Abstract
The HECT (homologous to E6AP C-terminus) ubiquitin ligases (E3s) are a small family of highly conserved enzymes involved in diverse cellular functions and pathological conditions. Characterised by a C-terminal HECT domain that accepts ubiquitin from E2 ubiquitin conjugating enzymes, these E3s regulate key signalling pathways. The activity and functional regulation of HECT E3s are controlled by several factors including post-translational modifications, inter- and intramolecular interactions and binding of co-activators and adaptor proteins. In this review, we focus on the regulation of HECT E3s by accessory proteins or adaptors and discuss various ways by which adaptors mediate their regulatory roles to affect physiological outcomes. We discuss common features that are conserved from yeast to mammals, regardless of the type of E3s as well as shed light on recent discoveries explaining some existing enigmas in the field.

Similar content being viewed by others
Facts
-
HECT E3s utilise several unique mechanisms to regulate their activity and function.
-
Adaptors can modulate HECT E3 activity by direct interaction or utilise different binding domains, affecting phosphorylation, ubiquitination or altering E2–E3 interaction.
-
By deubiquitinating both the adaptor and the E3, DUBs play a pivotal role in regulating HECT E3 activity.
Open questions
-
Is the dynamic exchange of adaptors and HECT E3 that is regulated by DUBs or sorting motifs within the substrate conserved in mammals?
-
What are the mechanisms that determine activation of a specific E3-adaptor in response to a stimulus? Understanding this will be critical in our ability to modulate specific E3 activity and improve strategies for developing better therapeutics.
-
What roles do adaptors, E3s or distinct adaptors-E3 combinations play in determining the type of ubiquitin chain linkages and thus the cellular fate of HECT E3 substrates?
Introduction
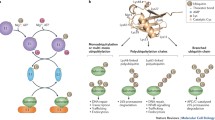
Post-translational regulation of proteins via ubiquitination determines their trafficking, signalling or degradation [1, 2]. Ubiquitination targets a large number of cellular proteins, thus affecting virtually all biological pathways including those involved in cell cycle, development, growth, differentiation, cell death, inflammation, autophagy, protein trafficking and inter-cellular communication [1, 3]. This evolutionarily conserved process involves attachment of a 8.5 kD protein, ubiquitin (Ub), to target proteins by the sequential actions of three enzymes. First, a Ub activating enzyme (E1) activates the Ub molecule in an ATP-dependent manner [4]. Next, activated Ub is transferred to E2, a Ub conjugating enzyme [5, 6], and finally Ub is transferred to a substrate via one of the many Ub protein ligases (E3). Substrates can be mono-, multi- or polyubiquitinated depending on the attachment of Ub at one or more lysine (Lys) residues in the substrate [1]. As Ub itself has seven Lys residues that can be ubiquitinated (K6, K11, K27, K29, K33, K48, K63) in addition to the N-terminus ubiquitination, multiple types of polyubiquitin chains can form, including linear Ub chains [7]. Each E3 can target and regulate several substrates, so their expression, activity and turnover are tightly regulated to prevent inappropriate ubiquitination and cellular dysfunction [8]. Some important factors that determine efficiency of E3 activity and how ubiquitination proceeds depend on E3 interaction with E2, substrate recognition and on the type of Ub linkage formed (mono- or polyubiquitin chains). Depending on the type of linkage, target proteins can have different fate. For example, K48-linked chains are most common and associated with proteasomal degradation, K63 linkages are involved in endocytotic trafficking, inflammation and DNA repair and K11 linkages are implicated in mitotic regulation and endoplasmic reticulum–associated degradation [9]. Furthermore, Ub can also be modified by Ub-like molecules, acetylation and phosphorylation, thus providing potentially numerous types of modifications for ubiquitinated substrates [7]. Deubiquitinases (DUBs) form another integral part of the Ub modification system as they remove Ub from the ubiquitinated proteins [10].
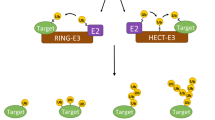
E3s comprise the largest family of enzymes in the Ub system, as they are responsible for recognising and ubiquitinating specific substrates. These enzymes belong to three groups: (i) HECT (homologous to the E6-AP C-terminus) E3s, which have an active site cysteine (Cys) that accepts Ub from an E2 before transferring it to a specific substrate, (ii) RING (really interesting new gene) domain-containing E3s, which act as a scaffold and allow the transfer of Ub directly from an E2 to a substrate and (iii) RBR (RING1-between-RING2) E3s, which function as hybrids of RING and HECT E3s [11]. Here Ub-conjugated E2 binds to one RING domain of a RBR E3 and is then transferred to the other RING domain enabling the formation of a E3∼Ub thioester bond before conjugation to respective substrate [11].
The HECT E3s
There are more than 600 E3s encoded by the human genome [12]. While most E3s belong to the RING family, HECT ligases form the second-largest group with 28 members in mammals. E6AP (E6-associated protein) and NEDD4 (Neural precursor cell–expressed developmentally downregulated 4) were the first identified E3s and the name HECT originated from E6AP that ubiquitinates p53 in cells expressing the human papilloma virus protein E6 [13, 14]. The HECT domain is the catalytic region comprising of ~350 residues with a N-terminal lobe, which binds an E2, and a C-terminal lobe harbouring an active Cys residue that receives Ub (Fig. 1a). Both lobes are connected by a flexible hinge region [15]. The N-lobe also contains a non-covalent Ub-binding site called Ub-exosite (Fig. 1a) that engages with a regulatory Ub molecule and is involved in elongation and stabilisation of the growing Ub chain [16]. Importantly, the substrate specificity of HECT E3s is determined by their respective N-terminal regions that often comprise one or more protein–protein interaction domains that facilitate binding to substrates or adaptor proteins [3].
a HECT domain contains a bulky N-terminal lobe (N lobe) and a smaller C-terminal lobe (C lobe) that contains the catalytic Cys. N lobe contains a site to receive the charged E2 that then transfers ubiquitin (Ub) to the Cys residue. A flexible linker region allows changes in orientation to occur easily. Ub-binding exosite that can non-covalently hold Ub is shown. b Schematic representation of domain structure of the S. cerevisiae HECT E3 Rsp5p as the prototypic NEDD4 family member, and various human HECT E3s. C2 calcium-binding domain, WW WW domains, RLD RCC1 like domain, SPRY B30.2 SPRY domain, WD40 W-D repeat domain, MIB MIB/HERC2 domain, CPH conserved domain within Cul7, PARC and HERC2 proteins, GBL galactose-binding like domain, ARM armadillo repeat containing domain, BH3 Bcl-2 homology 3 domain, UBA ubiquitin-associated domain, WWE WWE-containing domain, Znf zinc finger–containing domain, IQ IQ motif/ EF hand-binding site, DOC APC10/DOC domain, AZUL amino terminal Zn finger of Ube3a ligase, ANK ankyrin repeat-containing domain, PHD PHD-type zinc finger, CytB cytochrome-b5–like heme/steroid-binding domain.
HECT ligases are broadly divided in three groups/subfamilies according to the presence of distinct domains that they contain (Fig. 1b). The NEDD4 family of HECT E3s typically contain N-terminal C2 domain, 2–4 WW domains and a C-terminal HECT domain, the HERC (HECT and RCC1-like domain) E3s have at least one or more RCC1-like domains (RLDs), whereas the ‘other HECT' E3s may contain varied number and types of domains [3].
The NEDD4 family of E3s are the best studied of all HECTs. They are evolutionarily conserved from yeast to mammals and regulate several cellular processes including transcription, stability and trafficking of plasma membrane (PM) proteins and the degradation of misfolded proteins [17]. Several other NEDD4 family members are also involved in regulating virus egress from infected cells [18, 19]. The N-terminal C2 domain in NEDD4 family members is a calcium-responsive lipid binding motif, whereas the WW domains recognise PPxY (PY) or LPxY motifs in substrates or accessory proteins [12, 20, 21]. Mammals have nine NEDD4 members [3] including NEDD4, NEDD4-2 (NEDD4L), ITCH, SMURF1, SMURF2, WWP1, WWP2, NEDL1 (HECW1) and NEDL2 (HECW2) and domain organisation of the human proteins is shown in Fig.1b. Saccharomyces cerevisiae, which has provided pioneering functional information about HECT ligases, has a single NEDD4 E3, Rsp5p, that plays an essential function in cellular homeostasis [22].
The HERC family comprises six E3s, HERC1–HERC6. HERC1 and HERC2 are very large proteins (>500 kDa) with multiple domains (Fig. 1b), in addition to the RLD domains, whereas HERC3–6 in general only contain RLD and HECT domains [3]. Among the remaining HECT E3s, E6AP, HUWE1, EDD, TRIP12 and HACE1 have been studied to various extents [3]. These HECT E3s have a variety of different domains in their N-terminal region (Fig. 1b).
The ability of HECT E3s to synthesise linkage-specific polyubiquitin chain determines the fate of its target protein and, unlike RING E3s, is independent of the paired E2. Generally, NEDD4 family members form K63 linkages [23], but can also form K48 and other types of chains [23, 24]. HUWE1 generally forms K6-, K11- and K48-linked chains [25], whereas E6AP more often generates chains with K48 linkage [23]. While precise mechanisms underpinning Ub chain formation and its specificity remain unknown, remarkable progress has been made in understanding how this controls cellular processes [7, 23, 26,27,28].
Regulation of HECT E3s
Among the regulatory mechanisms that control E3 expression include tissue-specific transcription, cellular microenvironment and substrate concentration [29,30,31,32], while E3 catalytic activity can be regulated in multiple ways (Fig. 2). This includes post-translational modifications (PTM), intra- and intermolecular interactions, intrinsic catalytic activity (Ub exosite), strength of E2–E3 interaction, interaction with specific adaptors and DUBs [22, 33]. This review discusses the various modes regulating HECT E3 activity with a special focus on the crucial roles of adaptors.
(1) Post-translational modifications, e.g., phosphorylation; (2) intermolecular interactions, e.g., trimerisation of E6AP; (3) intramolecular interactions, e.g., NEDD4 autoinhibition by C2–HECT interaction; (4) intrinsic catalytic activity (Ub-exosite) mediated ubiquitination; (5) strength of E2–E3 interaction; (6) adaptor protein binding and (7) interaction with DUBs.
Post-translational modifications
Protein modifications such as phosphorylation, ubiquitination and neddylation can significantly alter the activity of an E3 ligase [34,35,36]. This is observed in case of ITCH where the self-inhibitory interaction is disrupted by JNK1-mediated phosphorylation of S199, T222 and S232 within its proline-rich region that alters the conformation of the WW domain and greatly enhances catalytic activity (Fig. 3a) [37]. The linker region L can also be targeted to relieve autoinhibition as it has tyrosine residues that when phosphorylated increase HECT activity [38]. Similarly, a c-Src kinase–dependent tyrosine phosphorylation of NEDD4 enhances its catalytic activity [34]. NEDD4 is phosphorylated when stimulated by fibroblast growth factor at Tyr43 in the C2 domain and Tyr585 in the HECT domain and this relieves autoinhibition, allowing NEDD4 to ubiquitinate fibroblast growth factor receptor 1, leading to its internalisation and degradation (Fig. 3b). Phosphorylation of E6AP by c-Abl also negatively regulates its catalytic activity [39]. Ubiquitination of Rsp5p [40], E6AP [41] and TRIP12, a Ub E3 ligase [42], modulates their activity, whereas NEDD8 binding to SMURF1 and 2 promotes their ligase activity [43]. Other HECTs that can be neddylated include NEDL1, NEDDL2 and ITCH [44]. SUMOylation of SMURF2 also enhances its ubiquitination activity and suppresses TGF signalling [36].
a JNK1 binds a region in HECT domain of ITCH and phosphorylates 3 sites in its N-terminal proline-rich region (PRR). This relieves the autoinhibition, allowing JunB to bind ITCH and its ubiquitination. Regulatory linker region (L) is shown. b Tyrosine phosphorylation of NEDD4 by c-Src both at C2 and HECT domain enhances its catalytic activity, fibroblast growth factor receptor 1 (FGFR1) ubiquitination, internalisation and degradation. c NEDD4-2 autoinhibition can be relieved by increasing Ca2+ concentration. High Ca2+ enables competition between HECT-binding and Ca2+-binding regions on C2 domain, and can also determine cellular relocalisation of NEDD4-2 via its interaction with membrane bound PIP2/IP3 (phosphatidylinositol 4,5 biphosphate)/IP3 (inositol 1,4,5 triphosphate). d Non-covalent binding of interferon stimulated gene 15 (ISG15) to NEDD4 prevents it interaction with E2 and attenuates catalytic activity.
Intermolecular interactions
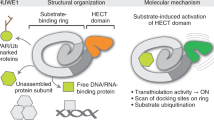
Homodimerisation and oligomerisation in some HECT E3s serve a regulatory role [45, 46]. For example, homodimerisation of HECT domains of HUWE1 leads to autoinhibition. However, a conserved activation segment at the N-terminal of HUWE1 interacts with the dimerisation to relieve autoinhibition [46]. In addition, binding of proteins such as p14ARF to this conserved activation region promotes dimerisation of HUWE1 changing its conformation to an inactive state [46]. Autoubiquitination of HECT E3s is crucial regulatory mechanism conserved from yeast to mammals. Rsp5p and NEDD4 display trimerisation after autoubiquitination that causes reversible inactivation and stabilisation of several targets [45]. In contrast, trimerisation of E6AP promotes its ligase activity [47]. SMURF1 C2 domain binds and inhibits HECT activity in trans [48] and is known to form homodimers via interaction between C2 and HECT domains [49].
Intramolecular interactions
One of the more commonly exploited mechanisms by HECT E3s is modulation of catalytic activity by strong intramolecular interactions between N- and C-terminal regions. Typically, this is achieved via C2 or WW and HECT domain interaction. For example, ITCH autoinhibition involves interaction between WW domains and the linker region between WW domains with HECT region [38, 50], while SMURF2 undergoes autoinhibition by interaction between its C2 and HECT domain [51]. The C2 domain binds close to Ub-exosite and interferes with Ub thioester formation [51]. WW1 domain in SMURF2 acts as an additional inhibitory region and its absence in SMURF1 promotes its constitutive open/active conformation [48]. A recent study revealed how some NEDD4 E3s are autoinhibited using a combination of regulatory sites named Re (right ear), Le (Left ear) and Hinge (H) in the HECT domain [52]. WWP1, WWP2 and ITCH utilise a highly similar multi-lock autoinhibition mechanism, whereas NEDD4, NEDD4-2 and SMURF2 have a slightly different process. Interaction between WW domains and PY-containing adaptor/substrate is sufficient to relieve autoinhibition of WWP1/2 and ITCH. However, for NEDD4, NEDD4-2 and SMURF2, both membrane targeting of C2 domain and substrate binding with WW domains are required for complete activation. Linker region in several HECT E3s have a significant role in regulating HECT activity [50]. Deletion of a linker region in NEDD4 (WW1–WW2) was found to result in increased autoubiquitination [50]. Overall, these observations suggest versatile autoinhibitory mechanisms that fine-tune the ligase activities of the HECT E3s [52]. As discussed later, adaptors play an important role in relieving this autoinhibition.
Phosphorylation of some E3s such as ITCH and NEDD4, nutrients availability and Ca2+ can also alter protein structure and ligase activity [53]. NEDD4-2 forms an autoinhibited state via interaction between its C2 and HECT domains. It was observed that the C2 domain of NEDD4-2 binds Ca2+ and inositol 1,4,5-trisphosphate (IP3) using the same interface that is used for binding HECT domain (Fig. 3c). Thus, changes in cytoplasmic Ca2+ can initiate transition from the closed to the active form as a result of competition between IP3 and Ca2+ and the HECT domain to bind C2 domain [53].
Ubiquitin-binding site (Ub-exosite)
Pioneering work in yeast [54] and mammals [16] led to the identification of a non-covalent Ub-binding site within HECT domain of NEDD4 E3s called Ub-binding site (Ub-exosite) that is involved in assembly and elongation of polyubiquitin chains on the substrate. Further studies indicated that Ub-exosite can modulate E3 activity [45]. Exosite-mediated autoubiquitination of specific lysine residues in α1-helix of Rsp5p HECT exerts a pull resulting a conformational change exposing the trimerisation interface and thus inhibiting activity [45]. A similar mechanism was also observed for mammalian NEDD4 [45]. Another study found that ubiquitinated Rsp5p adaptors locked at exosite were more active providing yet another function for this site [55]. An elegant screen for Ub variants that bind to exosite of various HECTs with varying affinity has provided more insight how the activity of different HECT E3s can be modulated via exosite [56].
E2–E3 interaction
E2–E3 interaction can be negatively regulated by inhibitors. Ub-like protein, ISG15 (interferon stimulated gene 15), when co-expressed with VP40, prevents NEDD4 interaction with its E2 (Fig. 3d) and attenuates viral budding [57, 58]. SMURF2 has low affinity for its E2 UbcH7 and depends on adaptors such as SMAD7 to promote E2–HECT interaction and enhance E3 activity [59].
Adaptors
Several HECT E3-binding proteins or ‘the adaptors’ have emerged as potent regulators of HECT enzymes. Adaptors can influence E3 localisation, catalytic activity and also regulate E3–substrate interaction [60]. In case of NEDD4 family members that are found in different subcellular locations and have several substrates lacking the classical proline-rich PY motif for interaction with the WW domains of NEDD4 protein [17], adaptors are vital since many contain PY motifs that facilitate the E3 to reach their substrates efficiently. Several other adaptors utilise different binding domains on E3 affecting phosphorylation, ubiquitination, alter E2–E3 interaction and thus regulate ligase activity. In the following section we discuss these and other recent ground-breaking discoveries in S. cerevisiae and mammals that enhance our understanding about regulatory functions of the HECT E3 adaptors.
Deubiquitinases
DUBs have crucial role in regulating E3 activity by removing Ub from both adaptor and E3s and recycling them [10]. These are discussed in the later sections.
Regulation of HECT E3s by adaptors and co-activators
Figure 4 shows some of the major HECT E3 adaptor types in yeast and mammals and Table 1 summarises their roles in regulating E3 activity.
Yeast Rsp5p and its adaptors
Investigations in S. cerevisiae have played a critical role in understanding how ubiquitination regulates membrane-associated nutrient transporters. Proteins synthesised in the endoplasmic reticulum are modified in Golgi and are released in membrane-bound vesicles to their respective destination, such as PM for transmembrane (TM) proteins [Fig. 5a(i)]. The endocytosis of many yeast transporters is triggered by nutrient ligand, a process that is dependent on Rsp5p, the only NEDD4 orthologous in S. cerevisiae. Initial studies revealed that several PY motif–containing adaptor such as Bsd2p protein [61] and arrestin-related proteins (ARTs) bind Rsp5p and downregulate amino acid transporters (AAT) [62].
a (i) Transmembrane (TM) proteins such as amino acid transporters (AAT; e.g., Mup1p) and metal ion transporters (e.g., Smf1p) are normally trafficked to the plasma membrane (PM). (ii) The adaptor protein Bsd2p (Ndfip orthologous) localised at Golgi recruits Rsp5p to ubiquitinate TM proteins (e.g., Smf1p) that can then be trafficked via endosome and degraded in the vacuole. Bsd2p–substrate interaction is assisted by PY motif–containing adaptors Tre1. (iii) Environmental stress can initiate adaptor-induced ubiquitination of other transporters to overcome stress. For example, under basal conditions, Chitin synthase3 (Chs3p) undergoes constitutive protein endocytosis (blue arrows). Stressed cells display increased intracellular calcium levels, resulting in enhanced Calcineurin activity, Rcr1 upregulation and sorting to the PM. As a result more Rsp5p is recruited to PM leading to increased Chs3p ubiquitination, exiting the retrograde trafficking and endocytosis, and degradation in the vacuole. (iv) Selective endocytosis is utilised in order to adapt to nutrient availability. This is brought about by the E3 recognising and complexing with different adaptors. For example, during amino acid and nutrient starvation, Art2 is upregulated and recognises C-terminal acidic sorting motifs in AATs and recruits Rsp5p to ubiquitinate proximal Lys residues (starvation-induced endocytosis). When amino acids are in excess, Rsp5p instead uses TORC1-activated Art1 to detect N-terminal acidic sorting motifs within the same AATs, to target Lys residues in the C-terminal thus facilitating substrate-induced endocytosis. b Adaptors also control HECT E3 activity by self-ubiquitination. Several adaptors compete for occupancy on Rsp5p. For example, the PY motif–containing protein Hua1 acts as an adaptor for Rsp5p. On binding, Hua1 undergoes ubiquitination by Rsp5p, binding Rsp5p more efficiently and increasing its catalytic activity. Ub-Hua1 also prevents other adaptors to bind to Rsp5p. DUBs such as Ubp2p then remove the ubiquitinated adaptor, allowing other adaptors to bind.
Bsd2p
Yeast Bsd2p is the homologous of mammalian NEDD4 family-interacting proteins (NDFIPs) [63, 64]. Bsd2p possesses a PPxY motif that binds to the WW domains of the Ub ligase Rsp5p, and this leads to formation of a substrate–Bsd2p–Rsp5p complex. Bsd2p contributes to the ubiquitination and sorting of membrane proteins into the multivesicular bodies (MVB) such as the vacuolar proteins Cps1p and Phm5p; the manganese transporters Smf1p and Smf2p [61]. In case of Smf1p, Bsd2p recruits another TM adaptor, Tre1 that also has a PY motif, to bridge the gap between Bsd2p and Smf1p [Fig. 5a(ii)] [65, 66].
Arrestins
Arrestins family of proteins comprise of three groups-α, β and visual arrestins [1]. α-arrestins or arrestin domain–containing proteins (ARRDC, in mammals) contain multiple PY motifs that recruit NEDD4/Rsp5p, to ubiquitinate many endocytic cargo proteins [62, 67]. Yeast contains 14 proteins with α-arrestin domains, namely Art1/Ldb19, Art2/Ecm21, Art3/Aly2, Art4/Rod1, Art5/Ygr068c, Art6/Aly1, Art7/Rog3, Art8/Csr2, Art9/Rim8, Art10 and Bul1–3 and Spo23 [68, 69]. All of these bind to Rsp5p and become ubiquitinated [70, 71]. Art1 and Art2 direct downregulation of distinct AAT by recruiting Rsp5p [62]. Endocytosis of the yeast pheromone G-protein–coupled receptor (GPCR), Ste2p, is mediated by Rsp5p and facilitated by ARTs [72]. Furthermore, it has been shown that Art1 is involved in selective regulation of mating events after receptor internalisation and cell cycle arrest, leading to zygote formation [73]. Some other arrestins such as Art8/Csr2 or Art2/Ecm21 are involved in facilitating the ubiquitination and endocytosis of Smf1p [67] as well as in regulating glucose transporters such as Hxt6p [74]. Glucose signalling in yeast also involves other transporters such as Jen1. Adaptor proteins Bul1 and Art4/Rod1 mediated the stepwise ubiquitination and endocytosis of Jen1 [75]. Thus, multiple adaptors can act sequentially along the endocytic pathway to control transporter homeostasis [75].
Rcr proteins
Additional yeast adaptors that can alter cellular response to nutrient stress have recently been identified. Rcr1/Rcr2 alter nutrient transport by modulating intracellular trafficking and determining whether membrane proteins will be targeted to the vacuole or the PM [76]. It was also observed that overexpression of Rcr1 results in increased resistant to Congo red by reducing the chitin content in the cell wall [77]. Rcr1 responds via calcineurin pathway to calcium levels and is upregulated during conditions of excess calcium that is often observed in cells experiencing environmental stress. Newly synthesised Rcr1 is sorted to the PM where it binds Rsp5p via its PY motifs and causes chitin synthase3 (Chs3p) downregulation [78]. Mechanistically, Chs3p is endocytosed and instead of getting recycled to the Golgi, ubiquitinated Chs3p gets sorted to the vacuole in a Ca2+-dependent manner [Fig. 5a(iii)]. This highlights that ubiquitinated cargoes can be trafficked to different intracellular locations following cellular stress. Importantly, this process relies on increasing the adaptor protein abundance at the PM, which in turn causes higher E3 ligase (Rsp5p) availability and activity [78].
Others proteins and deubiquitinases
Bmh1 and 2 are yeast homologous of 14-3-3 proteins involved in multiple signalling processes including pseudohyphal differentiation, TOR signalling, stress response, exocytosis and vesicle transport, protein degradation and glucose repression [79]. The yeast ART8/Csr2 contains a consensus 14-3-3 binding site (Rxx[pS/pT]xP) [80] that enables it to control Hxt6p endocytosis. During conditions of glucose deprivation, Csr2/Art8 is transcriptionally induced and becomes activated by ubiquitination causing receptor endocytosis. In contrast, glucose replenishment causes PKA-dependent Csr2 phosphorylation and Csr2–Bmh interaction results in Csr2 deubiquitination (possibly involving an unknown DUB) and inactivation [80]. Similarly, Bmh1 and Bmh2 control Bul arrestin-like adaptors that ubiquitinate general amino acid permease, GAP1 [81]. Rsp5p activity is regulated by DUBs such as Ubp2p [82]. The stability of ARTs also depends on DUBs. For example, Ubp2p and Ubp15p prevent hyper-ubiquitination and degradation of ARTs (Art4) by Rsp5p. Loss of these DUBs results in reduced Art4 and a defect in Hxt6p endocytosis [83].
Rup1, Sna3p, Ear1p, Ssh4p are some other proteins that can modulate either E3 or other adaptors and thus alter enzyme activity. Rup1 is a UBA domain protein with a PPPSY motif that binds to Rsp5p and controls Rsp5p–Ubp2p interaction [40]. Sna3p, Ear1p and Ssh4p were identified as Rsp5 adaptors crucial for cargo ubiquitination and in sorting them into endosome/MVBs [84, 85]. Interestingly, the ubiquitination of Sna3p was found to be critical in determining the efficacy of this process [84]. Ear1p and Ssh4p mediate recognition and vacuolar degradation of PM proteins that are misfolded/mistargeted and escape quality control systems [86].
The observations above suggest a dynamic relationship between an E3 and its cognate adaptors, and that E3s often partner with different adaptors depending on context, resulting in varied cellular responses. Two recent studies suggest alternate mechanisms explaining this. Ivashov et al. propose that the recruitment of different adaptors to an E3 in response to varied stimuli may depend on some recognition motifs within the substrate itself [87]. For example, Rsp5p complexes with Art2 during starvation and Art1 during periods of nutrient excess to control selective endocytosis of membrane transporters and adjust nutrient transport during these conditions. Importantly, while both Art1–Rsp5p and Art2–Rsp5p can promote substrate-induced and starvation-induced endocytosis, respectively, the mechanism of action varies considerably [Fig. 5a(iv)]. During amino acid and nutrient starvation, Art2 is upregulated and recognises C-terminal acidic sorting motifs in AATs and recruits Rsp5p to ubiquitinate proximal Lys residues (starvation-induced endocytosis). However, when amino acids are in excess, Rsp5p instead uses TORC1-activated Art1 to detect N-terminal acidic sorting motifs within the AATs, to target Lys residues in the C-terminal, thus facilitating substrate-induced endocytosis [87].
Another model explaining how the same ligand activates multiple adaptors and how they compete for E3 ligase was proposed by MacDonald et al. [55], wherein ubiquitination of the adaptors via the E3 promotes its catalytic activity that can be regulated by DUBs (Fig. 5b). Here, Rsp5p ubiquitinates the adaptor Hua1 and Ub–Hua1 docked on Rsp5p enhances its catalytic activity [55]. Rsp5p utilises multiple sites for binding different adaptors and they compete to occupy these sites. Adaptor switching involves DUBs such as Ubp2p that remove Ub from the docked adaptor, disengaging it from E3 and allowing other adaptors to bind (Fig. 5b). These studies provide additional insight on how different adaptors can target distinct regions of the same E3 and choose unique substrates.
Mammalian HECT E3s and their adaptors
In mammals the major group of adaptors for NEDD4 E3s include NDFIPs (NDFIP1 and 2) and arrestins (α and β arrestins) that interact with WW domains of NEDD4 via their PY motifs. Other HECT E3 adaptors interact with HECT E3s by binding to other regions and are discussed below.
NDFIP1 and NDFIP2
NDFIPs are the earliest known mammalian adaptors of HECT E3s [64]. NDFIP1 (originally named N4WBP5) was discovered in a screen designed to identify NEDD4 -interacting proteins that bind to its WW domains [64]. Subsequently, database searches identified a related member NDFIP2 (N4WBP5A) [63, 88]. NDFIPs are evolutionarily conserved with a single orthologous in yeast (Bsd2p) and Drosophila (dNdfip). As stated above, Bsd2p regulates Rsp5p [61, 65]. In Drosophila dNdfip promotes dNedd4- and Su(dx) (mammalian ITCH)-mediated Notch ubiquitination [89].
NDFIPs contain three N-terminal PY motifs, three TM domains and a short C-terminal region (Fig. 4). They localise to the Golgi, endosomes and MVBs [21, 63, 90]. Both NDFIP1 and NDFIP2 interact with multiple members of the NEDD4 family E3s. NDFIPs have been shown to mediate autoubiquitination of E3s while simultaneously also relieving their autoinhibition by binding to WW domains resulting in enhanced catalytic activity [91]. They use the PY motifs to interact with WW domains of ITCH and other HECT E3s to relieve their autoinhibition and this facilitates ubiquitination of substrates such as Jun and endophilin [91]. NDFIP1 and NDFIP2 are also involved in localising Robo1 receptors to endosomes. The Robo receptors are conserved in flies and mammals and mediate axon guidance at midline. NDFIPs recruit NEDD4 to promote Robo1 ubiquitination and degradation [92]. Furthermore, NDFIPs have been shown to recruit divalent metal transporter (DMT1, functional equivalent of yeast Smf1p) to WWP2 and NEDD4-2 to regulate the transporter abundance and activity [93, 94]. Increased DMT1 caused by loss of Ndfip1 in mice results in hemochromatosis [95] and metal toxicity in neurons [96]. Ischemia-induced rat brain injury also involves Ndfip1 and Nedd4-2 upregulation [97]. Most importantly, NDFIP1 has a critical role in suppressing inflammation and Ndfip1 KO mice die due to generalised inflammatory disease resulting from hyperactive T cells that induce T helper cell–mediated inflammation [98]. In one proposed mechanism NDFIP1 binds to ITCH and mediates JunB degradation thus preventing activation of the immune response [98]. NDFIP1, along with NEDD4-2, also acts as a negative regulator of IgE-mediated mast cell activation via targeting phosphorylated (p)-Syk [99] and NDFIP1 and NDFIP2 are necessary for NEDD4- and NEDD4L-mediated ubiquitination and degradation of water channel, aquaporin 2 (AQP2) in kidney cells [100].
Other proposed roles for NDFIPs include regulation of cell signalling via epidermal growth factor receptor by promoting NEDD4/ITCH-mediated ubiquitination of Phosphatase and tensin homologous (PTEN), c-Cbl and Src family kinases Lyn and Src ubiquitination. In this way by reducing cellular NDFIP levels, signalling via AKT pathway that is negatively regulated by PTEN is diminished, while JNK signalling is stimulated that results in cellular growth [101]. In another study, NDFIP1 was found to be involved in the Ub-mediated degradation of mitochondrial antiviral signalling (MAVS), which is a key adaptor in RIG-I-like receptor–mediated immune signalling [102]. When overexpressed, NDFIP1 severely impaired MAVS by enhancing SMURF1 activity and promoting interaction with MAVS leading to its degradation [102].
Interestingly, by interacting with a different E3, NDFIPs can alter the fate of a target protein. For example, NDFIP1 promotes NEDD4 binding to and ubiquitination of PTEN, which allows PTEN to move into the nucleus (Fig. 6a) [103]. On the other hand, recruitment of WWP2 by NDFIP1 leads to K48-linked Ub chain formation on PTEN and its degradation [104].
a NDFIPs bind to WW domains of NEDD4 via their PY motifs and this induces conformational alterations enabling NEDD4 to bind PTEN, undergo mono-ubiquitination and subsequent translocation to nucleus. However, recruitment of WWP2 by NDFIPs causes K48-linked polyubiquitination and degradation of PTEN. NUMB is another adaptor for NEDD4 that ubiquitinates PTEN and targets it for degradation. This suggests that adaptors can target the same substrate to form different type of ubiquitin linkages resulting in different outcomes. b and c Roles of the mammalian adaptor proteins NDFIPS and ARRDC in regulating ubiquitin-mediated protein trafficking. b Multistep regulation of G-coupled protein receptors (GCPR; e.g., β2-AR). Both α- and β-arrestins have been implicated in receptor trafficking. While both arrestins can act together at PM, others [114] propose their sequential recruitment. First, the activated receptor is phosphorylated and this allows β-arrestin2 to bind MDM2 (a RING E3), causing its ubiquitination and internalisation of the entire complex. Next, MDM2 is displaced by NEDD4 that ubiquitinates the receptor. ARRDC3 is activated at PM and, along with other ESCRT members, recognises and binds NEDD4–β2-AR complex leading to further sorting of the internalised ubiquitinated receptor to the lysosomes. c NDFIPs recruit NEDD4 E3s to ubiquitinate several TM proteins (e.g., DMT1) at Golgi that can be trafficked to the endosome and targeted for degradation in the lysosome. Some substrates are also released in exosomes (such as PTEN). Other adaptors such as ARRDC family are often localised near plasma membrane where they hijack a viral budding mechanism involving other proteins from the ESCRT pathway and an ATPase, to interact with an E3, promoting ubiquitination of TM proteins (such as DMT1) and their release into large EVs called microvesicles.
Arrestin family
α-arrestins
In mammals, α-arrestins or ARRDC contain multiple PY motifs that recruit NEDD4 family E3s to ubiquitinate many endocytic cargo proteins [62, 67]. Mammalian α-arrestins include ARRDC1–5 and TxNIP and all except ARRDC5 have PY motifs in their C-terminal region (Fig. 4) [105]. Among the PY-containing α-arrestins, ARRDC3 regulates the β2-AR by recruiting NEDD4 and trafficking receptor–NEDD4 complexes to endosomes [106]. ARRDC1 heterodimerises with β-arrestin1 and recruits ITCH to non-activated NOTCH, facilitating its ubiquitination and degradation [107]. In addition, ARRDC1 is involved in the budding of extracellular vesicles (EVs) from the PM [108]. ARRDC3 was also found to co-localise and facilitate ubiquitination of the endosomal sorting complex required for transport (ESCRT) protein ALIX by WWP2. Ubiquitinated ALIX then binds and targets protease-activated receptor-1, a GPCR, for lysosomal degradation [109]. Recent studies have identified ARRDC1 and ARRDC4 as adaptors for NEDD4 family of E3s that mediate DMT1 release in EVs [110]. In another study, α-arrestins were shown to mediate aggregation of unstable proteins leading to their ubiquitination by NEDD4 E3s [111].
β-arrestins
β-arrestins have been extensively studied for their role in regulating GPCRs where they act as adaptors for both RING and HECT ligases. Mammalian β-arrestins include arrestin1 (S-arrestin), arrestin2 (β-arrestin-1), arrestin3 (β-arrestin-2) and arrestin4 [105] (see Table 1 and Fig. 4). One of the best examples of β-arrestin–mediated regulation is that of two-step degradation for β-2-adrenergic receptor (β2-AR). Normally ligand stimulation phosphorylates the receptor and this triggers recruitment of β-arrestin-2 and the MDM2 (a RING E3). MDM2 then ubiquitinates β-arrestin-2, leading to receptor internalisation into endosomes. Here, NEDD4 replaces MDM2 and binds to β-arrestin-2–β2-AR complex and ubiquitinates the receptor [112]. As both ARRDC3 and β-arrestin-2 are involved in β2-AR trafficking, it is likely they work together (Fig. 6b). While there is some controversy over how this occurs, two distinct models have emerged, one stating that both ARRDC3 and β-arrestin-2 act at the PM co-ordinating their actions during early endocytosis of the receptor [113], while others suggest that they are sequentially recruited with β-arrestin-2 first internalising the receptor followed by NEDD4 recruitment and ubiquitination of receptor [114]. ARRDC3, constitutively activated at the PM and associated with ESCRT proteins, then binds NEDD4–β2-AR complex, and sorts the ubiquitinated receptor to lysosomes (Fig. 6b). This clearly indicates that arrestins can interact directly with Ub ligases, thereby stimulating the ubiquitination of PM receptors, and in some cases, the arrestin itself (Fig. 6b).
Ubiquitination is implicated in sorting proteins into MVBs and EV formation [115], and NDFIPs and ARRDCs are involved in this process (Fig. 6c) [108, 116, 117]. NDFIP1 was found to be released in exosomes and was crucial in targeting both NEDD4 E3s and PTEN to exosomes [116, 117]. Microvesicles (MVs) are defined as membrane-bound vesicles that directly bud off the PM. While NDFIPs can target membrane proteins such as DMT1 for internalisation into endosomes and lysosomal degradation [93], ARRDC1 and ARRDC4 act as adaptors between DMT1 and NEDD4-2 and facilitate the release of DMT1 in MVs (Fig. 6c) [110]. This process is aided by other proteins, for example, ARRDC1 requires ESCRT components Tsg101 and VPS4, while ARRDC4 requires Rab11a [110].
PRRG proteins
The family of mammalian proline-rich γ-carboxyglutamic acid (PRRG) proteins includes PRRG1–4 that were discovered about two decades ago [118]; however, their biological functions remain largely unknown. PRRG proteins are ~200 amino acids long TM proteins with an extracellular γ-carboxyglutamic (Gla) domain (characteristic of this family), PY motifs and a possible SH3 domain [118, 119]. This family of adaptors shares features similar to other small TM proteins with PY motifs that includes Rcr1 and Rcr2 in S. cerevisiae, and Comm (Drosophila commissureless).
In WAGR (Wilms tumor, aniridia, genitourinary malformations and mental retardation syndrome) PRRG4 is one of the major genes deleted [120]. Deletion of PRRG4 in this syndrome is associated with autism side of the disease [121]. Another interesting function for these proteins may be in response to food allergens. In a study on mesenteric lymph node gene expression profiles in mice exposed to three common food allergens, β-lactoglobulin, ovalbumin and peanut agglutinin, PRRG4 was identified as second most upregulated gene [122]. Furthermore, a genome-wide association study of genes influencing the onset of Parkinson disease with age showed that the most strongly associated SNP was in PRRG4 gene [123].
Interaction between NEDD4 and PRRG [64] and Yes-associated protein 1 (YAP1) and PRRG is known [119]. Yazicioglu et al. [124] identified new WW-binding motif in PRRG4 and confirmed interaction of PRRG4 cytosolic domain with membrane-associated guanylate kinase 1 transcriptional co-activator with PDZ motif (WWTR1/TAZ), YAP, NEDD4-2 and NEDD4. Furthermore, PRRG2 binds to most of PRRG4 interactors [124]. It was also observed that PRRG4 promoted migration and invasion of breast cancer cells [125], via binding to NEDD4 and inducing ubiquitination and degradation of Robo1. While other roles for mammalian PRRG proteins remain to be explored, given the recently discovered functions for Rcr1 and Rcr2 as Rsp5p adaptors, as discussed above, there is strong likelihood that PRRGs have analogous functions in mammals.
Others adaptors and co-activators/inhibitors
Viral proteins
Another PY motif–containing adaptor is the viral protein VP40 in Ebola virus and the Marburg virus [126] that interacts with WW domains of NEDD4. Importantly, the budding of these viruses depends on both NEDD4 WW domain–mediated VP40 interaction and NEDD4 catalytic activity [18] suggesting ubiquitination of some viral proteins by NEDD4 enhances release of virus particles (Fig. 7a). In HPV-infected cells oncoprotein E6 acts as an adaptor and binds the E3 E6AP via a LxxLL motif located at its N-terminus. This allows E6/E6AP to recruit p53 and promotes its proteasome-mediated degradation (Fig. 7b) [127]. E6/E6AP also targets other proteins that contribute to E6-induced cellular transformation such as the TERT (telomerase reverse transcriptase) repressor NFX1-91, E6TP1 (E6 targeted protein1), MCM7 (mini chromosome maintenance protein 7) and BAK (Bcl-2 homologous antagonist killer) [128]. Furthermore, it was also shown that E6 binding not only increases E6/E6AP ability to recruit substrates but also enhances the catalytic activity of E6AP [129].
a The viral protein VP40 contains a PY motif that binds to WW3 domain of NEDD4 to facilitate viral budding. b Binding of HPV E6 to a distinct site in E6AP allows it to recruit p53 and target it for ubiquitination, thus promoting HPV-induced carcinogenesis. c NUMB binds to ITCH and this allows Gli ubiquitination and degradation. d The adaptor SMAD7 has a dual role in activating its E3 SMURF2. Through its PY motif it binds to WW domains of SMURF2. It also binds to the E2, UbcH7, enhancing E2–E3 reaction and activation of the HECT ligase. e Adaptors can bind to sites other than WW domain such as in case of SMURF1 where CDH1 (Cadherin1) binds to the C2 domain and CKIP (casein kinase-interacting protein) binds to the WW domain linker region. Allosteric interactions of these adaptor proteins prevent homodimerisation of SMURF1 (C2-HECT) and increased ubiquitination of SMURF1 substrates. f 14-3-3 proteins bind NEDD4–2 phosphorylated by either PKA or SGK. This induces conformational change preventing the E3 from effectively assessing its substrates and results in reduced E3 activity. Regulatory linker region (L) between WW domains is shown.
NUMB
As mentioned above, NUMB, a proline-rich protein, is another binding partner of NEDD4 that regulates PTEN stability [130]. NUMB also regulates Gli, which is the major effector of Hedgehog signalling and its deregulation is often associated with medulloblastoma. NUMB binds to WW2 domain of ITCH and activates its catalytic activity by releasing it from an inhibitory intramolecular interaction (Fig. 7c). This allows Gli1 to interact with ITCH and stabilises the complex [131]. The WW domains of ITCH bind the PPXYs and pSP docking sites on Gli1 C-terminus resulting in ubiquitination and proteasomal-dependent degradation (Fig. 7c).
SMADs
SMADs regulate TGF-β signal transduction. The inhibitory SMAD7 inhibits TGF-β–induced transcriptional responses by interfering with SMAD2 and SMAD3 activation [132]. Acetylation of SMAD7 prevents its ubiquitination by SMURF1 and promotes its stability [133]. It was shown that SMAD7 is ubiquitinated in fibrotic kidneys with a concomitant increase in SMURF1 and SMURF2 suggesting a critical physiological role [134]. As mentioned above and shown in Fig. 7d, binding of SMAD7 to SMURF2 enhances the E2–E3 interaction (UbcH7 and HECT domain) and E3 activity [59].
CKIP/CDH1
Cadherin1 (CDH1), a well-known adaptor for the RING E3 anaphase promoting complex [135], also interacts with C2 domain of SMURF1 to relieve autoinhibited homodimer of this E3 [135]. Other adaptors, such as cerebral cavernous malformations 2 that interact with HECT domain [136] and casein kinase-2–interacting protein-1 (CKIP-1) that targets the linker between the WW domains [137], also prevent SMURF1 homodimerisation. It has been suggested that this occurs sequentially where CDH1 binding destabilises the homodimer, allowing CKIP1 to interact with the linker region (Fig. 7e) thereby enhancing substrate affinity and promoting ubiquitination [137].
14-3-3
Multiple members of the 14-3-3 family of proteins act as regulators of HECT E3s. 14-3-3 proteins are highly conserved and abundantly expressed adaptors that interact and regulate the cellular localisation and function of several proteins. A role for 14-3-3 proteins as adaptors for NEDD4-2 is well established [138]. In response to aldosterone or insulin signalling, SGK- or PKA-mediated phosphorylation of NEDD4-2 allows its binding to 14-3-3 that serves as an inhibitory interaction, preventing NEDD4-2 from ubiquitinating its substrates, such as the epithelial sodium channel (ENaC) (Fig. 7f) [138].
DUBs
As mentioned above, DUBs also constitute a group of modulators of HECT E3s [10] affecting their stability or by deubiquitination of adaptors that regulates E3 activity. As shown in Fig. 8a SMURF1 is autoubiquitinated through its intrinsic ligase activity. FAM/USP9X interacts with SMURF1 at WW2, resulting in deubiquitination and stabilisation of SMURF1 [139]. USP15 also deubiquitinates autoubiquitinated MDM2 (a RING E3 ligase) to stabilise it. This prevents T-cell activation by targeting the transcription factor NFATc2 [140]. Thus, downregulating USP15 is beneficial in promoting cancer cell apoptosis and T-cell response. Another regulatory role of DUBs is seen during NEDD4-2–mediated regulation of ENaC (Fig. 8b) where USP2-45 interacts with N-terminus of ENaC and deubiquitinates it. USP2-45 also interacts with NEDD4-2 both with its catalytic domain and cytoplasmic N-terminus and prevents NEDD4-2 from ubiquitinating ENaC [141].
a USP9x binds to WW2 of SMURF1 and prevents its autoubiquitination. b The epithelial sodium channel (ENaC) is normally targeted for degradation following NEDD4-2–mediated ubiquitination at the PM. However, USP2-45 can bind to ENaC, facilitating its deubiquitination and/or interact with NEDD4-2 and prevent ENaC channel degradation.
Other adaptors
HECT E3s can also modulate the enzyme activity of other E3s by acting as adaptors. This is observed in case of HERC2 that stimulates the E3 activity of E6AP [142]. Covalent modification of Parkin (RBR E3 ligase) by ISG15 is catalysed by HERC5 that involves two Lys residues of Parkin and relieves its autoinhibition [143]. Furthermore, in response to caspase-8–dependent inflammasome activation in intestinal epithelial cells, gasdermin D recruits NEDD4 and catalyses polyubiquitination of IL1β [144]. This serves as a signal for non-lytic release of IL1β in exosomes [144].
Perspectives
The precise regulatory dynamics of HECT E3 ligases are relatively poorly understood compared to the RING E3 ligase family. A major lacuna in this area has been lack of understanding of mechanisms that regulate their activities. In the last three decades the investigations in S. cerevisiae have provided crucial insights about major molecular pathways, modulators and their mode of action in this field. Most of these mechanisms are highly conserved across eukaryotes suggesting their critical importance. Recent findings uncover new insights on the role of deubiquitination of Rsp5p and its adaptors in controlling catalytic activity of this enzyme. By interacting with Rsp5p, and utilising its intrinsic Ub-binding site, adaptors are ubiquitinated that results in enhanced enzyme–substrate interaction. However, recruitment of DUBs with the E3 causes adaptor deubiquitination, releasing them from its E3 and allowing other adaptors to bind the E3. This highlights the critical role of ubiquitination in regulating adaptor function and thus overall E3 activity. Nutrient stress in yeast also upregulates other adaptors that in turn can increase E3 abundance and activity. Utilising a process of dynamic interchange between adaptors and E3, yeast cells are capable of turning on different pathways to adapt to periods of nutrient stress. These studies have far-reaching outcomes in mammals, since fine control of catalytic activity of E3s is vital in improved drug design and developing better therapeutics. Given the high conservation of HECT domain, it has been difficult to design specific drugs to target individual HECT E3s. However, the complexity of regulation of HECT activity by specific adaptors and PTM, including deubiquitination, allows rethinking specificity of targeting signalling pathways regulated by individual HECT ligases.
Change history
22 January 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41418-021-00737-8
References
Foot N, Henshall T, Kumar S. Ubiquitination and the regulation of membrane proteins. Physiol Rev. 2017;97:253–81.
Rotin D, Kumar S. Physiological functions of the HECT family of ubiquitin ligases. Nat Rev Mol Cell Biol. 2009;10:398–409.
Scheffner M, Kumar S. Mammalian HECT ubiquitin-protein ligases: biological and pathophysiological aspects. Biochim Biophys Acta. 2014;1843:61–74.
Haas AL, Warms JV, Rose IA. Ubiquitin adenylate: structure and role in ubiquitin activation. Biochemistry 1983;22:4388–94.
Jentsch S. The ubiquitin-conjugation system. Annu Rev Genet. 1992;26:179–207.
Hershko A, Ciechanover A. The ubiquitin system. Annu Rev Biochem. 1998;67:425–79.
Swatek KN, Komander D. Ubiquitin modifications. Cell Res. 2016;26:399–422.
Balaji V, Pokrzywa W, Hoppe T. Ubiquitylation pathways in insulin signaling and organismal homeostasis. Bioessays. 2018;40:e1700223.
Xu P, Duong DM, Seyfried NT, Cheng D, Xie Y, Robert J, et al. Quantitative proteomics reveals the function of unconventional ubiquitin chains in proteasomal degradation. Cell. 2009;137:133–45.
Komander D, Clague MJ, Urbe S. Breaking the chains: structure and function of the deubiquitinases. Nat Rev Mol Cell Biol. 2009;10:550–63.
Wenzel DM, Klevit RE. Following Ariadne’s thread: a new perspective on RBR ubiquitin ligases. BMC Biol. 2012;10:24.
Yang B, Kumar S. Nedd4 and Nedd4-2: closely related ubiquitin-protein ligases with distinct physiological functions. Cell Death Differ. 2010;17:68–77.
Huibregtse JM, Scheffner M, Beaudenon S, Howley PM. A family of proteins structurally and functionally related to the E6-AP ubiquitin-protein ligase. Proc Natl Acad Sci USA. 1995;92:5249.
Scheffner M, Nuber U, Huibregtse JM. Protein ubiquitination involving an E1-E2-E3 enzyme ubiquitin thioester cascade. Nature. 1995;373:81–83.
Huang L, Kinnucan E, Wang G, Beaudenon S, Howley PM, Huibregtse JM, et al. Structure of an E6AP-UbcH7 complex: insights into ubiquitination by the E2-E3 enzyme cascade. Science. 1999;286:1321–6.
Maspero E, Mari S, Valentini E, Musacchio A, Fish A, Pasqualato S, et al. Structure of the HECT:ubiquitin complex and its role in ubiquitin chain elongation. EMBO Rep. 2011;12:342–9.
Shearwin-Whyatt L, Dalton HE, Foot N, Kumar S. Regulation of functional diversity within the Nedd4 family by accessory and adaptor proteins. Bioessays. 2006;28:617–28.
Yasuda J, Nakao M, Kawaoka Y, Shida H. Nedd4 regulates egress of Ebola virus-like particles from host cells. J Virol. 2003;77:9987–92.
Ziegler CM, Dang L, Eisenhauer P, Kelly JA, King BR, Klaus JP, et al. NEDD4 family ubiquitin ligases associate with LCMV Z’s PPXY domain and are required for virus budding, but not via direct ubiquitination of Z. PLoS Pathog. 2019;15:e1008100.
Bernassola F, Karin M, Ciechanover A, Melino G. The HECT family of E3 ubiquitin ligases: multiple players in cancer development. Cancer Cell. 2008;14:10–21.
Harvey KF, Kumar S. Nedd4-like proteins: an emerging family of ubiquitin-protein ligases implicated in diverse cellular functions. Trends Cell Biol. 1999;9:166–9.
Sluimer J, Distel B. Regulating the human HECT E3 ligases. Cell Mol Life Sci. 2018;75:3121–41.
Kim HC, Huibregtse JM. Polyubiquitination by HECT E3s and the determinants of chain type specificity. Mol Cell Biol. 2009;29:3307–18.
Scialpi F, Malatesta M, Peschiaroli A, Rossi M, Melino G, Bernassola F. Itch self-polyubiquitylation occurs through lysine-63 linkages. Biochem Pharm. 2008;76:1515–21.
Michel MA, Swatek KN, Hospenthal MK, Komander D. Ubiquitin linkage-specific affimers reveal insights into K6-linked ubiquitin signaling. Mol Cell. 2017;68:233–46.e235.
Akutsu M, Dikic I, Bremm A. Ubiquitin chain diversity at a glance. J Cell Sci. 2016;129:875–80.
Sun H, Mali SM, Singh SK, Meledin R, Brik A, Kwon YT, et al. Diverse fate of ubiquitin chain moieties: the proximal is degraded with the target, and the distal protects the proximal from removal and recycles. Proc Natl Acad Sci USA. 2019;116:7805–12.
Ohtake F, Tsuchiya H, Saeki Y, Tanaka K. K63 ubiquitylation triggers proteasomal degradation by seeding branched ubiquitin chains. Proc Natl Acad Sci USA. 2018;115:E1401–8.
Rojas-Fernandez A, Plechanovova A, Hattersley N, Jaffray E, Tatham MH, Hay RT. SUMO chain-induced dimerization activates RNF4. Mol Cell. 2014;53:880–92.
Song R, Peng W, Zhang Y, Lv F, Wu HK, Guo J, et al. Central role of E3 ubiquitin ligase MG53 in insulin resistance and metabolic disorders. Nature. 2013;494:375–9.
Nagarajan A, Petersen MC, Nasiri AR, Butrico G, Fung A, Ruan HB, et al. MARCH1 regulates insulin sensitivity by controlling cell surface insulin receptor levels. Nat Commun. 2016;7:12639.
Mabb AM, Judson MC, Zylka MJ, Philpot BD. Angelman syndrome: insights into genomic imprinting and neurodevelopmental phenotypes. Trends Neurosci. 2011;34:293–303.
Weber J, Polo S, Maspero E. HECT E3 ligases: a tale with multiple facets. Front Physiol. 2019;10:370.
Persaud A, Alberts P, Mari S, Tong J, Murchie R, Maspero E, et al. Tyrosine phosphorylation of NEDD4 activates its ubiquitin ligase activity. Sci Signal. 2014;7:ra95.
Xie P, Zhang M, He S, Lu K, Chen Y, Xing G, et al. The covalent modifier Nedd8 is critical for the activation of Smurf1 ubiquitin ligase in tumorigenesis. Nat Commun. 2014;5:3733.
Chandhoke AS, Karve K, Dadakhujaev S, Netherton S, Deng L, Bonni S. The ubiquitin ligase Smurf2 suppresses TGFbeta-induced epithelial-mesenchymal transition in a sumoylation-regulated manner. Cell Death Differ. 2016;23:876–88.
Gallagher E, Gao M, Liu YC, Karin M. Activation of the E3 ubiquitin ligase Itch through a phosphorylation-induced conformational change. Proc Natl Acad Sci USA. 2006;103:1717–22.
Zhu K, Shan Z, Chen X, Cai Y, Cui L, Yao W, et al. Allosteric auto-inhibition and activation of the Nedd4 family E3 ligase Itch. EMBO Rep. 2017;18:1618–30.
Chan AL, Grossman T, Zuckerman V, Campigli Di Giammartino D, Moshel O, Scheffner M, et al. c-Abl phosphorylates E6AP and regulates its E3 ubiquitin ligase activity. Biochemistry. 2013;52:3119–29.
Kee Y, Lyon N, Huibregtse JM. The Rsp5 ubiquitin ligase is coupled to and antagonized by the Ubp2 deubiquitinating enzyme. EMBO J. 2005;24:2414–24.
Tomaic V, Pim D, Thomas M, Massimi P, Myers MP, Banks L. Regulation of the human papillomavirus type 18 E6/E6AP ubiquitin ligase complex by the HECT domain-containing protein EDD. J Virol. 2011;85:3120–7.
Cai JB, Shi GM, Dong ZR, Ke AW, Ma HH, Gao Q, et al. Ubiquitin-specific protease 7 accelerates p14(ARF) degradation by deubiquitinating thyroid hormone receptor-interacting protein 12 and promotes hepatocellular carcinoma progression. Hepatology. 2015;61:1603–14.
He S, Cao Y, Xie P, Dong G, Zhang L. The Nedd8 Non-covalent binding region in the smurf HECT domain is critical to its ubiquitn ligase function. Sci Rep. 2017;7:41364.
Li H, Zhu H, Liu Y, He F, Xie P, Zhang L. Itch promotes the neddylation of JunB and regulates JunB-dependent transcription. Cell Signal. 2016;28:1186–95.
Attali I, Tobelaim WS, Persaud A, Motamedchaboki K, Simpson-Lavy KJ, Mashahreh B, et al. Ubiquitylation-dependent oligomerization regulates activity of Nedd4 ligases. EMBO J. 2017;36:425–40.
Sander B, Xu W, Eilers M, Popov N, Lorenz S. A conformational switch regulates the ubiquitin ligase HUWE1. Elife. 2017;6:e21036. https://doi.org/10.7554/eLife.21036.
Ronchi VP, Klein JM, Edwards DJ, Haas AL. The active form of E6-associated protein (E6AP)/UBE3A ubiquitin ligase is an oligomer. J Biol Chem. 2014;289:1033–48.
Ruetalo N, Anders S, Stollmaier C, Jackl M, Schutz-Stoffregen MC, Stefan N, et al. The WW1 domain enhances autoinhibition in Smurf ubiquitin ligases. J Mol Biol. 2019;431:4834–47.
Wan L, Zou W, Gao D, Inuzuka H, Fukushima H, Berg AH, et al. Cdh1 regulates osteoblast function through an APC/C-independent modulation of Smurf1. Mol Cell. 2011;44:721–33.
Chen Z, Jiang H, Xu W, Li X, Dempsey DR, Zhang X, et al. A tunable brake for HECT ubiquitin ligases. Mol Cell. 2017;66:345–57.e346.
Wiesner S, Ogunjimi AA, Wang HR, Rotin D, Sicheri F, Wrana JL, et al. Autoinhibition of the HECT-type ubiquitin ligase Smurf2 through its C2 domain. Cell. 2007;130:651–62.
Wang Z, Liu Z, Chen X, Li J, Yao W, Huang S, et al. A multi-lock inhibitory mechanism for fine-tuning enzyme activities of the HECT family E3 ligases. Nat Commun. 2019;10:3162.
Escobedo A, Gomes T, Aragon E, Martin-Malpartida P, Ruiz L, Macias MJ. Structural basis of the activation and degradation mechanisms of the E3 ubiquitin ligase Nedd4L. Structure. 2014;22:1446–57.
French ME, Kretzmann BR, Hicke L. Regulation of the RSP5 ubiquitin ligase by an intrinsic ubiquitin-binding site. J Biol Chem. 2009;284:12071–9.
MacDonald C, Shields SB, Williams CA, Winistorfer S, Piper RC. A cycle of ubiquitination regulates adaptor function of the Nedd4-family ubiquitin ligase Rsp5. Curr Biol. 2020;30:465–79.e465.
Zhang W, Wu KP, Sartori MA, Kamadurai HB, Ordureau A, Jiang C, et al. System-wide modulation of HECT E3 ligases with selective ubiquitin variant probes. Mol Cell. 2016;62:121–36.
Okumura A, Pitha PM, Harty RN. ISG15 inhibits Ebola VP40 VLP budding in an L-domain-dependent manner by blocking Nedd4 ligase activity. Proc Natl Acad Sci USA. 2008;105:3974–9.
Malakhova OA, Zhang DE. ISG15 inhibits Nedd4 ubiquitin E3 activity and enhances the innate antiviral response. J Biol Chem. 2008;283:8783–7.
Ogunjimi AA, Briant DJ, Pece-Barbara N, Le Roy C, Di Guglielmo GM, Kavsak P, et al. Regulation of Smurf2 ubiquitin ligase activity by anchoring the E2 to the HECT domain. Mol Cell. 2005;19:297–308.
Leon S, Haguenauer-Tsapis R. Ubiquitin ligase adaptors: regulators of ubiquitylation and endocytosis of plasma membrane proteins. Exp Cell Res. 2009;315:1574–83.
Hettema EH, Valdez-Taubas J, Pelham HR. Bsd2 binds the ubiquitin ligase Rsp5 and mediates the ubiquitination of transmembrane proteins. EMBO J. 2004;23:1279–88.
Lin CH, MacGurn JA, Chu T, Stefan CJ, Emr SD. Arrestin-related ubiquitin-ligase adaptors regulate endocytosis and protein turnover at the cell surface. Cell. 2008;135:714–25.
Shearwin-Whyatt LM, Brown DL, Wylie FG, Stow JL, Kumar S. N4WBP5A (Ndfip2), a Nedd4-interacting protein, localizes to multivesicular bodies and the Golgi, and has a potential role in protein trafficking. J Cell Sci. 2004;117:3679–89.
Jolliffe CN, Harvey KF, Haines BP, Parasivam G, Kumar S. Identification of multiple proteins expressed in murine embryos as binding partners for the WW domains of the ubiquitin-protein ligase Nedd4. Biochem J. 2000;351:557–65.
Sullivan JA, Lewis MJ, Nikko E, Pelham HR. Multiple interactions drive adaptor-mediated recruitment of the ubiquitin ligase rsp5 to membrane proteins in vivo and in vitro. Mol Biol Cell. 2007;18:2429–40.
Stimpson HE, Lewis MJ, Pelham HR. Transferrin receptor-like proteins control the degradation of a yeast metal transporter. EMBO J. 2006;25:662–72.
Nikko E, Sullivan JA, Pelham HR. Arrestin-like proteins mediate ubiquitination and endocytosis of the yeast metal transporter Smf1. EMBO Rep. 2008;9:1216–21.
MacGurn JA, Hsu PC, Emr SD. Ubiquitin and membrane protein turnover: from cradle to grave. Annu Rev Biochem. 2012;81:231–59.
Becuwe M, Herrador A, Haguenauer-Tsapis R, Vincent O, Leon S. Ubiquitin-mediated regulation of endocytosis by proteins of the arrestin family. Biochem Res Int. 2012;2012:242764.
Kee Y, Munoz W, Lyon N, Huibregtse JM. The deubiquitinating enzyme Ubp2 modulates Rsp5-dependent Lys63-linked polyubiquitin conjugates in Saccharomyces cerevisiae. J Biol Chem. 2006;281:36724–31.
Gupta R, Kus B, Fladd C, Wasmuth J, Tonikian R, Sidhu S, et al. Ubiquitination screen using protein microarrays for comprehensive identification of Rsp5 substrates in yeast. Mol Syst Biol. 2007;3:116.
Dunn R, Hicke L. Domains of the Rsp5 ubiquitin-protein ligase required for receptor-mediated and fluid-phase endocytosis. Mol Biol Cell. 2001;12:421–35.
Choudhary P, Loewen MC. Evidence of a role for S. cerevisiae alpha-arrestin Art1 (Ldb19) in mating projection and zygote formations. Cell Biol Int. 2016;40:83–90.
Nikko E, Pelham HR. Arrestin-mediated endocytosis of yeast plasma membrane transporters. Traffic. 2009;10:1856–67.
Hovsepian J, Albanese V, Becuwe M, Ivashov V, Teis D, Leon S. The yeast arrestin-related protein Bul1 is a novel actor of glucose-induced endocytosis. Mol Biol Cell. 2018;29:1012–20.
Kota J, Melin-Larsson M, Ljungdahl PO, Forsberg H. Ssh4, Rcr2 and Rcr1 affect plasma membrane transporter activity in Saccharomyces cerevisiae. Genetics. 2007;175:1681–94.
Imai K, Noda Y, Adachi H, Yoda K. A novel endoplasmic reticulum membrane protein Rcr1 regulates chitin deposition in the cell wall of Saccharomyces cerevisiae. J Biol Chem. 2005;280:8275–84.
Zhu L, Sardana R, Jin DK, Emr SD. Calcineurin-dependent regulation of endocytosis by a plasma membrane ubiquitin ligase adaptor, Rcr1. J Cell Biol. 2020;219:e201909158. https://doi.org/10.1083/jcb.201909158.
Parua PK, Ratnakumar S, Braun KA, Dombek KM, Arms E, Ryan PM, et al. 14-3-3 (Bmh) proteins inhibit transcription activation by Adr1 through direct binding to its regulatory domain. Mol Cell Biol. 2010;30:5273–83.
Hovsepian J, Defenouillere Q, Albanese V, Vachova L, Garcia C, Palkova Z, et al. Multilevel regulation of an alpha-arrestin by glucose depletion controls hexose transporter endocytosis. J Cell Biol. 2017;216:1811–31.
Merhi A, Andre B. Internal amino acids promote Gap1 permease ubiquitylation via TORC1/Npr1/14-3-3-dependent control of the Bul arrestin-like adaptors. Mol Cell Biol. 2012;32:4510–22.
Lam MH, Emili A. Ubp2 regulates Rsp5 ubiquitination activity in vivo and in vitro. PLoS One. 2013;8:e75372.
Ho HC, MacGurn JA, Emr SD. Deubiquitinating enzymes Ubp2 and Ubp15 regulate endocytosis by limiting ubiquitination and degradation of ARTs. Mol Biol Cell. 2017;28:1271–83.
MacDonald C, Stringer DK, Piper RC. Sna3 is an Rsp5 adaptor protein that relies on ubiquitination for its MVB sorting. Traffic. 2012;13:586–98.
Leon S, Erpapazoglou Z, Haguenauer-Tsapis R. Ear1p and Ssh4p are new adaptors of the ubiquitin ligase Rsp5p for cargo ubiquitylation and sorting at multivesicular bodies. Mol Biol Cell. 2008;19:2379–88.
Sardana R, Zhu L, Emr SD. Rsp5 Ubiquitin ligase-mediated quality control system clears membrane proteins mistargeted to the vacuole membrane. J Cell Biol. 2019;218:234–50.
Ivashov V, Zimmer J, Schwabl S, Kahlhofer J, Weys S, Gstir R, et al. Complementary alpha-arrestin-ubiquitin ligase complexes control nutrient transporter endocytosis in response to amino acids. Elife. 2020;9:e58246. https://doi.org/10.7554/eLife.58246.
Harvey KF, Shearwin-Whyatt LM, Fotia A, Parton RG, Kumar S. N4WBP5, a potential target for ubiquitination by the Nedd4 family of proteins, is a novel Golgi-associated protein. J Biol Chem. 2002;277:9307–17.
Dalton HE, Denton D, Foot NJ, Ho K, Mills K, Brou C, et al. Drosophila Ndfip is a novel regulator of Notch signaling. Cell Death Differ. 2011;18:1150–60.
Konstas AA, Shearwin-Whyatt LM, Fotia AB, Degger B, Riccardi D, Cook DI, et al. Regulation of the epithelial sodium channel by N4WBP5A, a novel Nedd4/Nedd4-2-interacting protein. J Biol Chem. 2002;277:29406–16.
Mund T, Pelham HR. Control of the activity of WW-HECT domain E3 ubiquitin ligases by NDFIP proteins. EMBO Rep. 2009;10:501–7.
Gorla M, Santiago C, Chaudhari K, Layman AAK, Oliver PM, Bashaw GJ. Ndfip proteins target robo receptors for degradation and allow commissural axons to cross the midline in the developing spinal cord. Cell Rep. 2019;26:3298–312.e3294.
Foot NJ, Dalton HE, Shearwin-Whyatt LM, Dorstyn L, Tan SS, Yang B, et al. Regulation of the divalent metal ion transporter DMT1 and iron homeostasis by a ubiquitin-dependent mechanism involving Ndfips and WWP2. Blood. 2008;112:4268–75.
Foot NJ, Gembus KM, Mackenzie K, Kumar S. Ndfip2 is a potential regulator of the iron transporter DMT1 in the liver. Sci Rep. 2016;6:24045.
Duval M, Bedard-Goulet S, Delisle C, Gratton JP. Vascular endothelial growth factor-dependent down-regulation of Flk-1/KDR involves Cbl-mediated ubiquitination. Consequences on nitric oxide production from endothelial cells. J Biol Chem. 2003;278:20091–7.
Howitt J, Putz U, Lackovic J, Doan A, Dorstyn L, Cheng H, et al. Divalent metal transporter 1 (DMT1) regulation by Ndfip1 prevents metal toxicity in human neurons. Proc Natl Acad Sci USA. 2009;106:15489–94.
Lackovic J, Howitt J, Callaway JK, Silke J, Bartlett P, Tan SS. Differential regulation of Nedd4 ubiquitin ligases and their adaptor protein Ndfip1 in a rat model of ischemic stroke. Exp Neurol. 2012;235:326–35.
Oliver PM, Cao X, Worthen GS, Shi P, Briones N, MacLeod M, et al. Ndfip1 protein promotes the function of itch ubiquitin ligase to prevent T cell activation and T helper 2 cell-mediated inflammation. Immunity. 2006;25:929–40.
Yip KH, Kolesnikoff N, Hauschild N, Biggs L, Lopez AF, Galli SJ, et al. The Nedd4-2/Ndfip1 axis is a negative regulator of IgE-mediated mast cell activation. Nat Commun. 2016;7:13198.
Trimpert C, Wesche D, de Groot T, Pimentel Rodriguez MM, Wong V, van den Berg DTM, et al. NDFIP allows NEDD4/NEDD4L-induced AQP2 ubiquitination and degradation. PLoS One. 2017;12:e0183774.
Mund T, Pelham HR. Regulation of PTEN/Akt and MAP kinase signaling pathways by the ubiquitin ligase activators Ndfip1 and Ndfip2. Proc Natl Acad Sci USA. 2010;107:11429–34.
Wang Y, Tong X, Ye X. Ndfip1 negatively regulates RIG-I-dependent immune signaling by enhancing E3 ligase Smurf1-mediated MAVS degradation. J Immunol. 2012;189:5304–13.
Howitt J, Lackovic J, Low LH, Naguib A, Macintyre A, Goh CP, et al. Ndfip1 regulates nuclear Pten import in vivo to promote neuronal survival following cerebral ischemia. J Cell Biol. 2012;196:29–36.
Maddika S, Kavela S, Rani N, Palicharla VR, Pokorny JL, Sarkaria JN, et al. WWP2 is an E3 ubiquitin ligase for PTEN. Nat Cell Biol. 2011;13:728–33.
Puca L, Brou C. Alpha-arrestins—-new players in Notch and GPCR signaling pathways in mammals. J Cell Sci. 2014;127:1359–67.
Nabhan JF, Pan H, Lu Q. Arrestin domain-containing protein 3 recruits the NEDD4 E3 ligase to mediate ubiquitination of the beta2-adrenergic receptor. EMBO Rep. 2010;11:605–11.
Puca L, Chastagner P, Meas-Yedid V, Israel A, Brou C. Alpha-arrestin 1 (ARRDC1) and beta-arrestins cooperate to mediate Notch degradation in mammals. J Cell Sci. 2013;126:4457–68.
Nabhan JF, Hu R, Oh RS, Cohen SN, Lu Q. Formation and release of arrestin domain-containing protein 1-mediated microvesicles (ARMMs) at plasma membrane by recruitment of TSG101 protein. Proc Natl Acad Sci USA. 2012;109:4146–51.
Dores MR, Lin H, Grimsey NJ, Mendez F, Trejo J. The alpha-arrestin ARRDC3 mediates ALIX ubiquitination and G protein-coupled receptor lysosomal sorting. Mol Biol Cell. 2015;26:4660–73.
Mackenzie K, Foot NJ, Anand S, Dalton HE, Chaudhary N, Collins BM, et al. Regulation of the divalent metal ion transporter via membrane budding. Cell Disco. 2016;2:16011.
Mund T, Pelham HR. Substrate clustering potently regulates the activity of WW-HECT domain-containing ubiquitin ligases. J Biol Chem. 2018;293:5200–9.
Shenoy SK, Modi AS, Shukla AK, Xiao K, Berthouze M, Ahn S, et al. Beta-arrestin-dependent signaling and trafficking of 7-transmembrane receptors is reciprocally regulated by the deubiquitinase USP33 and the E3 ligase Mdm2. Proc Natl Acad Sci USA. 2009;106:6650–5.
Shea FF, Rowell JL, Li Y, Chang TH, Alvarez CE. Mammalian alpha arrestins link activated seven transmembrane receptors to Nedd4 family e3 ubiquitin ligases and interact with beta arrestins. PLoS One. 2012;7:e50557.
Han SO, Kommaddi RP, Shenoy SK. Distinct roles for beta-arrestin2 and arrestin-domain-containing proteins in beta2 adrenergic receptor trafficking. EMBO Rep. 2013;14:164–71.
MacDonald C, Buchkovich NJ, Stringer DK, Emr SD, Piper RC. Cargo ubiquitination is essential for multivesicular body intralumenal vesicle formation. EMBO Rep. 2012;13:331–8.
Putz U, Howitt J, Lackovic J, Foot N, Kumar S, Silke J, et al. Nedd4 family-interacting protein 1 (Ndfip1) is required for the exosomal secretion of Nedd4 family proteins. J Biol Chem. 2008;283:32621–7.
Putz U, Howitt J, Doan A, Goh CP, Low LH, Silke J, et al. The tumor suppressor PTEN is exported in exosomes and has phosphatase activity in recipient cells. Sci Signal. 2012;5:ra70.
Kulman JD, Harris JE, Xie L, Davie EW. Identification of two novel transmembrane gamma-carboxyglutamic acid proteins expressed broadly in fetal and adult tissues. Proc Natl Acad Sci USA. 2001;98:1370–5.
Kulman JD, Harris JE, Xie L, Davie EW. Proline-rich Gla protein 2 is a cell-surface vitamin K-dependent protein that binds to the transcriptional coactivator Yes-associated protein. Proc Natl Acad Sci USA. 2007;104:8767–72.
Xu S, Han JC, Morales A, Menzie CM, Williams K, Fan YS. Characterization of 11p14-p12 deletion in WAGR syndrome by array CGH for identifying genes contributing to mental retardation and autism. Cytogenet Genome Res. 2008;122:181–7.
Justice ED, Barnum SJ, Kidd T. The WAGR syndrome gene PRRG4 is a functional homologue of the commissureless axon guidance gene. PLoS Genet. 2017;13:e1006865.
Husain M, Boermans HJ, Karrow NA. Mesenteric lymph node transcriptome profiles in BALB/c mice sensitized to three common food allergens. BMC Genomics. 2011;12:12.
Latourelle JC, Pankratz N, Dumitriu A, Wilk JB, Goldwurm S, Pezzoli G, et al. Genomewide association study for onset age in Parkinson disease. BMC Med Genet. 2009;10:98.
Yazicioglu MN, Monaldini L, Chu K, Khazi FR, Murphy SL, Huang H, et al. Cellular localization and characterization of cytosolic binding partners for Gla domain-containing proteins PRRG4 and PRRG2. J Biol Chem. 2013;288:25908–14.
Zhang L, Qin Y, Wu G, Wang J, Cao J, Wang Y, et al. PRRG4 promotes breast cancer metastasis through the recruitment of NEDD4 and downregulation of Robo1. Oncogene. 2020. https://doi.org/10.1038/s41388-020-01494-7.
Harty RN, Brown ME, Wang G, Huibregtse J, Hayes FP. A PPxY motif within the VP40 protein of Ebola virus interacts physically and functionally with a ubiquitin ligase: implications for filovirus budding. Proc Natl Acad Sci USA. 2000;97:13871–6.
Huibregtse JM, Scheffner M, Howley PM. Localization of the E6-AP regions that direct human papillomavirus E6 binding, association with p53, and ubiquitination of associated proteins. Mol Cell Biol. 1993;13:4918–27.
Beaudenon S, Huibregtse JM. HPV E6, E6AP and cervical cancer. BMC Biochem. 2008;9:S4.
Mortensen F, Schneider D, Barbic T, Sladewska-Marquardt A, Kuhnle S, Marx A, et al. Role of ubiquitin and the HPV E6 oncoprotein in E6AP-mediated ubiquitination. Proc Natl Acad Sci USA. 2015;112:9872–7.
Shao C, Li Z, Ahmad N, Liu X. Regulation of PTEN degradation and NEDD4-1 E3 ligase activity by Numb. Cell Cycle. 2017;16:957–67.
Di Marcotullio L, Greco A, Mazza D, Canettieri G, Pietrosanti L, Infante P, et al. Numb activates the E3 ligase Itch to control Gli1 function through a novel degradation signal. Oncogene. 2011;30:65–76.
Nakao A, Afrakhte M, Moren A, Nakayama T, Christian JL, Heuchel R, et al. Identification of Smad7, a TGFbeta-inducible antagonist of TGF-beta signalling. Nature. 1997;389:631–5.
Gronroos E, Hellman U, Heldin CH, Ericsson J. Control of Smad7 stability by competition between acetylation and ubiquitination. Mol Cell. 2002;10:483–93.
Fukasawa H, Yamamoto T, Togawa A, Ohashi N, Fujigaki Y, Oda T, et al. Down-regulation of Smad7 expression by ubiquitin-dependent degradation contributes to renal fibrosis in obstructive nephropathy in mice. Proc Natl Acad Sci USA. 2004;101:8687–92.
Jia L, Yu H. Cdh1 is a HECT of an activator. Mol Cell. 2011;44:681–3.
Crose LE, Hilder TL, Sciaky N, Johnson GL. Cerebral cavernous malformation 2 protein promotes smad ubiquitin regulatory factor 1-mediated RhoA degradation in endothelial cells. J Biol Chem. 2009;284:13301–5.
Lu K, Yin X, Weng T, Xi S, Li L, Xing G, et al. Targeting WW domains linker of HECT-type ubiquitin ligase Smurf1 for activation by CKIP-1. Nat Cell Biol. 2008;10:994–1002.
Ichimura T, Yamamura H, Sasamoto K, Tominaga Y, Taoka M, Kakiuchi K, et al. 14-3-3 proteins modulate the expression of epithelial Na+ channels by phosphorylation-dependent interaction with Nedd4-2 ubiquitin ligase. J Biol Chem. 2005;280:13187–94.
Xie Y, Avello M, Schirle M, McWhinnie E, Feng Y, Bric-Furlong E, et al. Deubiquitinase FAM/USP9X interacts with the E3 ubiquitin ligase SMURF1 protein and protects it from ligase activity-dependent self-degradation. J Biol Chem. 2013;288:2976–85.
Zou Q, Jin J, Hu H, Li HS, Romano S, Xiao Y, et al. USP15 stabilizes MDM2 to mediate cancer-cell survival and inhibit antitumor T cell responses. Nat Immunol. 2014;15:562–70.
Oberfeld B, Ruffieux-Daidie D, Vitagliano JJ, Pos KM, Verrey F, Staub O. Ubiquitin-specific protease 2-45 (Usp2-45) binds to epithelial Na+ channel (ENaC)-ubiquitylating enzyme Nedd4-2. Am J Physiol Ren Physiol. 2011;301:F189–196.
Kuhnle S, Kogel U, Glockzin S, Marquardt A, Ciechanover A, Matentzoglu K, et al. Physical and functional interaction of the HECT ubiquitin-protein ligases E6AP and HERC2. J Biol Chem. 2011;286:19410–6.
Im E, Yoo L, Hyun M, Shin WH, Chung KC. Covalent ISG15 conjugation positively regulates the ubiquitin E3 ligase activity of parkin. Open Biol. 2016;6:160193. https://doi.org/10.1098/rsob.160193.
Bulek K, Zhao J, Liao Y, Rana N, Corridoni D, Antanaviciute A, et al. Epithelial-derived gasdermin D mediates nonlytic IL-1beta release during experimental colitis. J Clin Invest. 2020;130:4218–34.
Hatakeyama R, Kamiya M, Takahara T, Maeda T. Endocytosis of the aspartic acid/glutamic acid transporter Dip5 is triggered by substrate-dependent recruitment of the Rsp5 ubiquitin ligase via the arrestin-like protein Aly2. Mol Cell Biol. 2010;30:5598–607.
Wawrzycka D, Sadlak J, Maciaszczyk-Dziubinska E, Wysocki R. Rsp5-dependent endocytosis and degradation of the arsenite transporter Acr3 requires its N-terminal acidic tail as an endocytic sorting signal and arrestin-related ubiquitin-ligase adaptors. Biochim Biophys Acta Biomembr. 2019;1861:916–25.
Sen A, Hsieh WC, Hanna CB, Hsu CC, Pearson M II, Tao WA, et al. The Na(+) pump Ena1 is a yeast epsin-specific cargo requiring its ubiquitylation and phosphorylation sites for internalization. J Cell Sci. 2020;133:jcs245415. https://doi.org/10.1242/jcs.245415.
Becuwe M, Vieira N, Lara D, Gomes-Rezende J, Soares-Cunha C, Casal M, et al. A molecular switch on an arrestin-like protein relays glucose signaling to transporter endocytosis. J Cell Biol. 2012;196:247–59.
O’Donnell AF, Huang L, Thorner J, Cyert MS. A calcineurin-dependent switch controls the trafficking function of alpha-arrestin Aly1/Art6. J Biol Chem. 2013;288:24063–80.
Alvaro CG, O’Donnell AF, Prosser DC, Augustine AA, Goldman A, Brodsky JL, et al. Specific alpha-arrestins negatively regulate Saccharomyces cerevisiae pheromone response by down-modulating the G-protein-coupled receptor Ste2. Mol Cell Biol. 2014;34:2660–81.
Marques MC, Zamarbide-Fores S, Pedelini L, Llopis-Torregrosa V, Yenush L. A functional Rim101 complex is required for proper accumulation of the Ena1 Na+-ATPase protein in response to salt stress in Saccharomyces cerevisiae. FEMS Yeast Res. 2015;15:fov017.
Kaida D, Toh-e A, Kikuchi Y. Rsp5-Bul1/2 complex is necessary for the HSE-mediated gene expression in budding yeast. Biochem Biophys Res Commun. 2003;306:1037–41.
Crespo JL, Helliwell SB, Wiederkehr C, Demougin P, Fowler B, Primig M, et al. NPR1 kinase and RSP5-BUL1/2 ubiquitin ligase control GLN3-dependent transcription in Saccharomyces cerevisiae. J Biol Chem. 2004;279:37512–7.
Ren J, Kee Y, Huibregtse JM, Piper RC. Hse1, a component of the yeast Hrs-STAM ubiquitin-sorting complex, associates with ubiquitin peptidases and a ligase to control sorting efficiency into multivesicular bodies. Mol Biol Cell. 2007;18:324–35.
Myat A, Henry P, McCabe V, Flintoft L, Rotin D, Tear G. Drosophila Nedd4, a ubiquitin ligase, is recruited by Commissureless to control cell surface levels of the roundabout receptor. Neuron. 2002;35:447–59.
Foot NJ, Leong YA, Dorstyn LE, Dalton HE, Ho K, Zhao L, et al. Ndfip1-deficient mice have impaired DMT1 regulation and iron homeostasis. Blood. 2011;117:638–46.
Perry WL, Hustad CM, Swing DA, O’Sullivan TN, Jenkins NA, Copeland NG. The itchy locus encodes a novel ubiquitin protein ligase that is disrupted in a18H mice. Nat Genet. 1998;18:143–6.
Infante P, Faedda R, Bernardi F, Bufalieri F, Lospinoso Severini L, Alfonsi R, et al. Itch/beta-arrestin2-dependent non-proteolytic ubiquitylation of SuFu controls Hedgehog signalling and medulloblastoma tumorigenesis. Nat Commun. 2018;9:976.
Zhou F, Xie F, Jin K, Zhang Z, Clerici M, Gao R, et al. USP4 inhibits SMAD4 monoubiquitination and promotes activin and BMP signaling. EMBO J. 2017;36:1623–39.
Murakami G, Watabe T, Takaoka K, Miyazono K, Imamura T. Cooperative inhibition of bone morphogenetic protein signaling by Smurf1 and inhibitory Smads. Mol Biol Cell. 2003;14:2809–17.
Kavsak P, Rasmussen RK, Causing CG, Bonni S, Zhu H, Thomsen GH, et al. Smad7 binds to Smurf2 to form an E3 ubiquitin ligase that targets the TGF beta receptor for degradation. Mol Cell. 2000;6:1365–75.
Rauch S, Martin-Serrano J. Multiple interactions between the ESCRT machinery and arrestin-related proteins: implications for PPXY-dependent budding. J Virol. 2011;85:3546–56.
Mund T, Graeb M, Mieszczanek J, Gammons M, Pelham HR, Bienz M. Disinhibition of the HECT E3 ubiquitin ligase WWP2 by polymerized Dishevelled. Open Biol. 2015;5:150185.
Acknowledgements
Our work is supported by the National Health & Medical Research Council (NHMRC) project grant (1158366), a NHMRC Senior Principal Research Fellowship (1103006) and the University of South Australia internal research support to SK.
Author information
Authors and Affiliations
Contributions
Both authors contributed to the planning, writing and editing of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: There was an error in figure 2.
Edited by F. Pentimalli
Rights and permissions
About this article
Cite this article
Shah, S.S., Kumar, S. Adaptors as the regulators of HECT ubiquitin ligases. Cell Death Differ 28, 455–472 (2021). https://doi.org/10.1038/s41418-020-00707-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-020-00707-6
- Springer Nature Limited
This article is cited by
-
New insights into QTNs and potential candidate genes governing rice yield via a multi-model genome-wide association study
BMC Plant Biology (2024)
-
The E3 ubiquitin ligase Itch regulates death receptor and cholesterol trafficking to affect TRAIL-mediated apoptosis
Cell Death & Disease (2024)
-
WWP1 E3 ligase at the crossroads of health and disease
Cell Death & Disease (2023)
-
Global ubiquitinome profiling identifies NEDD4 as a regulator of Profilin 1 and actin remodelling in neural crest cells
Nature Communications (2022)
-
The ubiquitin system: from cell signalling to disease biology and new therapeutic opportunities
Cell Death & Differentiation (2021)