFs with a role in regulating ergosterol biosynthesis genes. Deletion mutants of the three TFs showed CO2 fitness that was similar to WT (Supplementary Data 2). IPC1 codes for the enzyme that generates inositol-phosphatidyl-ceramide which is the target of the antifungal molecule aureobasidin27; increased CO2, however, did not affect the antifungal activity of aureobasidin (Supplementary Fig. 2).
Indicated CO2-sensitive or -tolerant strains were cultured in RPMI 1640 medium with 165 mM MOPS, pH 7 for 24 hours at 37 °C in ambient air or 5% CO2. Total RNA was isolated from harvested cells, 100 ng of RNA was hybridized to a custom Nanostring probe set and quantified on a Nanostring Sprint nCounter. a Normalized counts for strains grown in 5% CO2 conditions are presented as a heat map hierarchically clustered in Morpheus. Gene names and functions for the subset of 13 genes expressed highly in CO2-sensitive strains compared to CO2-tolerant strains are described. Source data are reported in Supplementary Data 6. b Normalized Nanostring counts from total RNA for PDR9 in ambient or 5% CO2 conditions. c Overnight cultures of mNeon-Green labeled H99 and unlabeled H99 or PH3:PDR9 cells were combined in a 1:1 ratio in RPMI 1640 medium with 165 mM MOPS, pH 7 and incubated at 30 °C for 24 h in ambient air or with 5% CO2. Cell populations were characterized by flow cytometry and the percentage of mNeonGreen negative cells in 5% CO2 was normalized to those in ambient conditions to determine a competitive fitness score for each strain as indicated. Bars represent the average and SEM from two biological replicates. A two-tailed, unpaired t test was performed using GraphPad Prism. *P < 0.05; P = 0.0151. Source data are provided as a Source Data file.
CNAG_07799 is one of 10 PDR/ABCG family ABC transporters in the C. neoformans genome and has been named PDR9 by Winski et al.28. Phylogenetic analysis places PDR9 in Clade III of fungal PDR/ABC transporters and its closest C. neoformans homologue is AFR1 which is involved in azole resistance28. In addition to mediating drug efflux, PDR/ABC transporters have significant roles in lipid transport and membrane homeostasis29. We were, therefore, interested in its potential effect on CO2 tolerance.
The expression of PDR9 is increased by CO2 exposure in both tolerant and sensitive strains (Fig. 5b). However, the absolute expression of PDR9 in ambient and 5% CO2 conditions is much higher in sensitive strains. For tolerant strains, the expression of PDR9 following induction by 5% CO2 remains well below the baseline levels of intolerant strains. These data suggest that it is the expression level of PDR9 that is important for CO2 tolerance and not the fold change from ambient to 5% CO2. Nanostring profiling of TF mutants with altered CO2 tolerance also showed that PDR9 was downregulated in the resistant mutant rim101∆ but was upregulated in the hypersensitive mutant bwc2∆ (Supplementary Fig. 3a). These data suggested us that elevated expression of PDR9 may reduce CO2 fitness.
To test this hypothesis, we constructed a derivative of the CO2 tolerant strain H99 in which PDR9 was placed under the control of the Histone 3 promoter which has been used by So et al. to overexpress other genes14; importantly, these strains have increased expression of PDR9 by RT-PCR analysis at both ambient and 5% CO2 with the expression in CO2 slightly higher (<1.5 fold, Supplementary Fig. 3b). Consistent with our hypothesis, the PH3-PDR9 strain show reduced fitness in 5% CO2 relative to its H99 parental strain (Fig. 5c). Similar results were obtained on spot dilution assays (Supplementary Fig. 3c). We also introduced the PH3-PDR9 allele into a second C. neoformans background (A1-84-14) and observed an increase in CO2-susceptibilty, indicating that the phenotype is not dependent on strain background (Supplementary Fig. 3d). Thus, absolute expression of PDR9 correlates with reduced growth in host concentrations of CO2. It is likely that the increased expression of PDR9 in H99 and other relatively tolerant strains induced by CO2 contributes to their modest reduction in growth in 5% CO2 relative to ambient CO2 conditions (see Fig. 2a and Supplementary Fig. 3c). However, our data indicate that the absolute expression levels and not the difference in expression of PDR9 between ambient and elevated CO2 correlate with sensitivity to CO2.
We previously reported that CO2 increases C. neoformans susceptibility to the ergosterol inhibitor fluconazole and myriocin, an inhibitor of the first step of sphingolipid biosynthesis10. Since S. cerevisiae homologs of PDR9 affect susceptibility to these drugs, we tested the susceptibility of the PDR9 overexpression strain (Supplementary Fig. 3e). There was no difference in the zone of inhibition for fluconazole, indicating that PDR9 does not act as a fluconazole efflux pump and that it is unlikely that the strains have reduced ergosterol content28,30. In contrast, myriocin susceptibility was increased in the overexpression strain, suggesting that increased expression of PDR9 may alter sphingolipid homeostasis.
C. neoformans remodels phospholipid asymmetry through the TOR pathway in response to CO2 stress
The TOR pathway regulates phospholipid asymmetry at the plasma membrane (PM) by modulating the activity of aminoglycerolipid (AGL) flippases via Ypk1 in S. cerevisiae31. The flippases transport phosphatidylethanolamine (PE) and/or phosphatidylserine (PS) from the outer leaflet of the PM to the inner, cytosolic face of the PM and floppases transport PE/PS in the opposite direction32. Based on the phenotypes of TOR mutants and their function in S. cerevisiae, we hypothesized that PM asymmetry may play an important role in C. neoformans tolerance of host CO2 stress.
To test this hypothesis, we first examined the effect of CO2 and mutants with altered CO2 tolerance on susceptibility to the antifungal duramycin. Duramycin binds PE on the PM outer leaflet as part of its mechanism of action and cells with increased outer leaflet PE exposure show increased susceptibility to the drug31. Although the susceptibility of H99 to duramycin is not affected by 5% CO2, the ypk1∆ mutant is sensitive to duramycin, relative to H99 (Fig. 6a, b). In S. cerevisiae, the TOR pathway inhibits PE flippases though activation of Ypk1 which, in turn, is a negative regulator of Fpk1; accordingly, overexpression of YPK1 increases duramycin susceptibility while the fpk1∆ deletion mutant is resistant to duramycin31. These data suggest that Ypk1 has a distinct role in PE homeostasis in C. neoformans. Supporting this conclusion, the duramycin susceptibility of the fpk1∆ mutant is unchanged relative to H99 (Fig. 6a). Interestingly, deletion of SCH9, a second TOR dependent kinase that is also hypersusceptible to CO2, has increased susceptibility to duramycin. If Ypk1 and Fpk1 have the same lipid asymmetry functions in C. neoformans and S. cerevisiae, then the ypk1∆ mutant would be resistant to duramycin and the fpk1∆ mutant would be hypersusceptible. These results indicate that the Tor-Ypk1 axis has a completely different effect on PE asymmetry in C. neoformans compared to S. cerevisiae. Ypk1 is required for CO2 tolerance while deletion of FPK1 has no effect on this phenotype (Supplementary Data 1), further supporting the idea that the relationship between these two TOR-regulated kinases is different in C. neoformans
a, b Cells from overnight cultures of indicated strains were spread on solid RPMI 1640 medium with 165 mM MOPS, pH 7. Sterile disks were placed on plates and 200 μg duramycin was added to each disk. Cells were incubated at 30 °C for 48 h in ambient air or in 5% CO2 before images were acquired. The zone of clearance was measured and indicated in centimeters in the top right corner for each strain/condition. Plates are representative of three biological replicates. c, d Annexin V staining was performed on cells of indicated strains after 24 h incubation in RPMI 1640 medium with 165 mM MOPS, pH 7 in ambient air or in 5% CO2. Images were captured on a Leica confocal microscope (representative images (c)) and mean fluorescence intensity (MFI) was measured in ImageJ software (d). At least 100 cells were quantified for each condition. nH99- = 817; nH99+ = 128; nPDR9- = 263; nPDR9+ = 206; nypk1Δ- = 307; nypk1Δ+ = 130. An ordinary one-way ANOVA was performed using GraphPad Prism with a Sidak follow-up test adjusted for multiple comparisons. ****P < 0.0001. PH99-vsH99+ = 1.82-53; PPDR9-vs.PDR9+ = 3.13−18; PH99-vs.PDR9+ = 1.15−45; PH99+vsPDR9+ = 4.18−50; Pypk1Δ-vs.ypkΔ1+ = 1.29−9; PH99-vs.ypk1Δ- = 1.75−48; PH99+vs.ypk1Δ+ = 9.25−49. e, f Cells from overnight cultures of H99 were cultured in RPMI 1640 medium with 165 mM MOPS, pH 7 at 30 °C in ambient or 5% CO2 for 18 h to mid-log phase. Cells were washed and labeled with NBD-phosphatidylserine (e) or NBD-phosphatidylethanolamine (f) for 30 min before washing and assessment of fluorescence by flow cytometry. Bars represent MFI with SEM from two biological replicates. Graphs are representative of experiments performed on three independent days. A two-sided, unpaired t test was performed using GraphPad Prism. ***P < 0.001. e P = 0.0002 (f) P = 0.0038. g Overnight cultures of mNeon-Green labeled H99 and unlabeled H99 or cdc50Δ cells were combined in a 1:1 ratio in RPMI 1640 medium with 165 mM MOPS, pH 7 and incubated at 30 °C for 24 h in ambient air or with 5% CO2. Cell populations were characterized by flow cytometry and the percentage of mNeonGreen negative cells in 5% CO2 was normalized to those in ambient conditions to determine a competitive fitness score for each strain as indicated. Bars represent the average and SEM from three biological replicates. An ordinary one-way ANOVA with Tukey’s multiple comparisons test was performed using GraphPad Prism. ***P < 0.001. PH99vs. cdc50-11Δ = 0.0002; PH99vs. cdc50-21Δ = 0.0002. Source data are provided as a Source Data file.
One of the proposed roles of PDR/ABC transporters in lipid homeostasis is as floppases29. Because TOR-related mutants with increased PM outer leaflet PE exposure show increased CO2 susceptibility, we tested the duramycin susceptibility of the PH3-PDR9 strain. As shown in Fig. 6b, overexpression of PDR9 increases susceptibility to duramycin at both ambient and 5% CO2. The changes in zones of clearance observed with the PH3-PDR9 strain are similar to those observed in S. cerevisiae mutants involved in PE homeostasis31. These data are consistent with PDR9 having a possible PE floppase activity and that its effect on CO2 tolerance may involve a role in membrane lipid homeostasis. The definitive biochemical characterization of floppases has been technically difficult and somewhat controversial. In S. cerevisiae, Pdr5 has been proposed to be a floppase33. Pdr5 is primarily localized to the PM33. We, therefore, tagged PDR9 with mNeonGreen under both the endogenous and PH3 promoter; the latter strain was hypersusceptible to CO2, confirming the mNeonGreen-tagged allele is functional (Supplementary Fig. 4a). In both strains and under multiple growth conditions, Pdr9-mNeonGreen localizes to intracellular puncta and not to the plasma membrane (Supplementary Fig. 4b). Since validated organelle markers in C. neoformans are limited, we have not conclusively localized Pdr9-mNeonGreen, but the pattern of signal is characteristic of late- Golgi/endosomes and not the PM. Thus, Pdr9 affects PE homeostasis in a manner consistent with a floppase. However, Pdr9 does not localize to the PM, which is where most proposed floppases are found, raising the possibility that this effect on PE distribution may be indirect. As such, additional biochemical and cell biological studies will be required before a definitive lipid asymmetry-related function for Pdr9 can be made with confidence.
Flippases and floppases also affect the distribution of PS between the inner and outer leaflets of the PM32. To test the effect of CO2 on the distribution of PS, we used the cell impermeant molecule, annexin V, to stain outer leaflet PS following recently procedures reported by Huang et al. for the use of this assay with C. neoformans34. Consistent with the previous report34, outer leaflet PS is low but detectable under ambient air conditions (Fig. 6c, d). Exposure to 5% CO2 in RPMI at 37 °C significantly increases outer leaflet PS staining in H99. Overexpression of PDR9 reduced the level of outer leaflet PS induced by 5% CO2 as did deletion of YPK1. Together, these data clearly indicate that adaptation to 5% CO2 is associated with increased outer membrane PS relative to PE and that disruption of this balance by increased expression of PDR9 or by deletion of YPK1 leads to reduced fitness in 5% CO2.
To further test the hypothesis that elevated CO2 concentrations increase PM outer leaflet PS concentrations by altering phospholipid asymmetry, we performed PS uptake assays using fluorescently labeled NBD-PS following a previously reported approach with some modifications34. Mid-log phase H99 cells grown in ambient conditions or 5% CO2 were exposed to NBD-PS for 30 min and cellular uptake of NBD-PS quantitated by flow cytometry; intracellular uptake of NBD-PS was confirmed by microscopy (Supplementary Fig. 5a). As shown in Fig. 6e, cells exposed to 5% CO2 show ~50% reduction NBD-PS uptake relative to cells grown in ambient air conditions. Cdc50 regulates the activity of flippases and, consequently, PS asymmetry in S. cerevisiae. Multiple labs have shown that Cdc50 is also required for PS flip activity in C. neoformans34,35. We, therefore, tested the effect of a cdc50∆ mutant on CO2 susceptibility. Consistent with previous data showing that the cdc50∆ mutant has increased annexin V staining, it is resistant to 5% CO2 relative to WT (Fig. 6f).
An alternative possibility is that total cellular PS is increased in the presence of 5% CO2 leading to the apparent increase in outer leaflet PS localization. To test that possibility, we modulated the total cellular synthesis of PS using a strain containing a copper-repressible allele of CHO1, an enzyme required for the synthesis of PS36. If CO2 tolerance is dependent on an increase in total PS, then preventing that increase by incubating the PCTR4-CHO1 strain on media containing high copper concentrations should reduce CO2 tolerance. As expected, the PCTR4-CHO1 strain has a growth defect on copper-replete media compared to copper-deficient media at ambient air (Supplementary Fig. 6); however, there is no further reduction in the growth of the strain when it is incubated at 5% CO2. The results of this experiment are more consistent with CO2 tolerance being dependent on lipid asymmetry remodeling than with a mechanism involving a global increase in PS synthesis.
Finally, increased expression of PDR9 leads to increased duramycin sensitivity which indicates an increase in outer membrane PE (Fig. 6b). To test if this phenotype is associated with alterations in PE transport, we compared the uptake of fluorescently labeled PE as well; intracellular uptake of NBD-PE was also confirmed by microscopy (Supplementary Fig. 5b). As shown in Fig. 6g, overexpression of PDR9 leads to a 2.5-fold reduction in uptake of NBD-PE. These data further support, but do not conclusively establish, the possible function of Pdr9 as a phospholipid floppase. Taken together, these data strongly support a model in which host-relevant concentrations of CO2 lead to remodeling of PS lipid asymmetry and that disruption of this remodeling reduces tolerance of those conditions.
Overexpression of PDR9 reduces C. neoformans virulence
Environmental strains with elevated expression of PDR9 are not only intolerant of host-relevant concentrations of CO2 but are also less virulent in mouse models of cryptococcosis3,10. We, therefore, hypothesized that the reduced CO2 tolerance of the PH3-PDR9 strain would translate to reduced infectivity and virulence. To test this hypothesis, we used two infection models. First, we carried out a competitive fitness experiment in which a 1:1 mixture of H99 and PH3-PDR9 was used to infect the respiratory tract of mice. The mice were euthanized at DPI 14 and a ratio of the two strains in the lung tissue was determined by quantitative plating on YPD and YPD + NAT (selective for the overexpression strain). Consistent with in vitro studies, the PDR9 overexpression strain was 10-fold less fit in the lung (Fig. 7a). We also compared the virulence of H99 and the PH3-PDR9 strain using single-strain infection experiments. As shown in the survival curve in Fig. 7b, infection with H99 led to 70% moribundity by DPI 40 while no animals infected with the PH3-PDR9 strain showed signs of distress to day 86. The fungal burden at day of sacrifice was determined for five H99-infected animals and compared to the fungal burden of PH3-PDR9 strain-infected animals at DPI 86. Consistent with expectations, H99-infected animals showed high fungal burden in both the lung (Fig. 7c) and brain tissue (Fig. 7d). Interestingly, 4 out of five animals infected with the PH3-PDR9 strain had ~3 log10 organisms in the lung but none in the brain. This observation suggests that overexpression of PDR9 reduces lung burden and reduces dissemination to the brain.
a Five CD1 mice were infected intranasally with a 1:1 ratio of H99 and PH3:PDR9 cells (5 × 104 per mouse). At DPI 14, mice were sacrificed and the lung was dissected. For fungal burden quantifications, lungs were homogenized and serially diluted, then plated onto YNB and YNB with 100 μg/mL of nourseothricin (NAT) and incubated at 30 °C for two days before counting CFUs. The competitive fitness in vivo was determined by dividing the CFU of PH3:PDR9 cells (NAT resistant) by H99 cells. Data are represented with the average and SD. For the survival study (b), ten CD1 mice per group were infected intranasally with 1 × 104 fungal cells. Fungal burden was examined for five mice at the time of termination or at DPI 86. For fungal burden quantifications, lungs (c) or brains (d) were processed as above. e, f H99 and PH3:PDR9 cells were grown in RPMI 1640 + 165 mM MOPS for 24 h at 37 °C in ambient air or 5% CO2, then prepared for microscopy by counterstaining with India ink (representative images (e)). At least 50 cells were quantified per biological replicate, with three biological replicates per condition and processed in ImageJ software to measure capsule. Average capsule thickness at 24 h was normalized to 0 h for each biological replicate (f). Bars represent average and SEM. Two-way ANOVA with Tukey’s multiple comparisons test was performed in GraphPad Prism. **P < 0.01, ****P < 0.0001. PH99-vs.PH3:PDR9- = 0.0073; PH99-vs.H99+ = 0.0072; PPH3:PDR9-vs.PH3:PDR9+ = 2.49−5; PH99+vs.PH3:PDR9+ = 2.51−5. g J774 murine macrophage cells were incubated with opsonized H99 or PH3:PDR9 cells. Phagocytosis was assessed after 3 h by washing away non-phagocytosed yeast cells, lysing macrophages, and plating on solid YPD. Plates were incubated at 30 °C for 2 days before counting CFUs. CFU/ml were normalized to H99. Bars represent the average and SEM from three experimental replicates. A two-sided, ratio paired t test was performed using GraphPad Prism. **P < 0.01, P = 0.0044. Source data are provided as a Source Data file.
To determine if the reduced virulence of the PH3-PDR9 strain was due to alterations in the three canonical C. neoformans virulence traits2, we compared its growth at 37 °C, melaninization, and capsule formation to H99 under both ambient and 5% CO2 conditions. The PH3-PDR9 strain grew similar to H99 at 37 °C (Supplementary Fig. 7a) and showed no changes in melaninization (Supplementary Fig. 7b). Unexpectedly, the capsule formed by the PH3-PDR9 strain was 1.5-fold thicker relative to H99 in 5% CO2 (Fig. 7e, f). In mouse models, reduced capsule formation is generally associated with reduced virulence. One possible explanation for this discordance is that the capsule formed by PH3-PDR9 is quantitatively larger but may have qualitative or structural defects that affect its function. One of the best characterized functions of a capsule is to interfere with phagocytosis by macrophages37. Using the murine macrophage cell line J774, we compared the phagocytosis of the PH3-PDR9 strain to H99. Consistent with its increased capsule formation, phagocytosis of the PH3-PDR9 strain was reduced relative to H99 (Fig. 7g). This suggests that, to a first approximation, the capsule formed by this mutant functions as expected. Thus, we assert that the best explanation for the reduced fitness of the PH3-PDR9 strain in vivo is its reduced ability to tolerate host levels of CO2.
Discussion
Our group recently reported that the ability of C. neoformans to tolerate host levels of CO2 correlates with virulence in strains that express the three most well-characterized virulence traits10. Here, we identify the TOR pathway and remodeling of plasma membrane lipid asymmetry as key factors contributing to the ability of C. neoformans to tolerate host CO2 stress. The role of these processes as well as other PKs involved in stress responses support the conclusion that the transition from ambient CO2 concentrations in the environment to the 100-fold higher levels in the host represents an independent stress in C. neoformans; further supporting this conclusion is the large number of genes that are differentially expressed in cells replicating in host concentrations of CO2 compared to ambient conditions. Of the 15 mutants with reduced fitness in host CO2 conditions in vitro, twelve have reduced fitness in vivo as well, strongly supporting the concept that CO2 tolerance/adaptation is required for C. neoformans virulence10.
At the same time, two pathways that regulate C. neoformans adaptation to other features of the host environment, the CWI and Rim101 pathways, suppress CO2 tolerance. In addition, a transcription factor circuit that induces the expression of genes required for virulence, the Yap1-Gat201-Gat204 pathway, is also relatively maladaptive in the setting of CO2 stress. Host CO2 stress, therefore, appears to present C. neoformans with a conundrum in which pathways such as TOR must be activated for it to replicate while at the same time other pathways required for host adaptation must be suppressed. Very few Cryptococcus strains can adapt to mammalian physiology3,4 and we propose that the requirement for the fungus to balance contradictory physiological responses to host CO2, pH, and temperature may contribute to this bottleneck.
Our previous work has demonstrated that host temperature (37 °C) exacerbates CO2 stress13 and, therefore, it would be reasonable to expect the CWI pathway to be important for tolerance of CO2. Instead, CO2 suppresses activation of the CWI pathway, likely through the activation of the TOR pathway. There are, however, multiple signaling pathways that are required for temperature tolerance. We propose that suppression of CWI activity necessitates activation of the other pathways to mediate critical compensatory responses during adaptation to both the CO2 and temperature stresses of the host environment. Supporting this model is the fact that the RAM pathway is required for C. neoformans adaptation to host CO2 concentrations and temperature13. Furthermore, 10/14 PK mutants with reduced fitness in host CO2 are also hypersensitive to elevated temperature11. Therefore, our data suggest that CO2 and temperature are interrelated host stressors that require responses that must be finely balanced for C. neoformans to establish infection.
Although the TOR pathway plays pleiotropic roles in eukaryotic biology, we are unaware of previous reports linking it to CO2 responses. In fungi, however, studies of the biology of CO2 have focused almost entirely on the mechanisms by which these organisms generate sufficient bicarbonate in low CO2 environments such as the external air38. Two of the most well-studied functions of the TOR pathway are translational regulation and phospholipid homeostasis14,31. Our results indicate that both of these functions are critical for C. neoformans adaptation to host CO2 concentrations. The induction of canonical TOR target genes such as ribosomal components and translational machinery strongly supports this conclusion. Additional studies, however, will be needed to identify the specific transcription factors that mediate this transcriptional response because our screen of single gene deletion mutants has not identified candidate regulators. This suggests that there are likely to be overlapping TFs with redundant functions and that genetic interaction analysis will be needed to uncover those that are critical to the transcriptional response to CO2.
Our work has shed light on the role of the TOR pathway in maintaining lipid homeostasis at host CO2 concentrations in C. neoformans. Previous studies in S. cerevisiae39 as well as in C. neoformans40 have shown that TOR acts through its target Ypk1 to regulate both phospholipid asymmetry at the plasma membrane and sphingolipid biosynthesis. The ypk1∆ mutant is highly sensitive to CO2, indicating that this arm of the TOR pathway is also critical to high CO2 adaptation. We previously reported that CO2 increases C. neoformans susceptibility to the sphingolipid biosynthesis inhibitor myriocin10. The Del Poeta lab has also found that mutants (smt1∆ and gcs1∆) involved in sphingolipid biosynthesis have specific growth defects in host-like in vitro conditions containing 5% CO241. Taken together, these observations are consistent with the conclusion that TOR-Ypk1-regulated sphingolipid biosynthesis is an important cellular response to CO2 in C. neoformans.
In S. cerevisiae, TOR-Ypk1 regulates the function of phospholipid flippases responsible for maintaining the asymmetric distribution of PS and PE at the plasma membrane31,39. Our data suggests that the TOR-Ypk1 axis functions differently in C. neoformans compared to S. cerevisiae. The C. neoformans ypk1∆ mutant is sensitive to duramycin, an antifungal molecule that binds outer leaflet PE. The same mutant in S. cerevisiae is as sensitive to duramycin as WT while overexpression of YPK1 increases sensitivity31. In S. cerevisiae, Ypk1 inhibits the function of Fpk1 which activates flippases31. Consequently, deletion of FPK1 increases duramycin sensitivity in S. cerevisiae31; in C. neoformans, deletion of FPK1 has no effect on duramycin susceptibility or on C. neoformans CO2 tolerance. Thus, it appears that the function of the Ypk1-Fpk1 pair is distinct in C. neoformans compared to S. cerevisiae. Fpk1 contains a sequence that corresponds to the canonical Ypk1 substrate motif, suggesting it may still be a substrate for Ypk142. However, Bahn and co-workers, as part of their large-scale analysis of PKs in C. neoformans11, also observed phenotypes that were inconsistent with the conservation of the Ypk1-Fpk1 relationship between S. cerevisiae and C. neoformans. Additional work will be needed to understand the molecular mechanisms underlying these apparent changes in Tor-Ypk1-Fpk1 regulation of PE homeostasis.
The TOR-Ypk1 axis has also been implicated in the regulation of PS distribution at the plasma membrane in S. cerevisiae43. Whereas there appears to be little change in PE distribution in response to CO2 (Fig. 6b), PM outer leaflet PS exposure increases in H99 cells in host-relevant CO2 concentrations in a Ypk1-dependent manner (Fig. 6d). In the case of PS homeostasis, therefore, Ypk1 is functioning similarly in both C. neoformans and S. cerevisiae. It is somewhat surprising that the C. neoformans responds to CO2 by increasing outer membrane PS because this is most commonly an indication of apoptosis in eukaryotic cells32. It is important to note that 5% CO2 does not kill either tolerant or intolerant C. neoformans strains but is fungistatic10; therefore, increased PS exposure does not appear to be an indication of reduced viability. At this point, it is unclear how increased PM outer leaflet PS suppresses the fungistatic effects of elevated CO2. Our transcriptional and phenotypic data strongly support the notion that elevated CO2 concentrations affect plasma membrane and lipid homeostasis. In general, PS supports membrane fluidity whereas increased CO2 concentrations are reported to decrease the fluidity of microbial membranes44. Based on these opposing biophysical effects of PS and CO2 on biological membranes, one potential explanation for the increased CO2 fitness of C. neoformans cells is that the increased outer leaflet PS promotes membrane fluidity to counter the membrane rigidifying effects of elevated CO2.
Our data also indicate that the increased amount of PM outer leaflet PS in C. neoformans at host concentrations of CO2 is due to reduced uptake of PS and not due to a global increase in cellular PS. Previous studies have confirmed that Cdc5034,35, a conserved flippase subunit that regulates multiple flippases, is required for PS flippase function in C. neoformans. Consistent with that function, deletion of CDC50 increases CO2 tolerance, further supporting the conclusion that increased PM outer leaflet PS reduces the toxicity of host concentrations of CO2. Another possible mechanism for increased PM outer leaflet PS levels, would be an increase in transport from the inner leaflet to the outer leaflet also known as PS floppase activity. To date, no bona fide PS floppase has been described in fungi although Pdr5 appears to function as a PE floppase in S. cerevisiae33. Still, many ABC/PDR genes remain to be characterized and, in principle, could carry out this function28. Overall, our data and prior literature precedents are most consistent with a mechanism in which CO2 triggers a cellular response that dramatically reduces PS uptake and leads to a corresponding increase in outer leaflet PS.
Although annexin V staining is widely used as an assay for alterations in PS distribution between the inner and outer leaflets of the plasma membrane, some studies have indicated that annexin V is not specific for PS and can also bind to other anionic phospholipids such as PE, phosphatidylglycerol and phosphatidic acid45,46. Our uptake assays indicate that there is decreased transport of PS into the cells in the presence of CO2, supporting the notion that PS distribution is altered. However, it is also possible that the increased Annexin V staining observed in elevated CO2 concentrations is due to the accumulation of other anionic phospholipids on the outer leaflet of the plasma membrane. Since duramycin sensitivity does not change at elevated CO2, other anionic phospholipids would most likely be responsible for these effects. In either case, our data strongly support the notion that tolerance of host concentrations of CO2 requires a significant remodeling of outer membrane anionic phospholipid distribution; for simplicity, we will refer to PS in the remainder of the discussion with the understanding that other anionic phospholipids may also be involved.
The increased PM outer leaflet PS induced by host CO2 levels may also be linked to the increased fitness of RIM101 pathway mutants under those conditions. Brown et al. found that under neutral pH the cdc50∆ mutant has delayed proteolytic processing of Rim10147. This result indicates that increased PM outer leaflet PS (or reduced PM inner leaflet PS) negatively regulates Rim101 pathway activation. Our results suggest that both increased PM outer leaflet PS and reduced Rim101 pathway activity are involved in the adaptation of C. neoformans to host CO2 conditions. Based on the findings of Brown et al.47, these two observations may be mechanistically related.
Finally, our initial characterization of Pdr9 suggests that it is not a typical fungal ABC/PDR transporter. Overall, Pdr9 clearly affects plasma membrane lipid homeostasis based on the following PH3-PDR9 strain phenotypes: (1) increased susceptibility to the PE-targeted antifungal duramycin; (2) increased susceptibility to the sphingolipid biosynthesis inhibitor myriocin; (3) suppression of CO2-induced PM outer leaflet localized PS; and (4) reduced uptake of PE. As such, Pdr9 shows features of a PE floppase (phenotypes 1 and 4) and a PS flippase (phenotype 3). To date, all experimentally characterized flippases are P4-ATPases48. Therefore, the effect of Pdr9 on PS asymmetry is far more likely to be indirect rather than through a direct PS flippase activity.
ABC family transporters are proposed to function as lipid floppases32,33. The best characterized examples are ABCA group (not ABCG/PDR family) mammalian transporters of phosphatidylcholine and sphingomyelin (e.g., ABCA149,50,). Although the ABCG/PDR group S. cerevisiae Pdr5 has been proposed to be a PE floppase32,33 and our Pdr9 data are consistent with that function, it also affects sphingolipid biosynthesis which could contribute to its effect on PS and CO2 phenotypes. Taken together, the complex effect of elevated expression of PDR9 on lipid asymmetry suggests it does so through a combination of direct and indirect effects on lipid homeostasis.
Supporting the complex role that Pdr9 plays in lipid asymmetry and distinguishing it from a typical fungal ABC/PDR transporter is its apparent lack of PM localization. Instead, it localizes to intracellular puncta that show features of the late Golgi or endosome compartment. ABC transporters involved in lipid homeostasis have been localized to these compartments in mammalian systems51, providing some precedent for the correlation between the function and localization of Pdr9. A wide range of additional cell biological, biochemical and biophysical studies will be required to establish the molecular function of Pdr9. Regardless of its likely indirect and complex mechanism of action toward CO2 tolerance, the effect of PDR9 on multiple facets of lipid homeostasis and CO2 susceptibility has provided important insights into the mechanisms of the latter.
In summary, the ability of C. neoformans to tolerate host concentrations of CO2 is a trait that distinguishes low virulence strains from high virulence strains. Here, we have found a novel function of the TOR pathway as a critical mediator of adaptive CO2 responses in the CO2-tolerant strain H99. We also show that TOR-Ypk1 mediated remodeling of plasma membrane lipid asymmetry toward increased PS exposure is associated with CO2 tolerance.
Methods
Ethics statement
This study was performed according to the guidelines of the NIH and the University of Georgia Institutional Animal Care and Use Committee (IACUC). The animal models and procedures used have been approved by the IACUC (AUP protocol number: A2020 06-015).
Strains and growth conditions
Yeast extract-peptone-2% dextrose (YPD) and synthetic complete (SC) were prepared according to standard recipes52. RPMI 1640 without glutamine or sodium bicarbonate was buffered with 165 mM MOPS and pH adjusted to 7. Strains used in this work are described in Supplementary Data 7. C. neoformans kinase and transcription factor deletion libraries were acquired from the Fungal Genetic Stock Center (FGSC). H99 reference strain, environmental and clinical C. neoformans strains were generous gifts from A. Litvintseva, T. Mitchell, and J. Perfect. The rim13Δ, rim20Δ and rim101-GFP strains were generous gifts from J.A. Alspaugh. PCTR4:CHO1 strain was a generous gift from C. Xue. The cdc50∆ strains were a generous gift from J. Kronstad. Strains were stored at −80 °C in 20% glycerol. Frozen stocks were recovered on solid YPD medium at 30 °C for 2 days. To prepare for assays, 3 mL YPD liquid was inoculated per strain and grown overnight, 30 °C, shaking at 200 rpm.
Strain construction
For all strains generated in this work (Supplementary Data 7), CRISPR/Cas9 short-arm homology (SAH) and transient CRISPR-Cas9 coupled with electroporation (TRACE) methods were used as published53,54; CRISPR components were PCR purified and transformed into H99 via electroporation using the “Pic” setting on a Bio-Rad Micropulser.
For mNeonGreen-expressing H99, an intact pGWKS11 (a gift from James Fraser (Addgene plasmid #139418; http://n2t.net/addgene:139418; RRID: Addgene_13941855)) was transformed with Cas9 DNA and two sgRNAs targeting the Safe Haven 1 (SH1) location (generated with primers LCR075-LCR078, (oligonucleotide sequences in Supplementary Data 8)) into H99. Transformants were selected on YPD medium with 100 μg/mL nourseothricin (NAT) and fluorescent isolates were identified by flow cytometry. A single isolate was selected for use in this work by equivalent competition with wild-type H99 over 48 h at 30 °C and 37 °C in ambient or 5% CO2 conditions.
For gene deletion constructs, SAH Repair 5’ and 3’ oligos designed with a 50 bp sequence matching the flanks of the H99 target gene sequence and a 20 bp sequence matching the flanks of the NAT or hygromycin (HygB) resistance marker cassette were used to amplify a repair construct, which replaced the entire open reading frame for the target gene with the NAT or HygB cassette. For deletion of ssn3, vps15 and ypk1, primers L1 and R2 from the kinase deletion knockout collection11 were paired to amplify the resistance cassette from knockout specific genomic DNA and used as a repair construct. Repair constructs were transformed with Cas9 DNA and two gene-specific sgRNAs into H99. Transformants were selected on YPD medium with 100 μg/mL NAT or 400 μg/mL HygB and knockouts were PCR verified at the 5’ flank with LCR031 or AJ60 and gene specific “5’ KO confirmation primer”, at the 3’ flank with LCR323 or KA68 and gene specific “3’ KO confirmation primer” and for lack of the native locus with gene specific “orf confirmation” primer pairs (Supplementary Data 8).
For PH3:PDR9 strains, a NAT cassette-histone 3 promoter (CNAG_06745) fusion was synthesized commercially (Biomatik) and cloned into a pUC57 plasmid (NAT-PH3, Supplementary Data 9). LCR443 and LCR444 primers were used to amplify a repair construct targeted to replace the native promoter of PDR9 (CNAG_07799) with the NAT-PH3 fusion. Two promoter-region specific sgRNAs were generated with primers LCR451-LCR454. Repair constructs were transformed with Cas9 DNA and two sgRNAs into H99. Transformants were selected on YPD medium with 100 μg/mL NAT and integration was PCR verified at the 5’ flank with LCR031 and LCR460 and at the 3’ flank with LCR323 and LCR459.
For PDR9-mNeonGreen fusion proteins, the mNeonGreen cassette was amplified from pBHM2406 (a gift from Hiten Madhani (Addgene plasmid # 173442; http://n2t.net/addgene:173442; RRID:Addgene_17344254) with primers LCR480 and LCR481 targeting the 3’ end of CNAG_07799 and removing the stop codon. The presence of C-terminal tag was confirmed with primers LCR455 and LCR323 and mNeonGreen expression verified by fluorescence microscopy.
Competition assay
96-well plates were prepared with 200 μL YPD per well. Wells were inoculated with individual strains to be tested and the reference strain H99-mNeonGreen and grown statically overnight at 30 °C. Dilutions were performed to standardize input of unlabeled strains and H99-mNeonGreen in a 1:1 ratio, at 2 × 104 cells/mL final concentration. Co-cultures were grown at 30 °C for 24 h in ambient air or 5% CO2 before analyzing on an Attune NxT Flow Cytometer with CytKick autosampler and Attune Cytometric software. Gating strategy (Supplementary Fig. 8) was optimized for single cells. mNeonGreen positive and negative populations were identified by histogram plot with 100,000 cells counted per sample. The percent of mNeonGreen negative cells in CO2 conditions was divided by the percent of mNeonGreen negative cells in ambient conditions to determine the competitive fitness in CO2 relative to ambient conditions. Statistical significance was determined by CHI square test in Microsoft Excel. If mutant strains were deficient in ambient conditions (below 10% of the population at 24 h) a two-sided Student’s t-test was used to compare the population in CO2 to ambient.
Spot dilution assays
Cells from overnight cultures were washed twice with PBS prior to quantifying OD600. Strains were diluted to an OD600 :1, followed by ten-fold serial dilutions. 3 μL from each dilution was spotted on agar plates and grown inverted at 30 °C or 37 °C in ambient air or at 5% CO2 on RPMI 1640 with 165 mM MOPS, pH 7 plates alone or containing varying concentrations of rapamycin as indicated, with images captured at 48 h. For work with copper promoter-containing strains, 3 μL from each dilution was spotted on SC agar plates alone or containing copper sulfate or bathocuproinedisulfonic acid (BCS) as indicated and grown inverted at 37 °C with images captured at 96 hours.
RNA-Seq and differential expression analysis
Overnight cultures of indicated CO2 sensitive or tolerant strains were washed and diluted to 7.5 × 105 cells/mL, 3 mL cultured per condition in a 6-well plate in RPMI 1640 medium with 165 mM MOPS, pH 7 for 24 hours at 37 °C in ambient air or 5% CO2. Total RNA was isolated from harvested cells according to manufacturer instructions with an Invitrogen PureLink RNA mini-kit (catalog no. 12183018 A, ThermoFisher) with on-column DNAse treatment (catalog no. 12185010; ThermoFisher). Biological triplicates were harvested for each strain and condition. Total RNA (>2 μg per sample) was submitted to Azenta Life Sciences for standard RNA-Seq next-generation sequencing. The RNA samples were quantified using Qubit 2.0 Fluorometer (ThermoFisher) and RNA integrity was checked using TapeStation (Agilent Technologies). The RNA sequencing libraries were prepared using the NEB Next Ultra II RNA Library Prep Kit for Illumina using the manufacturer’s instructions (New England Biolabs). Briefly, mRNAs were initially enriched with Oligod(T) beads. Enriched mRNAs were fragmented for 15 min at 94 °C. First-strand and second-strand cDNA were subsequently synthesized. cDNA fragments were end repaired and adenylated at 3’ ends, and universal adapters were ligated to cDNA fragments, followed by index addition and library enrichment by PCR with limited cycles. The sequencing libraries were validated on the Agilent TapeStation (Agilent Technologies) and quantified by using Qubit 2.0 Fluorometer (ThermoFisher) as well as by quantitative PCR (KAPA Biosystems). The sequencing libraries were multiplexed and clustered onto a flowcell. After clustering, the flowcell was loaded onto the Illumina HiSeq instrument according to the manufacturer’s instructions. The samples were sequenced using a 2 × 150 bp Paired End configuration. Image analysis and base calling were conducted by the HiSeq Control Software. Raw sequence data (.bcl files) generated from Illumina HiSeq was converted into fastq files and de-multiplexed using Illumina bcl2fastq 2.20 software. One mismatch was allowed for index sequence identification. Paired-end Illumina sequence read files were evaluated for quality and the absence of adaptor sequence using FastQC (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/). Read files were mapped to C. neoformans reference genome H99 v48 (FungiDB) and gene transcript expression was quantified using HISAT2 and Stringtie56. Differential expression fold change, Wald test p values, and Benjamini-Hochberg adjustment for multiple comparisons were determined using DESeq2. Principle component analysis was performed on regularized log transformed gene counts to confirm the absence of batch effects57.
Protein extraction and Mpk1 western blot
Overnight cultures were diluted to an OD600:0.1 in YPD, then grown to mid-log phase (4 h) at 37 °C, shaking at 200 rpm, with samples taken each hour. Protein was extracted in extraction buffer (10 mM HEPES [pH 7.4 to 7.9], 1.5 mM MgCl2, 10 mM KCl, 1 mM dithiothreitol [DTT], 1× HALT protease and phosphatase inhibitor cocktail [catalog no. 1861280; ThermoFisher]) by five bead-beating cycles of 30 sec followed by 30 sec on ice per cycle. Debris and beads were pelleted before supernatant was recovered, and the protein concentration was quantified by Bradford assay. Protein (20 μg/lane) was loaded on a 10% SDS-PAGE gel and run at 80 V. Samples were transferred to nitrocellulose membrane for 1 h at 100 V, then the membrane was stained with Ponceau for 5 min at room temperature (RT), rinsed with distilled water and an image of loading acquired. The membrane was blocked with 5% bovine serum albumin (BSA) in tris-buffered saline with 0.1% tween 20 (TBST) for 1 h at RT, then incubated with 1:2,000 rabbit anti-p-p44 (phospho-p44/42 MAPK, catalog no. 4370; Cell Signaling) in 5% BSA/TBST overnight at 4 °C. The membrane was washed 3 times for 5 min with TBST, then incubated for 1 h at RT with 1:10,000 goat anti-rabbit horseradish peroxidase (HRP) (catalog no. STAR208P; Bio-Rad) in 5% BSA/TBST. The membrane was washed 3 times for 5 min with TBST, then developed with chemiluminescent substrate (catalog no. 1705060; BioRad) and imaged on a myECL imager (ThermoFisher).
Rim101-GFP pulldown and western blot
GFP-pulldown was performed as previously described26. Overnight cultures of H99 were washed twice in PBS and diluted to an OD600:0.2 into RPMI 1640 + 165 mM MOPS, pH 7 and grown at 37 °C in ambient air or 5% CO2 to an OD600:1. 20 mL of culture was harvested and washed twice with water. Pellets were resuspended in 1 mL of ice-cold lysis buffer (50 mM Tris-HCl pH 7.4, 150 mM NaCl, 1% NP-40, 5 mM EDTA, 1 mM phenylmethanesulfonylfluoride (PMSF), 1× HALT protease and phosphatase inhibitor cocktail [catalog no. 1861280; ThermoFisher]). Cells were lysed by bead beating, with cell suspensions combined with 0.2 mL of 0.5 mm glass beads in a FastPrep-24 (MP Biomedicals) at top speed with 4 rounds of 45 s bead beating followed by 1 min on ice. Supernatants were transferred to new tubes and glass beads washed 2 times with 0.4 mL lysis buffer. Lysates were cleared by centrifugation at 18,000 x g, 4 °C, for 10 min. GFP-TRAP resin (catalog no. gta; Chromotek) was equilibrated in lysis buffer, then 10 μL was added to the cleared lysate and rocked at 4 °C overnight. GFP-TRAP resin was washed with 1 mL lysis buffer 3 times and protein was eluted in 30 μL 4X Laemmli sample buffer (catalog no. 1610747; Bio-Rad) with 2-mercaptoethanol by boiling for 5 min. The entire sample was loaded on a 10% SDS-PAGE gel and run at 80 V. Samples were transferred to a nitrocellulose membrane for 1 h at 100 V. The membrane was blocked with 5% BSA in tris-buffered saline with tween 20 (TBST) for 1 h at RT, then incubated with 1:1,000 mouse anti-GFP clones 7.1 and 13.1 (catalog no. 118144600001; Sigma) in 5% BSA/TBST overnight at 4 °C. The membrane was washed 3 times for 5 min with TBST, then incubated for 1 h at RT with 1:10,000 goat anti-mouse horseradish peroxidase (HRP) (catalog no. STAR207P; Bio-Rad) in 5% BSA/TBST. The membrane was washed 3 times for 5 min with TBST, then developed with chemiluminescent substrate (catalog no. 1705060; Bio-Rad) and imaged on a myECL imager (ThermoFisher).
Nanostring and qRT-PCR
Overnight cultures of indicated strains were washed and diluted to 7.5 × 105 cells/mL, 3 mL cultured per condition in a 6-well plate in RPMI 1640 medium with 165 mM MOPS, pH 7 for 24 h at 37 °C in ambient air or 5% CO2. Total RNA was isolated from harvested cells according to manufacturer instructions with an Invitrogen PureLink RNA mini-kit with on-column DNAse treatment. Biological triplicates were harvested for each strain and condition. For Nanostring, 100 ng of RNA was hybridized to a custom Nanostring probe set (Supplementary Data 6) at 65 °C for 18 hours and quantified on a Nanostring Sprint nCounter. RCC files were imported into nSolver software to extract counts and evaluate quality control metrics. The mean of the negative control probes value plus 2 times the SD was subtracted from counts to provide a background threshold. Values below background were set to 1. After background subtraction, RNA counts were normalized by total count to the highest counts of any sample in comparison. Normalized counts were used to generate a heat map in Morpheus (https://software.broadinstitute.org/morpheus), hierarchically clustered by strain/condition and gene ID with one minus Pearson correlation, average linkage. For qRT-PCR, 1000 ng RNA was used for cDNA synthesis with iScript cDNA synthesis kit (catalog no. 1708891; Bio-Rad) according to manufacturer’s instructions, then cDNA was diluted 1:5 with ddH20. qRT analysis was performed in 20 μL reactions using 2 μL of dilute cDNA per reaction with iQ SYBR Green Supermix (catalog no. 1708882; Bio-Rad). pdr9 gene expression was measured with a Bio-Rad CFX Connect Real Time PCR Detection System (primers LCR461 and LCR462). Gene expression was normalized to actin expression (primers LCR225 and LCR226). Measurements were performed in technical duplicates with biological triplicates.
Disk diffusion assays
Cells from overnight cultures were washed twice with PBS prior to quantifying concentration on a Countess II FL automated cell counter. Cells were diluted to 1 × 107 cells/mL to plate on RPMI. A sterile q-tip was saturated in the cell solution before streaking an entire plate to generate a lawn. When plates had dried, a sterile filter paper dot was placed in the center of the plate for a single treatment per plate or evenly spaced for multiple treatments. Treatments were added in 20 μL volumes at indicated final concentrations per disk: myriocin 8 μg/disk, fluconazole 40 μg/disk, aureobasidin 100 μg/disk, duramycin 200 μg/disk.
Annexin V staining
Overnight cultures of Cryptococcus strains were washed in PBS and used to inoculate 25 mL cultures of RPMI 1640 medium with 165 mM MOPS, pH 7. Identical cultures for each strain were placed at 37 °C in ambient air or 5% CO2 shaking at 200 rpm overnight. The following day 1–2 mL of the culture were collected and spun down to be washed with 1 mL of Annexin V binding buffer (10 mM HEPES-NaOH, pH 7.4, 140 mM NaCl, and 2.5 mM CaCl2). The cell pellets were resuspended in 25 μl of Annexin V binding buffer and 1.5 μl of FITC-conjugated annexin V (catalog no. A13201, Life Technologies, Inc.) was added and incubated at RT for 15 min. The cells were pelleted and washed with 1 mL of Annexin V binding buffer. A final resuspension into 50 μl of Annexin V binding buffer was performed and cells were placed on a microscope slide with a coverslip. Confocal images (multiphoton laser scanning microscope (SP8; Leica Microsystem) LAS X software) of representative fields were taken for bright field and FITC fluorescence. All images were acquired using 488-nm excitation for the FITC signal with a 100X oil immersive objective lens. Image brightness and contrast were adjusted equivalently across samples in ImageJ for publication. Z-stacks were condensed into max projections and mean fluorescence intensity was calculated for a minimum of 100 cells for each condition using ImageJ.
Fluorescence microscopy
Overnight cultures were diluted to an OD600:0.1 and grown for 4 h to mid-log phase in YPD. Cells were washed, incubated with NucBlue Live ReadyProbes Reagent for 30 minutes according to manufacturer instructions (catalog no. R37605, ThermoFisher) and imaged on a confocal microscope (multiphoton laser scanning microscope (SP8; Leica Microsystem) LAS X software). All images were acquired using 488-nm excitation for mNeonGreen signal and the UV-laser for Hoescht signal with a 100X oil immersive objective lens. Image brightness and contrast were adjusted equivalently within each channel in ImageJ for publication.
NBD-lipid uptake assays
Uptake of NBD labeled phosphatidylserine or phosphatidylethanolamine was examined based on methods described in Viet et al.58. Briefly, overnight cultures were diluted to an OD600:0.2 in RPMI 1640 medium with 165 mM MOPS, pH 7 and grown for 18 hours at 30 °C in ambient or 5% CO2 shaking conditions. Cells were harvested, washed and resuspended to OD600:8 in PBS. 250 μL of cells were aliquoted into a round-bottom 96-well plate for each condition to be tested in triplicate, then incubated at 30 °C for 10 minutes. 1.5 μL of 10 mM NBD-PS (catalog no. 810192P, Avanti Polar Lipids) or 10 mM NBD-PE (catalog no. 810151P, Avanti Polar Lipids) was added to cells and incubated for 30 minutes at 30 °C. After incubation, cells were washed three times with PBS + 4% BSA to extract free NBD-lipids and examined by flow cytometry on an Attune NxT Flow Cytometer with CytKick autosampler and Attune Cytometric software. Gating strategy was optimized for single cells. 100,000 single cells were acquired per sample. Mean fluorescence intensity was determined for each sample.
Melaninization assay
Overnight cultures of H99 or PH3:PDR9 were washed twice in sterile PBS before resuspending in PBS and quantifying OD600. Strains were diluted to an OD600:1, followed by ten-fold serial dilutions. Three μL from each dilution was spotted onto solid L-DOPA media (7.6 mM L-asparagine monohydrate, 5.6 mM glucose, 22 mM KH2PO4, 1 mM MgSO47H20, 0.5 mM L-DOPA, 0.3 mM thiamine-HCl, 20 nM biotin, 2% agar), grown inverted at 30 °C and imaged at indicated time points.
Capsule production
Overnight cultures of H99 or PH3:PDR9 were washed twice in sterile PBS before resuspending in PBS and quantifying concentration on a Countess II FL automated cell counter. Cells were diluted to 1.25×105 cells/mL in RPMI 1640 with 165 mM MOPS, pH 7 and grown for 24 h at 37 °C in ambient air or 5% CO2, then prepared for microscopy by concentrating cells and counterstaining with India ink. Images were captured using a Rebel microscope at 60X magnification. At least 50 cells were quantified per condition and processed in ImageJ software to measure capsule.
Animal experiments
Six- to eight-week-old female CD1 mice were obtained from Charles River Laboratory. Animals were housed with 5 mice per cage, under a 12 hr dark/12 hr light cycle at ambient temperature (20–24 °C), 30–70% humidity. Prior to infection, overnight cultures of C. neoformans strains were washed with sterile saline three times and resuspended in saline. For the coinfection experiment, five CD1 mice per group were infected intranasally with a 1:1 ratio of WT and mutant cells (5 × 104 cells per mouse). The ratio of WT and mutant cells was confirmed by plating serial dilutions of the inoculums on YNB and YNB with 100 μg/mL of NAT to measure CFU. At DPI 14, the mice were sacrificed, and the lung, brain, and left kidney, were dissected. For fungal burden quantifications, dissected organs were homogenized in 2 mL of cold sterile PBS using an IKA-T18 homogenizer. Homogenized organs were serially diluted, plated onto YNB and YNB with 100 μg/mL of NAT, and incubated at 30 °C for two days before counting CFUs.
For the survival study, ten CD1 mice per group were infected intranasally with 1 × 104 fungal cells. All mice were used for the survival study, and fungal burden was examined for five mice at the time of termination or at DPI 86. Fungal burden quantification was performed as described above.
Phagocytosis assay
J774A.1 murine macrophage cells (catalog no. CB_91051511, ATCC) were seeded into a T25 flask at 40% confluence and incubated at 37 °C with 5% CO2 for 48 h until 80 to 90% confluent, approximately 7.8 × 106 cells/flask. Overnight cultures of H99 or PH3:PDR9 were washed twice with PBS, before quantifying concentration on a Countess II FL automated cell counter. Cells were diluted to 3.12 × 108 cells/mL in PBS. Each strain was opsonized with mAb 18b7 (a generous gift from A. Casadevall59) at 2 μg/mL final concentration at 37 °C for 1 hour with rotation. After opsonization, 250 μL of cells were added to each flask of J774 cells for an MOI of 10. Triplicate flasks were prepared for each strain, then incubated for 2 h and 40 min and washed to remove non-phagocytosed cells. Macrophage cells were lysed with ice-cold 0.1% triton X in PBS for 10 min while rocking. For plating, cells were serially diluted 10-fold down to 1:1000. Four dilutions (full down to 1:1000) were plated on YPD with 3 × 10 μL spots per dilution and grown for 48 h at 30 °C before counting CFU.
Statistical analysis
Data were graphed and analyzed in GraphPad Prism 9 software or Microsoft Excel. Statistical analysis and graph descriptions, including P values are provided in figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-Seq data discussed in this publication have been deposited in NCBI’s Gene Expression Omnibus60 and are accessible through GEO Series accession number GSE 241788. RNA-Seq data was analyzed with C. neoformans H99 genome v48 available on FungiDB. The data supporting the findings of this work are provided within the article and supplementary files. Source data are provided with this paper. Materials generated in this study will be made available upon request. Source data are provided with this paper.
References
Rajasingham, R. et al. Global burden of disease of HIV-associated cryptococcal meningitis: an updated analysis. Lancet Infect. Dis. 17, 873–881 (2017).
May, R. C., Stone, N. R., Wiesner, D. L., Bicanic, T. & Nielsen, K. Cryptococcus: from environmental saprophyte to global pathogen. Nat. Rev. Microbiol. 14, 106–17 (2016).
Litvintseva, A. P. & Mitchell, T. G. Most environmental isolates of Cryptococcus neoformans var. grubii (Serotype A) are not lethal for mice. Infect. Immun. 77, 3188–95 (2009).
Gerstein, A. C. et al. Identification of pathogen genomic differences that impact human immune response and disease during Cryptococcus neoformans infection. mBio 10, e01440–19 (2019).
Sabiiti, W. et al. Efficient phagocytosis and laccase activity affect outcome of HIV-associated cryptococcosis. J. Clin. Invest. 124, 2000–8 (2014).
Montoya, M. C., Magwene, P. M. & Perfect, J. R. Associations between Cryptococcus genotypes, phenotypes, and clinical parameters of human disease: a review. J. Fungi (Basel) 7, 260 (2021).
Kronstad, J., Jung, W. H. & Hu, G. Beyond the big three: systematic analysis of virulence factors in Cryptococcus neoformans. Cell Host Microbe 4, 308–10 (2008).
Sephton-Clark, P. et al. Genomic variation across a clinical Cryptococcus population linked to disease outcome. mBio 13, e0262622 (2022).
Mukaremera, L. et al. The mouse inhalational model of Cryptococcus neoformans infection recapitulates strain virulence in humans and shows that closely related strain can possess differential virulence. Infect. Immun. 87, e00046–19 (2019).
Krysan, D. J. et al. Host carbon dioxide concentration is an independent stress for Cryptococcus neoformans that affects virulence and antifungal susceptibility. mBio 10, e01410–19 (2019).
Lee, K. T. et al. Systematic functional analysis of kinases in the fungal pathogen Cryptococcus neoformans. Nat. Commun. 7, 12766 (2016).
Jung, K. W. et al. Systematic functional profiling of transcription factors networks in Cryptococcus neoformans. Nat. Commun. 6, 6757 (2015).
Chadwick, B. J. et al. The RAM signaling pathway links morphology, thermotolerance, and CO2 tolerance in the global fungal pathogen Cryptococcus neoformans. Elife 11, e82563 (2022).
So, Y. S., Lee, D. G., Idnurm, A., Ianiri, G. & Bahn, Y. S. The TOR pathway plays pleiotropic role is growth and stress responses of the fungal pathogen Cryptococcus neoformans. 2019. Genetics 212, 1241–1258 (2019).
Alcaide-Gavilán, M., Lucena, R., Banuelos, S. & Kellogg, D. R. Conserved Ark1-related kinases function in a TORC2 signaling network. Mol. Biol. Cell 31, 2057–2069 (2020).
Leskoske, K. L. et al. Phosphorylation by the stress-activated MAPK Slt2 down-regulates the yeast TOR complex 2. Genes Dev. 32, 1576–1590 (2018).
Idnurm, A. & Heitman, J. Light controls growth and development via a conserved pathway in the fungal kingdom. PLoS Biol. 3, e95 (2005).
Jang, E. H., Kim, J. S., Yu, S. R. & Bahn, Y. S. Unraveling capsule biosynthesis and signaling networks in Cryptococcus neoformans. Microbiol. Spectr. 10, e0286622 (2022).
Chun, C. D., Brown, J. C. S. & Madhani, H. D. A major role for capsule-independent phagocytosis-inhibitory mechanisms in mammalian infection by Cryptococcus neoformans. Cell Host Microbe 9, 243–251 (2011).
O’Meara, T. R., Holmer, S. M., Selvig, K., Dietrich, F. & Alspaugh, J. A. Cryptococcus neoformans Rim101 is associated with cell wall remodeling and evasion of the host immune response. mBio 4, e00522–12 (2013).
O’Meara, T. R. et al. Interaction of Cryptococcus neoformans Rim101 and protein kinase A regulates capsule. PLoS Pathog. 6, e1000776 (2010).
Heitman, J. On the discovery of TOR as the target of rapamycin. PLoS Pathog. 11, e1005245 (2015).
Gerik, K. J. et al. Cell wall integrity is dependent on the PKC1 signal transduction pathway in Cryptococcus neoformans. Mol. Microbiol 58, 393–408 (2005).
Kraus, P. R., Fox, D. S., Cox, G. M. & Heitman, J. The Cryptococcus neoformans MAP kinase Mpk1 regulates cell integrity responses to antifungal drugs and loss of calcineurin function. Mol. Microbiol 48, 1377–87 (2003).
Yang, D. H. et al. Rewiring of signaling networks modulating thermotolerance in the human pathogen Cryptococcus neoformans. Genetics 205, 201–219 (2017).
Ost, K. S., O’Meara, T. R., Huda, N., Esher, S. K. & Alspaugh, J. A. The Cryptococcus neoformans alkaline response pathway: identification of a novel rim pathway activator. PLoS Genet. 11, e1005159 (2015).
Luberto, C. et al. Roles for inositol-phosphoryl ceramide synthase 1 (IPC1) in pathogenesis of C. neoformans. Genes Dev. 15, 201–12 (2001).
Winski, C. J., Qian, Y., Mobashery, S. & Santiago-Tirado, F. H. An atypical ABC transporter is involved in antifungal resistance and host interactions in the pathogenic fungus Cryptococcus neoformans. mBio 13, e0153922 (2022).
Wu, A., Wojtowicz, K., Savary, S., Hamon, Y. & Trombik, T. Do ABC transporters regulate plasma membrane organization. Cell Mol. Biol. Lett. 25, 37 (2020).
Chang, M., Sionov, E., Khanal Lamichhane, A., Kwon-Chung, K. J. & Chang, Y. C. Roles of three Cryptococcus neoformans and Cryptococcus gattii efflux pump-coding genes in response to drug treatment. Antimicrob. Agents Chemother. 62, e01751–17 (2018).
Roelants, F. M., Baltz, A. G., Trott, A. E., Fereres, S. & Thorner, J. A protein kinase network regulates the function of aminophospholipid flippases. Proc. Natl Acad. Sci. USA 107, 34–9 (2010).
Shin, H. W. & Takatsu, H. Phosphatidylserine exposure in living cells. Crit. Rev. Biochem Mol. Biol. 55, 166–178 (2020).
Kumari, S., Kumar, M., Gaur, N. A. & Prasad, R. Multiple roles of ABC transporters in yeast. Fungal Genet. Biol. 150, 103550 (2021).
Huang, W. et al. Lipid flippase subunit Cdc50 mediates drug resistance and virulence in Cryptococcus neoformans. mBio 7, e00478–16 (2016).
Hu, G. et al. A P4-ATPase subunit of the Cdc50 family play a role in iron acquisition and virulence in Cryptococcus neoformans. Cell. Microbiol. 19, e12718 (2016).
Konarzewska, P. et al. Phosphatidylserine synthesis is essential for viability of the human fungal pathogen Cryptococcus neoformans. J. Biol. Chem. 294, 2329–2339 (2019).
Johnston, S. A. & May, R. C. Cryptococcus interactions with macrophages: evasion and manipulation of the phagosome by a fungal pathogen. Cell Microbiol. 15, 403–11 (2013).
Bahn, Y. S. & Mühlschlegel, F. A. CO2 sensing in fungi and beyond. Curr. Opin. Microbiol. 9, 572–8 (2006).
Thorner, J. TOR complex 2 is a master regulator of plasma membrane homeostasis. Biochem. J. 479, 1917–1940 (2022).
Lee, H., Khanal Lamichhane, A., Garraffo, H. M., Kwon-Chung, K. J. & Chang, Y. C. Involvement of PDK1, PKC, and TOR signaling pathways in basal fluconazole tolerance in Cryptococcus neoformans. Mol. Microbiol. 84, 130–46 (2012).
Singh, A. et al. Methylation of glycosylated sphingolipid modulates membrane lipid topography and pathogenicity of Cryptococcus neoformans. Cell Microbiol. 14, 500–16 (2012).
Muir, A., Ramachandran, S., Roelants, F. M., Timmons, G. & Thorner, J. TORC2-dependent protein kinase Ypk1 phosphorylates ceramide synthase to stimulate synthesis of complex sphingolipids. Elife 3, e03779 (2014).
Hatakeyama, R., Kono, K. & Yoshida, S. Ypk1 and Ypk2 maintain Rho1 at the plasma membrane by flippase-dependent lipid remodeling after membrane stress. J. Cell Sci. 130, 1169–1178 (2017).
Yeung, T. et al. Contribution of phosphatidylserine to membrane surface charge and protein targeting during phagosome maturation. J. Cell Biol. 185, 917–928 (2009).
Weingärtner, A. et al. Leishmania promastigotes lack phosphatidyl serine but bind Annexin V upon permeabilization or miltefosine treatment. PLoS ONE 7, e42070 (2012).
Kolbeck, S., Kienberger, Kleigrewe, K., Hilgarth, M. & Vogel, R. F. Effect of high levels of CO2 and O2 on membrane fatty acid profile and membrane physiology of meat spoilage bacteria. Eur. Food Res Tech. 247, 999–1011 (2021).
Brown, H. E. et al. Identifying a novel connection between the plasma membrane and pH-sensing. Mol. Microbiol. 109, 474–493 (2018).
Hankins, H. M., Baldridge, R. D., Xu, P. & Graham, T. R. Roles of flippases, scramblases, and transfer proteins in phosphatidylserine subcellular distribution. Traffic 16, 35–47 (2015).
Clarke, R. J., Hossain, K. R. & Cao, K. Physiological roles of transverse lipid asymmetry of animal membranes. Biochim Biophys. Acta Biomembr. 1862, 183382 (2020).
Hamon, Y. et al. ABC1 promotes engulfment of apoptotic cells and transbilayer redistribution of phosphatidylserine. Nat. Cell Biol. 2, 399–406 (2000).
Bevers, E. M. & Williamson, P. L. Getting to the outer leaflet: physiology of phosphatidylserine exposure at the plasma membrane. Physiol. Rev. 96, 605–645 (2016).
Sherman, F. Getting started with yeast. Methods Enzymol. 350, 3–41 (2002).
Fan, Y. & Lin, X. Multiple applications of a transient CRISPR-Cas9 coupled with electroporation (TRACE) system in the Cryptococcus neoformans species complex. Genetics 208, 1357–1372 (2018).
Huang, M. Y. et al. Short homology-directed repair using optimized Cas9 in the pathogen Cryptococcus neoformans enables rapid gene deletion and tagging. Genetics 220, iyab180 (2022).
Spencer, G. W. K. et al. Broadening the spectrum of fluorescent protein tools for use in the encapsulated human fungal pathogen Cryptococcus neoformans. Fungal Genet. Biol. 138, 103365 (2020).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Love, M. I., Anders, S., Kim, V. & Huber, W. RNA-Seq workflow: gene-level exploratory analysis and differential expression. F1000Res 4, 1070 (2015).
Veit, S., Laerbusch, S., Lopez-Marques, R. L. & Pomorski, T. G. Functional analysis of the P-type ATPases Apt2-4 from Cryptococcus neoformans by heterologous expression in Saccharomyces cerevisiae. J. Fungi 9, 202 (2023).
Mukherjee, J., Casadevall, A. & Scharff, M. D. Molecular characterization of the humoral responses to Cryptococcus neoformans infection and glucuronoxylomannan-tetanus toxoid conjugate immunization. J. Exp. Med. 177, 1105–1116 (1993).
Edgar, R., Domrachev, M. & Lash, A. E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–10 (2002).
Acknowledgements
This work was supported by the following grants from NIAID: 5R01AI147541 (D.J.K. and X.L.) and T32AI007511 (A.J.J.). The authors thank Andy Alspaugh and Connie Nichols (Duke) for sharing protocols and strains for experiments with Rim101. We thank Chaoyang Xue (Rutgers) for providing the PCTR4-CHO1 strain and Jim Kronstad (British Columbia) for providing the cdc50∆ strains. We also thank Scott Moye-Rowley (Iowa) for helpful discussions regarding ABC transporter biology and Rohan Wakade for assistance with fluorescence microscopy.
Author information
Authors and Affiliations
Contributions
L.C.R.: experimentation, data analysis, figure construction, writing of draft and editing manuscript. A.J.J.: experimentation, data analysis, figure construction. B.J.C.: experimentation, data analysis. M.A.S.: supervision, data analysis. X.L.: conception, data analysis, supervision, funding, editing of manuscript. D.J.K.: conception, data analysis, supervision, funding, writing and editing of manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interest.
Peer review
Peer review information
Nature Communications thanks Thomas Günther Pomorski and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ristow, L.C., Jezewski, A.J., Chadwick, B.J. et al. Cryptococcus neoformans adapts to the host environment through TOR-mediated remodeling of phospholipid asymmetry. Nat Commun 14, 6587 (2023). https://doi.org/10.1038/s41467-023-42318-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-023-42318-y
- Springer Nature Limited