Abstract
Histone acetylation is essential for initiating and maintaining a permissive chromatin conformation and gene transcription. Dysregulation of histone acetylation can contribute to tumorigenesis and metastasis. Using inducible cre-recombinase and CRISPR/Cas9-mediated deletion, we investigated the roles of the histone lysine acetyltransferase TIP60 (KAT5/HTATIP) in human cells, mouse cells, and mouse embryos. We found that loss of TIP60 caused complete cell growth arrest. In the absence of TIP60, chromosomes failed to align in a metaphase plate during mitosis. In some TIP60 deleted cells, endoreplication occurred instead. In contrast, cell survival was not affected. Remarkably, the cell growth arrest caused by loss of TIP60 was independent of the tumor suppressors p53, INK4A and ARF. TIP60 was found to be essential for the acetylation of H2AZ, specifically at lysine 7. The mRNA levels of 6236 human and 8238 mouse genes, including many metabolism genes, were dependent on TIP60. Among the top 50 differentially expressed genes, over 90% were downregulated in cells lacking TIP60, supporting a role for TIP60 as a key co-activator of transcription. We propose a primary role of TIP60 in H2AZ lysine 7 acetylation and transcriptional activation, and that this fundamental role is essential for cell proliferation. Growth arrest independent of major tumor suppressors suggests TIP60 as a potential anti-cancer drug target.
Similar content being viewed by others
Introduction
Chromatin modifications affect DNA accessibility and are essential for the activation of gene expression [1,2,3,4,5,6]. Histone lysine acetylation mediated by lysine acetyltransferases (KATs) is one of the most abundant histone modifications and is generally associated with actively transcribed gene loci [7,8,9]. Genetic alterations of genes encoding histone acetyltransferases are found in a wide range of cancers [10]. Targeting the histone acetyltransferases KAT6A and KAT7, members of the MYST family of histone acetyltransferases has emerged as a possible avenue for the treatment of cancer [11, 12].
HIV tat interacting protein, 60 kDa (TIP60) is another member of the MYST family of histone acetyltransferases. TIP60 has been proposed to be involved in a range of cellular processes, including embryonic stem (ES) cell self-renewal [13], DNA damage response [14, 15], transcriptional co-activation, acetylating both histones [16], and transcription factors, including the MYC oncoprotein [17, 18], steroid hormone receptors [19] and the tumor suppressor p53 [20, 21].
TIP60 has been reported to acetylate histone H2A on lysines 5 and 15 (H2AK5, H2AK15) as well as H2Av, H4K5, H4K8, H4K12, H4K16, and H3K14, largely in cell-free and RNAi experiments [16, 22,23,24,25]. In contrast, other MYST histone acetyltransferases appear to have a more specific histone acetylation activity in vivo [26,27,28,29,30,31,32].
In the context of HIV infection, TIP60 appears to promote cell death, and HIV-1 tat binding to TIP60 facilitates its degradation resulting in impaired TIP60-mediated apoptosis [33, 34]. Overexpression of catalytically inactive TIP60 leads to defects in DNA repair and decreased apoptosis upon γ-irradiation [14]. Similarly, siRNA knockdown of TIP60 impairs the UV-induced DNA damage response and apoptosis [35]. Relevant to the regulation of apoptosis, TIP60 has been reported to acetylate p53 at lysine 120, which is thought to be required for p53-induced apoptosis [20, 21]. In contrast, germline deletion of Tip60 in mice causes developmental arrest and apoptotic cell death in peri-implantation mouse embryos [36], indicating that in this instance TIP60 is required for cell survival. The conflicting reports on the role of TIP60 in promoting apoptosis or cell survival have not been resolved.
TIP60 appears to either promote or retard cancer progression. Heterozygous loss of Tip60 accelerates tumor progression in mouse models of breast cancer [37] and lymphoma [38]. Reduced levels of TIP60 are associated with metastatic colorectal cancer [39], metastatic gastric cancer [40], and with poor outcomes in melanoma patients [41]. In contrast, in prostate cancer, high levels of nuclear TIP60 correlate with hormone treatment-refractory disease [42]. The seemingly contradictory reports of TIP60 in various cancers may reflect context-specific functions, affected by parameters such as cancer type and stage, or they may result from changes in TIP60 levels/function that are a consequence rather than a cause of certain oncogenic processes.
Here we used inducible genetic deletion of the TIP60 gene in human and mouse cells to investigate its role in cell proliferation, cell death, histone acetylation, and transcriptional regulation.
Materials and methods
Mice
Tip60fl/+ mice were generated by CRISPR/Cas9 on a C57BL/6 J background, using techniques previously described [43, 44]. Briefly, two sgRNAs of the sequence GTACGGAGATGATCCGGGCG and TGGAAGCTACGCCTGCAACT were used to insert loxP sites flanking exons 3 and 4 of the Tip60 gene (Figure S1). The deletion of exon 3 and 4 is predicted to result in a frameshift and disruption of proper TIP60 protein synthesis. Successful targeting was confirmed by sequencing. Tip60fl/+ mice were crossed to Rosa26-CreERT2 transgenic mice [45] on a C57BL/6 background and used to isolate Tip60fl/fl;ERT2 MEFs.
Trp53–/– mice [46] and Cdkn2a–/– mice [47] were used to isolate MEFs. All mice were on a C57BL/6 inbred background.
Male and female young adult mice were used. Mice were genotyped by PCR; details are described in the supplement. Genotyping primers are listed in the supplement Methods Table 1.
Experiments were performed with the approval of the Walter and Eliza Hall Institute for Medical Research Animal Ethics Committee and according to the Australian code of practice for the care and use of animals for scientific purposes.
Cells
Murine embryonic fibroblasts (MEFs) were cultured in MEF medium (DMEM, 10% FBS, and 2 mM GlutaMAXTM) at 37 °C, in 5% CO2, and 3% O2 (near-physiological tissue oxygen concentration). HEK293T, HEK293, and U2OS cells were cultured in MEF medium at 37 °C and 5% CO2. To induce recombination of the Tip60fl locus, Tip60fl/fl;ERT2 and Tip60+/+;ERT2 control MEFs were treated with 250 nM 4-OHT for a maximum of 5 days, if not indicated otherwise, and consequently grown in 4-OHT free MEF medium.
MEF isolation
Briefly, E13.5 embryos were dissected and washed in PBS. Head, extremities, and liver were removed, and the torso was collected in MEF medium, supplemented with 1% (v/v) penicillin and streptomycin (Gibco, 15140), and minced by pipetting several times consecutively through 18 G, 21 G, and 24 G needles. The cells were centrifuged with low speed (500 × g) for 5 min and resuspended in warm MEF medium supplemented with penicillin and streptomycin and plated onto gelatin-coated 10 cm Petri dishes.
Inducible Tip60 CRISPR knockout cell lines
Dox-inducible CRISPR knockout cell lines were generated as described [48, 49]. Two single guide RNAs, sgRNA1 GCCGGCACCGCCTCAAGCCG and sgRNA2 CTGCCTCCCTACCAGCGCCG, were used to target exons 8 and 11 of the human TIP60 gene and murine Tip60 gene. Cells were treated with 1 μg/ml dox for the indicated times. CRISPR indels were analyzed using the primers displayed in the supplement Methods Table 2. Indel frequencies were determined by NGS and calculated as a proportion of sequences with indels of all sequences obtained. Only sequences with at least 25 reads were included in this calculation.
SDS-PAGE and immunoblotting
Detailed protocols for acid histone extraction and quantification, cell fractionation protein extraction, and SDS-PAGE and immunoblotting can be found in the supplement. Histones were acid extracted, as described [50] with slight modifications. Cells were fractionated using a subcellular protein fractionation kit for cultured cells (Thermo Scientific, 78840) according to the manufacturer’s instructions. SDS-PAGE and immunoblotting were performed using the Li-Cor system (LI-COR Biosciences), details are described in the supplement. Antibodies are listed in the supplement Methods Table 3.
SA-β-galactosidase staining by X-gal
SA-β-galactosidase staining was performed with a senescence β-galactosidase staining kit (CST, 9860) according to the manufacturer’s instructions. SA-β-galactosidase positive and negative cells were counted in triplicate in three images with at least 300 nuclei per replicate using ImageJ. Cells were irradiated with 32 Gy and cultured for 5 days prior to staining as positive controls.
RT-qPCR
RT-qPCRs were performed in triplicate. RNA was isolated using RNeasy Kit (Qiagen, 74106) according to the manufacturer’s instructions. Details of the RT-qPCR procedure can be found in the supplement. RT-qPCR primers are listed in the supplement Methods Table 4.
RNA-seq
RNA-seq was performed on Tip60fl/fl;ERT2 and Tip60+/+;ERT2 MEFs induced with 4-OHT for 3 days and 5 days, as well as HEK293g1/C9, HEK293g2/C9, and guide-only HEK293g1 and HEK293g2 controls induced with dox for 3 days. RNA-seq was performed on 2–3 replicates each of U2OSg1/C9, U2OSg2/C9, and guide-only U2OSg1 and U2OSg2 control cells, in total 6 replicates of TIP60 deleted cells and 5 replicates of control cells, induced with dox for 4 days and with D. melanogaster S2 cell spike-in. Details of the RNA-seq procedure can be found in the supplement.
CUT&Tag sequencing
CUT&Tag was performed on replicate cultures and 100,000 cells per sample, 43,000 human cells each of U2OSg1/C9 and U2OSg2/C9 either treated with doxycycline for 4 days (KO) or untreated (CTL) and 57,000 D. melanogaster cells as a spike-in, in total two replicates of TIP60 deleted cells and two replicates of control cells, as described by the Henikoff laboratory [51] with slight modifications and described in detail in the supplement. The CUT&Tag method produces results similar to ChIP-seq. Briefly, cells are bound by magnetic beads, permeabilized, and subjected with intervening wash steps to incubation with a primary antibody against the target of interest (e.g., H2AZ), followed by a secondary antibody incubation and then pAG-conjugated transposase enzyme (pAG-Tn5). pAG-Tn5 binds to the antibody (target) and is activated. Tn5 cuts the DNA and inserts DNA oligomers (previously loaded on the Tn5) site-specifically into the genome and thereby produces genomic fragments near the position of the target protein. Oligomer-tagged (“tagmented) fragments are recovered, amplified, and sequenced. D. melanogaster S2 cells as a spike-in was used to enable normalization and quantification of potential global changes in read coverage between control and TIP60 deleted cells.
Flow cytometry analysis
For flow cytometry analysis, including BrdU incorporation, live/cell death cell assay, and γH2AX staining, cells were collected, stained, and analyzed as described in the supplement.
Live-cell imaging
Cells were imaged using a live-cell imaging system (IncuCyte S3) at 37 °C and 5% CO2 with a CMOS camera and ×10 objective for up to 4 days. For 3D confocal live-cell time-lapse imaging, cells were grown in DKSH Ibidi 8-well chamber (DKSH, 80826) and stained with 1 μM SiR-DNA stain and 1 μM verapamil (Spirochem, SC-007). Cells were imaged every 5 min for a total of 24 h using a Leica SP8 and images were analyzed using ImageJ.
Statistical analysis
Statistical analysis was carried out using the Prism v8.0a software. Statistical methods are stated in the figure legends. Data are presented as mean ± SEM, unless indicated otherwise.
Results
To assess the role of TIP60 we selected cell types to capture human and mouse cells, normal cells (mouse embryonic fibroblasts, MEFs), immortalized cells (HEK293), and cancer cells (U2OS). U2OS cells are osteosarcoma cells, that can be considered as a cancerous equivalent of primary isolates of fibroblasts [52]. Like MEFs, HEK293 and U2OS cells are of mesodermal origin.
Tip60 deletion in mouse fibroblasts causes cell cycle arrest
Conditional Tip60 knockout mice were generated (Figure S1A) and crossed to Rosa26-CreERT2 (ERT2) mice [45]. Tip60fl/fl;ERT2 mouse embryonic fibroblasts (MEFs), Tip60fl/+;ERT2 MEFs, and ERT2 control MEFs were isolated and induced with 4-hydroxy-tamoxifen (4-OHT) for 1–5 days resulting in deletion of exon 3 and 4 and a frameshift of Tip60 (Tip60iKO/iKO;ERT2 Figure S1B). No Tip60 mRNA could be detected in Tip60iKO/iKO;ERT2 MEFs after 3 days of 4-OHT treatment (Fig. 1A).
A RT-qPCR assessment of Tip60 mRNA levels normalized to the housekeeping (Hk) gene Hsp90ab1 in Tip60fl/fl;ERT2 MEFs induced with 4-hydroxy-tamoxifen (4-OHT) for 3 days, or treated with ethanol (EtOH) vehicle control (means ± SEM, n = 3 cell isolates from individual embryos per genotype, unpaired two-tailed t test). B Proliferation of Tip60fl/fl;ERT2 MEFs. Knockout of Tip60 was induced (Tip60iKO) by treatment for 3 days with 4-OHT, or cells were treated with EtOH vehicle control before culture in normal growth medium (means ± SEM, n = 3, unpaired two-tailed t test on slopes of log-transformed values). C Representative phase-contrast images of Tip60fl/fl;ERT2 MEFs 5 days after 4-OHT induction vs. heterozygous and EtOH controls. D Quantification of indel frequencies after doxycycline (dox) induction of sgRNA expression in iC-Tip60 MEFs for sgRNA#1 (g1/C9) and sgRNA#2 (g2/C9) and sgRNA-only controls (g-only CTL) (means ± SEM, n = 3, one-way ANOVA). E Proliferation of iC-Tip60 MEFs induced to deleted Tip60 with dox treatment vs. untreated cells (expressing Cas9 only) and g-only CTL controls (means ± SEM, n = 3, unpaired two-tailed t test on slopes of log-transformed values). F Representative phase-contrast images of iC-Tip60 MEFs 5 days after dox induction vs. untreated cells (expressing Cas9 only) and g-only CTL controls. Circles represent individual data points of cell isolates from individual embryos (A, D).
Treatment of Tip60fl/fl;ERT2 and Tip60fl/+;ERT2 MEFs with 4-OHT for 3 days followed by passaging in culture for 2 weeks in the absence of 4-OHT resulted in complete growth arrest in Tip60iKO/iKO;ERT2 MEFs, whereas heterozygous Tip60+/iKO;ERT2 control MEFs continued to proliferate normally (Fig. 1B). Tip60iKO/iKO;ERT2 MEFs retained the ability to adhere to the cell culture plastic after passaging and no excessive floating cells were observed, suggesting an absence of cell death. Tip60iKO/iKO;ERT2 MEFs displayed a large and flat morphology typical of senescent MEFs (Fig. 1C; compare to [53, 54].
In parallel, we deleted Tip60 by inducible CRISPR/Cas9 genome editing, involving doxycycline (dox) inducible sgRNA expression marked with GFP, constitutively expressed mCherry-tagged Cas9 [48], using two sgRNAs targeting exons 8 and 11 of TIP60 in human and mouse cells (Figure S1C, D; referred to as g1/C9 and g2/C9 or g/C9 for merged data). Cell lines containing only sgRNA, lacking Cas9 (referred to as g1 and g2 or g-only for merged data) and either treated with dox or not treated (g-only dox or g-only CTL), and cell lines containing the sgRNA and Cas9 constructs, but not induced to express the sgRNA (referred to as g/C9 CTL), were used as controls. After dox treatment, insertion and deletion (indels) mutations of Tip60 were analyzed in the bulk population via high-throughput sequencing (Figure S1E). After 48 h of doxycycline induction, the indel frequency for both sgRNAs plateaued at around 85% in MEFs (Fig. 1D). sgRNA-only controls showed no indel activity. Assessment of the top four predicted off-target loci for each sgRNA, identified by WTSI Genome Editing (WGE) [55], showed no indel activity (Figure S1F). RT-qPCR analysis showed a reduction of Tip60 mRNA in bulk MEFgC/9 cultures after 3 days of induction (Figure S1G); however, small indels are expected to cause a loss of function even without a reduction in mRNA. Currently commercially available TIP60 antibodies (Invitrogen (PA5–23290), Sigma (SAB4500118), Abcam (ab135490), SantaCruz (sc-166323), CST (12058 S)) did not detect endogenous TIP60 protein in our experiments, preventing us from assessing TIP60 protein levels.
The inducible CRISPR Tip60 mutated MEFs (iC-Tip60) were continuously cultured in dox supplemented medium for 9 days. iC-Tip60 cells displayed growth arrest after 3 days of dox treatment compared to controls (Fig. 1E). After 5 days of doxycycline treatment, iC-Tip60 MEF morphology closely resembled ERT2-mediated Tip60 knockout MEFs (Fig. 1F).
The slight outgrowth of cells observed after 9 days of dox treatment was consistent with an approximately 85% efficiency of the CRISPR/Cas9 genome editing system (Fig. 1D). Analysis of cells grown for an extended period in dox supplemented medium revealed a partial loss of sgRNA or Cas9 expression (Figure S2). Cells that either did not integrate or lost or silenced the sgRNA or Cas9 constructs would be expected to grow similarly fast as controls.
Although TIP60 has been reported to play a central role in the DNA damage response [14, 15], we did not observe an increase in γH2AX staining, used as an indication of DNA damage, in Tip60iKO/iKO;ERT2 MEFs 3 days after 4-OHT induction (Figure S1H).
TIP60 deletion in human cells causes cell cycle arrest and failure to progress to metaphase
TIP60 was mutated in U2OS and HEK293 cells using dox-inducible CRISPR/Cas9 genome editing. Continuously culturing the cells in dox resulted in a strong reduction in TIP60 mRNA after 3 days (Fig. 2A) and a reduction in cell growth (Fig. 2B).
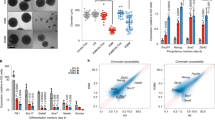
A RT-qPCR assessment of TIP60 mRNA levels normalized to the Hk gene Hsp90ab1 in iC-TIP60 U2OS and HEK293 cells induced with dox for 3 days vs. untreated control cells expressing Cas9 only (means ± SEM; n = 2, unpaired two-tailed t test). B Proliferation of iC-TIP60 U2OS and HEK293 cells vs. untreated control (g/C9 CTL, Cas9 expressing only) and g-only CTL controls (means ± SEM, n = 4, unpaired two-tailed t test on slopes of log transformed values). C Representative phase-contrast images of iC-TIP60 U2OS and HEK293 cells 5 days after dox induction vs. untreated control (Cas9 expressing only) and g-only CTL control. D Flow cytometric cell cycle analysis of BrdU incorporation and DNA content of iC-TIP60 U2OS cells vs. untreated control cells (g/C9 CTL, Cas9 expressing only). Cells with >4n DNA content suggested endoreplication cells. (Means ± SEM, n = 4, two-way ANOVA). E Representative maximum intensity projection still images of 3D confocal live-cell time-lapse imaging of iC-TIP60 U2OS cells after 3 days of dox treatment vs. untreated control (Cas9 expressing only). Further IncuCyte live-cell time-lapse imaging is displayed in Figure S3 and Movies 1A−C. Confocal live-cell imaging is displayed in Movies 2A, B. Assessment of cell death is displayed in Figure S5. Circles represent individual data points of replicate experiments using two sgRNAs (A, D).
Live-cell imaging and tracking of individual cell divisions on iC-TIP60 U2OS and control cells revealed that most dox-treated iC-TIP60 U2OS cells underwent a final cell division before complete proliferation arrest (Figure S3A and Movies 1A–C). Some cells maintained normal division rates, consistent with <100% efficiency of CRISPR/Cas9 genome editing (Fig. 1D, S2A).
Brightfield microscopy of iC-TIP60 U2OS cells revealed a senescent cell morphology. In contrast, iC-TIP60 HEK293 cells displayed reduced adherence to the tissue culture surface and largely lost contact after 4–5 days of dox treatment (Fig. 2C). TIP60 deletion led to a 2.4-fold (TIP60iKO/iKO;ERT2 MEFs) to fivefold (iC-TIP60 U2OS cells) increase in senescence-associated-β-galactosidase (β-gal) positive cells (Figure S3B). Further analysis of senescence markers revealed only a partial senescence footprint, with increased levels of p21 and p53 in the nucleus of iC-TIP60 U2OS cells, while other previously reported senescence-associated markers, such as TNF-α, lamin B1, MMP-3, RB, γ-H2AX, and macroH2A remained unchanged and IL-6 was not expressed in U2OS cells (Figure S4A–D). CDKN2A (coding for p16 and p19) is deleted in U2OS cells [56] and therefore could not be used as a senescence marker in these cells.
Three different tests assessing cell death (live/dead cell assay, TMRE, and annexin V flow cytometric analysis), showed that cell survival was not affected in U2OS or adherent and floating HEK293 cells after 3 days of doxycycline treatment (Figure S5A–C).
Loss of TIP60 in iC-TIP60 U2OS cells led to a twofold decrease in S-phase, a 47% increase in G2/M-phase cells, a 34% increase in G1 phase cells, and an eightfold increase in cells with a DNA content >2 N, indicating endoreplication as assessed by DNA content and BrdU incorporation (Fig. 2D). High-resolution 3D confocal live-cell imaging of iC-TIP60 U2OS cells revealed severely disturbed mitoses with chromosome condensation prolonged from the 25 min (control) to 4 h 45 min during mitosis, failure to arrange chromosomes in a metaphase plate and ultimate failure to divide (Fig. 2E and Movies 2A, B).
TIP60 is essential for histone H2AZ lysine 7 acetylation
We assessed histone acetylation at lysines that had previously been proposed to be the histone acetylation targets of TIP60, including H2A-K5, H2A-K15, H4K5, H4K8, H4K12, H4K16 and H3K14 [16, 22,23,24,25], as well as lysines on H2AZ, a homolog of Drosophila melanogaster H2Av and S. cerevisiae Htz1 [57, 58], which were reported to be TIP60 acetylation targets [24, 59, 60]. Cell-free acetylation assays with recombinant human TIP60 and histone H2AZ confirmed that the anti- H2AZK7ac antibody used recognized acetylated H2AZ and did not cross-react with unmodified H2AZ (Fig. 3A). Western blotting of acid extracted proteins from Tip60iKO/iKO;ERT2 and control MEFs 3 days post 4-OHT induction (Fig. 3B) showed that loss of TIP60 caused a 5-fold reduction in pan-H2AZK4-7-11ac, 5-fold in H2AZK7ac, and 2-fold reduction in H2AZK4ac levels (Fig. 3C). Total H2AZ protein levels remained unchanged. Other proposed TIP60 histone acetylation targets showed no changes, apart from a 20% reduction in H4K5ac (Fig. 3C, S6A). No differences were observed between heterozygous Tip60+/iKO;ERT2 MEFs and wild-type controls.
A Cell-free acetylation assay analyzed by Western blot shows full-length recombinant H2AZ histone, with (lane 1–4) and without (lane 5) recombinant TIP60 (rec-Tip60). B Representative Western blots of acid extracted proteins from Tip60fl/fl;ERT2 MEFs treated with vehicle (EtOH, −) or 4-OHT (+) for 3 days and probed for histone and histone acetylation levels as indicated. Total H3 was used as a loading control. C Fold change of histone lysine acetylation levels normalized to total H3 protein after 3 days of EtOH or 4-OHT treatment. Shown are Tip60+/iKO;ERT2 control MEFs and Tip60iKO/iKO;ERT2 MEFs. D Fold change of histone lysine acetylation levels normalized to H3 between iC-Tip60 and control MEFs. E Fold change of histone lysine acetylation levels normalized to H3 between iC-TIP60 and control HEK293 cells after 3 days of dox treatment and time-course of H2AZK7ac decline. Data from sgRNA#1 and sgRNA#2 were combined. Means ± SEM of 2 to 8 independent experiments are shown (C–E). Circles represent individual data points of replicates (C–E). Data were log-transformed and analyzed by unpaired two-tailed t test (C, E left panel) or one-way ANOVA with Benjamini and Hochberg correction (D, E right panel). Experiments assessing acetylation of histones H2A, H3, and H4 at specific lysine residues are displayed in Figure S6 and full-size Western blots are displayed in the supplemental information.
iC-Tip60 MEFs showed a progressive decrease in H2AZK7ac over 3 days to sevenfold and a twofold decline in H2AZK4ac (Fig. 3D). The rapid onset of reduction in H2AZ acetylation after genomic deletion of Tip60 supports a direct dependence on TIP60. We further observed a ~29% reduction in H4K12ac, but no other changes (Fig. 3D, S6B).
Similarly, TIP60 mutation in HEK293 cells caused a 4-fold reduction in H2AZK7ac, a 2-fold decrease in H2AZK4ac, and a 13-fold reduction H2AZK4-7-11ac (Fig. 3E). H4K16ac was reduced by 37% (Fig. 3E), but other histone lysine acetylation levels were unchanged (Figure S6C).
Analysis of H2AZ, H2AZK4–7ac, H2AZK4-7-11ac, H4K8ac, and H4K16ac, levels and distribution in the genome of iC-TIP60 U2OS cells by CUT&Tag sequencing with D. melanogaster S2 cell spike-in revealed a global reduction of H2AZ acetylation by 80–90%, using two separate antibodies against H2AZK4–7ac and H2AZK4-7–11ac (Fig. 4A and S7A). H2AZ and acetylated H2AZ were observed to form peaks over the TSS of genes (Figure S7B), consistent with the previously described patterns [61,62,63,64,65,66,67,68,69]. Differential occupancy analysis at the TSS of genes (±1000 bp) revealed a global reduction with an average reduction of 90% and 92% in H2AZK4-7-11ac and H2AZK4–7ac, respectively (Fig. 4B). Occupancy of unmodified H2AZ, H4K8ac, and H4K16ac at the TSS remained unchanged. The data point distribution of fold changes for the TSS regions of all genes suggested that histone acetylation changes were not locus-specific, but rather occurred at all or most genes that were decorated with the respective mark (Fig. 4B).
A Total genome-wide CUT&Tag sequencing reads in iC-TIP60 U2OS and control cells as a fraction of spike-in S2 D. melanogaster cells normalized to control U2OS cells using antibodies detecting H2AZK4-7-11ac, H2AZK4–7ac, total H2AZ, H4K8ac, and H4K16ac (means, n = 2, circles represent individual data points of replicates). B CUT&Tag sequencing reads quantified in a window ±1000 bp of the TSS of all genes. The distributions of log2 fold changes in read counts (logFC) between iC-TIP60 U2OS and control cells are displayed as violin plots, with respective box plots inside. Boxes show median and quartiles with boxplot whiskers extending 1.5 inter-quartile ranges beyond the quartiles. P values of fry gene set tests for global shifts in occupancy (two-sided). Related data are displayed in Figure S7.
Western blotting of histones and modified histones on the cytosolic, soluble nuclear, and chromatin-bound fractions revealed that chromatin-bound total H2AZ levels remained unchanged in iC-TIP60 U2OS cells 3 days after dox treatment, while H2AZac levels were reduced by 85% (Figure S8A, B).
TIP60 is required for normal gene expression in mouse and human cells
Since H2AZ acetylation is reported to play an important role in gene expression in yeast [59], we examined the effect of TIP60 mutation on transcription. RNA-seq analysis of Tip60+/+;ERT2 control and Tip60fl/fl;ERT2 MEFs 3 days and 5 days after 4-OHT treatment revealed 8238 differentially expressed genes in Tip60iKO/iKO;ERT2 cells compared to control cells (Table S1, Fig. 5A). Differentially expressed genes after 3 days and 5 days were highly correlated (p = 5×10–5; Figure S9A). For the subsequent analysis, we focused on genes differentially expressed 3 days after induction of Tip60 deletion to avoid secondary effects of loss of TIP60, at which time a total of 6311 genes were differentially expressed (Table S1). Among the top 50 differentially expressed genes ranked by FDR, 92% were downregulated (Fig. 5B) and among the top 500 differentially expressed genes 77.6% were downregulated, suggesting a bias towards transcriptional repression in the absence of TIP60 (Table S2).
A–D RNA-seq results of Tip60iKO/iKO;ERT2 vs. Tip60+/+;ERT2 control MEFs after 3 days and 5 days (A, combined analysis) or after 3 days of 4-OHT treatment B–D to induced Tip60 deletion (n = 3 cell isolates from individual embryos per genotype). A Mean-difference plot showing the log2 fold change vs. average log2 counts-per-million of each mRNA in Tip60iKO/iKO;ERT2 MEFs vs. Tip60+/+;ERT2 control MEFs after treatment for 3 and 5 days with 4-OHT. Differentially expressed genes (FDR < 0.05) are indicated. B Heatmap of the top 50 differentially expressed (DE) genes by FDR in Tip60iKO/iKO;ERT2 vs. Tip60+/+;ERT2 control MEFs after 3 days of 4-OHT treatment. C Expression level changes of the Trp53 (p53) gene and p53 target genes and cyclin-dependent kinase inhibitor genes Cdkn2b and Cdkn2c in the absence of TIP60. D Top 10 biological processes (GO terms) and gene expression and transcription processes (GO terms) enriched for genes differentially expressed (DE) in the absence of TIP60 in MEFs. RNA-seq results of Tip60iKO/iKO and control MEFs, including genes encoding other histone acetyltransferases, TIP60 complex proteins, cell adhesion proteins, extracellular matrix proteins, and gene expression regulators, are displayed in Figure S9.
No compensatory upregulation of another KAT gene was seen in the absence of TIP60 (Figure S9B). Only minor changes in mRNA levels of the TIP60 protein complex subunits were seen (Figure S9B).
p53 and its target genes, including Cdkn1a (encoding p21CIP1/WAF1), were generally downregulated in the absence of TIP60 in MEFs (Fig. 5C), suggesting that the p53 pathway was not involved in the cell cycle arrest mechanism caused by the absence of TIP60. Interestingly, Cdkn2b and Cdkn2c (encoding p15INK4B and p18INK4C respectively) were upregulated (Fig. 5C). Cell adhesion genes and collagen genes were up to 25-fold upregulated (Figure S9C), consistent with a senescent cell phenotype. This notably included cell-substrate adhesion genes, such as integrin and collagen genes, with 30% of the genes within this biological process upregulated (p = 0.0002; Table S3).
Among the top 10 enriched biological processes, 8 were downregulated, including gene expression, protein translation, and associated processes (Fig. 5D; Table S3), consistent with cell cycle arrest and reduced requirement for metabolism and protein synthesis after the loss of TIP60. Transcription and gene expression processes overall were downregulated in the absence of TIP60 (Fig. 5D; Table S3). Downregulated pathways included p53-signaling, cell cycle regulation, apoptosis, and necroptosis (Table S4), congruent with the observation that TIP60 knockout MEFs neither proliferate, nor undergo apoptosis or other forms of cell death, but rather enter cell cycle arrest.
In HEK293 cells, the use of sgRNA#1 or sgRNA#2 resulted in highly correlated expression changes 3 days after induction of TIP60 deletion (Figure S10A) and therefore the data from the two different sgRNAs were combined for all subsequent analyses.
In the absence of TIP60, 6236 genes were differentially expressed between iC-TIP60 and control HEK293 cells (Fig. 6A; Table S5).
A–D RNA-seq results from iC-TIP60 vs. control HEK293 cells after 3 days of 4-OHT treatment to induce Tip60 deletion (n = 4). A Volcano plot displaying genes significantly differentially expressed (DE) in iC-TIP60 vs. control HEK293 cells (FDR < 0.05). Results from both sgRNAs were similar and were thus combined. B Expression level changes of protocadherin genes in iC-TIP60 vs. control HEK293 and schematic drawing of cell-cell adhesion interactions upregulated in the absence of TIP60. FDR values are indicated in the graph. Gray bars represent genes not significantly changed. C Top 10 biological processes (GO terms) enriched among genes differentially expressed (DE) in the absence of TIP60. D Top 10 gene expression and transcription processes (GO terms) enriched among genes differentially expressed (DE) in iC-TIP60 vs. control HEK293 cells. Further RNA-seq data of iC-TIP60 vs. control HEK293 cells, including genes encoding histone acetyltransferases, TIP60 complex proteins, cell cycle regulators, cell senescence genes, apoptosis regulators, and cell-cell adhesion proteins are displayed in Figure S10.
Similar to MEFs, TIP60 complex subunit and KAT genes were not greatly dysregulated in HEK293 cells lacking TIP60 (Figure S10B). Cell cycle and mitosis processes and genes were predominantly downregulated in the absence of TIP60 (Figure S10C; Table S6). Similar to MEFs, cyclin-dependent kinase genes CDKN2B and CDKN2C were upregulated in HEK293 cells lacking TIP60 (Figure S10D).
TIP60 deleted HEK293 displayed a large proportion of cells that lifted off the cell culture plastic. However, the proportion of dead cells was not increased (Figure S5B), suggesting a primary failure in cell-substrate adhesion. Genes encoding cell-cell adhesion proteins, in particular of the protocadherin, cadherin, and ephrin receptor families were upregulated in the absence of TIP60 (Fig. 6B and S10E, Table S5), suggesting cell-cell adhesion increased as cell-substrate adhesion decreased. Overall, 33% of genes with the annotation “cell-cell adhesion” were upregulated in the absence of TIP60 (Table S6).
Six of the top 10 biological processes were downregulated in HEK293 cells lacking TIP60. These included metabolic processes, whereas organ development processes were upregulated (Fig. 6C, Table S6). Similarly, among the pathways affected, metabolic pathways and the cell cycle were downregulated (Table S7). Gene expression and transcription processes were comprehensively downregulated (Fig. 6D).
We performed RNA-seq on iC-TIP60 U2OS cells 4 days after induced TIP60 mutation adding in a D. melanogaster spike-in cells to enable detection of global changes in mRNA, as previously described [70]. We found a total of 12716 expressed genes, 2707 genes were downregulated, and 1555 genes were upregulated (Table S8; Figure S11A, S11B), representing a shift towards downregulated genes compared to RNA-seq results from HEK293 cells and MEFs without D. melanogaster spike-in normalization, while total mRNA levels remained similar. The directional change and the amplitude of change in gene expression in iC-TIP60 U2OS cells correlated strongly positively with gene expression changes in iC-TIP60 HEK293 cells (Figure S11C) and Tip60iKO/iKO MEFs (Figure S11D).
We integrated the iC-TIP60 U2OS RNA-seq dataset and CUT&Tag dataset to examine correlations between gene expression and H2AZ occupancy and H2AZ acetylation. H2AZ and H2AZac occupancy was detected in wild-type cells at the start of transcription of more than 99% of expressed genes (Table S9). Of the genes decorated with H2AZ and H2AZac, 35% were differentially expressed in the absence of TIP60, whereas the RNA levels of 65% were not affected by the loss of TIP60. Of the differentially expressed genes, 64% were downregulated in RNA levels (Table S9). In TIP60 wild-type U2OS cells, RNA levels correlated significantly with H2AZ occupancy (p < 10−6) and H2AZK4-K7 acetylation levels (p < 10−6; Figure S12A). Furthermore, downregulation of genes in iC-TIP60 vs. control U2OS cells correlated significantly with H2AZ (p < 10−6) occupancy and H2AZK4-K7 acetylation levels in genes that were differentially expressed in iC-TIP60 vs. control U2OS cells (p < 10−6; Figure S12B).
Loss of TIP60 causes cell cycle arrest even in the absence of the cell regulators p53, INK4A, and ARF
Our RNA-sequencing results suggested that the ARF-p53 tumor suppressor pathway (Fig. 7A) was not engaged in the cell cycle arrest induced by loss of TIP60 in MEFs. This finding could potentially have bearing on the suitability of TIP60 as a target for cancer therapeutics, as the TP53 gene is mutated in 42% of cancers [71], and CDKN2A, encoding INK4A and ARF, is mutated in 3.6% in a pan-cancer study [71] and in up to 78% in specific types of pediatric leukemia [72]. To formally test if p53 or INK4A-ARF were required for cell cycle arrest induced by loss of TIP60, we deleted the Tip60 gene in MEFs lacking p53 (Trp53–/– MEFs) or INK4A-ARF (Cdkn2a–/– MEFs). In Trp53–/– MEFg1/C9 cells indel frequency reached 80% after only 24 h of doxycycline induction (Fig. 7B). TIP60 deletion in Trp53–/– MEFs or in Cdkn2a–/– MEFs resulted in growth arrest, indicating a p53 and INK4A-ARF-independent mechanism of cell cycle arrest (Fig. 7C). The morphology was indistinguishable from Tip60 only deleted MEFs (Figs. 7D vs. 1C, F).
A Simplified schematic displaying the regulatory interplay between ARF, MDM2, and p53. B Quantification of indel frequencies in iC-Tip60 Trp53–/– MEFs. (Means ± SEM, n = 3, one-way ANOVA). C Proliferation of iC-Tip60 Trp53–/– MEFs and iC-Tip60 Cdkn2a–/– MEFs vs. untreated controls (expressing Cas9 only) and g-only controls (means ± SEM, n = 6, unpaired two-tailed t test on slopes of log-transformed values). D Representative phase-contrast images iC-Tip60 Trp53–/– MEFs and iC-Tip60 Cdkn2a–/– MEFs 5 days after dox treatment vs. untreated controls. E Fold change of total H2AZ and histone acetylation levels normalized to H3 in iC-Tip60 Trp53–/– and iC-Tip60 Cdkn2a–/– vs. g-only control MEFs after 3 days of dox treatment (means ± SEM, n = 2, unpaired two-tailed t test on log-transformed values). Experiments assessing histones H2A, H3, and H4 lysine acetylation levels are displayed in Figure S6D.
Similar to wild-type MEFs, deletion of Tip60 in Cdnk2a–/– or Trp53–/– MEFs resulted in a reduction in H2AZK4, H2AZK7, and H2AZK4–11 acetylation (Fig. 7E). In addition to the profound effects on H2AZ acetylation, we found a modest reduction in H4K12ac and H4K16ac in Tip60 deleted Cdkn2–/– MEFs compared to control MEFs (Fig. 7E), but not in Tip60 deleted Trp53–/– MEFs (Figure S6D).
Discussion
In this study, we induced TIP60 deletion in normal mouse cells, immortalized human cells, and human cancer cells, as well as cells lacking the tumor suppressors p53, INK4A, and ARF to investigate the role of TIP60. TIP60 deletion caused cell growth arrest, but cell survival was not affected.
In our study loss of TIP60 caused a p53-independent cell cycle arrest. In contrast, TIP60 has previously been reported to induce cell cycle arrest, either as a co-activator of p53 at target genes such as CDKN1A (p21CIP1/WAF1) [73], or in a p53-independent manner [74] after genotoxic stress in the context of DNA repair. The active role of TIP60 in inducing cell cycle arrest in the context of genotoxic stress appears to involve specific molecular mechanisms, including interaction with DNA damage repair proteins [15]. Our data, showing that the loss of TIP60 causes cell cycle arrest in a range of cell cycle phases, suggest that the observed cell cycle arrest is not due to effects on a single cell cycle regulatory mechanism, but is rather mediated through a broad transcriptional dysregulation of multiple genes required in different phases of the cell cycle for a range of cellular mechanisms, in a manner independent of tumor suppressor and cell cycle regulators. The effects of TIP60 deletion might differ depending on the phase of the cell cycle when individual cells first experience critically low levels of TIP60 and deficiencies in gene expression. Thus, only a small proportion of cells undergo endoreplication due to the lack of TIP60, while the majority undergo cell cycle arrest without endoreplication. The same effect might be the reason for our observation that not all cells display cellular senescence. γH2AX staining indicative of DNA damage was not increased in Tip60 deleted MEFs, excluding accumulation of DNA damage as a factor causing cell cycle arrest.
Our data identify the specific lysine residues that rely on TIP60 for acetylation. Loss of TIP60 results in a marked decline in H2AZK7 acetylation and a reduction in H2AZK4 acetylation, consistent with previous reports proposing that TIP60 is required for the acetylation of H2A.Z and gene activation in human and mouse cells [75, 76]. Apart from the profound reduction in H2AZ acetylation, we observed only a modest reduction in acetylation levels at some H4 lysine residues in the absence of TIP60. However, the effects on H4 acetylation were not present in all cell types and the specific H4 residue affected was not consistent between cell types. Various other HATs are known to acetylate H4 lysines [77]. Redundancy between those HATs might conceal a potential effect of loss of TIP60 on global H4 acetylation.
Global chromatin-bound H2AZ levels were not reduced upon TIP60 deletion in U2OS cells, indicating either no effect of TIP60 deletion on H2AZ dynamics or a complete halt of H2AZ exchange.
The mitotic defect and increased endoreplication indicate that Tip60 deleted cells were able to synthesize DNA, but failed to arrange chromosomes in a metaphase plate, and a failure of chromosome segregation. These observations may relate to the knockout phenotype of the yeast homolog of TIP60, esa1. Esa1 null mutant or catalytic mutant Saccharomyces cerevisiae display a budding phenotype characterized by incomplete DNA segregation and aberrant spindle orientation [78]. In this context, it is interesting that TIP60-mediated acetylation of aurora B kinase, the catalytic subunit of the chromosome passenger complex, promotes accurate chromosome segregation in mitosis [79].
H2AZ has been shown to play an important role in centromere structure formation [80] and H2AZ depletion has been linked to abnormal nuclear chromatin structure, as well as lagging chromosomes and chromosome bridges during cell division in mammalian cells [81]. The mitosis defect upon loss of TIP60 (and 80% of H2AZac) suggests that TIP60-dependent acetylation of H2AZ may be important in centromere and spindle formation.
Our RNA-seq data on TIP60-deficient cells support a broad role of TIP60 in gene transcription, congruent with the role of H2AZ acetylation. Thousands of genes relied on TIP60 for normal expression levels in MEFs, including genes related to protein translation, ribosome biogenesis, and general gene expression. The TIP60 mutant cells phenotype is likely to be due to widespread failure of transcription rather than a small subset of genes being responsible for the phenotype. While more genes are downregulated than upregulated in response to the loss of TIP60, the amplitude of change is generally larger in the upregulated genes, suggesting that we also observed gene expression changes secondary to cellular events such as cell cycle arrest. H2AZ acetylation has been correlated with active transcription, consistent with a role of TIP60 in H2AZ acetylation and gene activation, albeit less locus-specific than previously reported [82,83,84]. H2AZ is found broadly at the transcription start site (TSS) of active and inactive genes as well as in gene bodies of inactive genes [63]. In contrast, acetylated H2AZ is shown to exclusively accumulate at the TSS of active genes [63]. Our data suggest that TIP60 is responsible for the acetylation of H2AZK7 at active genes. A comparison of the RNA-seq and the Cut&Tag data revealed a positive correlation of RNA levels with H2AZ occupancy and H2AZ acetylation. High H2AZ acetylation and H2AZ occupancy correlated with stronger downregulation in gene expression upon TIP60 mutation. However, RNA levels of 65% of genes displaying decreased H2AZ acetylation at their TSS were not significantly changed in their expression levels after TIP60 mutation. Residual H2AZ acetylation, i.e., at lysine 4, might be sufficient to keep genes active in the absence of TIP60. Furthermore, high expression levels and thus more open chromatin structure combined with active RNAPII transcription through the locus might facilitate continued transcription even upon loss of H2AZ acetylation. Lastly, H2AZ might only be more important for the onset of gene transcription but not for maintaining open chromatin structure and active gene expression.
In conclusion, we propose a mechanism by which TIP60 acts as a co-activator of gene transcription through H2AZ acetylation. Identifying H2AZK7 and H2AZK4 as endogenous histone acetylation targets of TIP60 in a number of human and mouse cell types is an important step in understanding the role of TIP60 and its histone acetylation function. Furthermore, the finding that loss of TIP60 induces cell cycle arrest independent of the tumor suppressors p53, INK4A, and ARF, and identification of H2AZK7 acetylation as a potential biomarker are essential steps for the assessment of TIP60 as a potential therapeutic target.
Data availability
Complete results from our RNA-seq studies, including complete lists of differentially expressed genes, differentially enriched GO terms and KEGG pathways, are included in the in the supplementary tables. Raw data NCBI Short Read Archive accession number GSE208383.
References
Allfrey VG, Faulkner R, Mirsky AE. Acetylation and methylation of histones and their possible role in the regulation of rna synthesis. Proc Natl Acad Sci USA. 1964;51:786–94.
Wang L, Liu L, Berger SL. Critical residues for histone acetylation by Gcn5, functioning in Ada and SAGA complexes, are also required for transcriptional function in vivo. Genes Dev. 1998;12:640–53.
Kuo MH, Zhou J, Jambeck P, Churchill ME, Allis CD. Histone acetyltransferase activity of yeast Gcn5p is required for the activation of target genes in vivo. Genes Dev. 1998;12:627–39.
Hebbes TR, Thorne AW, Crane-Robinson C. A direct link between core histone acetylation and transcriptionally active chromatin. EMBO J. 1988;7:1395–402.
Hassig CA, Tong JK, Fleischer TC, Owa T, Grable PG, Ayer DE, et al. A role for histone deacetylase activity in HDAC1-mediated transcriptional repression. Proc Natl Acad Sci USA. 1998;95:3519–24.
Kadosh D, Struhl K. Histone deacetylase activity of Rpd3 is important for transcriptional repression in vivo. Genes Dev. 1998;12:797–805.
Wang Z, Zang C, Rosenfeld JA, Schones DE, Barski A, Cuddapah S, et al. Combinatorial patterns of histone acetylations and methylations in the human genome. Nat Genet. 2008;40:897–903.
Wang Z, Zang C, Cui K, Schones DE, Barski A, Peng W, et al. Genome-wide mapping of HATs and HDACs reveals distinct functions in active and inactive genes. Cell. 2009;138:1019–31.
Stasevich TJ, Hayashi-Takanaka Y, Sato Y, Maehara K, Ohkawa Y, Sakata-Sogawa K, et al. Regulation of RNA polymerase II activation by histone acetylation in single living cells. Nature. 2014;516:272–5.
Hu Z, Zhou J, Jiang J, Yuan J, Zhang Y, Wei X, et al. Genomic characterization of genes encoding histone acetylation modulator proteins identifies therapeutic targets for cancer treatment. Nat Commun. 2019;10:733.
Baell JB, Leaver DJ, Hermans SJ, Kelly GL, Brennan MS, Downer NL, et al. Inhibitors of histone acetyltransferases KAT6A/B induce senescence and arrest tumour growth. Nature. 2018;560:253–7.
MacPherson L, Anokye J, Yeung MM, Lam EYN, Chan Y-C, Weng C-F, et al. HBO1 is required for the maintenance of leukaemia stem cells. Nature. 2020;577:266–70.
Fazzio TG, Huff JT, Panning B. An RNAi screen of chromatin proteins identifies Tip60-p400 as a regulator of embryonic stem cell identity. Cell. 2008;134:162–74.
Ikura T, Ogryzko VV, Grigoriev M, Groisman R, Wang J, Horikoshi M, et al. Involvement of the TIP60 histone acetylase complex in DNA repair and apoptosis. Cell. 2000;102:463–73.
Sun Y, Jiang X, Chen S, Fernandes N, Price BD. A role for the Tip60 histone acetyltransferase in the acetylation and activation of ATM. Proc Natl Acad Sci USA. 2005;102:13182–7.
Kusch T, Mei A, Nguyen C. Histone H3 lysine 4 trimethylation regulates cotranscriptional H2A variant exchange by Tip60 complexes to maximize gene expression. Proc Natl Acad Sci USA. 2014;111:4850–5.
Patel JH, Du Y, Ard PG, Phillips C, Carella B, Chen CJ, et al. The c-MYC oncoprotein is a substrate of the acetyltransferases hGCN5/PCAF and TIP60. Mol Cell Biol. 2004;24:10826–34.
Frank SR, Parisi T, Taubert S, Fernandez P, Fuchs M, Chan HM, et al. MYC recruits the TIP60 histone acetyltransferase complex to chromatin. EMBO Rep. 2003;4:575–80.
Brady ME, Ozanne DM, Gaughan L, Waite I, Cook S, Neal DE, et al. Tip60 is a nuclear hormone receptor coactivator. J Biol Chem. 1999;274:17599–604.
Sykes SM, Mellert HS, Holbert MA, Li K, Marmorstein R, Lane WS, et al. Acetylation of the p53 DNA binding domain regulates apoptosis induction. Mol cell. 2006;24:841–51.
Tang Y, Luo J, Zhang W, Gu W. Tip60-dependent acetylation of p53 modulates the decision between cell-cycle arrest and apoptosis. Mol cell. 2006;24:827–39.
Kimura A, Horikoshi M. Tip60 acetylates six lysines of a specific class in core histones in vitro. Genes Cells. 1998;3:789–800.
Jeong KW, Kim K, Situ AJ, Ulmer TS, An W, Stallcup MR. Recognition of enhancer element-specific histone methylation by TIP60 in transcriptional activation. Nat Struct Mol Biol. 2011;18:1358–65.
Kusch T, Florens L, Macdonald WH, Swanson SK, Glaser RL, Yates JR 3rd, et al. Acetylation by Tip60 is required for selective histone variant exchange at DNA lesions. Science. 2004;306:2084–7.
Jacquet K, Fradet-Turcotte A, Avvakumov N, Lambert JP, Roques C, Pandita RK, et al. The TIP60 complex regulates bivalent chromatin recognition by 53BP1 through direct H4K20me binding and H2AK15 acetylation. Mol cell. 2016;62:409–21.
Thomas T, Dixon MP, Kueh AJ, Voss AK. Mof (MYST1 or KAT8) is essential for progression of embryonic development past the blastocyst stage and required for normal chromatin architecture. Mol Cell Biol. 2008;28:5093–105.
Kueh AJ, Dixon MP, Voss AK, Thomas T. HBO1 is required for H3K14 acetylation and normal transcriptional activity during embryonic development. Mol Cell Biol. 2011;31:845–60.
Voss AK, Collin C, Dixon MP, Thomas T. Moz and retinoic acid coordinately regulate H3K9 acetylation, Hox gene expression, and segment identity. Dev Cell. 2009;17:674–86.
Taipale M, Rea S, Richter K, Vilar A, Lichter P, Imhof A, et al. hMOF histone acetyltransferase is required for histone H4 lysine 16 acetylation in mammalian cells. Mol Cell Biol. 2005;25:6798–810.
Gupta A, Guerin-Peyrou TG, Sharma GG, Park C, Agarwal M, Ganju RK, et al. The mammalian ortholog of Drosophila MOF that acetylates histone H4 lysine 16 is essential for embryogenesis and oncogenesis. Mol Cell Biol. 2008;28:397–409.
Akhtar A, Becker PB. Activation of transcription through histone H4 acetylation by MOF, an acetyltransferase essential for dosage compensation in Drosophila. Mol cell. 2000;5:367–75.
Kueh AJ, Eccles S, Tang L, Garnham AL, May RE, Herold MJ, et al. HBO1 (KAT7) does not have an essential role in cell proliferation, DNA replication, or histone 4 acetylation in human cells. Mol Cell Biol. 2020;40:e00506−19.
Kamine J, Elangovan B, Subramanian T, Coleman D, Chinnadurai G. Identification of a cellular protein that specifically interacts with the essential cysteine region of the HIV-1 Tat transactivator. Virology. 1996;216:357–66.
Col E, Caron C, Chable-Bessia C, Legube G, Gazzeri S, Komatsu Y, et al. HIV-1 Tat targets Tip60 to impair the apoptotic cell response to genotoxic stresses. EMBO J. 2005;24:2634–45.
Kranz D, Dohmesen C. Dobbelstein MBRCA1 and Tip60 determine the cellular response to ultraviolet irradiation through distinct pathways. J Cell Biol. 2008;182:197–213.
Hu Y, Fisher JB, Koprowski S, McAllister D, Kim MS, Lough J. Homozygous disruption of the Tip60 gene causes early embryonic lethality. Dev Dyn. 2009;238:2912–21.
Bassi C, Li YT, Khu K, Mateo F, Baniasadi PS, Elia A, et al. The acetyltransferase Tip60 contributes to mammary tumorigenesis by modulating DNA repair. Cell Death Differ. 2016;23:1198–208.
Gorrini C, Squatrito M, Luise C, Syed N, Perna D, Wark L, et al. Tip60 is a haplo-insufficient tumour suppressor required for an oncogene-induced DNA damage response. Nature. 2007;448:1063.
Sakuraba K, Yasuda T, Sakata M, Kitamura YH, Shirahata A, Goto T, et al. Down-regulation of Tip60 gene as a potential marker for the malignancy of colorectal cancer. Anticancer Res. 2009;29:3953–5.
Sakuraba K, Yokomizo K, Shirahata A, Goto T, Saito M, Ishibashi K, et al. TIP60 as a potential marker for the malignancy of gastric cancer. Anticancer Res. 2011;31:77–9.
Chen G, Cheng Y, Tang Y, Martinka M, Li G. Role of Tip60 in human melanoma cell migration, metastasis, and patient survival. J Invest Dermatol. 2012;132:2632–41.
Halkidou K, Gnanapragasam VJ, Mehta PB, Logan IR, Brady ME, Cook S, et al. Expression of Tip60, an androgen receptor coactivator, and its role in prostate cancer development. Oncogene. 2003;22:2466–77.
Kueh AJ, Pal M, Tai L, Liao Y, Smyth GK, Shi W, et al. An update on using CRISPR/Cas9 in the one-cell stage mouse embryo for generating complex mutant alleles. Cell Death Differ. 2017;24:1821–2.
Yang H, Wang H, Shivalila CS, Cheng AW, Shi L, Jaenisch R. One-step generation of mice carrying reporter and conditional alleles by CRISPR/Cas-mediated genome engineering. Cell. 2013;154:1370–9.
Seibler J, Zevnik B, Kuter-Luks B, Andreas S, Kern H, Hennek T, et al. Rapid generation of inducible mouse mutants. Nucleic Acids Res. 2003;31:e12.
Jacks T, Remington L, Williams BO, Schmitt EM, Halachmi S, Bronson RT, et al. Tumor spectrum analysis in p53-mutant mice. Curr Biol. 1994;4:1–7.
Serrano M, Lee H, Chin L, Cordon-Cardo C, Beach D, DePinho RA. Role of the INK4a locus in tumor suppression and cell mortality. Cell. 1996;85:27–37.
Aubrey BJ, Kelly GL, Kueh AJ, Brennan MS, O’Connor L, Milla L, et al. An inducible lentiviral guide RNA platform enables the identification of tumor-essential genes and tumor-promoting mutations in vivo. Cell Rep. 2015;10:1422–32.
Kueh AJ, Herold MJ. Using CRISPR/Cas9 technology for manipulating cell death regulators. Methods Mol Biol. 2016;1419:253–64.
Sidoli S, Bhanu NV, Karch KR, Wang X, Garcia BA. Complete workflow for analysis of histone post-translational modifications using bottom-up mass spectrometry: from histone extraction to data analysis. J Vis Exp. 2016:e54112.
Kaya-Okur HS, Wu SJ, Codomo CA, Pledger ES, Bryson TD, Henikoff JG, et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat Commun. 2019;10:1930.
Pontén J, Saksela E. Two established in vitro cell lines from human mesenchymal tumours. Int J Cancer. 1967;2:434–47.
Sheikh BN, Phipson B, El-Saafin F, Vanyai HK, Downer NL, Bird MJ, et al. MOZ (MYST3, KAT6A) inhibits senescence via the INK4A-ARF pathway. Oncogene. 2015;34:5807–20.
Salama R, Sadaie M, Hoare M, Narita M. Cellular senescence and its effector programs. Genes Dev. 2014;28:99–114.
Hodgkins A, Farne A, Perera S, Grego T, Parry-Smith DJ, Skarnes WC, et al. WGE: a CRISPR database for genome engineering. Bioinformatics. 2015;31:3078–80.
Miller CW, Aslo A, Campbell MJ, Kawamata N, Lampkin BC, Koeffler HP. Alterations of the p15, p16,and p18 genes in osteosarcoma. Cancer Genet Cytogenet. 1996;86:136–42.
Baldi S, Becker PB. The variant histone H2A.V of Drosophila-three roles, two guises. Chromosoma. 2013;122:245–58.
Talbert PB, Henikoff S. Histone variants-ancient wrap artists of the epigenome. Nat Rev Mol Cell Biol. 2010;11:264–75.
Millar CB, Xu F, Zhang K, Grunstein M. Acetylation of H2AZ Lys 14 is associated with genome-wide gene activity in yeast. Genes Dev. 2006;20:711–22.
Mehta M, Braberg H, Wang S, Lozsa A, Shales M, Solache A, et al. Individual lysine acetylations on the N terminus of Saccharomyces cerevisiae H2A.Z are highly but not differentially regulated. J Biol Chem. 2010;285:39855–65.
Guillemette B, Bataille AR, Gévry N, Adam M, Blanchette M, Robert F, et al. Variant histone H2A.Z is globally localized to the promoters of inactive yeast genes and regulates nucleosome positioning. PLOS Biol. 2005;3:e384.
Barski A, Cuddapah S, Cui K, Roh TY, Schones DE, Wang Z, et al. High-resolution profiling of histone methylations in the human genome. Cell. 2007;129:823–37.
Valdes-Mora F, Song JZ, Statham AL, Strbenac D, Robinson MD, Nair SS, et al. Acetylation of H2A.Z is a key epigenetic modification associated with gene deregulation and epigenetic remodeling in cancer. Genome Res. 2012;22:307–21.
Hayashi-Takanaka Y, Maehara K, Harada A, Umehara T, Yokoyama S, Obuse C, et al. Distribution of histone H4 modifications as revealed by a panel of specific monoclonal antibodies. Chromosome Res. 2015;23:753–66.
Lu L, Chen X, Sanders D, Qian S, Zhong X. High-resolution mapping of H4K16 and H3K23 acetylation reveals conserved and unique distribution patterns in Arabidopsis and rice. Epigenetics. 2015;10:1044–53.
Li QL, Wang DY, Ju LG, Yao J, Gao C, Lei PJ, et al. The hyper-activation of transcriptional enhancers in breast cancer. Clin Epigenetics. 2019;11:48.
Boyer LA, Plath K, Zeitlinger J, Brambrink T, Medeiros LA, Lee TI, et al. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature. 2006;441:349–53.
Mikkelsen TS, Ku M, Jaffe DB, Issac B, Lieberman E, Giannoukos G, et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 2007;448:553–60.
Raisner RM, Hartley PD, Meneghini MD, Bao MZ, Liu CL, Schreiber SL, et al. Histone variant H2A.Z marks the 5’ ends of both active and inactive genes in euchromatin. Cell. 2005;123:233–48.
Orlando DA, Chen MW, Brown VE, Solanki S, Choi YJ, Olson ER, et al. Quantitative ChIP-Seq normalization reveals global modulation of the epigenome. Cell Rep. 2014;9:1163–70.
Kandoth C, McLellan MD, Vandin F, Ye K, Niu B, Lu C, et al. Mutational landscape and significance across 12 major cancer types. Nature 2013;502:333–9.
Ma X, Liu Y, Liu Y, Alexandrov LB, Edmonson MN, Gawad C, et al. Pan-cancer genome and transcriptome analyses of 1,699 paediatric leukaemias and solid tumours. Nature 2018;555:371–6.
Legube G, Linares LK, Tyteca S, Caron C, Scheffner M, Chevillard-Briet M, et al. Role of the histone acetyl transferase Tip60 in the p53 pathway. J Biol Chem. 2004;279:44825–33.
Eymin B, Claverie P, Salon C, Leduc C, Col E, Brambilla E, et al. p14ARF activates a Tip60-dependent and p53-independent ATM/ATR/CHK pathway in response to genotoxic stress. Mol Cell Biol. 2006;26:4339–50.
Giaimo BD, Ferrante F, Vallejo DM, Hein K, Gutierrez-Perez I, Nist A, et al. Histone variant H2A.Z deposition and acetylation directs the canonical Notch signaling response. Nucleic Acids Res. 2018;46:8197–215.
Dalvai M, Bellucci L, Fleury L, Lavigne AC, Moutahir F, Bystricky K. H2A.Z-dependent crosstalk between enhancer and promoter regulates cyclin D1 expression. Oncogene 2013;32:4243–51.
Voss AK, Thomas T. Histone Lysine and Genomic Targets of Histone Acetyltransferases in Mammals. Bioessays 2018;40:e1800078.
Clarke AS, Lowell JE, Jacobson SJ, Pillus L. Esa1p is an essential histone acetyltransferase required for cell cycle progression. Mol Cell Biol. 1999;19:2515–26.
Mo F, Zhuang X, Liu X, Yao PY, Qin B, Su Z, et al. Acetylation of Aurora B by TIP60 ensures accurate chromosomal segregation. Nat Chem Biol. 2016;12:226–32.
Greaves IK, Rangasamy D, Ridgway P, Tremethick DJ. H2A.Z contributes to the unique 3D structure of the centromere. Proc Natl Acad Sci USA. 2007;104:525–30.
Rangasamy D, Greaves I, Tremethick DJ. RNA interference demonstrates a novel role for H2A.Z in chromosome segregation. Nat Struct Mol Biol. 2004;11:650–5.
Cao X, Sudhof TC. A transcriptionally [correction of transcriptively] active complex of APP with Fe65 and histone acetyltransferase Tip60. Science 2001;293:115–20.
Baek SH, Ohgi KA, Rose DW, Koo EH, Glass CK, Rosenfeld MG. Exchange of N-CoR corepressor and Tip60 coactivator complexes links gene expression by NF-kappaB and beta-amyloid precursor protein. Cell. 2002;110:55–67.
Hattori T, Coustry F, Stephens S, Eberspaecher H, Takigawa M, Yasuda H, et al. Transcriptional regulation of chondrogenesis by coactivator Tip60 via chromatin association with Sox9 and Sox5. Nucleic Acids Res. 2008;36:3011–24.
Acknowledgements
We thank F. Dabrowski and L. Johnson, and N. Senzo for technical support and A. Kueh for discussions. We thank S. Henikoff for providing us with pAG-Tn5 with adapters. We thank T. Jacks and M. Serrano for making Trp53–/– and Cdkn2a–/– mice, respectively, available to the scientific community. This work was funded by the Australian Government through NHMRC project grants to A.K.V. and T.T. (1084248, 1084352, 1143612, 1160518), Research Fellowships (1081421 to A.K.V., 1003435 to T.T., 1058892 to G.K.S.) and an Investigator grant to A.K.V (1176789), the NHMRC IRIISS and the Victorian State Government OIS Grants to WEHI.
Author information
Authors and Affiliations
Contributions
J.W. planned and performed experiments, analyzed and interpreted data; C.P., S.E., R.M., E.A., S.W. performed experiments; A.L.G., C.S.N.L.-W.-S. and G.K.S. analyzed RNA-seq and CUT&Tag sequencing data; M.H. supplied the inducible CRISPR/Cas9 system; B.M. contributed to interpretation of data; A.K.V. and T.T. conceived the project and contributed to experimental design and data interpretation. J.W., T.T. and A.K.V. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
BJM and EA have received salary support from the Cancer Therapeutics CRC and have worked on a KAT5 drug discovery project for that organization. TT and AKV received funding for research from the CRC for Cancer Therapeutics and CTxOne, not for the project described here, but for research on other MYST family histone acetyltransferases. All other authors have no competing interest to declare.
Ethics statement
All animal experiments were approved by the Walter and Eliza Hall Institute Animal Ethics Committee and conducted in accordance with the Australian code of practice for the care and use of animals for scientific purposes.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Professor Massimiliano Agostini
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wichmann, J., Pitt, C., Eccles, S. et al. Loss of TIP60 (KAT5) abolishes H2AZ lysine 7 acetylation and causes p53, INK4A, and ARF-independent cell cycle arrest. Cell Death Dis 13, 627 (2022). https://doi.org/10.1038/s41419-022-05055-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-022-05055-6
- Springer Nature Limited
This article is cited by
-
Multifunctional histone variants in genome function
Nature Reviews Genetics (2024)
-
The Tip60/Ep400 chromatin remodeling complex impacts basic cellular functions in cranial neural crest-derived tissue during early orofacial development
International Journal of Oral Science (2023)