Abstract
Trib2 pseudokinase is involved in the etiology of a number of cancers including leukaemia, melanoma, ovarian, lung and liver cancer. Both high and low Trib2 expression levels correlate with different types of cancer. Elevated Trib2 expression has oncogenic properties in both leukaemia and lung cancer dependent on interactions with proteasome machinery proteins and degradation of transcription factors. Here, we demonstrated that Trib2 deficiency conferred a growth and survival advantage both at steady state and in stress conditions in leukaemia cells. In response to stress, wild type leukaemia cells exited the cell cycle and underwent apoptosis. In contrast, Trib2 deficient leukaemia cells continued to enter mitosis and survive. We showed that Trib2 deficient leukaemia cells had defective MAPK p38 signalling, which associated with a reduced γ-H2Ax and Chk1 stress signalling response, and continued proliferation following stress, associated with inefficient activation of cell cycle inhibitors p21, p16 and p19. Furthermore, Trib2 deficient leukaemia cells were more resistant to chemotherapy than wild type leukaemia cells, having less apoptosis and continued propagation. Trib2 re-expression or pharmacological activation of p38 in Trib2 deficient leukaemia cells sensitised the cells to chemotherapy-induced apoptosis comparable with wild type leukaemia cells. Our data provide evidence for a tumour suppressor role of Trib2 in myeloid leukaemia via activation of p38 stress signalling. This newly identified role indicates that Trib2 may counteract the propagation and chemotherapy resistance of leukaemia cells.
Similar content being viewed by others
Introduction
The Tribbles pseudokinases (Trib1, Trib2 and Trib3) are multifaceted signalling mediators controlling fundamental biological processes, including cell proliferation and survival, in both physiological and disease conditions1. Tribbles are inducible genes, modulated by a wide range of mitogens and stressors, and associated with downstream regulation of key signalling pathways, including AKT, ATF4, NF-kB and the MAPKs2,3,4,5,6. TRIB2 protein oscillates during cell cycle phases and induces the nuclear protein turnover of the dual specificity phosphatase and positive cell cycle mitotic regulator CDC25C7. Tribbles are newly recognised regulators of normal and malignant haemopoiesis8,9. Whilst TRIB2 levels are low in myeloid cells at steady state8, TRIB2 has been shown to regulate activation and inflammatory functions of human monocytes and macrophages3,10,11. Recent investigations showed that Trib2 is required for normal T-cell and erythroid development12,13. Trib2 has been shown to interact with different MAPK kinases (MAPKK), such as MEK1 and MKK7, and to either promote or inhibit MAPK cascade activation in distinct cellular contexts4,11.
Trib2 was first identified as an oncogene in acute myeloid leukaemia14. We have previously shown that Trib2 leukaemogenic potential relies on the ability to promote proteasomal dependent degradation of the tumour suppressor transcription factor CCAAT/enhancer-binding protein α (C/EBPα) and reported elevated TRIB2 expression in a subset of human myeloid leukaemia patients with dysregulated C/EBPα profile and mixed myeloid/T-lymphoid phenotype14,15. Further studies have associated high and low levels of TRIB2 with leukaemia subtypes with distinct genetic backgrounds. TRIB2 expression is positively associated with leukaemia patients that have FLT3 mutated t(15;17) genetics8, and with patients with elevated BCL2 expression46, and is negatively associated with leukaemia patients that have NPM1 and FLT3 mutations16. It has also been shown that the absence of Trib2 accelerated NOTCH1-driven T-cell leukaemia development12,17. Both high and low TRIB2 expression levels were shown to be associated with distinct human T-cell leukaemia phenotypes12.
The physiological role of Trib2 in myeloid leukaemia is not well understood. We previously showed that ectopic Trib2 expression cooperates with Homeobox transcription factor Hoxa9 to accelerate myeloid leukaemia development in mice18. We and others have shown that expression of TRIB2 is driven by several transcription factors including NOTCH119,20, TAL121, PITX122, MEIS123,24 and E2F125. The knockdown of TRIB2 in leukaemia cells led to leukaemia cell death21,25. However, low TRIB2 expression is associated with subgroups of myeloid leukaemia. It is not understood how the absence of Trib2 expression affects myeloid leukaemia.
Deregulation of the HOX genes occurs in ~70% of myeloid leukaemias. Indeed HOXA9 alone is overexpressed in over 50% of acute myeloid leukaemia patients and correlates with poor prognostic outcome26,27. Here we used the oncofusion gene NUP98/HOXA9 (NH9) as a deregulated HOX myeloid leukaemia model28,29 to investigate the effects of Trib2 deficiency in leukaemia cells. We showed that the absence of Trib2 does not impede the ability of NH9 to drive transformation. However, Trib2 deficiency enhanced myeloid leukaemia cell proliferation and survival in both steady state and stress conditions. Trib2 deficient leukaemia cells had impaired MAPK stress responses, evaded cell cycle checkpoint control mechanisms, and resisted chemotherapy-induced apoptosis. Our data identify Trib2 as a central regulator of p38-mediated stress signalling pathways and leukaemia cell cycle control.
Results
Trib2’s dispensability for NH9-initiated myeloid leukaemia
The impact of Trib2 deficiency in myeloid leukaemia is not well understood. To address this, we investigated the ability of NH9 oncofusion to transform wild type (WT) and Trib2 knockout (Trib2−/−) haemopoietic stem and progenitor cells (HSPCs) (S1A). HSPCs were isolated from WT and Trib2−/−mice and transduced with NH9 or empty vector control MigR1 retrovirus (S1B). In vitro myeloid cell transformation was assessed by serial replating ability in a colony forming cell (CFC) assay (S1A). WT and Trib2−/− MigR1 control groups failed to replate after the second round of CFC, whereas WT and Trib2−/− NH9 HSPCs formed colonies up to the fourth replating indicative of cell transformation (Fig. 1a and S1C). Moreover, both WT and Trib2−/− HSPCs transduced with NH9 and maintained in liquid culture (LC) conditions outgrew MigR1 controls and untransduced cells, as indicated by the increase in the fraction of GFP-expressing cells over time (S1D-E). To assess the self-renewal capability of NH9 in the absence of Trib2, a feature of leukaemic stem cells, we analysed the mRNA expression of Hoxa9, Hoxa7, Meis1, Runx1, Bm1, Sox4 and Smad730,31,32, in MigR1 or NH9 transduced WT and Trib2−/− HSPCs 48 h post-transduction (Fig. 1b). Equal transduction levels and self-renewal profiles of NH9 expressing cells was confirmed by upregulation of the known target genes Hoxa7 and Hoxa9 in the WT and Trib2−/− cells. Gene expression analysis of Runx1, Bm1, Sox4 and Smad7 revealed no differences between the WT and Trib2−/− NH9 HSPCs or compared to MigR1 controls (Fig. 1b). Moreover, basal expression of Hox genes was equivalent between WT and Trib2−/− MigR1 control cells. These data indicate that Trib2 is not required for NH9-mediated transformation of HSPCs into myeloid leukaemia cells. To confirm leukaemic stem cell properties of the WT and Trib2−/− NH9 expressing cells we analysed leukaemic cell surface marker expression33by flow cytometry (Fig. 1c–g and S2A). Interestingly, the proportion of c-Kit+ cells, and the Lineage(Lin)-Sca1-CD11b+c-Kit+ fraction were higher in the Trib2 deficient cells compared to the WT NH9 cells (Fig. 1c–e). Similarly, Lin-Sca1-CD127-c-Kit+CD34+/lowCD16/32+ cells, previously identified as L-GMP (leukaemic granulocyte macrophage progenitors34), were increased in the Trib2−/− NH9 compared to the WT NH9 cells (Fig. 1f, g and S2B). Overall these results suggest that Trib2 deficient cells retain higher leukaemic stem cell potential.
a CFC assay of WT and Trib2−/− HSPCs retrovirally transduced with MigR1 or NH9 expressing vectors. Graph shows total colony number at CFC1-4. Data are mean of duplicate cultures ± SD; representative of 2 independent experiments. b Gene expression analysis in WT and Trib2−/− HSPCs retrovirally transduced with MigR1 or NH9 at 48 h post transduction. Data are means of 3 biological samples (from 1 to 2 mice per biological replicate) ± SD and representative of 2 independent experiments. *P < 0.05, **P < 0.005, ***P < 0.001, using unpaired t-test. c, d Representative dot plots of the CD11b+c-Kit+ fraction (c) and c-Kit expression histograms (d) of WT and Trib2−/− NH9 cells transformed by CFC assay (top panels) or LC condition (bottom panels), gated through GFP+DAPI-Lin(CD4, CD8, B220, Ter119)-Sca1- gates. e Graphed percentages are average of 3 independent measurements, as shown (c, d) ±SD. *P ≤ 0.05, **P < 0.005, using unpaired t-test. f Representative flow cytometry analyses identifying the L-GMP population in WT and Trib2−/− NH9 cells transformed by CFC assay (top panels) or LC condition (bottom panels), gated through GFP+DAPI-Lin(CD3, CD4, CD8, CD19, B220, Ter119, Gr1)-CD127- and Sca1-c-Kit+(LK) gates. g Graph shows percentages of the L-GMP cells, as shown (f), in the GFP+DAPI-Lin-fraction. Data are mean of 3 independent measurements ± SD. *P ≤ 0.05, **P < 0.005, using unpaired t-test
Trib2 deficiency promotes the propagation and survival of leukaemia cells
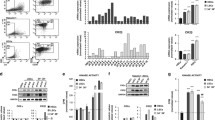
To determine the impact of Trib2 deficiency on the growth properties of myeloid transformed cells, we compared the cell division using the viable dye cell trace violet (CTV) of MigR1 or NH9 transduced WT and Trib2−/− HSPCs (from two independent transduction experiments) that had expanded by CFC assay (GFP sorted, Fig. 2a, top panels) or in liquid culture (LC) for 2 weeks (unsorted for GFP, Fig. 2a, bottom panels). Our results showed a higher number of cell divisions in the Trib2−/− NH9 cells compared to the WT NH9 cells (Fig. 2a), indicating that Trib2 negatively regulates myeloid leukaemia cell proliferation. WT and Trib2-/- NH9 transduced cells were expanded in LC for several months. Growth curves confirmed faster proliferation rate and significantly shorter doubling time in Trib2-/- NH9 cells compared to WT NH9 cells (Fig. 2b, c). To assess the impact of Trib2 deficiency on myeloid leukaemia cell survival, we measured the level of apoptosis following growth factor deprivation (GFD) and myeloid leukaemia-based chemotherapy (daunorubicin (DNR)) in the WT and Trib2-/- NH9 cells by flow cytometry using AnnexinV/DAPI staining (Fig. 2d–g). Our analyses showed that basal levels of apoptosis in Trib2-/- NH9 cells were significantly lower compared to the WT NH9 cells (~5 vs. ~20% respectively, Fig. 2d–g 0 h GFD and DMSO treated samples). Following GFD or DNR treatment, Trib2-/- NH9 cells exhibited lower levels of cell death compared to the WT NH9 cells (Fig. 2d, e 24 h GFD and Fig. 2f, g DNR treated samples). These data show that Trib2 deficiency promotes myeloid leukaemia cell proliferation and survival.
a CTV assay of WT and Trib2-/- NH9 cells derived from CFC3 (CFC, top) and CTV assay of WT and Trib2-/- MigR1/NH9 cells derived from 2 weeks in LC, in presence of recombinant SCF, IL3, IL6 (LC, bottom). Samples were analysed by flow cytometry at 96 h after staining with the CTV dye, and gated through GFP+ gates. Undivided, stained control cells are shown in red (colcemid ctrl). Cell division gates are indicated and % of cells in each cell division for the transformed WT and Trib2-/- NH9 cells is graphed. Data are representative of at least 3 independent experiments. b Growth curves (right) of WT and Trib2-/- NH9 cell lines, grown in presence of IL3 (NH9 medium). In green and pink are the projected non-linear regression curves that best fit the exponential growth models of WT and Trib2-/- NH9 cells, respectively (calculated with the GraphPad software). c Growth rate and doubling time of WT and Trib2-/- NH9 cells, as measured in B. Data shown are means of 3 independent experiments ± SD. *P < 0.05, using unpaired t-test. d–g Flow cytometric analysis of apoptotic levels by means of AnnexinV/DNA DAPI staining in WT and Trib2-/- NH9 cell lines after GFD (d) and 24 h DNR treatment (f). f, g Graphed percentages of Late Apoptotic cells (AnnexinV+/DNA DAPI+), at the indicated time points, as measured in (d) and (f), respectively. Data are representative of at least 3 independent experiments, graphs show mean ± SD. *P ≤ 0.05, **P < 0.005, ***P < 0.001, using unpaired t-test
Impaired MAPK activation in the absence of Trib2
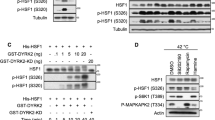
Trib2 modulates MAPK signalling3,4,11,12, and MAPKs are linked to cell proliferation and cancer35,36. Thus, we interrogated the activation status of the MAPKs ERK, p38 and JNK in WT and Trib2-/- myeloid leukaemia cells. Western blot analyses showed that Trib2-/- NH9 cells had lower levels of the phosphorylated (active) forms of ERK1-2 (Thr202/Tyr204, p-ERK), p38 (Thr180/Tyr182, p-p38) and JNK (Thr183/Tyr185, p-JNK), compared to the WT NH9 cells (Fig. 3b). WT and Trib2-/- NH9 cells showed IL3-dependent growth (Fig. 2b), therefore we tested MAPK following GFD and re-stimulation with IL3-containing medium (NH9 medium). We showed that MAPKs phosphorylation was impaired in Trib2-/- NH9 compared to WT NH9 thus indicating inefficient MAPK responses in Trib2-/- NH9 (Fig. 3c). Conversely, the phosphorylation of AKT (Ser473) was similarly activated following IL3 stimulation in both WT and Trib2-/- NH9 cells (Fig. 3c). We tested the activation of p38 in response to anisomycin (Ans), a pharmacological activator of the p38 and JNK pathway, that directly activates MKK6 and p38 kinases37 (Fig. 3d). The fold increase in p-p38 after Ans treatment measured by phosphoflow cytometry was reduced in the Trib2-/- NH9 cells compared to the WT NH9 cell (Fig. 3d). These data showed that Trib2 is necessary for efficient activation of the p38 kinase. Overall, these results showed that Trib2 deficient myeloid leukaemia cells have compromised basal- and induced-MAPK signalling activation.
a Schematic of MAPK signalling cascade. b Western blot analysis of MAPKs total and phosphorylated levels (ERK1-2 (Thr202/Tyr204), p38 (Thr180/Tyr182) and JNK (Thr183/Tyr185)) in WT and Trib2-/- NH9 cells. Data are representative of at least 3 independent experiments. c Western blot analysis of MAPKs and AKT (Ser473) activation in WT and Trib2-/- NH9 cells after 1 h of GFD, followed by re-stimulation with IL3-containing medium (NH9 medium) for 0 and 15 min. d Ans binds and activates the p38 signalling pathway. Model of TRIB2-mediated regulation of MKK3/6 or p38 kinases (top). The p-p38 signal was measured by flow cytometry following stimulation of the WT and Trib2-/- NH9 cells with 25 and 50 ng/ml Ans, and normalised by DMSO treated control cells (bottom). Data shown is average of 2 independent experiments ± SD, the genotype significantly affects the result, P < 0.05 using two-way ANOVA
Trib2 deficiency enables cell cycle progression following stress
To gain further insight into the mechanism involved in the enhanced growth and survival of Trib2 deficient leukaemic cells, we assessed the cell cycle control and check point activation in response to genotoxic stress in WT and Trib2-/- NH9 cells. Using Ki67/DNA DAPI flow cytometry staining, we assessed the cell cycle profile of WT and Trib2-/- NH9 cells following DNR treatment. No significant difference in the cell cycle profiles was observed in untreated WT and Trib2-/- NH9 cells, where the majority of the cells were in the G1 phase of the cell cycle (Fig. 4a, b). Following DNR treatment, the majority of WT NH9 cells were lost from the active G1, S, G2, M phases and with remaining cells in the GO quiescent phase of the cell cycle. In contrast, a significant proportion of Trib2-/- NH9 cells were still cycling and with less cells in the GO quiescent state (Fig. 4a, b). We next assessed the proportion of cells in the mitotic phase (mitotic index) by p-HH3/DNA PI staining. In untreated cells, no significant differences were observed between the WT and Trib2-/- NH9 cells. However, a significantly higher fraction of p-HH3+ cells was observed in Trib2-/- NH9 cells compared to WT NH9 cells, following 16, 20 and 24 h post DNR treatment (Fig. 4c, d). Similarly, a higher fraction of p-HH3+ cells was detected upon GFD condition in Trib2-/- NH9 (Figure S3). We analysed metaphase spreads of DNR-treated WT and Trib2-/- NH9 cells by microscopy to validate the role of Trib2 in mitotic cell cycle control following stress stimuli. Similar to p-HH3 mitotic indices, there was an equivalent percentage of mitotic spreads in the untreated WT and Trib2-/- NH9 cells (Fig. 4e, f). DNR treatment virtually abolished mitotic entry in the WT NH9 cells, as no mitotic spreads were observed in any field/slide analysed. Conversely, a significantly higher proportion of DNR-treated Trib2-/- NH9 were in metaphase compared to WT NH9 cells (Fig. 4e, f). These data show that Trib2-/- NH9 cells can actively cycle and undergo mitosis in the presence of stress. These results indicate a role for Trib2 in the prevention of cell cycle progression in stress conditions.
a Cell cycle flow cytometric analysis by means of Ki67/DNA staining in WT and Trib2-/- NH9 cells in response to DNR treatment. Plots are representative of 6 independent experiments with similar trend. b Graphed percentages of G0, G1 and SG2M fractions, as measured in (A). Data are means ± SD of 3 independent experiments. *P < 0.05, using unpaired t-test. c Flow cytometric analyses of the mitotic index in WT and Trib2-/- NH9 cells after 24 h DNR treatment, as measured by p-HH3/PI DNA levels. d Graphed percentages, at the indicated time points, as measured (c). e Representative pictures of DAPI stained metaphase spreads from WT and Trib2-/- NH9 cells after 24 h DNR treatment. f Graph showing percentages of mitotic spreads from WT and Trib2-/- NH9 cells after 24 h DNR/DMSO treatment as measured e. Data are representative of at least 2 independent experiments, graphs show mean ± SD. *P < 0.05, **P < 0.005, using unpaired t-test
Trib2 deficiency compromises stress response signalling in myeloid leukaemia cells
As we have demonstrated a role for Trib2 in leukaemia cell cycle control in response to stress, we hypothesized that Trib2 is an inducible component of the cellular stress response pathways, contributing to check point activation and survival or death decision. To address this, we assessed Trib2 mRNA expression levels following GFD and DNR treatment. In WT NH9 cells, a significant upregulation of Trib2 mRNA expression was observed in response to DNR treatment (Fig. 5a) and GFD (Figure S4A). Gene expression analyses of DNR treated WT and Trib2-/- NH9 cells showed that the activation of cell cycle inhibitors and regulators, including Cdkn1a (p21), Cdkn1b (p27), INK4A (p16), ARF (p19) and Gadd45a/b, was abolished or significantly lowered in the absence of Trib2. Indeed, levels of Cdkn1a (p21), INK4A (p16), ARF (p19) and Gadd45b were lower in untreated Trib2-/- NH9 cells compared to the WT NH9 cells (Fig. 5a). Similar results were observed following GFD (Figure S4A). Levels of Cdk4, a positive regulator of cell cycle progression, were higher in Trib2-/- NH9 cells compared to the WT NH9 cells, at both steady state and upon GFD (Figure S4A), consistent with the highly proliferating phenotype of Trib2-/- NH9 cells. In addition, Trib2-/- NH9 cells failed to upregulate Egr1 and Nfatc4 (apoptotic mediators), Jun and Fos (stress response TFs downstream of MAPK signalling), mRNA in both vehicle control and DNR treated samples (Figure S4B). These results demonstrate that Trib2 expression is required in myeloid leukaemia cells for efficient activation of cell cycle check point genes and DNA damage response signalling pathways. Assessment of key cell cycle checkpoint regulator proteins by western blotting showed that induction of p-p38 and p21 was impaired after 16, 20 and 24 h DNR treatment in Trib2-/- NH9 compared to WT NH9 cells. Importantly, the total protein levels of p38 level were unaffected (Fig. 5b). Furthermore, mRNA levels of several MAPKs genes, including MAPK3 (ERK1), MAPK9 (JNK2), MAPK11 (p38β), MAPK12 (p38γ), MAPK13 (p38δ), and the MKNK1 (MNK1), were significantly lower in Trib2-/- NH9 cells in response to DNR treatment compared to the treated WT NH9 cells (Figure S4C). Reduced activation of the MAPK p-p38 was also confirmed by flow cytometry in WT and Trib2-/- NH9 cells in response to DNR treatment (Fig. 5c, d). The fraction of p-p38+ cells and the MFI of p-p38 (which correlates with the number of p38 molecules activated per cell) were significantly lower in the Trib2-/- NH9 treated cells compared to the WT NH9 cells. Levels of p-Chk1 and γ-H2Ax (Ser139), key regulators of DNA damage response signalling, were significantly reduced in Trib2-/- NH9 treated cells compared to the WT NH9 treated cells (Fig. 5c, d). Together these data demonstrate a role for Trib2 as a major orchestrator of cell cycle check point and stress signalling pathways in leukaemia, required for efficient activation of p38, Chk1 and γ-H2Ax, and stress-induced p21 expression.
a Trib2, Cdkn1a (p21), Cdkn1b (p27), INK4A (p16), ARF (p19), Gadd45a and Gadd45b relative mRNA levels in WT and Trib2-/- NH9 cells after 16 h DNR treatment. Data are representative of 2 (p16 and Gadd45a) or 3 (Trib2, p21, p27, p19 and Gadd45b) independent experiments with similar trend, graphs show mean of technical replicates ± SD. *P < 0.05, **P < 0.005, ***P < 0.001 using unpaired t-test. b Western blot analysis of the indicated proteins in WT and Trib2-/- NH9 cells in response to DNR treatment for the indicated time points. Data is representative of at least 3 independent experiments, with similar trend. c Flow cytometric analysis of p-p38, p-Chk1 and γH2Ax levels in the live fraction of WT and Trib2-/- NH9 cells, after 24 h DNR treatment. Histograms are representative of at least 3 independent experiments, with similar trend. d Graphed percentages (c) show average ± SD of three independent measurements. MFI Median Fluorescent Intensity (indicated in the top right panel of the representative histograms). *P < 0.05, **P < 0.005, ***P < 0.001 using unpaired t-test
Trib2 deficiency enables propagation of drug resistant leukaemic cells
To test whether the inappropriate stress response of Trib2 deficient leukaemia cells results in the propagation of drug-resistant myeloid leukaemia cells, we performed DNR washout (wo) experiments. WT and Trib2-/- NH9 cells were treated with the drug for 24 h, then washed and replated in either LC or CFC assays (Fig. 6a). Trib2-/- NH9 cells displayed reduced apoptotic rate in LC, and higher clonogenic potential compared to the WT NH9 cells in CFC (Fig. 6b-f). Moreover, cell surface marker analyses revealed that drug resistant Trib2-/- NH9 cells continued to express CD11b myeloid marker, higher levels of the immature marker c-Kit (Fig. 6g, left and central panels), and lower levels of the inflammatory surface marker Sca1 compared to WT NH9 cells in both untreated and drug resistant cells (Fig. 6g, right panel). These data indicate the Trib2 deficiency contributes to a drug resistant phenotype in myeloid leukaemia.
a Schematic showing experimental strategy of the DNR wo experiments in LC or IL3-supplemented MC. b Apoptosis analysis of the WT and Trib2-/- NH9 cells after 24 h DMSO/DNR treatment, and at 24, 48 h and 7 days after drug wo. c Day 7 and d day 5 colony pictures (4×) of WT and Trib2-/- NH9 samples treated with DMSO/DNR for 24 h and plated in MC, after wo, at the indicated cell range, in independent experiments. e, f Colony counts at day 7, as per experiment in (c and d), respectively. Graphs show mean of duplicate (c, e) or triplicates (d, f) cultures ± SD. *P < 0.05, **P < 0.005, ***P < 0.001 using unpaired t-test. g Flow cytometric analysis of the indicated myeloid and primitive surface markers of WT and Trib2-/- NH9 cells in (d, f), grown in MC for 7 days, after removal of the drug. Graphs show mean of triplicate cultures ± SD. *P < 0.05, **P < 0.005, ***P < 0.001 using unpaired t-test
Re-expression of Trib2 reverts the drug resistant phenotype of Trib2-deficient myeloid leukaemia cells
To validate the drug resistant phenotype as a result of the Trib2 deficiency, we reintroduced Trib2 protein expression in the Trib2-/- NH9 cells. WT and Trib2-/- NH9 were retrovirally transduced with NGFR and NGFR Trib2, andTrib2 mRNA or protein expression confirmed in NGFR+ sorted cells (Figure S5). NH9-expressing WT NGFR, Trib2-/- NGFR, and Trib2-/- NGFR Trib2 were treated with DNR and apoptosis and cell cycle profiles were assessed. Our results showed that reintroduction of Trib2 in the Trib2-/- NH9 cells increased apoptosis in untreated and DNR treated cells to levels comparable with WT NH9 cells (Fig. 7a). NH9-expressing Trib2-/- NGFR Trib2 cells showed a higher proportion of cells in G0 and less cells in G1 compared to Trib2-/- NGFR (Fig. 7b). This was supported by the significant decrease in the percentage of p-HH3+ cells in NH9-expressing Trib2-/- NGFR Trib2 treated cells compared to NH9-expressing Trib2-/- NGFR treated cells (Fig. 7c). Together, these results demonstrate that Trib2 expression is required for the appropriate cell cycle stress response and the effective killing of myeloid leukaemia cells.
Flow cytometric analysis of apoptotic levels by means of AnnexinV/DNA DAPI (a), Cell cycle Ki67/DNA DAPI (b), and mitotic index by means of p-HH3/DNA DAPI staining (c) in WT and Trib2-/- NH9 MigR1 NGFR/NGFR Trib2 transduced cells, after 24 h DNR treatment. d Shows graphed percentages of Live (AnnexinV-/DNA DAPI-) and Late Apoptotic (AnnexinV+/DNA DAPI+) cells, as measured (a). Data are representative of 4 independent experiments, graphs show mean ± SD. *P < 0.05, **P < 0.005, ***P < 0.001, using unpaired t-test. e Graphed percentages of p-HH3+ cells, as measured (c). Data are representative of 2 independent experiments, graphs show mean ± SD. **P < 0.005 using unpaired t-test
Pharmacological activation of p38 reverts the drug resistant phenotype of Trib2-deficient myeloid leukaemia cells
Next we tested whether pharmacological activation of p38 by Ans could revert the DNR resistant phenotype of the Trib2-/- NH9 cells similar to Trib2 re-expression (Fig. 8a). Ans pre-treatment resulted in increased basal and DNR-induced apoptosis in the Trib2-/- NH9 cells (+Ans Trib2-/- NH9) (Fig. 8b top and bottom panels). In addition, Ans pre-treatment restored drug-induced expression of p38 target genes in the Trib2-/- NH9 cells to levels comparable (INK4A, ARF) or higher (Cdkn1a, Cdkn1b, Gadd45a) than those of the WT NH9 treated cells (Fig. 8c). Overall, these data suggest that TRIB2 controls stress signalling responses via p38 activation. We next tested the direct binding of Trib2 and p38 as a mechanism for Trib2-mediated p38 regulation. Using a co-immunoprecipitation (co-IP) assay in Hek293T cells overexpressing myc-tagged Trib2 (PHMA Trib2) or transfected with the empty control plasmid (PHMA) (S6), we did not detect any binding between p38 and Trib2. This result does not exclude the possibility that p38 MAPK and its upstream MAPKK activators could interact with Trib2 in myeloid leukaemic cells transiently under stress conditions.
a Ans/DMSO pre-treatment strategy in WT and Trib2-/- NH9 cells, followed by DNR treatment. Representative histogram of p-p38 expression following Ans pre-treatment and after DNR treatment are shown and MFIs are indicated. b Flow cytometric analysis of apoptotic levels by means of AnnexinV/DNA DAPI after 24 h DNR treatment (top panel) and graph percentages of Late Apoptotic (AnnexinV+/DNA DAPI+) cells (bottom panel). Data are representative of 3 independent experiments, graphs show mean ± SD. **P < 0.005, ***P < 0.001, using unpaired t-test. c Trib2, Cdkn1a (p21), Cdkn1b (p27), INK4A (p16), ARF (p19), and Gadd45a relative mRNA levels in WT, Trib2-/- NH9 cells and +Ans Trib2-/- NH9 cells after 16 h DNR treatment. Data are representative of 2 independent experiments with similar trend, graphs show mean of technical replicates ± SD. *P < 0.05, **P < 0.005, ***P < 0.001 using unpaired t-test. d Schematic representation of the Trib2-p38 regulatory axis. Trib2 promotes damage response pathways via p38 activation (left), and its absence results in impaired signalling response, uncontrolled cell cycle progression and chemoresistance (right)
Discussion
In this study, we investigated the impact of Trib2 deficiency in myeloid leukaemia. We demonstrate that Trib2 is not required for the initiation of myeloid leukaemia, but is required for p38 and stress signalling, induction of cell cycle checkpoint response and apoptosis. This tumour suppressive role of the TRIB2-p38 axis is also supported by the evidence that Trib2 deficiency accelerates the onset of acute lymphoid leukaemia12. We show here the defective stress response manifests a chemotherapy resistant phenotype and a growth advantage to myeloid leukaemia revealing, for the first time, TRIB2 as a tumour suppressor in myeloid leukaemia.
p38 is a regulator of key cell cycle regulators (p21, p16, p19, GADD45), signalling molecules (Chk1, γH2Ax), TFs (Egr1, Fos and Jun) which are inappropriately activated in Trib2 deficient leukaemia cells. These, and others, are involved in proliferation, survival and stress response pathways38,39,40,41, including the phosphatase CDC25(Ref7), which is known to be regulated by Trib2. Therefore, impaired p38 activation due to the absence of Trib2 most likely accounts for the defective signalling response and chemoresistant phenotype of myeloid leukaemia cell (Fig. 8d). Indeed, using IL3 stimulation and the p38 pharmacological activator Ans, p38 phosphorylation/activation was still lower in Trib2 deficient cells compared to treated WT NH9 cells, demonstrating that Trib2 functions in the activation of p38. The re-expression of Trib2 or the activation of p38 in DNR treated Trib2 deficient cells further confirms the Trib2-p38 axis as a regulator of cell cycle progression and damage signalling pathways. As it is a pseudokinase, Trib2 would not lead to phosphorylation of p38 upon binding (whether indirect or direct), but act as a kinase inhibitor/competitive inhibitor/allosteric regulator. It would be interesting to obtain structural data on Trib2 necessary to support for this. To date, no structural data for Trib2 is available.
Our data is consistent with the finding that Trib2 deletion does not impede MLL/AF9-driven myeloid leukaemia17, as we did not observe any defect in the ability of NH9 to transform Trib2 deficient cells. Hoxa9-dependent transactivation of the Trib2 promoter was shown to be efficient only in the presence of the cofactors Meis1 and Pbx323,24. Overexpression of NH9, enhancing self-renewal potential of normal cells, did not induce Trib2 gene upregulation in WT HSPCs. Moreover, no differences were observed in Hox genes expression in presence or absence of Trib2, suggesting that Trib2 cooperativity in myeloid leukaemia18 is independent on the HOX transcriptional program. This is supported by the observation that cooperating lesions in myeloid leukaemia usually affect non-redundant pathways, e.g., proliferation/survival and self-renewal/differentiation42. Indeed, we have demonstrated that Trib2 modulates the expression or activation of several signalling molecules and cell cycle inhibitors, whereas NH9 confers aberrant self-renewal potential to the cells.
We demonstrated that Trib2 is required in response to stress to efficiently inhibit cell cycle progression and mitosis, in order to prevent the propagation of damaged cells. These data suggest that Trib2 may be an important regulator in myeloid leukaemia drug resistance and relapse. We showed that Trib2 is transcriptionally upregulated in the WT myeloid leukaemia cells after either starvation or genotoxic challenge. Our results are in line with previous studies, reporting TRIB2 upregulation in response to survival factor withdrawal in primary T-cells and erythroleukaemia cells43, miR155 induction in acute myeloid leukaemia cells44 and cisplatin treatment in ovarian cancer cells6, which all associated with activation of the apoptotic pathway. Conversely, ectopic or intrinsic high TRIB2 expression confers chemoresistance in solid cancers and leukaemia, by modulating the AKT/FOXO/p53 signalling pathway45 and upregulation of BCL2 expression46. Therefore, similar to other renowned cancer related proteins (e.g. E2Fs, p21, GADD45A, MYC and RUNX1), Trib2 obeys the Goldilocks principle in cancer biology; too little or too much are both pro-tumorigenic. Data from us and others would conclude that Trib2 exerts a dual role in myeloid leukaemia and other cancers (e.g., T cell leukaemia, liver), in a stage- and context-specific manner.
Summarising, this work identified a novel Trib2-p38 regulatory pathway important for cell cycle checkpoint and stress responses, with implications on drug resistance and relapse in myeloid leukaemia (Fig. 8d).
Materials and methods
Mice
Trib2 knock out mice (B6; 129S5-Trib2tm1Lex, referred to as Trib2-/-), backcrossed onto C57B6 as previously described12, were housed in the university of Glasgow. All animals were handled in accordance with good animal practice as defined by the Animals (Scientific Procedures) Act 1986. Mice used in the study were from age matched adult (7-14 week old) WT and Trib2-/- mice of either sex.
Primary cell culture, transduction and generation of WT and Trib2 -/- NH9 cell lines
Retroviral supernatants were generated by transient transfection of Hek293T cells with the retroviral constructs (together with pCMV-Gag-Pol packaging vector and pHIT123 envelope vector) with the calcium-phosphate method and titered with NIH-3T3 cell as previously described15. For transduction experiments, total bone marrow cells were isolated and were enriched for c-Kit expression, using anti-CD117 (c-Kit) microbeads and MS MACS columns (Miltenyi Biotec) according to manufacturer’s instructions. c-Kit+ cells are an enriched HSPC population. They were cultured o/n in prestimulation medium (DMEM, 15% FBS, 2 mM l-glutamine, 100 U/mL Penicillin/Streptomycin (Pen/Strep), 10 ng/ml IL3, 10 ng/ml IL6, 100 ng/ml SCF). WT and Trib2-/- HSPCs were transduced with MigR1 or MigR1 NH9 GFP tagged retroviral vectors31. WT and Trib2-/- NH9 immortalized lines were generated by expansion in liquid culture (LC) in prestimulation medium for 9 weeks. Afterwards, the culture was continued in NH9 medium (DMEM, 10% FBS, 10% WEHI-3B conditioned medium, 2 mM l-glutamine, 100 U/mL Pen/Strep). NH9 samples outgrew MigR1 controls and 100% GFP + NH9 cells were immortalised. The growth of the NH9 cell lines was monitored by trypan blue counts. The cells were routinely tested for mycoplasma contamination before and after freezing and thawing. For growth factor deprivation (GFD) experiments, NH9 cell lines were seeded at 0.1 × 106 cells/ml 24 h before starvation. The cells were incubated in starvation medium (DMEM, 2 mM l-glutamine, 100 U/ml Pen/Strep), for the indicated time points. For daunorubicin (DNR) and anisomycin (Ans) treatment experiments, NH9 cell lines were seeded at 0.2 × 106 cells/ml 24 h before treatment with either 50 nM DNR or 25 and 50 ng/ml Ans or DMSO as vehicle control. The DNR inhibitory concentration 50 (IC50) was determined in WT NH9 overexpressing cells (46.07 nM) (S3A). Cells transductions were performed centrifuging 2–5 × 106 cells/ml cells with the required vectors for 90 min at 1290xg in either prestimulation medium (HSPCs) or NH9 medium (NH9 cell lines), supplemented with 4 µg/ml of polybrene.
Colony forming cell (CFC) assay
For CFC assays of freshly transduced HSPCs (isolated by c-Kit enrichment), equal numbers of sorted GFP-expressing cells (0.5 or 10 × 103) were seeded in MethoCult™ GF M3434 (Stem Cell Technologies) (MC) medium, containing 15% FBS, 1% BSA, insulin (10 μg/ml), transferrin (200 μg/ml), IL3 (10 ng/ml), IL6 (10 ng/ml), SCF (50 ng/ml) and Erythropoietin (3 U/mll), and supplemented with Pen/Strep (100 U/ml). For DNR wash out CFC assays, equal numbers of WT and Trib2-/- NH9 cells (0.1 or 30 × 103) were treated with DNR for 24 h and remaining cells plated in MC medium (M3231, Stem Cell Technologies), containing 30% FBS and 1% BSA, supplemented with IL3 (10 ng/ml) and Pen/Strep 100 U/ml. Colonies were scored after 1 to 2 weeks.
Fluorescent activated cell sorting (FACS) analyses
Flow cytometry experiments were performed using BD FACSCanto™ II and BD FACSAria™ (BD biosciences, UK). Cell trace™ violet (CTV, Invitrogen) staining was performed following manufacturer instructions and the fluorescent signal assessed in the proliferating samples after 4 days of culture. Cells treated for 24 h with 100 ng/ml demecolcine solution (Sigma) were used as undivided control (colcemid control) and used to calculate the undivided generation peak (division 0). Apoptosis was measured by AnnexinV (eBioscience) / 4′,6-diamidino-2-phenylindole (DAPI, Sigma) staining, performed in HBSS solution (Gibco). For staining of the intracellular antigens, the cells of interest were fixed and permeabilised using the BD Cytofix/Cytoperm™ Fixation/Permeabilization Kit (BD biosciences), subsequently incubated with anti-Ki67 (eBioscience), anti-phospho(p)-p38 (eBioscience), anti-phospho(p)-Chk1 (New England Biolab) or anti-γH2Ax (BD Biosciences) antibodies (Table S1). For mitotic index measurement, the cells were fixed using the BD Fixation/Permeabilization solution (BD biosciences) and permeabilised with 90% methanol, followed by incubation with anti-phospho-Histone H3 (p-HH3) antibody (Cell Signaling Technology, Table S1). DNA staining was performed on fresh or fixed samples using PI/RNase Staining Solution (BD biosciences), or DAPI, for cell cycle or viability analyses. For surface antigens detection, fresh samples were incubated with anti-CD3, anti-CD4, anti-CD8, anti-CD19, anti-B220, anti-Ter119, anti-Gr1, anti-CD127 (IL7R), anti-CD34, anti-CD16/32, anti-CD11b, anti-CD117(c-Kit) and anti-Sca1 antibodies (eBioscience), and DAPI+ dead cells exclusion was performed. In all flow cytometry analyses, the desired populations were gated through FSC-Area/SSC-Area and doublets were excluded based on FSC-Height/FSC-Area.
Real time PCR (RT-PCR)
Up to 1 µg of total RNA was reverse transcribed with the high capacity cDNA reverse transcription kit (Applied Biosystems), following manufacturers’ instruction. In case of low cell number samples, Specific Target Amplification (STA) reaction was performed with the Qiagen® Multiplex PCR Kit (Qiagen), following manufacturers’ instructions. RT-PCR was performed using the Fast SYBR™ Green 2× Master Mix (Applied Biosystems), according to manufacturers’ instruction. All primers, listed in Table S2, were designed to target murine sequences. Failed reactions/outliers were excluded from the analyses.
Co-immunoprecipitation (Co-IP)
Hek293T cells were transiently transfected with the myc-tagged PHMA Trib2 or PHMA control vectors as previously described15. The transfected cells were harvested at 24 h and whole cell lysates were prepared using ice-cold Hepes buffer (50 mM Hepes pH 7.4, 150 mM NaCl, 1 mM EDTA, 0.5% NP-40, 5% glycerol, with protease and phosphatase inhibitors). Crosslinking was performed using dithiobis-succinimidylpropionate (DSP) (Thermo Fisher Scientific) at a concentration of 1.5 mM, and the reaction quenched using Tris (pH 7.4) (50 mM). One milligram of precleared lysates were incubated with Myc9E10 antibody (Santa Cruz Biotechnology) or normal mouse IgG overnight at 4 °C, followed by incubation with Protein A/G UltraLink™ Resin (Thermo Fisher Scientific) for 1.5 h at 4 °C. The samples were washed in Tris buffer (50 mM Tris pH 7.4, 150 mM NaCl, 1 mM EDTA, 0.5% NP-40, 5% glycerol, with protease and phosphatase inhibitors), eluted in Laemmli buffer and analysed by western blotting.
Western Blotting
Whole cell lysates were prepared using ice-cold modified RIPA buffer (50 mM Tris, pH 8.0, containing 0.5% NP-40, 0.25% sodium deoxycholate, 150 mM NaCl, 1 mM EDTA, with protease and phosphatase inhibitors). Up to 25 µg of protein samples were resolved on hand-casted SDS-PAGE gels, transferred to 0.45 μm nitrocellulose membrane (Whatman) and analysed by immunoblotting with the antibodies listed in Table S3. SuperSignal™ West Pico or Femto Chemoluminescent Substrates (Thermo Fisher Scientific) were used for signal detection on CL-XPosure™ radiography films. The films were developed using the Medical film processor SRX-101A (Konika Minolta, Tokyo, Japan).
Metaphase spreads
WT and Trib2-/- NH9 cells were seeded at 0.2 × 106 cells/ml and treated with DNR/ DMSO from WT and Trib2-/- NH9 cells after 24h DNR treatment from WT and Trib2-/- NH9 cells after 24h DNR treatment for 24h. Demecolcine solution (Sigma) was added to the cell suspension (100 ng/ml) 45 min prior the end of the treatment. The cells were incubated at 37 °C for 15 min in 75 mM KCl (swelling). The cell were fixed in 3:1 methanol:acetic acid. Fixed cells were dropped on microscope slides and let dry at RT. Mounting medium with DAPI (Vector shield) was applied to each sample. The slides were examined at an Axio Imager M1 Epifluorescence and Brightfield Microscope (Zeiss) and pictures captured to show representative mitotic cells (Axio Vision software).
Statistics
Graphs and statistical analyses were performed using GraphPad Prism 5 (GraphPad Software, La Jolla California USA). Flow cytometry data were analysed using FlowJo (Tree Star, Ashland OR USA). Unpaired, two-tailed Student’s t-test was used to test statistical significance whenever comparing two experimental groups with similar variance. Two-way ANOVA was used to evaluate the effect of the genotype (WT and Trib2-/-) on the fold change p-p38 expression over two doses of Ans stimulation. The statistical test used, the P value (*P < 0.05, **P < 0.005, ***P < 0.001) and the number of times each experiment was biologically and technically replicated was indicated in the related figure legends and graphs.
References
Eyers, P. A., Keeshan, K. & Kannan, N. Tribbles in the 21st Century: the evolving roles of tribbles pseudokinases in biology and disease. Trends Cell. Biol. 0, 686–693 (2016).
Wilkin, F. et al. Characterization of a phosphoprotein whose mRNA is regulated by the mitogenic pathways in dog thyroid cells. Eur. J. Biochem. 248, 660–668 (1997).
Wei, S.-C. et al. Tribbles 2 (Trib2) is a novel regulator of Toll-like receptor 5 signaling. Inflamm. Bowel. Dis. 18, 2432–2437 (2012).
Yokoyama, T. et al. Trib1 links the MEK1/ERK pathway in myeloid leukemogenesis. Blood 116, 2768–2775 (2010).
Yokoyama, T. & Nakamura, T. Tribbles in disease: signaling pathways important for cellular function and neoplastic transformation. Cancer Sci. 102, 1115–1122 (2011).
Kritsch, D. et al. Tribbles 2 mediates cisplatin sensitivity and DNA damage response in epithelial ovarian cancer. Int. J. Cancer 141, 1600–1614 (2017).
Liang K. L. et al. Human TRIB2 oscillates during the cell cycle and promotes ubiquitination and degradation of CDC25C. Int. J. Mol. Sci. 17, 1378 (2016).
Liang, K. L., Rishi, L. & Keeshan, K. Tribbles in acute leukemia. Blood 121, 4265–4270 (2013).
Stein, S. J., Mack, E. A., Rome, K. S. & Pear, W. S. Tribbles in normal and malignant haematopoiesis. Biochem. Soc. Trans. 43, 1112–1115 (2015).
Deng, J. et al. Human tribbles homologue 2 is expressed in unstable regions of carotid plaques and regulates macrophage IL-10 in vitro. Clin. Sci. 116, 241–248 (2009).
Eder, K. et al. Tribbles-2 is a novel regulator of inflammatory activation of monocytes. Int. Immunol. 20, 1543–1550 (2008).
Liang, K. L., O’Connor, C., Veiga, J. P., McCarthy, T. V. & Keeshan, K. TRIB2 regulates normal and stress-induced thymocyte proliferation. Cell Discov. 2, 15050 (2016).
Lin, K. R. et al. Murine tribbles homolog 2 deficiency affects erythroid progenitor development and confers macrocytic anemia on mice. Sci. Rep. 6, 31444 (2016).
Keeshan, K. et al. Tribbles homolog 2 (Trib2) inactivates C/EBPalpha and causes acute myelogenous leukemia. Cancer Cell 10, 401–411 (2006).
O’Connor, C. et al. The presence of C/EBPα and its degradation are both required for TRIB2-mediated leukaemia. Oncogene 35, 1–10 (2016).
Gilby, D. C. et al. Tribbles-1 and -2 are tumour suppressors, down-regulated in human acute myeloid leukaemia. Immunol. Lett. 130, 115–124 (2010).
Stein, S. J. et al. Trib2 suppresses tumor initiation in Notch-driven T-ALL. PLoS One 11, e0155408 (2016).
Keeshan, K., Shestova, O., Ussin, L. & Pear, W. S. Tribbles homolog 2 (Trib2) and HoxA9 cooperate to accelerate acute myelogenous leukemia. Blood Cells Mol. Dis. 40, 119–121 (2008).
Wouters, B. J. et al. Distinct gene expression profiles of acute myeloid/T-lymphoid leukemia with silenced CEBPA and mutations in NOTCH1. Blood 110, 3706–3714 (2007).
Hannon, M. M. et al. Elevated TRIB2 with NOTCH1 activation in paediatric/adult T-ALL. Br. J. Haematol. 158, 626–634 (2012).
Sanda, T. et al. Core transcriptional regulatory circuit controlled by the TAL1 complex in human T cell acute lymphoblastic leukemia. Cancer Cell 22, 209–221 (2012).
Nagel, S. et al. Activation of paired-homeobox gene PITX1 by del(5)(q31) in T-cell acute lymphoblastic leukemia. Leuk. Lymphoma 52, 1348–1359 (2011).
Argiropoulos, B. et al. Linkage of Meis1 leukemogenic activity to multiple downstream effectors including Trib2 and Ccl3. Exp. Hematol. 36, 845–859 (2008).
Garcia-Cuellar, M.-P., Steger, J., Fuller, E., Hetzner, K. & Slany, R. K. Pbx3 and Meis1 cooperate through multiple mechanisms to support Hox-induced murine leukemia. Haematologica 100, 905–913 (2015).
Rishi, L. et al. Regulation of Trib2 by an E2F1-C/EBPα feedback loop in AML cell proliferation. Blood 123, 2389–2400 (2014).
Collins, C. T. & Hess, J. L. Role of HOXA9 in leukemia: dysregulation, cofactors and essential targets. Oncogene 35, 1090–1098 (2016).
Alharbi, R., Pettengell, R., Pandha, H. & Morgan, R. The role of HOX genes in normal hematopoiesis and acute leukemia. Leukemia 27, 1000–1008 (2012).
Gough, S. M., Slape, C. I. & Aplan, P. D. NUP98 gene fusions and hematopoietic malignancies: common themes and new biologic insights. Blood 118, 6247–6257 (2011).
Rio-Machin A. et al. The molecular pathogenesis of the NUP98-HOXA9 fusion protein in acute myeloid leukemia. Leukemia 31, 2000–2005 (2017).
Calvo, K. R., Sykes, D. B., Pasillas, M. P. & Kamps, M. P. Nup98-HoxA9 immortalizes myeloid progenitors, enforces expression of Hoxa9, Hoxa7 and Meis1, and alters cytokine-specific responses in a manner similar to that induced by retroviral co-expression of Hoxa9 and Meis1. Oncogene 21, 4247–4256 (2002).
Kvinlaug, B. T. et al. Common and overlapping oncogenic pathways contribute to the evolution of acute myeloid leukemias. Cancer Res. 71, 4117–4129 (2011).
Moore, M. A. S. et al. NUP98 dysregulation in myeloid leukemogenesis. Ann. N. Y. Acad. Sci. 1106, 114–142 (2007).
Kirstetter, P. et al. Modeling of C/EBPα mutant acute myeloid leukemia reveals a common expression signature of committed myeloid leukemia-initiating cells. Cancer Cell 13, 299–310 (2008).
Krivtsov, A. V. et al. Cell of origin determines clinically relevant subtypes of MLL-rearranged AML. Leukemia 27, 852–860 (2013).
Wagner, E. F. & Nebreda, A. R. Signal integration by JNK and p38 MAPK pathways in cancer development. Nat. Rev. Cancer 9, 537–549 (2009).
Platanias, L. C. Map kinase signaling pathways and hematologic malignancies. Blood 101, 4667–4679 (2003).
Hazzalin, C. A. et al. p38 / RK is essential for stress-induced nuclear responses: JNK / SAPKs and c-Jun / ATF-2 phosphorylation are insufficient. Curr. Biol. 6, 1028–1031 (1996).
Thornton T. M. & Rincon M. Non-classical P38 map kinase functions: cell cycle checkpoints and survival. Int. J. Biol. Sci. 5, 44–52 (2009).
Dong, Y. et al. H2AX phosphorylation regulated by p38 is involved in Bim expression and apoptosis in chronic myelogenous leukemia cells induced by imatinib. Apoptosis 19, 1281–1292 (2014).
Wu, X. et al. Resveratrol induces apoptosis of human chronic myelogenous leukemia cells in vitro through p38 and JNK-regulated H2AX phosphorylation. Acta Pharmacol. Sin. 36, 353–361 (2015).
Gao, F. & Liu, W. Advance in the study on p38 MAPK mediated drug resistance in leukemia. Eur. Rev. Med. Pharmacol. Sci. 20, 1064–1070 (2016).
Renneville, A. et al. Cooperating gene mutations in acute myeloid leukemia: a review of the literature. Leukemia 22, 915–931 (2008).
Lin, K. R. et al. Survival factor withdrawal-induced apoptosis of TF-1 cells involves a TRB2-Mcl-1 axis-dependent pathway. J. Biol. Chem. 282, 21962–21972 (2007).
Palma, C. A. et al. MicroRNA-155 as an inducer of apoptosis and cell differentiation in acute myeloid leukaemia. Mol. Cancer 13, 79 (2014).
Hill, R. et al. TRIB2 confers resistance to anti-cancer therapy by activating the serine/threonine protein kinase AKT. Nat. Commun. 8, 14687 (2017).
O'Connor, C. et al. Trib2 expression in granulocyte-monocyte progenitors drives a highly drug resistant acute myeloid leukaemia linked to elevated Bcl2. Oncotarget 9, 14977–14992 (2018).
Acknowledgements
We acknowledge the technical assistant of Jennifer Cassels and the flow cytometry core funding from Kay Kendall Leukaemia Fund and The Howat Foundation. We thank Helen Wheadon, Chinmay Munje, Karen Dunn and all the technical staff at the Paul O’Gorman Leukaemia research Center. We acknowledge the Cancer Research UK Glasgow Center (C596/A18076) and the BSU facilities at the Cancer Research UK Beatson Institute (C596/A17196). KK was supported by The Howat Foundation and Children with Cancer UK grants. KY was supported by Bloodwise (13011). MS was supported by Friends of Paul O’Gorman. SC and ES were supported by Yorkhill Leukaemia & Cancer Research Fund. RJC was supported by the Medical Research Council (MR/M010694/1) and the Biotechnology and Biological Sciences Research 552 Council (BB/M003671/1).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by H.-U. Simon
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Salomé, M., Magee, A., Yalla, K. et al. A Trib2-p38 axis controls myeloid leukaemia cell cycle and stress response signalling. Cell Death Dis 9, 443 (2018). https://doi.org/10.1038/s41419-018-0467-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-018-0467-3
- Springer Nature Limited