Abstract
Glioblastoma (GBM) is the most common and lethal brain tumor in adults. Due to its fast proliferation, diffusive growth and therapy resistance survival times are less than two years for patients with IDH-wildtype GBM. GBM is noted for the considerable cellular heterogeneity, high stemness indices and abundance of the glioma stem-like cells known to support tumor progression, therapeutic resistance and recurrence. Doublesex- and mab-3–related transcription factor a2 (DMRTA2) is involved in maintaining neural progenitor cells (NPC) in the cell cycle and its overexpression suppresses NPC differentiation. Despite the reports showing that primary GBM originates from transformed neural stem/progenitors cells, the role of DMRTA2 in gliomagenesis has not been elucidated so far. Here we show the upregulation of DMRTA2 expression in malignant gliomas. Immunohistochemical staining showed the protein concentrated in small cells with high proliferative potential and cells localized around blood vessels, where it colocalizes with pericyte-specific markers. Knock-down of DMRTA2 in human glioma cells impairs proliferation but not viability of the cells, and affects the formation of the tumor spheres, as evidenced by strong decrease in the number and size of spheres in in vitro cultures. Moreover, the knockdown of DMRTA2 in glioma spheres affects the stabilization of the glioma stem-like cell-dependent tube formation in an in vitro angiogenesis assay. We conclude that DMRTA2 is a new player in gliomagenesis and tumor neovascularization and due to its high expression in malignant gliomas could be a biomarker and potential target for new therapeutic strategies in glioblastoma.
Similar content being viewed by others
Introduction
Malignant gliomas (WHO grades 3 and 4) are the most common primary tumors of the central nervous system (CNS) in adults [1]. Glioblastoma (GBM, classified as a WHO grade 4) due to its aggressiveness, highly diffusive growth and vascularization is a deadly tumor with average survival of patients less than 15 months [2]. The intratumoral cellular heterogeneity and diffusive growth of GBM make complete surgical resection difficult. Particularly, the presence of tumor stem-like cells contributes to therapeutic resistance and quick tumor recurrence after surgery combined with radio and chemotherapy [3].
Due to their fatality and limited therapeutic options, malignant gliomas are intensively studied to identify biomarkers and pathological mechanisms. The numerous studies have characterized different glioma groups and subtypes with regard to DNA mutations, DNA copy number alterations, gene expression, DNA methylation, miRNA or protein expression profiles [4, 5]. Common oncogenic drivers of pediatric and high and lower grade adult gliomas were found [6], but up to now standard and personalized therapies were not effective. Intensive attempts at understanding the underlying pathobiology are undertaken to find new targets that would expand therapeutic options for GBM patients.
Doublesex and mab-3-related transcription factor a2 (Dmrta2) is the DM domain transcription factor involved in gonadal differentiation and CNS development. The expression of Dmrta2/DMRTA2 in neural tissues during development, and its role in neurogenesis and normal brain development have been documented in Zebrafish [7], Xenopus [8], mice [9,10,11,12] and humans [13]. Studies of Dmrta2 null mice indicate that the genetic ablation of Dmrta2 causes the hypoplasia or loss of the medial structures in the telencephalic cortex, thus implying that Dmrta2 plays a pivotal role in the regulation of neocortical patterning and in the formation of the signaling center for brain differentiation [7, 9]. In zebrafish, Dmrta2 controls neurogenin1 expression by repressing her6 in the posterior-dorsal telencephalon and the lack of Dmrta2 function affects the neuronal differentiation gene—neurod expression [7]. In mouse, Dmrta2 plays pivotal roles in the early development of the telencephalon and in the maintenance of neural progenitors as a downstream component of the Wnt pathway [9]. Loss of Dmrta2 results in decreased Wnt and Bmp expression in the dorsomedial telencephalon and in a reduction of the number of Cajal–Retzius cells [14]. Furthermore, Dmrta2 is known as a regulator of the neuron-glia cell-fate switch in the developing hippocampus. Dmrta2 reciprocally regulated Lhx2: loss of either factor promoted gliogenesis; overexpression of either factor suppressed gliogenesis and promoted neurogenesis; while each could substitute for the loss of the other [15]. Moreover, Dmrta2 plays an important role in maintaining neural precursor cells (NPC) in the cell cycle. Loss of Dmrta2 accelerates the cell cycle exit and differentiation of NPCs into postmitotic neurons, while its forced expression suppresses NPC differentiation [16]. Compelling evidence from studies in animal models indicates that neural stem or precursor cells accumulating genetic alterations are glioma-initiating cells and may contribute to the biological and genomic phenotypes of glioblastoma [17]. Nevertheless, to date, there is no data evaluating the role of DMRTA2 in gliomagenesis.
Here, we report that DMRTA2 is highly upregulated in human gliomas, in particular in WHO grades 3 and 4 gliomas and its expression inversely correlated with patient survival. The protein is detected around blood vessels and in a subset of small tumor cells with high proliferative potential. DMRTA2 expression is also elevated in glioma cell lines compared to normal human astrocytes and is even higher in glioblastoma-derived spheres enriched for cancer stem cells. Knock-down of DMRTA2 in human glioma cells impairs their proliferation without affecting cell viability and reduces formation of tumor spheres in in vitro cultures. Moreover, DMRTA2-depleted glioma spheres demonstrate reduced pro-angiogenic competences and faster tube disruption in an in vitro tube formation assay. Altogether, the results show an important role of DMRTA2 in the maintenance of the stemness phenotype of glioma cells and aberrant angiogenesis.
Materials and methods
RNA and DNA purification from glioma samples
The glioma biopsies were obtained from the Canadian Brain Tumor Tissue Bank in London (London Health Sciences Centre, Ontario CA), Children’s Memorial Health Institute, Warsaw, Poland and The Maria Sklodowska-Curie Memorial Cancer Center and Institute, Warsaw, Poland and informed consents were obtained from all subjects. The use of patient tissues was permitted by the Ethical Committee of the respective hospitals.
Total RNA was isolated using Tri Reagent extraction from snap-frozen tissues followed by the RNeasy Mini Kit isolation as previously described [18]. RNA quantity, quality and integrity were verified using the NanoDrop 2000 spectrophotometer (Thermo Scientific) and the Agilent Bioanalyzer 2100 and RNANanoChip assay (Agilent Technologies). DNA precipitation from the interphase and organic phase was performed with 100% ethanol. Samples were centrifuged at 5000g for 10 min at 4 °C. After 3 washes in 0.1 M trisodium citrate, 10% ethanol for 30 min and a final wash in 75% ethanol the DNA pellet was air dried and resuspended in H2O.
TCGA data analysis and Kaplan-Meier survival analyses
DMRTA2 expression within TCGA data was performed in GlioVis—data portal for visualization and analysis of brain tumor expression datasets (http://gliovis.bioinfo.cnio.es). Statistical analysis was performed with Tukey’s Honest Significant Difference (HSD) test. Survival plots were generated in GlioVis using the Rembrandt dataset (http://gliovis.bioinfo.cnio.es).
Quantitative determination of the DMRTA2 expression in human samples
Real time PCR amplifications with the primer sets: DMRTA2 - Hs00294890_m1; GAPDH - Hs02753991_g1 (Life Technologies) were performed in triplicates. The reference brain RNA was a mixture obtained from 23 normal brains (FirstChoice® Human Brain Reference RNA, Ambion, Austin, TX, USA), 5 samples were from single donors. The relative quantification of gene expression was determined with ABI PRISM 7700 using the comparative CT method.
Preparation of DNA for bisulfide conversion and DNA methylation analyses
Tri Reagent extracted DNA was additionally digested with 600 µg/mL proteinase K and purified by phenol:chloroform extraction followed by ethanol precipitation and re-suspended in H2O. Next, DNA was subjected to bisulfite modification using EpiTect Kit (Qiagen) according to the manufacturer’s instructions. DNA methylation was assessed using methylation-specific PCR (MS-PCR). Primers recognizing methylated and unmethylated DNA sequences were designed using Methyl Primer Express software and sequences were as follows: DMRTA2mF: TTAAGGAGTCGTTAAGGTGC, DMRTA2mR: TCAATCAAAACGTTTAACGA, DMRTA2umF: TTTTTAAGGAGTTGTTAAGGTGT, DMRTA2umR: CCTTCAATCAAAACATTTAACAA. Commercially available methylated and unmethylated DNA (Qiagen) were used as controls. PCR products were resolved by electrophoresis on 1% agarose gel containing ethidium bromide and DNA was visualized with UV light. Densitometry analysis of the picture was performed with ImageJ.
Immunohistochemical staining
Staining for DMRTA2 was performed on 5-μm paraffin-embedded PA and GBM tissue sections. Sections were deparaffinised at 60 °C for 2 h followed by incubation in xylene, ethanol (100, 90, 70%) and rehydration. Epitopes were retrieved by microwave boiling in pH 6.0 citrate buffer for 20 min. Endogenous peroxidase was blocked in 0.3% H2O2 in methanol for 30 min followed by blocking with 1% swine/5% horse serum. Sections were incubated overnight at 4 °C with rabbit anti-DMRTA2 antibody (Abcam cat# ab156244, dilution 1:200, in 3% horse serum), washed in PBS, incubated with a biotinylated horse anti-mouse immunoglobulin (50 pg/mL PBS), then with avidin-DH-biotinylated-HP (horseradish peroxidase) (90 μg/mL PBS) (Vector Labs., Burlingame, CA, USA) for 60 min and with 3,3′-diaminobenzidine (DAB). As a control, staining with the primary antibody was omitted. Sections were stained with hematoxylin (Sigma-Aldrich, Munich, Germany), dehydrated through ethanol, cleared in xylene and mounted. Images were obtained using Leica DM4000B microscope and DAB intensity was measured by image deconvolution using ImageJ according to modified protocol [19].
Immunofluorescent staining
For immunofluorescent staining specimens from the Brain Tumor Tissue Bank were used. Paraffin-embedded sections were incubated in 60 °C, deparaffinised in xylene, rehydrated in ethanol (100, 90, 70%) and washed with water. Epitopes were retrieved by boiling in a pH 6.0 citrate buffer for 30 min. Endogenous peroxidase was blocked in 0.3% H2O2 in 10% methanol for 30 min followed by blocking in PBS containing 10% donkey serum in 0.1% Triton X-100 solution for 1 h and incubated overnight at 4 °C with rabbit anti-DMRTA2, and mouse anti-OLIG2 or mouse anti-CD45 or mouse anti-Smooth Muscle Actin or anti-NESTIN and sheep anti-von Willebrand Factor antibodies. All antibodies were diluted in 0.1% Triton X-100/PBS solution containing 3% of donkey serum. Next, sections were washed in PBS and incubated with corresponding secondary antibodies for 1 h at room temperature. Nuclei were counter-stained with DAPI (1µg/mL) and specimens were mounted. Images were obtained on Zeiss LSM800 Airyscan microscope. For reagent specifications, catalogue numbers and concentrations, see Supplementary Table 1. To estimate DMRTA2+ and NESTIN co-localization, images were analyzed with ImageJ software. Cells showing protein co-localization were quantified. Only cells with a visible nucleus in the DAPI staining were counted.
Cell lines, GBM patient-derived cultures and their maintenance
Human LN18, U87-MG, LN229 and T98G glioma cells were purchased from American Type Culture Collection (ATCC, Manassas, VA, USA). Glioma cells were authenticated using the Multiplex Cell Authentication protocol (Multiplexion GmbH, Heidelberg, Germany) and tested for mycoplasma contamination once a month. Patient-derived glioma primary cultures WG1, WG4 and WG14 (GBM WHO grade 4) were generated as previously described [20, 21]. Normal human astrocytes (NHA) were purchased from Lonza (Walkersville, MD, USA) and cultured as previously described [22]. Human Umbilical Vein Endothelial Cells (HUVEC) were purchased from Gibco (Angiogenesis Starter Kit Cat. No. A14609-01, Life Technologies, Rockville, MD, USA) and cultured according to the manufacturer’s protocol in Medium 200 supplemented with Large Vessel Endothelial Supplement (LVES).
Glioma sphere cultures
For sphere formation, LN18 glioma cells were cultured as described [21]. Briefly, cells were seeded at a low density (1500 cells/cm2) on non-adherent plates and cultured in DMEM/F-12 GlutaMAX™ supplemented with 2% B27, 20 ng/mL rhuman bFGF, 20 ng/mL rhuman EGF, 0.0002% heparin and antibiotics. Cells were fed every 3 days by adding 1 mL of the fresh medium. After 7 days of culture, the pictures of the spheres were taken using light microscope and spheres above 100 µm in diameter were counted. Then, the spheres were collected by centrifugation at 1200 rpm at 4 °C for total RNA isolation and protein isolation for Western blotting. WG4 and WG14 spheres were obtained as described [20].
Preparation of protein extracts and Western blot analysis
Cells were lysed in the buffer containing phosphatase and protease inhibitors (20 mM Tris HCl, pH 6.8, 137 mM sodium chloride, 25 mM β-glycerophosphate, 2 mM sodium pyrophosphate, 2 mM EDTA, 1 mM sodium orthovanadate, 1% Triton X-100, 10% glycerol, 5 µg/mL leupeptin, 5 µg/mL aprotinin, 2 mM benzamidine, 0.5 mM DTT, 1 mM PMSF). The protein concentration was determined with the Pierce BCA Protein Assay Kit (Thermo Scientific). Protein extracts were separated on SDS-PAGE before electrophoretic transfer onto a nitrocellulose membrane (Amersham Biosciences, Germany) as described [23]. After blocking with 5% non-fat milk in TBS-T (Tris-buffered saline pH 7.6/0.15% Tween 20) the membranes were incubated with primary antibody recognizing DMRTA2 (Abcam #ab156244) diluted in a TBS-T overnight at 4 °C and then with horseradish peroxidase-conjugated anti-rabbit IgG (#PI-1000, Vector Laboratories) for one hour at RT. Immunocomplexes were visualized by using SuperSignal West Pico PLUS Chemiluminescent Substrate (Thermofisher Scientific). The membranes were stripped and re-probed with horseradish peroxidase-conjugated anti-β-Actin antibody (Sigma-Aldrich) to verify total protein loading.
Immunofluorescent staining of cells
LN18 spheres were collected by cytospin. LN18 cells were grown on coverslips at the density of 2 × 105/well in 24 well plates. Cells were fixed with 4% paraformaldehyde, postfixed with 100% methanol at −20 °C and blocked with 5% donkey serum, 1% BSA and 0.3% Triton X-100 for 1 h at room temperature. Primary antibody to DMRTA2 (Abcam cat# ab156244) was diluted 1:200 in PBS 1% BSA, 0.3% Triton X-100 containing 3% donkey serum and incubated with cells at 4 °C overnight. The cells were then incubated with anti-rabbit Alexa-555-conjugated secondary antibody (Invitrogen). After several washes with PBS containing 0.1% BSA, 0.3% Triton-X100, the cells were counterstained with 1 μg/mL DAPI (Sigma). The coverslips were dried, mounted on slides and visualized by fluorescence microscopy.
Knock-down of DMRTA2 in human LN18 glioma adherent cells
LN18 glioma cells were seeded on 96-well microplates (at the density of 4 ×103/well) for MTT metabolism and BrdU incorporation assays and 6-well plates (at the density of 2.25 × 105/well) for total RNA and protein isolation and left overnight to attach. The next day, the culture medium was removed and cells were transfected with 25 nM control or DMRTA2-specific siRNA (Dharmacon) using Viromer® BLUE transfection reagent (Lipocalyx GmbH, Halle, Germany). The culture medium was replaced after 4 h. The cells were assessed for cell viability and proliferation or collected for total RNA and protein isolation for Western Blot or harvested and seeded at a low density for the sphere formation assay. The efficiency of gene knock-down was determined by qPCR and Western blot analysis.
Knock-down of DMRTA2 in glioma-derived spheres
LN18 spheres cultured for 10 days or WG14 spheres cultured for 21 days were collected by centrifugation, washed with PBS and dissociated to single-cell suspension using TrypLe Express (Thermo Fisher Scientific). 1 × 105 cells were transfected with 50 nM of control or DMRTA2-specific siRNA (Dharmacon) in 20 μl of SF buffer (SF Cell Line 4D-Nucleofector TM X Kit) using 4D-Nucleofector system (Lonza). After transfection, cells were immediately resuspended in a fresh medium for spheres without antibiotics, plated on 60 mm plates for cell suspension and incubated for 24 h.
Tube formation assay
Pre-chilled 48-well culture plates were coated with Geltrex® LDEV-Free Reduced Growth Factor Basement Membrane Matrix (Angiogenesis Starter Kit # A14609-01, Gibco, Life Technologies) and incubated at 37 °C for 30 min. Meanwhile, the LN18 or WG14 spheres were dissociated to single-cell suspension using TrypLe Express (Thermo Fisher Scientific) and cells were counted. Then, 5 × 104 cells were stained with Vybrant™ DiI Cell-Labeling Solution (Thermo Fisher Scientific) following manufacturer instructions, washed with PBS and re-suspended in Medium 200 without a supplement. Human umbilical vein endothelial cells (HUVEC) were trypsinized, washed with PBS and counted. 7 × 104 HUVEC cells in 200 μL and 1 × 104 LN18 cells from spheres in 100 μL Medium 200 without a supplement were plated on 48-well plate previously coated with Geltrex® LDEV-Free Reduced Growth Factor Basement Membrane Matrix. Large Vessel Endothelial Supplement (LVES) was added as a control. The cells were visualized 16 h after seeding with fluorescent microscopy for endothelial tube formation and pictures were taken for angiogenesis analysis with ImageJ.
scRNA-seq data analysis
Published single-cell RNA-seq count matrices for GBM patients were obtained [24]. Data analysis was performed in R using Seurat v4 [25]. All samples (N = 14, mean number of cells per sample = 3829) were first filtered to remove dying cells (> 8% Unique Molecular Identifiers to mitochondrial genes, <5,5% UMI to ribosomal genes) and low complexity cells (<200 genes expressed per cell). Subsequently, MALAT1, ribosomal genes and genes with less than 20 counts have been discarded according to 10X Genomics recommendations. Samples were joined together using the “merge” function from the Seurat package. Next, raw counts were normalized to log2(RPKM+1). Expression levels were scaled for each gene by dividing the centered feature expression levels by their standard deviations. Multiplets (two or more cells with the same 10X barcode) were identified with DoubletFinder [26] and next discarded from the analysis. All major cell populations (with the exception of neural cells) have been identified: myeloid cells, lymphoid cells, oligodendrocytes, astrocytes, pericytes, and endothelial cells (markers used for identification are in Supplemental Table 2). The SOX2+ clusters were regarded as neoplastic and further analyzed.
Statistical analysis
All biological experiments were performed on 3–4 independent cell passages. Results are expressed as means ± standard deviation (SD). P-values were calculated using two-tailed t test or one-way ANOVA followed by appropriate post-hoc test using GraphPad Prism v6 (GraphPad Software, USA). Differences were considered statistically significant for p values < 0.05. Moreover, we calculated the effect size (Hedge’s ‘g’ between the groups) [27]. A commonly used interpretation for effect size is as follows: small (0.2), medium (0.5) and large (0.8); however, these values are arbitrary and should not be considered rigidly [27, 28].
Results
DMRTA2 is highly expressed in human glioblastoma and predicts poor survival
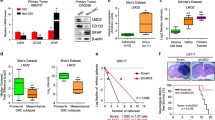
To explore DMRTA2 expression in brain tumors we took advantage of GlioVis, the open access web application for data visualization and analysis. Using RNA-sequencing data from the TCGA dataset, the levels of DMRTA2 mRNA were evaluated in a large group of gliomas of different WHO grades (G2-3-4). The DMRTA2 expression increased in tumors with higher malignancy grade and was significantly higher in GBM when compared to lower grade gliomas (Fig. 1A). In order to assess if DMRTA2 expression affects patient outcome we used the Rembrandt (REpository of Molecular BRAin Neoplasia DaTa) database [29]. Expression of DMRTA2 was strongly associated with patient survival in all gliomas (Fig. 1B), which is also evident among GBMs, highlighting the DMRTA2 involvement in GBM malignant phenotype (Fig. 1C). We confirmed high expression of DMRTA2 in our cohort of GBM samples in comparison to juvenile pilocytic astrocytoma (PA) and normal brain tissue (NB) (Fig. 1D). PA is a slow growing, not diffusive tumor [30], which rarely undergoes progression to high-grade tumors [31]. Immunohistochemistry staining confirmed high expression of DMRTA2 protein in malignant gliomas compared to benign tumors (Fig. 1E, F). In GBM sections positive DMRTA2 staining was evident in two distinct cell populations: first are small, highly proliferative cells present in tumor parenchyma, and the other are cells localized around blood vessels.
A Analysis of DMRTA2 mRNA expression in the TCGA data set using GlioVis. B, C Kaplan–Meier survival plots of all gliomas (B), and glioblastoma (C), patients. Patients were assigned to groups according to DMRTA2 expression levels. D Quantification of DMRTA2 mRNA levels in pilocytic gliomas (20 patients) and glioblastomas (23 patients) determined by qRT-PCR. E Quantification of immunohistochemistry staining for DMRTA2 in high and low-grade gliomas. F Representative images of immunohistochemical staining for DMRTA2 in high and low grade gliomas, black arrows point to positive staining around vessels, white arrows mark small cells with high proliferation potential, scale bar—100 µm. GBM glioblastoma, PA pilocytic astrocytoma, NB normal brain, NC negative control; *p < 0.05, **p < 0.01, ***p < 0.001.
DMRTA2 positive cells exhibit certain stem-like and pericyte-like properties
To characterize the functions of DMRTA2 positive cells, we performed the Gene Ontology (GO) biological processes analysis of glioma samples in the Rembrandt database. Tumors expressing high levels of DMRTA2 were distinguished by the enrichment in terms such as extracellular structure/matrix organization, blood vessel morphogenesis and angiogenesis (Supplemental Fig. 1). Further, we analyzed public scRNA-seq data from GBMs [24]. To characterize the cell identity of the obtained clusters, we applied the panel of markers identifying specific cells present in the brain and analyzed their co-expression with DMRTA2. All major cell populations have been identified, and non-tumor cells were removed from the further analysis. The remaining clusters expressed SOX2 and were regarded as neoplastic cells (Supplemental Fig. 2A). We analyzed these cells for the expression of neural/stem precursor cell markers (OLIG1, OLIG2, CSPG4/NG2, NESTIN), which are also transcription factors and lineage markers associated with glioma stem cells (GSC) [32] (Supplemental Fig. 2C), and DMRTA2 (Supplemental Fig. 2B). DMRTA2 expressing cells within SOX2 expressing clusters co-expressed also oligodendrocyte (OLIG1, OLIG2, CSPG4) and/or neural (NESTIN) progenitor cell genes (Supplemental Fig. 2C, E). Considering these data and immunohistochemistry staining of DMRTA2 in GBM, we focused on determining DMRTA2 co-expression with markers of stem/progenitor cells (OLIG2 [33] and NESTIN [34]), immune cells (CD45 [35]) and blood vessels (von Willebrant factor—VWF [34]). We found that neither OLIG2 nor CD45 co-localize with DMRTA2 staining (Fig. 2A, B), but we observed co-localization of DMRTA2 with NESTIN (Fig. 2D and Supplemental Fig. 3). Moreover, we noticed DMRTA2 positive cells in a close proximity to those positive for an endothelial marker—VWF (Fig. 2C). Many of DMRTA2+NESTIN+ cells were found around VWF+ endothelial cells. Vascular pericytes stretch around endothelial cells (ECs) and are functionally associated with regulating vessel stabilization, vessel diameter and EC proliferation. Some pericyte cells, which participate in angiogenesis, express NESTIN [36]. Therefore, we performed staining for another pericyte marker—α-Smooth Muscle Actin (αSMA), together with DMRTA2 and VWF. αSMA was found in pericytes supporting vessel formation in GBM [37]. We detected αSMA+ cells among DMRTA2+ cells (Fig. 2E), and these cells were localized next to VWF+ cells forming vessels. This suggests that DMRTA2+ cells contain a subpopulation of cells which acquire pericyte-specific features (Fig. 2E). The analysis of scRNA-seq data demonstrated a subset of DMRTA2 expressing cells having pericyte markers: out of which ACTA2 (coding for αSMA) and/or SIPR3 (coding for Sphingosine-1-Phosphate Receptor 3) and/or MCAM (coding for Melanoma Cell Adhesion Molecule, known as cluster of differentiation 146) were most frequently expressed (Supplemental Fig. 2D, F).
DMRTA2 is overexpressed in glioma sphere-forming cells and supports their self-renewal
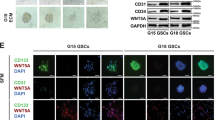
We analyzed DMRTA2 expression in the panel of primary and established glioma cell lines. The highest level of DMRTA2 protein was detected in two established cell lines: LN18 and T98G and primary GBM-derived WG4 cell cultures (Fig. 3A, B and Supplemental Fig. 4A). LN18 and WG4 cells have been previously described as forming spheres enriched in cells with cancer stem cell properties when cultured under proper conditions [21]. Increased DMRTA2 expression was indeed detected in spheres at both mRNA and protein level, when compared with adherent cells (Fig. 3C–E and Supplemental Figs. 4B, 5). These results are in line with scRNA-seq and immunofluorescence results, and implicate DMRTA2 in GSC functions.
A Western blot showing DMRTA2 protein levels in established glioma cells (LN18, U87MG, T98G, LN229), patient-derived (WG4, WG1) glioma cell cultures and normal human astrocytes (NHA). B Densitometry analysis of Western blot for DMRTA2 protein expression in normal human astrocytes (NHA), established (LN18, U87MG, T98G, LN229) and patient-derived (WG4, WG1) glioma cell lines (2–3 biological replicates for each cell line). C Quantification of DMRTA2 expression in LN18 adherent cells and LN18-derived spheres by qRT-PCR (n = 3 biological replicates). *p < 0.05. D Representative immunoblot showing the DMRTA2 protein levels in LN18 cells and LN18-derived spheres. E Immunofluorescence for DMRTA2 in LN18 adherent cells and LN18-derived spheres (n = 3 biological replicates). Nuclei visualized with DAPI, scale bar—50 µm. Of note is uniform, nuclear expression of DMRTA2 in most LN18 glioma sphere-forming cells, while in adherent cells DMRTA2 is visible in a cytoplasm and rare cells.
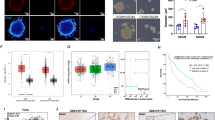
To investigate the role of DMRTA2 in glioma stem-like cells, we efficiently silenced DMRTA2 expression in LN18 cells (Fig. 4A, B and Supplemental Fig. 6). Loss of DMRTA2 expression impaired proliferation but did not affect viability of LN18 cells (Fig. 4C, D, respectively). Furthermore, DMRTA2-depleted cells had reduced potential to form spheres in vitro (Fig. 4E, F). The corresponding effect size of sphere number reduction in DMRTA2-depleted cells was high (g = 2.5) emphasizing its role in GSC self-renewal.
A The efficacy of DMRTA2 knock-down was evaluated by qRT-PCR 48 h after siRNA transfection (n = 4 biological replicates). B Western blot showing reduction of the DMRTA2 protein level in LN18 cells after DMRTA2 knock-down. C Cell proliferation of LN18 cells decreased in DMRTA2-depleted cells as verified with BrdU incorporation test (n = 4 biological replicates in triplicates). D Viability of LN18 cells was not affected by DMRTA2 knock-down as evaluated using MTT metabolism test (n = 2 biological replicates in triplicates). E Sphere formation capacities decrease in LN18 cells depleted of DMRTA2. Representative photos of LN18-derived spheres untreated, or transfected with siCTRL or siDMRTA2 and cultured for 7 days under sphere-forming conditions, scale bar—200 µm. F Number of spheres formed by LN18 cells was reduced after DMRTA2 knock-down (n = 4 biological replicates). G Representative photos of vascular nets formed by HUVEC alone or in co-culture with control LN18 or DMRTA2 depleted cells, scale bar—200 µm. H Quantification of selected properties of the vascular net formed by HUVEC alone or in co-culture with control or DMRTA2 depleted LN18 cells (n = 4 biological replicates). UNT—control untreated cells; *p < 0.05; **p < 0.01; ***p < 0.001; Hedge’s ‘g’ stands for effect size.
Loss of DMRTA2 disrupts vascular net created in vitro by endothelial cells and stabilized by glioma stem-like cells
Immunofluorescent staining revealed the presence of the DMRTA2 protein in the cells surrounding blood vessels. Most of these cells were also positive for αSMA, a pericyte marker. Pericytes stabilize and monitor the maturation of endothelial cells by direct communication with those cells as well as through paracrine signaling [38]. Some reports suggested glioblastoma stem cells may acquire a pericyte-like phenotype and this way contribute to tumor neovascularization [37]. We detected DMRTA2 positive cells with αSMA expression around endothelial cells (Fig. 2E). To study a potential role of DMRTA2 in tumor neovascularization, we employed an in vitro tube formation assay which measures the ability of endothelial cells, plated at sub-confluent densities to form capillary-like structures (tubes) provided with the appropriate extracellular matrix support [39]. Tube formation occurs quickly with endothelial cells aligning on a matrix within 1 h and is completed within 12–20 h following endothelial cell plating. After that time, the in vitro formed net starts to collapse without further support from angiogenic factors [39]. We performed a tube formation assay with human HUVEC cells in the presence of LN18 or WG14 cells grown as spheres. We selected the WG14 cell cultures due to the high expression of DMRTA2 (RNA-seq data in [20]) and the ability to form spheres. HUVEC cells plated on matrigel formed tubules within 12 h and at 16 h we observed disruption of the vascular net (Supplemental Figure 7). When HUVEC cells were plated in co-cultures with LN18 or WG14 cells derived from spheres, we observed stabilization of the formed vessels and the vascular net was still present at 16 h after plating (Fig. 4G, Supplemental Fig. 7, 8A). When HUVEC cells were co-cultured with sphere-derived, DMRTA2 depleted LN18 or WG14 cells (Fig. 4G, Supplemental Fig. 8A), we observed the decrease of mesh area, and the reduced numbers of nodes, junctions and segments in the vascular network formed by HUVEC (Fig. 4G, H and Supplementary Fig. 8A, B). Moreover, the corresponding effect size was high in each performed comparison between groups of interest, underlining the role of DMRTA2 in sustaining the vascular network stability.
Increased expression of DMRTA2 in high-grade gliomas is associated with DNA methylation
Epigenetic changes are considered to be among the earliest and most comprehensive genomic aberrations occurring during carcinogenesis [40]. The contribution of epigenetic mechanisms to glioblastoma pathology has been broadly studied and numerous alterations in DNA methylation patterns were detected [41]. To decipher a potential mechanism responsible for aberrant DMRTA2 expression in GBM, we sought to determine if DNA methylation alterations are responsible for the upregulation of DMRTA2 expression. We analyzed the DMRTA2 coding sequence for the presence of CpG island using EMBOSS Cpgplot. The whole DMRTA2 coding sequence appeared to be enriched for CpG dinucleotides. We took advantage of the DNA methylation data from Stepniak et al., [42] and looked at the DNA methylation within the DMRTA2 gene in GBM and PA. We observed differences in DNA methylation in the gene body and at the 3′ UTR of DMRTA2 [42]. Using methyl-specific PCR we found both methylated and unmethylated DMRTA2 sequences in DNA isolated from GBM, whereas no or very low amplification of methylated sequences was detected in PA samples (Fig. 5A, B). Furthermore, both methylated and unmethylated sequences were detected in DNA isolated from non-cancerous brain tissues (5 samples) (Fig. 5A); however, no or very low levels of DMRTA2 transcripts were found in contrast to GBM highly expressing DMRTA2 (Fig. 1D). Correlation analysis revealed a positive correlation between the DNA methylation level and DMRTA2 expression in the analyzed cohort of samples (Fig. 5C). These results suggest that DNA methylation may contribute to DMRTA2 expression in GBM.
A Representative results of MS-PCR for the CpG sites located in the 3′ UTR of the DMRTA2 gene. B Densitometry analysis of MS-PCR results relative to methylated control DNA (15 GBM and 15 PA samples). C Analysis of correlation of DNA methylation and DMRTA2 gene expression in high (14 samples) and low (15 samples) grade gliomas—p < 0.0001, r = −0.7150. GBM glioblastoma, PA pilocytic astrocytoma, NB normal brain, M methylated control, UM unmethylated control.
Discussion
The transcription factor DMRTA2 is involved in gonadal differentiation and CNS development taking part in determining cell fate decisions. The potential role of DMRTA2 as a prognostic marker was indicated in several non-CNS cancers as this gene appeared deregulated in gene expression [43, 44] or DNA methylation-based signatures [45, 46]. Tumor cell origin studies indicated glioblastoma originates from neural stem/progenitor cells. Therefore, any factors controlling cell lineage fate or differentiation are of particular interest, but the role of DMRTA2 in gliomagenesis has been unknown.
In this study, we determined the expression of DMRTA2 in human gliomas of different WHO grades, primary GBM-derived cultures and established glioma cell lines, and we found upregulation of DMRTA2 in high-grade gliomas. The DMRTA2 protein accumulated around blood vessels and in small cells with high proliferative potential in GBM sections. Such cells are consistent with primitive, quiescent stem cells residing in adult tissues suggesting a correlation between small cell sizes with the stemness [47]. ScRNA-seq data revealed that DMRTA2 expressing cells exhibited high expression of markers characteristic for stem and progenitor cells: SOX2, OLIG1, OLIG2, Nestin, CSPG4 (NG2) (Supplemental Fig. 2) supporting a link between DMRTA2 expression and the GSC phenotype. Furthermore, expression of DMRTA2 was increased in cultured glioma cells compared with normal human astrocytes. GBM-derived spheres enriched for cancer stem cells had higher DMRTA2 expression at mRNA and protein levels than adherent cells. Knock-down of DMRTA2 in glioma cells did not affect cell viability but affected proliferation and the GCS self-renewal. These results suggest a role of DMRTA2 in stemness maintenance and dysregulation of the differentiation program. DMRTA2 has been demonstrated as a factor in maintaining NPC in the cell cycle and preventing differentiation during brain development [16]. The presence of GCS is an important hallmark of GBM [32, 48].
We found DMRTA2-positive cells expressing pericyte markers around blood vessels in GBMs. Vascular pericytes play critical roles in supporting vascular structure and function, maintaining the blood-brain barrier, facilitating vessel maturation, and initiation of vessel sprouting [37]. However, the pericyte-endothelial cells interactions change substantially in tumors [49, 50]. The tumor vasculature frequently exhibits structural and functional abnormalities with irregular pericytes on endothelial tubules. In vivo cell lineage tracing demonstrated that GSC may generate vascular pericytes [37]. Our results suggest that DMRTA2 could be a marker of such GSC-derived pericytes as these cells co-express pericyte-specific markers as indicated by scRNA-seq data and immunofluorescence staining. Of particular interest is the presence of NESTIN as NESTIN+ (but not NESTIN-) pericytes were recruited to the blood vessels during tumor angiogenesis in the murine intracerebral glioblastoma [36]. We demonstrate the supportive role of cells derived from LN18 and WG14 spheres in stabilization of the vascular net created by endothelial cells in vitro. This support is lost when DMRTA2 is depleted in GSCs. The blood-tumor barrier (BTB) is a major obstacle to drug delivery to GBM but could be also a new attractive target [51]. Targeting GSC-derived pericytes may potentially enhance drug delivery, improving GBM therapy.
We explored DNA methylation as a potential mechanism through which DMRTA2 expression could be deregulated in GBMs. The DMRTA2 methylation level was described as a poor prognosis factor in Clear Cell Renal Cell Carcinoma [45] and as prognostic factor in bladder cancer [46]. However, the analysis of DNA methylation within CpG islands in the 3’ UTR of the DMRTA2 gene showed that despite a similar methylation pattern of the DMRTA2 in normal brain and GBM, the gene was expressed only in malignant samples. On the other hand, the DMRTA2 gene was not methylated at the tested sites in PA samples and the gene was not expressed. DNA methylation in different genomic regions may exert a different influence on gene activity, and DNA methylation of the gene body was associated with a higher level of gene expression in dividing cells [52, 53]. In slowly dividing and non-dividing cells such as the cells of the brain, gene body methylation was not associated with increased gene expression [53, 54]. These data are in agreement with our findings of high expression of the DMRTA2 gene in proliferating, malignant cells and not detectable expression in normal brain tissue. However, the presence of DNA methylation in the 3’ UTR of DMRTA2 may also affect the binding of repressors and this effect could result in high expression of DMRTA2 in GBMs. Further studies to elucidate the role of DNA methylation in DMRTA2 expression regulation in glioblastoma and its implication in glioma angiogenesis are necessary, as some DNA methylation inhibitors markedly decrease vessel formation in different tumor in vitro and in vivo models [55].
Altogether, our findings point to a role of DMRTA2, highly overexpressed in GBM, in glioma pathogenesis. Differential DMRTA2 expression in low and high-grade gliomas and in cultured glioma spheres enriched in GSC suggests dysregulation of the differentiation program. Moreover, the involvement of DMRTA2 in glioblastoma neovascularization opens a new possibility of targeting the brain-tumor barrier, which may facilitate drug delivery to GBM to effectively block tumor progression and improve therapy.
Data availability
The datasets analyzed during the current study are available in the GlioVis web application for data visualization and analysis, http://gliovis.bioinfo.cnio.es/. ScRNA-seq data are avialable from the authors of the original study on special request.
References
Miller KD, Ostrom QT, Kruchko C, Patil N, Tihan T, Cioffi G, et al. Brain and other central nervous system tumor statistics, 2021. CA Cancer J Clin. 2021;71:381–406.
Delgado-Martín B, Medina M. Advances in the knowledge of the molecular biology of glioblastoma and its impact in patient diagnosis, stratification, and treatment. Adv Sci (Weinh). 2020;7:1902971.
Valdebenito S, D’Amico D, Eugenin E. Novel approaches for glioblastoma treatment: Focus on tumor heterogeneity, treatment resistance, and computational tools. Cancer Rep. (Hoboken). 2019;2:e1220.
Ceccarelli M, Barthel FP, Malta TM, Sabedot TS, Salama SR, Murray BA, et al. Molecular profiling reveals biologically discrete subsets and pathways of progression in diffuse glioma. Cell. 2016;164:550–63.
Neftel C, Laffy J, Filbin MG, Hara T, Shore ME, Rahme GJ, et al. An integrative model of cellular states, plasticity, and genetics for glioblastoma. Cell. 2019;178:835–49.e21.
Sturm D, Bender S, Jones DT, Lichter P, Grill J, Becher O, et al. Paediatric and adult glioblastoma: multiform (epi)genomic culprits emerge. Nat Rev Cancer. 2014;14:92–107.
Yoshizawa A, Nakahara Y, Izawa T, Ishitani T, Tsutsumi M, Kuroiwa A, et al. Zebrafish Dmrta2 regulates neurogenesis in the telencephalon. Genes Cells. 2011;16:1097–109.
Parlier D, Moers V, Van Campenhout C, Preillon J, Leclère L, Saulnier A, et al. The Xenopus doublesex-related gene Dmrt5 is required for olfactory placode neurogenesis. Dev Biol. 2013;373:39–52.
Konno D, Iwashita M, Satoh Y, Momiyama A, Abe T, Kiyonari H, et al. The mammalian DM domain transcription factor Dmrta2 is required for early embryonic development of the cerebral cortex. PLoS ONE. 2012;7:e46577.
De Clercq S, Keruzore M, Desmaris E, Pollart C, Assimacopoulos S, Preillon J, et al. DMRT5 together with DMRT3 directly controls hippocampus development and neocortical area map formation. Cereb Cortex. 2018;28:493–509.
Konno D, Kishida C, Maehara K, Ohkawa Y, Kiyonari H, Okada S, et al. Dmrt factors determine the positional information of cerebral cortical progenitors via differential suppression of homeobox genes. Development. 2019;146:174243.
Ratié L, Desmaris E, García-Moreno F, Hoerder-Suabedissen A, Kelman A, Theil T, et al. Loss of Dmrt5 affects the formation of the subplate and early corticogenesis. Cereb Cortex. 2020;30:3296–312.
Urquhart JE, Beaman G, Byers H, Roberts NA, Chervinsky E, O’Sullivan J, et al. DMRTA2 (DMRT5) is mutated in a novel cortical brain malformation. Clin Genet. 2016;89:724–7.
Saulnier A, Keruzore M, De Clercq S, Bar I, Moers V, Magnani D, et al. The doublesex homolog Dmrt5 is required for the development of the caudomedial cerebral cortex in mammals. Cereb Cortex. 2013;23:2552–67.
Muralidharan B, Keruzore M, Pradhan SJ, Roy B, Shetty AS, Kinare V, et al. Dmrt5, a novel neurogenic factor, reciprocally regulates Lhx2 to control the neuron-glia cell-fate switch in the developing hippocampus. J Neurosci. 2017;37:11245–54.
Young FI, Keruzore M, Nan X, Gennet N, Bellefroid EJ, Li M. The doublesex-related Dmrta2 safeguards neural progenitor maintenance involving transcriptional regulation of Hes1. Proc Natl Acad Sci USA. 2017;114:E5599–E607.
Alcantara Llaguno SR, Parada LF. Cell of origin of glioma: biological and clinical implications. Br J Cancer. 2016;115:1445–50.
Sielska M, Przanowski P, Wylot B, Gabrusiewicz K, Maleszewska M, Kijewska M, et al. Distinct roles of CSF family cytokines in macrophage infiltration and activation in glioma progression and injury response. J Pathol. 2013;230:310–21.
Crowe AR, Yue W. Semi-quantitative determination of protein expression using immunohistochemistry staining and analysis: an integrated protocol. Bio Protoc. 2019;9:e3465.
Ciechomska IA, Wojnicki K, Wojtas B, Szadkowska P, Poleszak K, Kaza B, et al. Exploring novel therapeutic opportunities for glioblastoma using patient-derived cell cultures. Cancers (Basel). 2023;15:1562.
Ciechomska IA, Przanowski P, Jackl J, Wojtas B, Kaminska B. BIX01294, an inhibitor of histone methyltransferase, induces autophagy-dependent differentiation of glioma stem-like cells. Sci Rep. 2016;6:38723.
Was H, Krol SK, Rotili D, Mai A, Wojtas B, Kaminska B, et al. Histone deacetylase inhibitors exert anti-tumor effects on human adherent and stem-like glioma cells. Clin Epigenetics. 2019;11:11.
Sliwa M, Markovic D, Gabrusiewicz K, Synowitz M, Glass R, Zawadzka M, et al. The invasion promoting effect of microglia on glioblastoma cells is inhibited by cyclosporin A. Brain 2007;130:476–89.
Couturier CP, Ayyadhury S, Le PU, Nadaf J, Monlong J, Riva G, et al. Single-cell RNA-seq reveals that glioblastoma recapitulates a normal neurodevelopmental hierarchy. Nat Commun. 2020;11:3406.
Hao Y, Hao S, Andersen-Nissen E, Mauck WM, Zheng S, Butler A, et al. Integrated analysis of multimodal single-cell data. Cell 2021;184:3573–87.e29.
McGinnis CS, Murrow LM, Gartner ZJ. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 2019;8:329–37.e4.
Lakens D. Calculating and reporting effect sizes to facilitate cumulative science: a practical primer for t-tests and ANOVAs. Front Psychol. 2013;4:863.
Thompson B. Effect sizes, confidence intervals, and confidence intervals for effect sizes. Psychol Sch. 2007;44:423–32.
Gusev Y, Bhuvaneshwar K, Song L, Zenklusen JC, Fine H, Madhavan S. The REMBRANDT study, a large collection of genomic data from brain cancer patients. Sci Data. 2018;5:180158.
Collins VP, Jones DT, Giannini C. Pilocytic astrocytoma: pathology, molecular mechanisms and markers. Acta Neuropathol. 2015;129:775–88.
Collins KL, Pollack IF. Pediatric low-grade gliomas. Cancers (Basel). 2020;12:1152.
Suvà ML, Tirosh I. The glioma stem cell model in the era of single-cell genomics. Cancer Cell. 2020;37:630–6.
Ohnishi A, Sawa H, Tsuda M, Sawamura Y, Itoh T, Iwasaki Y, et al. Expression of the oligodendroglial lineage-associated markers Olig1 and Olig2 in different types of human gliomas. J Neuropathol Exp Neurol. 2003;62:1052–9.
Prosniak M, Kenyon LC, Hooper DC. Glioblastoma contains topologically distinct proliferative and metabolically defined subpopulations of Nestin- and Glut1-expressing cells. J Neuropathol Exp Neurol. 2021;80:674–84.
Ochocka N, Segit P, Walentynowicz KA, Wojnicki K, Cyranowski S, Swatler J, et al. Single-cell RNA sequencing reveals functional heterogeneity of glioma-associated brain macrophages. Nat Commun. 2021;12:1151.
Birbrair A, Zhang T, Wang ZM, Messi ML, Olson JD, Mintz A, et al. Type-2 pericytes participate in normal and tumoral angiogenesis. Am J Physiol Cell Physiol. 2014;307:C25–38.
Cheng L, Huang Z, Zhou W, Wu Q, Donnola S, Liu JK, et al. Glioblastoma stem cells generate vascular pericytes to support vessel function and tumor growth. Cell 2013;153:139–52.
Bergers G, Song S. The role of pericytes in blood-vessel formation and maintenance. Neuro Oncol. 2005;7:452–64.
Ponce ML. Tube formation: an in vitro matrigel angiogenesis assay. Methods Mol Biol. 2009;467:183–8.
Dawson MA, Kouzarides T. Cancer epigenetics: from mechanism to therapy. Cell 2012;150:12–27.
Maleszewska M, Kaminska B. Is glioblastoma an epigenetic malignancy? Cancers (Basel). 2013;5:1120–39.
Stępniak K, Machnicka MA, Mieczkowski J, Macioszek A, Wojtaś B, Gielniewski B, et al. Mapping chromatin accessibility and active regulatory elements reveals pathological mechanisms in human gliomas. Nat Commun. 2021;12:3621.
Jin Y, Qin X. Significance of TP53 mutation in treatment and prognosis in head and neck squamous cell carcinoma. Biomark Med. 2021;15:15–28.
Zhao H, Wang F, Wang X, Zhao X, Ji J. HPV-related prognostic signature predicts survival in head and neck squamous cell carcinoma. J Oncol. 2022;2022:7357566.
Zhou J, Liu G, Wu X, Zhou Z, Li J, Ji Z. A risk score model based on nine differentially methylated mrnas for predicting prognosis of patients with clear cell renal cell carcinoma. Dis Markers. 2021;2021:8863799.
Deng L, Chao H, Deng H, Yu Z, Zhao R, Huang L, et al. A novel and sensitive DNA methylation marker for the urine-based liquid biopsies to detect bladder cancer. BMC Cancer. 2022;22:510.
Li Q, Rycaj K, Chen X, Tang DG. Cancer stem cells and cell size: a causal link? Semin Cancer Biol. 2015;35:191–9.
Gimple RC, Bhargava S, Dixit D, Rich JN. Glioblastoma stem cells: lessons from the tumor hierarchy in a lethal cancer. Genes Dev. 2019;33:591–609.
Morikawa S, Baluk P, Kaidoh T, Haskell A, Jain RK, McDonald DM. Abnormalities in pericytes on blood vessels and endothelial sprouts in tumors. Am J Pathol. 2002;160:985–1000.
Winkler EA, Bell RD, Zlokovic BV. Central nervous system pericytes in health and disease. Nat Neurosci. 2011;14:1398–405.
Zhou W, Chen C, Shi Y, Wu Q, Gimple RC, Fang X, et al. Targeting glioma stem cell-derived pericytes disrupts the blood-tumor barrier and improves chemotherapeutic efficacy. Cell Stem Cell. 2017;21:591–603.e4.
Ball MP, Li JB, Gao Y, Lee JH, LeProust EM, Park IH, et al. Targeted and genome-scale strategies reveal gene-body methylation signatures in human cells. Nat Biotechnol. 2009;27:361–8.
Aran D, Toperoff G, Rosenberg M, Hellman A. Replication timing-related and gene body-specific methylation of active human genes. Hum Mol Genet. 2011;20:670–80.
Guo JU, Ma DK, Mo H, Ball MP, Jang MH, Bonaguidi MA, et al. Neuronal activity modifies the DNA methylation landscape in the adult brain. Nat Neurosci. 2011;14:1345–51.
Hellebrekers DM, Jair KW, Viré E, Eguchi S, Hoebers NT, Fraga MF, et al. Angiostatic activity of DNA methyltransferase inhibitors. Mol Cancer Ther. 2006;5:467–75.
Acknowledgements
This work was supported by grant IP2010 052170 from the Ministry of Science and Higher Education (Poland). We thank Marcela White (Brain Tumor Tissue Bank, London Health Sciences Centre, London, Ontario, Canada) for her assistance with the procurement of high-quality PA and GBM specimens.
Author information
Authors and Affiliations
Contributions
MM designed the study, performed research, interpreted the data and wrote the manuscript; JM designed the study and performed data analysis; KW, SKK, MS, MK, IAC performed research, KJ performed data analysis; MB, JS, KK, and WG analyzed and interpreted the patient data, MZ performed data analysis and interpreted data, BK designed the study, interpreted the data and wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
J. Mieczkowski is a co-founder of and shareholder in Genegoggle. The rest of the authors declare no competing interests.
Ethics approval and consent to participate
The use of patient tissues was permitted by the Ethical Committee of the respective hospitals (14/KBE/2012 and 8/2012).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Professor Bertrand Joseph
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Maleszewska, M., Wojnicki, K., Mieczkowski, J. et al. DMRTA2 supports glioma stem-cell mediated neovascularization in glioblastoma. Cell Death Dis 15, 228 (2024). https://doi.org/10.1038/s41419-024-06603-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-024-06603-y
- Springer Nature Limited