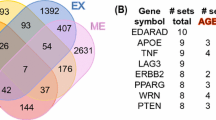
Abstract
Aging is a complex biological process that is far from being completely understood. Analyzing transcriptional differences across age might help uncover genetic bases of aging. In this study, 1573 differentially expressed genes, related to chronological age, from the Genotype-Tissue Expression (GTEx) project, were categorized as upregulated age-associated genes (UAGs) and downregulated age-associated genes (DAGs). Characteristics in evolution, expression, function and molecular networks were comprehensively described and compared for UAGs, DAGs and other genes. Analyses revealed that UAGs are more clustered, more quickly evolving, more tissue specific and have accumulated more single-nucleotide polymorphisms (SNPs) and disease genes than DAGs. DAGs were found with a lower evolutionary rate, higher expression level, greater homologous gene number, smaller phyletic age and earlier expression in body development. UAGs are more likely to be located in the extracellular region and to occur in both immune-relevant processes and cancer-related pathways. By contrast, DAGs are more likely to be located intracellularly and to be enriched in catabolic and metabolic processes. Moreover, DAGs are also critical in a protein–protein interaction (PPI) network, whereas UAGs have more influence on a signaling network. This study highlights characteristics of the aging transcriptional landscape in a healthy population, which may benefit future studies on the aging process and provide a broader horizon for age-dependent precision medicine.
Similar content being viewed by others
Introduction
Aging is considered to be a dominating risk factor for many fatal diseases, including cancer, cardiovascular diseases and neurodegenerative diseases1,2,3,4,5,6. A large number of studies have found that aging is associated with telomere attrition, mitochondrial dysfunction, DNA damage, immune system impairment etc., and can be inhibited by calorie restriction7,8,9,10,11. However, the detailed mechanisms involved in aging remain unclear. In recent years, rapidly developing high-throughput omics have provided a broader insight, with the identification of a number of longevity-relevant loci based on genome-wide association studies(GWAS) and epigenome analyses12,13. As previous studies have shown, aging is distinct at molecular, cellular and tissue levels14, which indicates that the relatively dynamic transcriptome might also provide important clues for the study of aging. A large number of human age-associated genes have been identified in previous transcriptomic studies, based on specific tissues like muscle, blood, skin, adipose, brain etc., and have been compared across tissues15,16,17,18,19,20,21,22,23,24. The identified age-associated genes in these studies vary from each other, which could be partly due to differences in the health condition of donors, sample size, sample quality, tissue, platform and the method of identifying age-associated genes. Also, the divergent results may be accounted for by the generally low repeatability of microarray data that most of the analyses were based on. In addition, given that aging is likely to confound with other factors, profiling the aging factor alone can be difficult, which may lead to biases in screening age-associated genes. Among these studies, the age-associated gene set from the Genotype-Tissue Expression (GTEx) project25 is of high quality. This set of aging-associated genes was screened out by using a regression model on large-sample RNA-Seq data, collected from >40 tissues from hundreds of healthy individuals; whereas sex, race, and tissue were controlled as a covariate to avoid biases24. However, characteristics of these genes still remain unexplored. A comprehensive analysis of these genes might help to improve the understanding of aging process and provide valuable clues for strategies in anti-aging interventions. Here, we comprehensively explored characteristics of the age-associated genes derived from GTEx. The results revealed that differences and interactions exist in evolution, expression, function, associated diseases and molecular network between the upregulated age-associated genes (UAGs) and downregulated age-associated genes (DAGs).
Result
The human transcriptional age-associated genes from GTEx
To characterize the age-associated genes, we extracted a protein-coding transcriptional age-associated gene set from a GTEx transcriptional analysis (see Materials and methods section). The gene set analysis procedure is summarized in Fig. 1a. Ultimately, we extracted 710 UAGs and 863 DAGs across tissues. The age-associated genes account for 7.71% of the protein-coding genes (Fig. 1b and Supplementary Table 1). Genomic information of UAGs and DAGs is shown in Supplementary Tables 2 and 3, respectively.
a The workflow for characterizing age-associated genes. b A pie chart showing the proportion of age-associated genes within all protein-coding genes. c Chromosomal distribution of the 1573 age-associated genes. d Circos54 plot displaying the age-associated genes. e Genomic distances between age-associated genes. UDs distances of each pair of UAGs on the same chromosome, DDs distances of each pair of DAGs on the same chromosome, UDDs distances of any pair of genes on the same chromosome between UAGs and DAGs groups. **P < 0.01, ***P < 0.001 from Wilcoxon test
Distribution of the age-associated genes across chromosomes
To characterize the age-associated genes at the genomic level, we investigated the chromosomal distribution pattern of the age-associated genes. Results showed age-associated genes are widespread across chromosomes. Over 97% of the age-associated genes are located on autosomes, where the proportion of DAGs is higher than that of UAGs, with the exception of five autosomes. Although no age-associated genes were found on the Y chromosome, possibly due to the adjustment for sex in the regression model used to discover age-associated genes, some UAGs (2.86%) and a smaller number of DAGs (1.83%) were observed on the X chromosome (Figs. 1c, d).
Genes with similar functions are likely to locate adjacently on chromosomes26. To study whether the age-associated genes are close to each other on chromosomes, we calculated the genomic distances of each pair of genes on the same chromosome, within one age-associated gene group only (UDs, DDs), and between groups (UDDs). Results showed these three distances are significantly different from each other (median values of UDs, DDs and UDDs are 3.52e + 7, 4.67e + 7 and 4.42e + 7, respectively). Of the three distances, UDs possess the smallest value compared with DDs (P = 7.52e-56, Wilcoxon test) and UDDs (P = 1.17e-57, Wilcoxon test). Surprisingly, UDDs show smaller value than DDs (P = 0.0038, Wilcoxon test) (Fig. 1e). These results suggest that on the same chromosome, UAGs are more clustered than DAGs and non-age-associated genes (NAGs); DAGs on the other hand are relatively dissociated, whereas some of them tend to be more adjacent to UAGs than other DAGs.
Evolutionary characteristics and expression profile of age-associated genes
To gain a better insight into the evolutionary background of genes, the evolutionary characteristics of age-associated genes were investigated in three aspects: evolutionary rate, homologous gene number and phyletic age. Results showed that DAGs have the lowest dN/dS ratio (P = 1.43e-50, Wilcoxon test), the highest homologous gene number (P = 5.73e-55 Wilcoxon test) and have a relatively earlier origin in phyletic age compared with UAGs (P = 7.37e-24, chi-squared test). The corresponding values for NAGs are P = 5.02e-45, P = 3.39e-60 and P = 4.74e-32. Comparatively, UAGs evolves most rapidly, with the smallest homologous gene number, and originated later than DAGs but earlier than NAGs (Figs. 2a–c; Table 1). These results revealed that DAGs are more conserved in evolution, suggesting the DAGs are more crucial to fundamental functions in humans. However, UAGs are less stable and have a shorter history, which implies they are likely to function in more specific and advanced functions.
Box plots and bar graphs show UAGs, DAGs and NAGs compared in the following characteristics: a dN/dS ratio of each human–mouse homolog gene. b Number of homologous genes. c Origin in phyletic evolution. d Average expression level across tissues. e Tissue specificity of gene expression. f Earliest expression stage. *P < 0.05, ***P < 0.001 from Wilcoxon test
Essential genes are those that function in basic biological processes and must exist for an organism to survive27. As expected, DAGs are significantly more enriched in human essential genes (15.3% of DAG genes), compared with NAGs (11.8%; P = 0.0035, chi-squared test) (Table 1). UAGs (11.8%) and NAGs (11.9%) have no significant difference.
The expression profile also provides important characteristics for a gene and often gives valuable clues to potential gene function. Therefore, we explored the expression characteristics of age-associated genes from three aspects: average expression level across tissue, tissue specificity and the earliest expression stage. We found DAGs have the highest expression level and lowest tissue specificity compared with UAGs and NAGs, suggesting DAGs have a more global function, providing basic support to the whole body. UAGs are significantly higher in expression level and tissue specificity than NAGs, which suggests UAGs are also important in function, but more tissue specific (Figs. 2d, e; Table 1). As for the earliest expression stage, although the stage of embryoid body is predominant for expression in all three groups, DAGs are far more expressed than UAGs and NAGs at this stage. UAGs on the other hand outnumber DAGs and NAGs in the following stages of blastocyst and fetus (Fig. 2f; Table 1). Altogether the age-associated genes tend to express in earlier stages than other genes, underlining they are likely to play important roles in early development.
Functional annotation for age-associated genes
Functional enrichment analyses were carried out to explore the functions of UAGs and DAGs. First, we investigated the subcellular locations of UAGs and DAGs to see if they are located differently. Genes located in nucleus (NU), cytoplasm (CY), membrane (ME) and extracellular region (ER) were filtered from gene ontology (GO) terms and the age-associated genes were mapped into these cellular locations. The number of UAGs and DAGs in these four locations were, 78 versus 118 (10.99% versus 13.67%, NU), 163 versus 231 (22.96% versus 26.77%, CY), 67 versus 123 (9.44% versus 14.25%, ME) and 85 versus 15 (11.97% versus 1.74%, ER), respectively, see Fig. 3a. The distributions of UAGs and DAGs in subcellular locations are significantly different (P = 1.971e-16, chi-squared test). Noted that there is a striking difference in the fraction of UAGs and DAGs in ER, suggesting the UAG group includes far more secretory protein genes.
a The number of UAGs and DAGs in four subcellular locations is shown. ER extracellular region, ME membrane, CY cytoplasm, NU nucleus. UAGs and DAGs are presented in red and blue, respectively. b KEGG enrichment analysis for UAGs with the threshold of P < 0.05. c KEGG enrichment analysis for DAGs with the threshold of P < 0.05
Next, we focused on the biological processes and pathways. GO biological process analysis was carried out for 626 UAGs and 772 DAGs mapped in DAVID. A total of 759 and 549 enriched terms were filtered for UAGs and DAGs with the threshold of P-value < 0.05. Most terms enriched with UAGs are relevant to the immune system, such as processes related to immune cells, ‘response to stimulus’ (54.8%, P = 1.10e-10) or ‘defense response’ (15.10%, P = 1.20e-10). In all, 178 UAGs are most significantly enriched in the term ‘immune system process’ (25.1%, P = 1.40e-19), whereas 292 are enriched in the term ‘cell communication’, with a fairly high number (41.1%, P = 1.10e-6) (Supplementary Fig. 1). Apart from processes related to cell migration, the data showed that cell proliferation, cell adhesion and cell differentiation are also highly enriched with UAGs, hinting that UAGs are likely to be involved in processes relevant to tumorigenesis.
As for DAGs, terms related to catabolic processes are enriched most significantly (with 31 terms in total); whereas metabolic process-relevant terms appear in most genes (with 80 terms in total). In total, 87 DAGs (10.1%, P = 5.20e-19) are most significantly enriched in the term ‘proteolysis involved in cellular protein catabolic process’, whereas 571 genes are enriched in the term ‘metabolic process’ (66.2%, P = 6.40e-7). In addition, 37 terms are relevant to mitochondria, and 19 terms are associated with cell cycle (111 genes are enriched in term ‘cell cycle’, P = 6.30e-5). Moreover, 11 terms with the keyword ‘immune’ were also observed in DAGs enriched processes, 5 terms have the keyword ‘innate immune’ and 37 DAGs are enriched in the term ‘positive regulation of innate immune response’ (3.8%, P = 4.80e-6) (Supplementary Fig. 2).
Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis revealed 26 and 32 enriched terms (with the threshold of P < 0.05) for 257 UAGs and 355 DAGs (Figs. 3b, c). Notably, apart from being enriched in pathways related to inflammation and infection, several pathways enriched with UAGs are oncogenic pathways. DAGs are enriched in foundational metabolic pathways, which is similar with the result of the enrichment analysis for biological processes. In addition, there are pathways related to neurodegenerative diseases enriched with DAGs, as a relevant study mentions28. Considering DAGs are downregulated across age, this result indicates that healthy old individuals may suffer from a functional decline in relevant pathways, but this result may not serve as direct evidence for the relation between healthy aging and these diseases.
UAGs have higher SNP density and are more enriched in disease genes
In the human genome, single-nucleotide polymorphism (SNP) is the most common genetic variation. To investigate the difference between SNP in age-associated gene and other genes, we calculated the SNP density (the number of SNPs in a gene divided by the gene’s length) for UAGs, DAGs and NAGs, and compared their distributions across chromosomes (Fig. 4a). As expected, DAGs bear the lowest SNP density compared with UAGs and NAGs (P = 0.0012 and 0.0011, respectively, Wilcoxon test, the median value of UAGs, DAGs and NAGs are 0.037, 0.012 and 0.025, respectively; Fig. 4b), which is consistent with the evolutionary characteristic analysis that DAGs tend to be more conserved.
a Distribution of average SNP density value for UAGs, DAGs and NAGs across chromosomes. The standard error of mean (SEM) in every chromosome is presented by error bars. b Cumulative distribution plot for the average SNP density across chromosomes. c Distribution for the fraction of disease-related SNP (dSNP) in the total number of SNPs across chromosomes among UAGs, DAGs and NAGs. d Cumulative distribution plot for the proportion of dSNP compared with total SNP. e The proportion of disease-related genes contained in UAGs, DAGs and NAGs across chromosomes. UAGs presented a much closer relationship with diseases. dGenes: disease-related genes; U, D, N: short for UAG, DAG, NAG, respectively. f Cumulative distribution plot for the proportion of disease-related genes. *P < 0.05, **P < 0.01, ***P < 0.001 from Wilcoxon test
To further study whether UAGs have more disease-relevant SNPs (dSNPs) compared with other groups, we mapped the dSNPs to each gene, and calculated the fraction of the number of dSNPs, versus total number of SNPs, in each gene group (Fig. 4c). Surprisingly, no significant difference was found between the age-associated genes (data not shown), whereas NAGs have a higher dSNP fraction than UAGs and DAGs (P = 0.011 and 0.0045, respectively, Wilcoxon test, the median value for the fraction of dSNP to total SNP for UAGs, DAGs and NAGs is 0.31%, 0.29% and 0.52%, respectively; Fig. 4d). Given that the individuals contributing age-associated genes are healthy, it is not likely that they have many fatal mutations as reflected in the UAGs and DAGs.
In addition, 7291 disease genes from DisGeNET database were used to analyze the enrichment of the three gene groups to see whether there are differences between groups (Fig. 4e). Results showed that UAGs are enriched the most in disease genes, with the median proportion of 47.69% across chromosomes, compared with DAGs (38.36%, P = 0.0031, Wilcoxon test) and NAGs (35.82%, P = 1.02e-4, Wilcoxon test). No significant difference was observed between DAGs and NAGs (Fig. 4f). Disease genes in UAGs and DAGs are listed in Supplementary Table 4.
Moreover, for 495 UAGs, 93 items are significantly enriched in Genetic Association Database (GAD) diseases (with threshold P < 0.05). In all, 134 UAGs are most significantly enriched in term ‘Type 2 Diabetes|edema|rosiglitazone’ (18.9%, P = 9.30e-9), whereas ‘hypertension’, ‘multiple sclerosis’, various infectious diseases and cancers were also observed. As for the GAD disease class, 162 UAGs are significantly enriched in ‘pharmacogenomics’ (22.8%, P = 5.10e-10), also there are 172, 153, 106 and 182 UAGs enriched in terms ‘immune’, ‘cancer’, ‘infection’ and ‘cardiovascular’, respectively. DAGs are only enriched in one GAD item ‘infection’, with the proportion of 19.2% (P = 1.10e-14) (Supplementary Fig. 3).
Age-associated genes in PPI and signaling networks
Genes cooperate with each other to maintain homeostasis in the human body. To investigate the network characteristics of age-associated genes, we first estimated the centrality difference for each gene node in a non-directional protein–protein interaction (PPI) network through three metrics: degree, betweenness centrality and closeness centrality. These serve as important characteristics to evaluate whether a gene node in the network is ‘well-connected’ to other nodes. Among the three gene groups, DAG nodes have the highest degree, betweenness centrality and closeness centrality, compared with UAG nodes and NAG nodes, whereas UAG nodes have lower degrees than NAG nodes (Figs. 5a–c). Statistical results are shown in Table 2. To further explore how closely UAGs and DAGs are linked to each other in the PPI network, first we performed a permutation test, which revealed that the number of direct DAG-DAG links (DLs) and UAG-DAG links (UDLs) tend to be greater than when gene nodes are randomly distributed to the three gene groups (P = 0 and 6e-4, respectively) (Figs. 5e, f), whereas no significant results were observed for UAG-UAG links (ULs). Similar results, revealed by network distance analysis, are that DAGs have the shorter distances from each other than NAG nodes (median value of average D-D distances and D-N distances are 2.68 and 2.88, respectively), UAGs have the longest distances within the group, with no significant difference between the distance to NAGs (median value of average U-U distance and U-N distance are 3.07 and 3.06, respectively), and the UAG-DAG distances remain shorter than UAG-UAG distances (Fig. 5d and Table 3). These results suggest DAGs are of great connectivity and are cross-functioning, whereas UAGs are likely to stay in smaller groups and be more specialized in function. The fact that these two groups are fairly connected to each other in the PPI network, implies some of the UAGs and DAGs function similarly and may be attached to the same pathways.
a-c: Box plots display the difference among UAGs, DAGs and NAGs for the following characteristics in the PPI network: a Degree. b Betweenness centrality. c Closeness centrality. d Average length of the shortest paths of each group is shown in boxplot. **P < 0.01, ***P < 0.001 from Wilcoxon test. e The actual link number between DAG nodes, compared with link number in 10,000 randomized DAG node groups. The arrow shows link number in actual DAG node groups. f The actual link number between UAG and DAGs nodes compared with link number in 10,000 randomized UAG and DAG node groups. ***P < 0.001 from permutation test
In addition, we explored the characteristics of UAGs and DAGs in a cellular signaling network, in which 299 UAGs, 319 DAGs were mapped, with 5506 other genes considered as NAG nodes29. Centrality and distance analysis for UAGs and DAGs, in the signaling network, revealed different results, compared to PPI network, which UAGs have higher betweenness and closeness centrality value than DAGs and NAGs (Figs. 6c, d and Table 2). Concomitantly, DAGs have higher values than NAGs. Besides, UAGs have more positive links and negative links, compared with DAGs, respectively in out-degree and in-degree (Figs. 6a, b and Table 2). As for the distance analysis, more U-U links, D-D links and U-D/D-U links were observed in the UAG and DAG node groups compared with randomized groups (P = 0, 0, 0.021, respectively, permutation test) (Supplementary Fig. 4), whereas U-U distances remain the shortest when compared with U-D, D-D and U-N. D-D and U-D distances are also shorter, in comparison with distances to other nodes (Fig. 6e and Table 3). These results showed UAGs and DAGs both have high connectivity in the signaling network; nonetheless UAGs are more clustered and of great centrality, suggesting that UAGs are more interconnected in the signaling network.
a The proportion of positive and negative links in out-degree. UAGs have greater number of positive links than DAGs. b Greater number of negative links in in-degree of UAGs as compared with DAGs. *P < 0.05, ***P < 0.001 from chi-squared test. c, d Box plots present the difference in c betweenness centrality and d closeness centrality between groups in signaling networks. e The network distances revealed by average of the shortest paths are shown in boxplot. *P < 0.05, **P < 0.01, ***P < 0.001 from Wilcoxon test
Discussion
Aging is a process characterized by progressive loss of physiological integrity, which leads to impaired function and increased vulnerability to death30. By analyzing the aging differences revealed by gene expression, important clues can be gained to better understand the process of aging at the transcriptional level. Here, we systematically characterized the characteristics of transcriptional age-associated genes in multiple aspects. To sum up, genes that increase expression with age are less conserved in evolution, more tissue specific in expression and more enriched in SNPs and disease genes. They are more active in the signaling network and are located more in the extracellular region, with clustered functions mostly involving the immune system and are likely to play crucial roles in various cancer-related pathways. Genes with a decreased level of expression with age have a longer phyletic history, tend to be stable over time, expressed globally across tissues and are expressed early in a human lifetime. These genes function mostly in basic metabolic or catabolic process that are vital for human survival, and play important roles in gene networks, especially in the PPI network.
Notably, these age-associated genes, in the up and downregulated directions, tend to be close to each other in genomic distances and distances in both PPI and signaling networks. To further study potentially aging-related processes, 323 interacting UAGs and DAGs nodes were extracted from the signaling network, visualized (Supplementary Fig. 5) and functionally annotated (results for GO BP and pathway enrichment analysis with threshold FDR < 0.05 are shown in Supplementary Tables 5 and 6, respectively) by Cytoscape31. Examples for the interacting age-associated gene nodes enriched in process ‘T-cell receptor signaling pathway’ and enriched in pathway ‘pathways in cancers’ are shown in Supplementary Figure 6.
Apart from this, some DAGs are overlapped in function with UAGs, other DAGs are more dispersive with functions in multiple aspects, whereas UAGs are consistently more clustered and co-functioned. These dispersive distributed DAGs are enriched in terms related to catabolic, metabolic, mitochondrion-related and innate immune processes, which is consistent with previous studies that aging is a process with mitochondrial and immune dysfunction9,32. UAGs are mostly clustered in immune-related function and more located in the extracellular region, indicating the tendency of senescent cells to secrete pro-inflammatory cytokines, which lead to a chronic inflammatory state in healthy aging individuals. Besides, these immune-relevant UAGs and some of their interacted DAGs are also highly enriched in oncogenic signaling pathways, including Ras, nuclear factor kappa-light-chain-enhancer of activated B cells(NF-kappa B), phosphatidylinositol 3-kinase(PI3K)/AKT, Notch and P53 signaling pathways, which implies that senescent cells resemble cancer cells. This needs to be further verified by comparing aging profiles to specifically precancerous profiles. When mapping the 1020 cancer genes from OncoKB33 to all gene groups, UAGs were observed to bear the highest cancer genes proportion (7.75%) compared with DAGs (4.17%, P = 0.0036, chi-squared test) and NAGs (4.97%, P = 0.0013, chi-squared test). DAGs have the lowest proportion and no significant difference is shown between DAGs and NAGs. The cancer genes mapped in UAGs and DAGs are listed in Supplementary Table 7. In addition, we observed 15 and 14 cancer driver genes, respectively, from a 299 pan-cancer driver gene set provided by a recent study of TCGA’s Pan-Cancer34 in UAGs and DAGs, such as ERBB2 (level 1 driver gene in OncoKB33), TNFAIP3 in UAGs, and MAPK1, PIK3CB in DAGs (driver genes mapped in UAGs and DAGs are listed in Supplementary Table 8). As these driver genes are mostly increased in expression in tumorigenesis process, the driver genes found in DAGs may serve as a protective factor in this driving process. However, a previous study showed some driving processes are not likely to co-occur in cancer35, therefore the decreased level in these driver genes may accompany an increased level of other driver genes.
Aging is widely considered to be caused by accumulated cellular damage36,37, which is led by random mutations. SNP enrichment analysis shows more variations in UAGs, which means these genes are likely to be the positions that random mutation occurred. There is a possibility that the mutation in some UAGs cause them to increase in expression, and interfere with other interactive DAGs and UAGs. These changes may contribute to part of the aging transcriptional landscape. Considering DAGs experience less selection with lower evolutionary rates, mutations in DAGs are likely to be fatal, which would cause more disruptive outcomes that would be hard to observe in healthy individuals. Besides, as previous studies noted13,38,39, the decreased expression in DAGs may be mainly caused by age-associated epigenetic drift. The analyses in evolutionary characteristics, expression pattern and connectivity in the PPI network, showed consistent results with previous studies that UAGs are younger genes than DAGs, as studies showed that younger genes evolve more rapidly40, are more likely to present different temporal and spatial expression patterns41 and have fewer interactions in the PPI network42. Although UAGs and DAGs are both of high connectivity in the signaling network, UAGs remain more interactive, which suggests aging may accompany more accumulated in signaling crucial proteins.
Moreover, when performing the functional enrichment analysis, it was of note that the number of enriched terms that have a positive and a negative regulating function are about the same. This suggests those differentially expressed genes in healthy old individuals are not likely to cause imbalances that lead to conspicuously disruptive outcomes. As for an effort in precise medicine, 82 and 88 FDA-approved target genes from DrugBank43, mapped in UAGs and DAGs (Supplementary Table 9), are provided, respectively, which hopefully can help future research in different drug use for the young and old. Overall, our findings provide multiple biological implications for further study in healthy aging.
Materials and methods
The age-associated gene set
The age-associated gene set was extracted from a dataset of genes expressed differentially across age, as given in the GTEx project. These genes were screened out using a linear mixed model where sex, race and tissue were controlled to avoid the biases24. Our study focused on protein-coding genes, thus noncoding RNA genes were excluded from the gene set and the remaining genes were then categorized into two groups according to the regression coefficient. Genes with positive and negative coefficients were described as UAGs and DAGs, respectively. The original Ensembl IDs in the dataset were transformed into Entrez gene IDs and official gene symbols for further analysis. Apart from the age-associated genes, the rest of the protein-coding genes were regarded as NAGs.
Chromosomal distribution and genomic distance analyses
Chromosomal distribution of age-associated genes was revealed by calculating and comparing the proportion of both UAGs and DAGs in the totality of genes on each chromosome. The genomic distances of every pair of UAGs/DAGs (UDs/DDs), as well as the distances between a UAG and a DAG (UDDs) on the same chromosome, were calculated. The Wilcoxon rank-sum test was used to compare the distribution of UDs, DDs and UDDs across chromosomes.
Evolutionary and expression characteristic analyses
The average expression level across tissues and tissue specificity of each gene were calculated based on a gene atlas database from Su et al.44. The dN/dS ratio dataset of each human–mouse homolog was derived from the Ensembl database (release 83) to illustrate gene evolutionary rate. The homologous gene number dataset was obtained from the Homologene database45 (build 68), the phyletic evolution and earliest expression stage datasets were obtained from Online Gene Essentiality database46 and the essential gene dataset was obtained from the DEG database (version 10.6)27. The Wilcoxon test was performed to statistically compare the evolutionary rate, expression level, expression specificity and homologous gene number of UAGs, DAGs and NAGs, whereas a chi-squared test was used to compare the proportion of earliest expression stage, phyletic evolution and the essential genes between the three gene sets.
Functional enrichment analysis for age-associated genes
We calculated the number of genes in four terms of the GO: NU, CY, ME and ER, and compared the numbers between UAGs, DAGs and NAGs by performing chi-squared tests. Meanwhile, functional enrichment analysis of GO biological process and KEGG pathway annotations were both performed using the DAVID Web server47.
Analysis of SNP densities
A dataset of SNPs in human protein-coding genes (SNPs and indels, excluding flagged variants) (GRCh38.p2) with genome coordinates was derived from the Ensembl database48. SNP density of each gene was defined as the total number of mapped SNPs, divided by gene length. SNPs were mapped to all protein-coding genes, and the average SNP densities on each chromosome of UAGs, DAGs and NAGs were compared using the Wilcoxon test.
The enrichment analysis of disease-related SNP and disease genes
The disease-related SNPs (dSNPs) were obtained from the ClinVar database49 and Human Gene Mutation Database;50 data without a dbSNP ID or labeled as ‘protective’, ‘(likely) benign’, ‘uncertain significance’, ‘conflicting data from submitters’, ‘other’ and ‘not provided’ were excluded. dSNPs were mapped to each chromosome and the fractions of dSNPs to total SNPs in UAGs, DAGs and NAGs were calculated and compared using the Wilcoxon test. As for disease gene analysis, the DAVID Web server47 was used to perform functional enrichment analysis for terms of specific disease and disease class in the GAD, and the curated gene–disease association dataset downloaded from the DisGeNET database51 was also used to compare the fraction of disease genes for UAGs, DAGs and NAGs across chromosomes by performing the Wilcoxon test.
Network analysis
The human PPI network was downloaded from the BioGRID database (build 3.4.140) with the deletion of links that include non-human protein52. The human cellular signaling network was taken from our previous study29. The Wilcoxon test was performed to investigate the degree, betweenness centrality and closeness centrality differences for the UAG, DAG and NAG nodes in the PPI network. For the degree analysis in the signaling network, the proportion of positive links to the sum of positive links and negative links in UAGs and DAGs was compared for both in-degree and out-degree by performing chi-squared tests. As for network distance analysis, permutation tests were performed to compare the numbers of UAG-UAG links (ULs), DAG-DAG links (DLs) and UAG-DAG/DAG-UAG links (UDLs) in our identified networks with the numbers of those three types of links for 10,000 randomized node groups. The P-value was calculated as the frequency of times when link number in randomized groups was greater than that in identified groups. In addition, the average length of the shortest paths for UAG and DAG nodes in their own group, between each group and with the NAG nodes were also compared by performing the Wilcoxon test. The degree, betweenness centrality, closeness centrality and length of the shortest path values were calculated by using the python package NetworkX53. For the signaling network, distances were counted from both directions in each distance group. Network visualization and functional enrichment analysis for interacting age-associated gene nodes in the signaling network were carried out by Cytoscape31.
Change history
10 July 2019
Due to a technical error, content intended for publication in Volume 4 (2018) published in Volume 5 (2019). The content has been moved into the correct volume, and the citation information was updated accordingly.
References
Niccoli, T., & Partridge, L. Ageing as a risk factor for disease. Curr. Biol. 22, R741–R752 (2012).
Finkel, T., Serrano, M. & Blasco, M. A. The common biology of cancer and ageing. Nature 448, 767–774 (2007).
Jousilahti, P., Vartiainen, E., Tuomilehto, J. & Puska, P. Sex, age, cardiovascular risk factors, and coronary heart disease. Circulation 99, 1165 (1999).
North, B. J. & Sinclair, D. A. The intersection between aging and cardiovascular disease. Circ. Res. 110, 1097–1108 (2012).
Lindsay, J. et al. Risk factors for Alzheimer’s disease: a prospective analysis from the Canadian Study of Health and Aging. Am. J. Epidemiol. 156, 445–453 (2002).
Phillipson, O. T. Management of the aging risk factor for Parkinson’s disease. Neurobiol. Aging 35, 847–857 (2014).
Anderson, R. M. & Weindruch, R. The caloric restriction paradigm: implications for healthy human aging. Am. J. Human. Biol. 24, 101–106 (2012).
Blackburn, E. H., Epel, E. S. & Lin, J. Human telomere biology: a contributory and interactive factor in aging, disease risks, and protection. Science 350, 1193 (2015).
Theurey, P. & Pizzo, P. The aging mitochondria. Genes 9, 22 (2018).
Hoeijmakers, J. H. J. DNA damage, aging, and cancer. New Engl. J. Med. 361, 1475–1485 (2009).
Bandaranayake, T., & Shaw, A. C. Host resistance and immune aging. Clin. Geriatr. Med. 32, 415–432 (2016).
Jeck, W. R., Siebold, A. P., & Sharpless, N. E. Review: a meta-analysis of GWAS and age-associated diseases. Aging Cell 11, 727–731 (2012).
Sen, P., Shah, P. P., Nativio, R., & Berger, S. L. Epigenetic mechanisms of longevity and aging. Cell 166, 822–839 (2016).
Kenyon, C. J. The genetics of ageing. Nature 464, 504–512 (2010).
Welle, S. et al. Skeletal muscle gene expression profiles in 20-29 year old and 65-71 year old women. Exp. Gerontol. 39, 369–377 (2004).
Park, S. K., & Prolla, T. A. Gene expression profiling studies of aging in cardiac and skeletal muscles. Cardiovasc. Res. 66, 205–212 (2005).
Harries, L. W. et al. Human aging is characterized by focused changes in gene expression and deregulation of alternative splicing. Aging Cell 10, 868–878 (2011).
Peters, M. J. et al. The transcriptional landscape of age in human peripheral blood. Nat. Commun. 6, 8570 (2015).
Glass, D. et al. Gene expression changes with age in skin, adipose tissue, blood and brain. Genome. Biol. 14, R75 (2013).
Somel, M. et al. MicroRNA, mRNA, and protein expression link development and aging in human and macaque brain. Genome Res. 20, 1207–1218 (2010).
Tan, Q. et al. Genetic dissection of gene expression observed in whole blood samples of elderly Danish twins. Hum. Genet. 117, 267–274 (2005).
Berchtold, N. C. et al. Gene expression changes in the course of normal brain aging are sexually dimorphic. Proc. Natl. Acad. Sci. USA 105, 15605–15610 (2008).
de Magalhaes, J. P., Curado, J. & Church, G. M. Meta-analysis of age-related gene expression profiles identifies common signatures of aging. Bioinformatics 25, 875–881 (2009).
Mele, M. et al. Human genomics. The human transcriptome across tissues and individuals. Science 348, 660–665 (2015).
Consortium, G. Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Cohen, B. A., Mitra, R. D., Hughes, J. D. & Church, G. M. A computational analysis of whole-genome expression data reveals chromosomal domains of gene expression. Nat. Genet. 26, 183 (2000).
Zhang, R., Ou, H. Y. & Zhang, C. T. DEG: a database of essential genes. Nucleic Acids Res. 32, D271–D272 (2004).
Yang, J. et al. Synchronized age-related gene expression changes across multiple tissues in human and the link to complex diseases. Sci. Rep. 5, 15145 (2015).
Cui, Q. et al. A map of human cancer signaling. Mol. Syst. Biol. 3, 152 (2007).
Lopez-Otin, C., Blasco, M. A., Partridge, L., Serrano, M., & Kroemer, G. The hallmarks of aging.Cell 153, 1194–1217 (2013).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Shaw, A. C., Goldstein, D. R. & Montgomery, R. R. Age-dependent dysregulation of innate immunity. Nat. Rev. Immunol. 13, 875–887 (2013).
Chakravarty, D. et al. OncoKB: a precision oncology knowledge base. JCO Precision Oncol. 1, 1–16, https://doi.org/10.1200/PO.17.00011 (2017).
Bailey, M. H. et al. Comprehensive characterization of cancer driver genes and mutations. Cell 173, 371–385 (2018).
Sanchez-Vega, F. et al. Oncogenic signaling pathways in the cancer genome atlas. Cell 173, 321–337 (2018).
Gems, D. & Partridge, L. Genetics of longevity in model organisms: debates and paradigm shifts. Annu. Rev. Physiol. 75, 621–644 (2013).
Vijg, J. & Campisi, J. Puzzles, promises and a cure for ageing. Nature 454, 1065–1071 (2008).
Teschendorff, A. E., West, J. & Beck, S. Age-associated epigenetic drift: implications, and a case of epigenetic thrift? Hum. Mol. Genet. 22, R7–R15 (2013).
Booth, L. N., & Brunet, A. The aging epigenome. Mol. Cell 62, 728–744 (2016).
Wolf, Y. I., Novichkov, P. S., Karev, G. P., Koonin, E. V. & Lipman, D. J. The universal distribution of evolutionary rates of genes and distinct characteristics of eukaryotic genes of different apparent ages. Proc. Natl. Acad. Sci. USA 106, 7273–7280 (2009).
Popadin, KonstantinY. et al. Gene age predicts the strength of purifying selection acting on gene expression variation in humans. Am. J. Hum. Genet. 95, 660–674 (2014).
Zhang, W., Landback, P., Gschwend, A. R., Shen, B. & Long, M. New genes drive the evolution of gene interaction networks in the human and mouse genomes. Genome Biol. 16, 202 (2015).
Wishart, D. S. et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res. 46, D1074–D1082 (2018).
Su, A. I. et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl. Acad. Sci. USA 101, 6062–6067 (2004).
NCBI. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 44, D7–D19 (2016).
Chen, W. H., Minguez, P., Lercher, M. J. & Bork, P. OGEE: an online gene essentiality database. Nucleic Acids Res. 40, D901–D906 (2012).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Kinsella, R. J. et al. Ensembl BioMarts: a hub for data retrieval across taxonomic space. Database 2011, bar030, https://doi.org/10.1093/database/bar030 (2011).
Landrum, M. J. et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 44, D862–D868 (2016).
Stenson, P. D. et al. The Human Gene Mutation Database: building a comprehensive mutation repository for clinical and molecular genetics, diagnostic testing and personalized genomic medicine. Hum. Genet. 133, 1–9 (2014).
Pinero, J. et al. DisGeNET: a discovery platform for the dynamical exploration of human diseases and their genes. Database 2015, bav028 (2015).
Chatr-aryamontri, A. et al. The BioGRID interaction database: 2017 update. Nucleic Acids Res. 45, D369–D379 (2017).
Hagberg, A., Swart, P. & S Chult, D. Exploring Network Structure, Dynamics, and Function Using NetworkX. (Los Alamos National Lab (LANL), Los Alamos, NM, USA 2008).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Acknowledgements
This work has been supported by the grants from the Special Project on Precision Medicine under the National Key R&D Program (2016YFC0903003), the Natural Science Foundation of China (81670462).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by N. Barlev
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jia, K., Cui, C., Gao, Y. et al. An analysis of aging-related genes derived from the Genotype-Tissue Expression project (GTEx). Cell Death Discov. 4, 91 (2018). https://doi.org/10.1038/s41420-018-0093-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-018-0093-y
- Springer Nature Limited
This article is cited by
-
Cognitive and immunological effects of yoga compared to memory training in older women at risk for alzheimer’s disease
Translational Psychiatry (2024)
-
OXR1 maintains the retromer to delay brain aging under dietary restriction
Nature Communications (2024)
-
Tissue-specific impacts of aging and genetics on gene expression patterns in humans
Nature Communications (2022)
-
Functional conservation in genes and pathways linking ageing and immunity
Immunity & Ageing (2021)
-
Molecular evolution and the decline of purifying selection with age
Nature Communications (2021)