Abstract
As two diseases with rapidly increasing incidence, the molecular linkages between obesity and breast cancer (BC) are intriguing. Overall, obesity may be a negative prognostic factor for BC. Single-cell RNA-sequencing (scRNA-seq) was performed on tumor tissues from 6 obese and non-obese BC patients. With 48,033 cells analyzed, we found heterogeneous tumor epithelium and microenvironment in these obese and lean BC patients. Interestingly, the obesity-associated epithelial cells exhibited specific expression signatures which linked tumor growth and hormone metabolism in BC. Notably, one population of obesity-specific macrophage up-regulated the nuclear receptor subfamily 1 group H member 3 (NR1H3), which acted a transcription factor and regulated FABP4 expression through its interaction with the DNA of SREBP1, and further increased the proliferation of tumor cells in BC. Using single-cell signatures, our study illustrate cell diversity and transcriptomic differences in tumors from obese and non-obese BC patients, and sheds light on potential molecular link between lipid metabolism and BC.
Similar content being viewed by others
Introduction
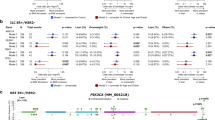
Female breast cancer (BC) has surpassed lung cancer as the most commonly diagnosed cancer with an estimated 2.3 million new cases (11.7% of total cases), and ranks the fifth in the leading causes of cancer death (6.9%) in 2020 [1]. Obesity has become a major public health issue in China, which increased rapidly in the past four decades [2]. The WHO criteria define obesity in adults as a BMI ≥ 30.0 kg/m2 [3]. Besides metabolic, cardiovascular and neurogenic diseases, obesity increases the risk of multiple types of cancer, including liver, colorectal and breast cancer [4,5,6,7]. Studies showed that obesity was associated with increased BC risk in postmenopausal women [7, 8], while an inverse association between obesity and premenopausal BC risk has been also reported [9, 10]. Interestingly, obesity was significantly associated with poorer overall survival whether in premenopausal or postmenopausal BC [11], and increased risk of recurrence and death of approximately 35–40% [12]. Therefore, the molecular links between obesity and BC are poorly understood, and the regulatory mechanisms underlying the obesity-associated breast cancer (OABC) remain to be further investigated.
Some cancer-associated or obesity-associated factors have been reported to regulate metabolic pathways in both cancer cells and cells from the breast microenvironment, which explained the mechanism of obesity-promoted breast tumor growth in some degree. For example, circulating adipose fatty acid binding protein increased in obesity and promoted breast tumor stemness and aggressiveness through activation of the IL-6/STAT3/ALDH1 pathway [13]. Leptin-STAT3 axis increased oxidation of fatty acids within CD8+ effector T cells in BC, leading to inhibition of antitumor immune responses and promotion of obesity-associated breast tumorigenesis [14]. However, BC is a highly heterogeneous disease, which hinders the discovery of other potential mechanisms involved in the OABC. Moreover, few studies have been conducted on the molecular and cellular heterogeneity between obese and non-obese BC.
Advances in single-cell RNA sequencing (scRNA-seq) now offer remarkable new opportunities to explore cellular heterogeneity on a single cell level and reveal new insights into cell biology in multiple carcinomas [15, 16]. Several studies have successfully applied scRNA-seq to understand the complex microenvironment and cell heterogeneity in BC [17,18,19]. However, cellular diversity and transcriptome differences have not been evaluated systematically on the single cell level between obese and non-obese BC.
Here, we sought to further probe cellular and molecular diversity within obese and non-obese BC through single-cell transcriptome analysis. The results revealed tremendous intra- and inter-heterogeneity in tumor samples from three obese and lean BC patients. The obesity-specific epithelial cells showed distinct transcriptional signatures which linked tumor growth and hormone metabolism in BC. The obesity-specific macrophage preferentially expressed transcription factor NR1H3, which regulated the expression level of fatty acid binding protein FABP4, and further increased the proliferation of tumor cells in BC. Our single-cell data provided new insights for uncovering the molecular link between lipid metabolism and BC.
Results
Single-cell gene expression atlas of the tumor microenvironment in obese and non-obese BC
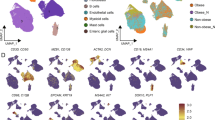
An increase in T stage based on the tumor maximum diameter can significantly increase the hazard of death in BC patients [20]. All patients diagnosed with invasive BC from January 2015 to December 2021 were included in the analysis, the results revealed that the maximum diameter and sectional area of obese BC patients were significantly larger than that of non-obese BC patients (Fig. 1a, b). However, other clinical parameters including number of patients with different positive lymph nodes, the proportions of lympho-vascular invasion, perineural invasion and histologic grade were not significantly different between obese and lean BC patients (Supplementary Fig. S1). To depict the tumor ecosystem in OABC, we collected surgical tumor specimens from 3 obese (BMI > 30) and 3 lean (BMI < 30) BC patients for scRNA-seq using 10X Genomics platform (Fig. 1c, Supplementary Table S1). After quality controls, we obtained 48,033 high-quality cells from these 6 individuals (6382–10,989 cells, average 8005 cells per patient) for further analysis. These cells consisted of 30 clusters (Supplementary Fig. S2a) and 8 major cell types (Fig. 1d) by the T-distributed stochastic neighbor embedding (t-SNE) method, including B cell (n = 190, marked by CD79A, MS4A1), plasma cell (n = 216, JCHAIN), mast cell (n = 206, CPA3, TPSAB1), myeloid cell (n = 4056, LYZ, TYROBP), T/natural killer (NK) cell (n = 2566, CD3D, CD3E), epithelial cell (n = 32,440, KRT19, EPCAM), endothelial cell (n = 1363, PECAM1, CLDN5) and cancer-associated fibroblast (CAF, n = 6996, COL1A1, PDGFRB, Fig. 1e–i, Supplementary Fig. S2b). The obese and lean samples shared most immune and non-immune cell types except some epithelium, which showed tremendous heterogeneity among BC tumors (Fig. 1j, k, Supplementary Fig. S2c). Moreover, most epithelial cells were separated by patients (Supplementary Fig. S2c), demonstrated huge inter-tumor heterogeneity among obese and lean individuals. The epithelial cells were dominated in most samples, indicating their tumor identity of all obtained samples (Supplementary Fig. S2d).
Scatter plot showing the differences of maximum diameter (a) and sectional area (b) between obese and lean BC patients. A total of 396 and 364 BC patients were included for the analysis, respectively. Mann–Whitney U test was used for significance comparison, *p-value < 0.05, **p-value < 0.01. c Schematic showing the study design of scRNA-seq in tumors from 3 obese and 3 lean BC patients. d T-distributed stochastic neighbor embedding (t-SNE) plot of 48,033 cells color-coded for different cell types. NK natural killer, CAF cancer-associated fibroblast. e Dot plot showing the expression of selected marker genes for each major cell type. f–i t-SNE plot showing the expression of different marker genes. t-SNE visualization of all cells, colored by group (j) or cell type (k). See also Supplementary Figs. S1 and S2.
Obesity-specific epithelium showed distinct transcriptomic and regulatory signatures
The re-clustering of all epithelial cells (n = 32,440) could identified 13 sub-populations based on their highly expressed genes (Fig. 2a). These sub-clusters of epithelium showed obvious sample specificity among BC patients, for example, AREG+ Epi and MUCL1+ Epi were enriched in the lean3 patient of lean group, CPB1+ Epi and CGA+ Epi were mainly enriched in obese3 patient of obese group (Fig. 2b, c, Supplementary Fig. S2e). Different sub-clusters showed significantly differential expression signatures, in which multiple epithelial-related genes or markers had specific expression in some sub-clusters, including KRT15, MUCL1, AREG, CXCL14 and CGA (Fig. 2d).
The t-SNE plot showing 32,440 epithelial cells colored by sub-population (a), sample (b) or group (c) based on their differential transcriptome signatures, Epi, Epithelium. d Heatmap showing the relative expression of up-regulated markers in 13 sub-populations of epithelium. e InferCNV heatmap of all epithelial cells grouped by clinical samples compared with referenced T/NK cells, red and blue represented amplifications and deletions on the respective chromosomes, respectively. O, obese, L, lean. f The relative CNV score of all epithelium in different samples compared to T/NK cells. g The correlation of BMI and CNV of BC patients in TCGA dataset. h The t-SNE plot showing CNV score of all epithelial sub-populations by obese and lean group.
We then performed Single-Cell Regulatory Network Inference and Clustering (SCENIC) analysis to correlate transcription factors (TFs) with gene expression differences among these cell subtypes in epithelium. This analysis revealed a lot of TFs as the underlying regulators in different epithelial cell populations of BC. For example, PSMD12, NR2F6 and POU2F1 showed specific regulation with higher regulon specificity scores (RSS) in APOD+ Epi, while basal-like epithelial cells had significant up-regulation patterns of SOX9, FOXC1 and SOX10. Some TFs were identified with similarly regulatory roles in different epithelial subtypes, like FOXD1, FOXO6, HOXB4 and HOXB7 (Supplementary Figs. S3, S4).
We intended to distinguish malignant from non-malignant epithelial cells by inferring large-scale chromosomal copy number variations (CNVs). The hierarchical clustering showed that the epithelium in most samples, except sample L3, had relatively more deletions or amplifications of entire chromosomes compared with T/NK cells (Fig. 2e). Moreover, the epithelium of obese samples exhibited higher CNV scores than lean samples (Fig. 2f). The bulk samples of TCGA database also demonstrated that the BMI was positively correlated with CNV in breast cancer (Fig. 2g). The subtype CPB1+ Epi, CGA+ Epi and APOD+ Epi had relatively higher CNV scores compared to other cells, which might represent malignant cells enriched in obese samples (Fig. 2h). These results suggested that the CNV analysis could be used to differentiate tumors of breast cancer with different BMI.
Obesity-specific macrophages up-regulated key regulator NR1H3 underlying lipid metabolism in OABC
Myeloid cells (n = 4056) were re-clustered into 8 sub-populations consisting of 13 sub-clusters, including 5 macrophage populations (marked by APOE and C1QC), 1 monocyte population (S100A8) and 2 dendritic cell (DC) populations (CD1C, CLEC9A) (Fig. 3a, b, Supplementary Fig. S5a). These two DC subsets exhibited conventional dendritic cell (cDC) expression signatures with high levels of CLEC9A (DC2), CD1C and CLEC10A (DC1) [21, 22]. We noticed that macrophage subsets had different distribution in obese and lean samples, the Macro3 was enriched in lean samples, while Macro4 had higher proportion in obese BC patients (Fig. 3c, Supplementary Fig. S5b). Macro3 expressed Fc gamma binding protein (FCGBP) and solute carrier family 1 member 3 (SLC1A3), Macro4 preferentially expressed fatty acid binding protein 4 (FABP4) and CD36 molecule (Fig. 3d). FABP4 has been demonstrated to promote obesity-associated breast cancer development [23]. In order to validate the expression of FABP4 in specific macrophage, we performed the immunofluorescent staining on tumor resection specimens (Supplementary Table S1), and observed the increased proportion of Macro4 (IBA-1+FABP4+) in obese versus lean BC patients (Fig. 3e, Supplementary Fig. S6). There was evidence suggesting that the interactions between FABP4 and CD36 regulated fatty acid import, transport, and metabolism in breast cancer [24]. The GO analysis of highly-expressed genes in Macro4 also showed lipoprotein, lipid and cholesterol-related pathways were significantly enriched (Fig. 3f). Therefore, we speculated this obesity-enriched Macro4 population might have a vital role in lipid metabolism of OABC.
t-SNE plot showing 4056 myeloid cells colored by cluster (a), cell type (b) and group (c). d Heatmap showing the relative expression of up-regulated markers in 8 sub-populations of myeloid cells. e Immunofluorescence staining of the sub-population Macro4 in one representative obese and lean tumor sample, IBA-1 is a marker of macrophages, scale bars, 200 μm. f The GO analysis of up-regulated genes in Macro4 population based on biological process terms. g Heatmap of TF regulon activity for myeloid sub-populations. T-test was used for these significance comparison, ***p-value < 0.001.
The SCENIC analysis revealed potential TFs underlying the regulation in myeloid subtypes, for example, nuclear receptor subfamily 1 group H member 3 (NR1H3) was identified as a potential TF regulating gene expression in Macro4 specifically (Fig. 3g). The NR1 family members have been reported as key regulators of macrophage function, including NR1H3 [25, 26]. We noticed that NR1H3 was significantly up-regulated in Macro4 compared to other myeloid subsets (Supplementary Fig. S5). The multiplexed immunofluorescent staining verified that NR1H3 was highly expressed in Macro4 of obese samples compared with lean ones (Fig. 3e, Supplementary Fig. S6). FABP4 was identified as a potential target gene of transcription factor NR1H3 by SCENIC analysis, the staining showed that the expression of NR1H3 and FABP4 was colocalized in Macro4 (Fig. 3e, Supplementary Figs. S6, S7). It is noteworthy that the independent samples used for validation included not only ER+ BCs previously used for scRNA-seq, but also Her-2 enriched BCs and triple negative breast cancers (TNBCs). Macro4 was surprisingly consistent in the samples presented, and it appears that this alteration in the tumor microenvironment is not dependent on hormone receptor status but is a common phenotype in OABCs. Obviously this requires a larger sample size for validation, but it does give us some encouragement, especially for triple-negative and obese BC patients with a poor prognosis, and provides additional potential therapeutic direction.
To explore the possible mechanism of inducing the high expression of NR1H3 in Macro4, we first co-cultured ZR-751 with human macrophages (THP-1) to detect the NR1H3 expression level of THP-1 by qPCR and found that co-culture with BC cells did not stimulate the high expression of NR1H3 in THP-1 (Fig. 4a). We obtained the same results using MCF7, MDA-MB-231 and SKBR3 cells co-cultured with THP-1, respectively (Supplementary Fig. S8a). Then we collected the peri-cancerous adipose tissue of 3 BC patients with BMI > 30 and 3 BC patients with BMI < 25 (Supplementary Table S1), and added the adipose tissue homogenate of the two groups to the THP-1 culture medium, respectively. The qPCR results showed that the adipose tissue homogenates from patients with BMI > 30 stimulated NR1H3 expression in THP-1 more strongly compared to patients with BMI < 25 (Fig. 4b). These results demonstrate that the high expression of NR1H3 of Macro4 in OABCs is not influenced by BC cells, but may be induced by the adipose tissue of obese patients.
a The qPCR results of NR1H3 in PMA-stimulated THP-1 cells co-cultured with ZR751 cells or THP-1 cells alone served as controls. b The qPCR results of NR1H3 in PMA-stimulated THP-1 cells co-cultured with peri-cancerous adipose tissue homogenates from patients with BMI > 30 (n = 3) or patients with BMI < 25 (n = 3). c Heatmap showing the relative expression of lipoprotein metabolism related genes and NR1H3 by bulk RNA sequencing in THP-1 cells, with a control (siCtrl, n = 3) or NR1H3 siRNA (siNR1H3, n = 3). The qPCR results of NR1H3 (d) and FABP4 (e) mRNA in siNR1H3 and control cells (n = 3). f, g The relative protein expression of NR1H3 and FABP4 by western blot in siNR1H3 compared with control. h The ELISA results of FABP4 in supernatants of cultures of THP-1 cells after knockdown of NR1H3 compared with control. The CCK-8 proliferation assay of MCF7 (i) and ZR751 (j) cells co-cultured with THP-1 medium with or without siNR1H3. T-test or Mann–Whitney U test was used for these significance comparison, *p-value < 0.05, **p-value < 0.01, ***p-value < 0.001.
To validate the regulation of NR1H3 on FABP4, we interfered the expression of NR1H3 in PMA-stimulated THP-1 by siRNA. The RNA sequencing of THP-1 cells revealed the changed transcriptome upon NR1H3 knockdown, the lipoprotein metabolism-related genes including FABP4 were significantly dysregulated (Fig. 4c), qPCR results also showed the levels of NR1H3 and FABP4 were significantly down-regulated compared with the control (Fig. 4d, e;Supplementary Fig. S8b, c). The western blot revealed the protein levels of NR1H3 and FABP4 were decreased after NR1H3 knockdown (Fig. 4f, g;Supplementary Fig. S8d, e). However, the functions of NR1H3 have not been comprehensively revealed in BC. Therefore, we co-cultured the human BC cell line MCF7 or ZR751 with 30% conditioned medium from THP-1. Before this, we verified that THP1 can exocrine FABP4 and be regulated by NR1H3. ELISA results showed that the content of FABP4 in the supernatant of THP-1 culture medium decreased significantly after knocking down NR1H3 (Fig. 4h; Supplementary Fig. S8f). The CCK-8 assay revealed that the proliferation abilities of both MCF7, ZR-751, MDA-MB-231 and SKBR3 cells were significantly declined via NR1H3 knockdown (Fig. 4i, j; Supplementary Fig. S8g, h). The transwell assay indicated the migration of MCF7 and ZR-751 cells were not significantly different between the siNR1H3 and negative control (Supplementary Fig. S8i, j). These results suggested that the expression of NR1H3 in macrophages affected the proliferation of BC cells, not the migration. The NR1H3 expression level might be related to different tumor volume between obese and lean BC patients, while had little effects on lymph node metastasis (Fig. 1a, b, Supplementary Fig. S1).
Since NR1H3 is a crucial transcription factor in macrophages, our initial approach involved using ChIP-qPCR to validate whether NR1H3 directly binds to FABP4 to regulate its transcriptional expression. We employed an NR1H3 antibody to bind the DNA in THP-1 cells, and qPCR was used to detect FABP4. Surprisingly, the NR1H3 antibody failed to pull down the DNA of FABP4, indicating that NR1H3 cannot directly bind to FABP4 to regulate its expression (Fig. 5a). It is widely recognized that FABP4 plays a role in lipid metabolism as a fatty acid binding protein, and SREBP1 serves as a critical regulator of lipid metabolism. To clarify the relationship between FABP4 and SREBP1, we referred to gene expression data of macrophages from the Gene Expression Omnibus database (GSE100521). The probe identifiers were converted to gene symbols based on the annotation file of the Illumina HumanHT-12 V4.0 expression beadchip. A correlation analysis was conducted using the pearson method, and the results demonstrated a positive correlation between the expressions of FABP4 and SREBP1 in macrophages. Consequently, we treated THP-1 cells with 20 μM Fatostatin, an inhibitor of SREBP1, and conducted qPCR, which clearly demonstrated a significant downregulation of FABP4 (Fig. 5b, c). Notably, the results of ChIP-qPCR revealed that the use of NR1H3 antibody successfully pulled down the DNA of SREBP1 (Fig. 5d). To further elucidate the relationship between NR1H3, SREBP1, and FABP4, we proceeded with the transfection of the NR1H3 overexpression plasmid into THP-1 cells. Subsequently, qPCR and Western blotting revealed that NR1H3 upregulated the expression of both SREBP1 and FABP4. However, after treatment with 20 μM Fatostatin, the expression of FABP4 was significantly downregulated (Fig. 5e, f). Moreover, the RNA sequencing of THP-1 cells revealed the changed transcriptome upon NR1H3 knockdown, the SREBP1 were significantly dysregulated (Supplementary Fig. S8k). In conclusion, these findings strongly suggest that NR1H3 regulates FABP4 expression through its interaction with the DNA of SREBP1.
a The ChIP-qPCR results of interaction between NR1H3 and FABP4. b, c The relationship between FABP4 and SREBP1 in bioinformatics analysis and the relative expression of FABP4 by qPCR in Fatostatin compared with control. d The ChIP-qPCR results of interaction between NR1H3 and SREBP1. e, f The relative expression of SREBP1 and FABP4 by qPCR and western blot in GV-NR1H3 transfected THP-1 cells treated with or without Fatostatin. g Monitoring body weight of C57BL/6 mice either an LFD or an HFD for 13 weeks (n = 8/group). h Measurement of the tumor volume in lean and obese mice (n = 8/group) post-orthotopical injection of E0771 cells (5 * 105) at the indicated time points. i Surgically removed tumor tissues from lean and obese mice 21 days post-inoculation. j Tumor weight of lean and obese mice (n = 8/group) post implantation of E0771 cells for 21 days. k Immunofluorescence staining of the sub-population Macro4 in one representative obese and lean mice tumor sample, 100 μm. T-test or Mann–Whitney U test was used for these significance comparison, *p-value < 0.05, **p-value < 0.01.
Subsequently, we conducted an experiment in which C57BL/6 mice were divided into two groups: one group was fed a low-fat diet (LFD), while the other group was fed a high-fat diet (HFD) for a duration of 13 weeks. Furthermore, we orthotopically injected an equal number of E0771 mammary tumor cells into both lean and obese mice (Fig. 5g). Notably, tumor growth in the obese mice was significantly accelerated compared to the lean mice, with the tumors in obese mice weighing twice as much as those in lean mice (Fig. 5h–j).To further investigate the underlying mechanisms, we surgically removed the tumors from the mice and conducted immunofluorescence analysis. This analysis confirmed the presence of FABP4+ Macro4 in the obese mice (Fig. 5k). When combined with the aforementioned results, our findings successfully validate the presence of Macro4 in in vivo experiments and provide confirmation that obesity can promote the development of breast cancer.
Differential expression profiles of cancer-associated fibroblast sub-populations
Cancer-associated fibroblasts (CAF) contained 6 sub-clusters of inflammatory fibroblasts (iCAFs) and myofibroblasts (myCAFs, Fig. 6a, Supplementary Fig. S9a), the iCAFs showed elevated expression of decorin (DCN) and C-X-C motif chemokine ligand 12 (CXCL12), and myCAFs up-regulated actin alpha 2, smooth muscle (ACTA2) and transgelin (TAGLN, Fig. 6b, c). The classification of CAFs was similar in BC [27] and other cancers, like pancreatic ductal adenocarcinoma [28] and ovarian cancer [29]. However, we also found potential cell markers between iCAFs and myCAFs by differential analysis (Fig. 6d). The iCAFs up-regulated CFD, APOD and CXCL14, while myCAFs over-expressed RGS5, MCAM and MYH11.
a t-SNE plot showing 6996 fibroblasts colored by sub-population. b, c t-SNE plot showing the expression of marker genes of iCAFs and myCAFs. d Scatter plot showing the differentially expressed genes between iCAFs and myCAFs, red represented differential genes (DIFF). e t-SNE plot showing all fibroblasts colored by group. f Heatmap showing the relative expression of up-regulated genes in 6 sub-populations of CAF. g The GO analysis of up-regulated genes in myCAF2 population based on biological process terms.
The obese samples had higher proportions of iCAF2 and myCAF2 compared to lean samples (Fig. 6e, Supplementary Fig. S9b), collagen-related genes (COL10A1, COL11A1), INHBA and POSTN were specifically expressed in iCAF2, CC cytokine genes (CCL19, CCL21), CTSC, FABP4 and CD36 were highly expressed in myCAF2 (Fig. 6f, Supplementary Fig. S9c, d). We noticed that myCAF2 up-regulated many genes (log2FC > 1) which were also over-expressed in Macro4 compared with other corresponding cell subsets, especially FABP4, these genes were enriched in lipid or fatty acid transport-related pathways (Fig. 6g). It is suggested that these two sub-populations may play synergistic roles in lipid metabolism in OABC. Both myCAF2 and Macro4 may be involved in regulating the biological function of breast cancer through exocrine FABP4. Building on our previous finding that NR1H3 affects the expression of FABP4 in Macro4 through SREBP1, we have reason to believe that Macro4 may have a mechanism for regulating the activation of myCAF2 and the expression of FABP4. Our research group aims to further explore the role of Macro4 in the breast cancer microenvironment to provide additional evidence.
Discussion
Despite extensive research on the associations between obesity and the risk of breast cancer in recent years, there has been a lack of understanding regarding cellular heterogeneity in obesity-associated and non-obesity associated breast cancer. In this study, we conducted scRNA-seq analysis on 6 BC patients with different BMIs. Our analysis resulted in a comprehensive single-cell transcriptomic atlas to characterize tumor ecosystems in both obese and non-obese BC. Our work revealed distinct expression signatures of tumor cells in these two types of BC. The highly expressed genes of tumor epithelium in OABC were found to be involved in the estrogen response pathway. There is certainly evidence that the local production of estrogen in adipose tissue drives the growth of breast cancer after menopause [30]. Therefore, the tumor cells of OABC are more closely related to estrogen signaling compared to non-obese BC. The chemokine CXCL14, which was up-regulated in CPB1+ Epi and CGA+ Epi, has been reported to secrete brown fat and promote enhanced brown adipose tissue activation, thereby providing protection against obesity [31]. However, several contradictory findings from studies have shown that CXCL14 was positively associated with obesity [32, 33]. In our study, we found high expression of CXCL14 in different cell types of the obesity group, including epithelium, macrophages, and iCAFs. Additionally, it was observed that brown adipocytes-secreted CXCL14 could recruit alternatively activated (M2) macrophages [31]. Therefore, we speculated that highly expressed CXCL14 in obese patients might also recruit other types of cells, such as specific macrophages, to regulate metabolic processes that link obesity and breast cancer.
Previous studies have found that genomic CNVs contribute to the genetic susceptibility of common childhood obesity or severe early-onset obesity [34, 35]. However, few studies have reported the associations between obesity and CNV on the genome-wide in cancers. Although we analyzed CNVs of tumor epithelium between obese and lean patients using bioinformatic methods in currently limited samples, further validation from more bulk samples from the TCGA dataset confirmed the correlation with BMI in breast cancer. Of course, additional clinical samples are needed to evaluate the effects of BMI on genomic CNVs.
Although sporadic reports have depicted the molecular function of NR1H3 in breast cancer [36], few research linked it with OABC. In this study, we demonstrated that transcription factor NR1H3 was up-regulated in a subset of obesity-specific macrophages, Macro4, which further regulated the expression of fatty acid binding protein FABP4 and increased the proliferation of tumor cells in BC (Fig. 7). It is well known that SREBP1 plays a crucial role in regulating lipid homeostasis by modulating cholesterol and fatty acid metabolism [37]. Additionally, it has been shown that SREBP1 is critical in the development of cancers such as hepatocellular carcinoma, pancreatic carcinoma, and prostate cancer [38,39,40]. Through bioinformatics analysis, a positive correlation between SREBP1 and FABP4 in macrophages was found. Therefore, we further explored the relationship between NR1H3 and SREBP1. Through cellular experiments, we found that NR1H3 could bind to the DNA of SREBP1 and promote the expression of SREBP1. Singnificantly, SREBP1 inhibitor Fatostatin abrogated the increased FABP4 expression caused by overexpression of NR1H3. These results demonstrate that NR1H3 regulates FABP4 by directly binding to the DNA of SREBP1. Studies suggested that liver X receptors α and β (LXRα and LXRβ), namely NR1H3 and NR1H2, were nuclear receptors with critical roles in the transcriptional programs of cholesterol homeostasis and lipid metabolism [41]. Nelson et al. have shown that 27-hydroxycholesterol (27HC) acted through the LXR in macrophages and possibly other cells, stimulated the growth and metastasis of tumors in several models of breast cancer [42, 43]. These data provided singnifical support for the function of NR1H3 in macrophages of BC and its link between lipid metabolism and cancer.
Our analysis revealed that fatty acid binding protein FABP4 was significantly up-regulated in both Macro4 and myCAF2, which has been demonstrated to be highly expressed in adipocytes and macrophages [44]. The study reported that expression of adipocyte/macrophage FABP (A-FABP or FABP4) in tumor-associated macrophages (TAM) promoted breast cancer progression through IL-6/STAT3 signaling [45]. However, it is necessary to explore further the role of FABP4 in specific cell types, including macrophage and fibroblast in breast cancer. A recent study identified a vital role for circulating adipose fatty acid binding proteins in OABC as a new link between obesity and breast cancer risk [13]; it provided necessary experimental evidence for our single-cell findings that FABP4 was a cell type specifically expressed in OABC.
In conclusion, our single-cell atlas from obese and non-obese BC patients uncovered cell diversity and transcriptomic difference in the ecosystem of breast tumors, the signal axis NR1H3-FABP4 up-regulated in obesity-specific macrophage might be related to the development of OABC.
Methods
Sample collection
The primary untreated breast cancers used utilized in this study (Supplementary Table S1) were obtained following collected with written informed consent from all patients. For scRNA-seq analysis, three obese patients (BMI > 30) and three normal-weight BC patients (BMI < 30) were randomly selected, while four obese and four normal-weight BC patients were randomly selected for subsequent laser confocal immunofluorescence staining. Tumor tissues from 3 obese (BMI > 30) and 3 normal-weight BC patients (BMI < 30) were randomly selected for scRNA-seq, and tumor from 5 obese and 5 normal-weight BC patients were randomly selected for subsequent laser confocal immunofluorescence staining. Detailed clinical data were shown in Supplementary Table S1. In addition, clinical data of BC patients with primary invasive ductal carcinoma were collected to analyze the correlation between BMI and tumor-related clinical parameters. These patients underwent surgical resection in Qilu Hospital (Qingdao), Shandong University from January 2015 to December 2021 with written informed consent for their clinical data.
Tissue dissociation
Samples were obtained from fresh surgical resections. BC tumors were dissociated using the human Tumor Dissociation Kit (Miltenyi Biotec, Germany) following the manufacturer’s protocol. The dissociated cells were then resuspended to remove dead cells using Dead Cell Removal Kit (Miltenyi Biotec, Germany). Subsequently, the overall cell viability was confirmed to be above 85%. The cells were further resuspended for cell counting and diluted to an appropriate concentration.
Single-cell RNA-seq library preparation and sequencing
Single-cell suspensions were loaded onto 10X Chromium chip in order to capture 5000-10,000 single cells, following the manufacturer’s instructions of 10X Genomics Chromium Single-Cell 3’ kit (V3). The captured single cells underwent cDNA amplification and library construction using the standard protocol of the 3’ kit (V3). The single-cell libraries were sequenced on Illumina NovaSeq 6000 sequencing system (paired-end multiplexing run, 150 bp) by LC-Bio Technology Co., Ltd. (Hangzhou, China).
Single-cell data preprocessing and quality control
To generate a raw gene expression matrix for each sample, CellRanger (10X Genomics, version 3.1.0) was employed. The STAR algorithm (https://github.com/alexdobin/STAR) was used for mapping the reads to the human reference genome GRCh38. Subsequently, the Seurat package (version 3.1.5) was utilized for quality control (QC) of the single cells. Cells meeting the criteria of having >500 detected genes and <25% mitochondrial genes were considered to have passed QC. A total of 48,033 high-quality single cells were included for subsequent bioinformatic analysis using Seurat, which included cell normalization, principal component analysis (PCA), and cell clustering.
After removing low quality cells, we applied a global-scaling normalization method “LogNormalize” that normalized the gene expression measurements for each cell by default. To overcome the extensive technical noise in any single gene for scRNA-seq data, we then performed PCA and determined the number of PCs was used for the cell clustering. To further correct for batch effects originating from different patients, we applied the Harmony algorithm immediately after completing PCA. Harmony utilized the combined matrix of informative genes to correct for batch effects.
Identification of the major cell types and their subtypes
The clustering of major cell types and subtypes was performed with the “FindClusters” function of the Seurat. In order to identify the subtypes within each major cell type, the cells belonging to each major type were reanalyzed individually. Subsequently, major cell clusters and their sub-clusters were visualized using t-distributed Stochastic Neighbor Embedding (t-SNE), and the annotation of cell types was based on the average expression of well-recognized marker genes or highly expressed genes from related literatures of breast cancer [21, 22]. To identify genes that are specifically expressed in certain clusters or differentially expressed between clusters, the Seurat “FindAllMarkers” function was utilized. This function tested genes with a > 0.25-fold difference (log-scale) on average between the two groups of cells and detectable expression in more than 25% of cells in either of the two groups.
Single-Cell Regulatory Network Inference and Clustering (SCENIC) analysis
To investigate gene regulatory networks, we performed SCENIC analysis using the pySCENIC pipeline (v0.12.1), a fast implementation of the SCENIC pipeline as described by the developers [46, 47]. SCENIC utilizes transcription factors and cis-regulatory sequences to reconstruct gene regulatory networks [46]. Activated transcription factors (TFs) were identified in the Binary matrix, and the top differentially activated TFs in each cell population were selected using the Wilcoxon test based on the AUC matrix.
CNV analysis of epithlium
To distinguish the epithelium in obese and lean BC samples, we calculated the copy number variations (CNVs) of all epithelial cells in each sample based on the current scRNA-seq data using R package inferCNV (version 1.3.3) as described previously [48]. This package inferred the genetic profiles of each epithelial cell based on the average expression of large genes sets (101 genes) in each chromosomal region (sex chromosomes were excluded) compared to the control cells (T/NK cells). Other parameters were set as default.
In addition, the CNV and BMI data of 765 BC patients (including 711 patients with BMI > 30) from TCGA database were employed to study their correlation.
Multiplexed immunofluorescent staining
For immunofluorescence staining, breast cancer tissues were fixed in 4% paraformaldehyde, embedded in paraffin and then sectioned. Standardized procedures for dewaxing and antigen retrieval were applied to the sections. After treatment with 3% H2O2 for 15 min and blocking with 5% normal serum for 30 min at room temperature. The samples were incubated with primary antibodies for 1.5 h at room temperature and subsequently with secondary antibodies for 30 min at room temperature. Nuclei were stained with 4’, 6 diamidino-2-phenylindole (DAPI) for 15 min. Confocal microscopy was performed using a Leica ATELLARIS 5 laser scanning confocal microscope. The following antibodies were used for immunofluorescence staining: anti-IBA-1 antibody (1:200, Proteintech, 66827-1-Ig), anti-DAP12 antibody (1:200, Santa Cruz, sc-133174), anti-FABP4 antibody (1:200, Abcam, ab92501), anti-LXR alpha antibody (1:100,Abcam, ab41902), Goat-Anti-Mouse IgG H&L (1:1000, Abcam, ab150115), Goat-Anti-Rabbit IgG H&L (1:1000, Abcam, ab150077), DAPI (Solarbio, C0065).
Cell lines and siRNAs
Breast cancer cell line ZR-751 and human leukemia monocytic cell line THP-1 were cultured in RPMI 1640 (Invitrogen, Shanghai, China). Breast cancer cell line MCF7 and MDA-MB-231 was cultured in DMEM (Viva Cell, Shanghai, China). Breast cancer cell line SKBR3 was cultured in McCoy’s (Viva Cell, Shanghai, China). All the cells were purchased from Shanghai Institute of Cell Biology, Chinese Academy of Sciences, and they were authenticated and tested for mycoplasma contamination. All the media were supplemented with 10% fetal bovine serum (Viva Cell, Shanghai, China), 100 units/mL penicillin, and 100 μg/mL streptomycin (Procell, Wuhan, China). THP-1 cells (1 × 106/ml) were differentiated using 200 ng/ml phorbol 12-myristate 13-acetate (PMA, Sigma-Aldrich, Shanghai, China) for 8 h. Differentiation of PMA treated cells was enhanced by removing the PMA-containing media then incubating the cells in fresh RPMI 1640 (10% FBS) for a further 16 h, before being used in the indicated experiments. The siRNA against NR1H3 (siRNA#1, Sense: 5’-GGAGUGUCGGCUUCGCAAA-3’ Antisense: 5’-UUUGCGAAGCCGACACUCC-3; siRNA#2, Sense: 5’-CACAGAGAUCCGUCCACAA-3’ Antisense: 5’-UUGUGGACGGAUCUCUGUG-3) was synthesized by TSINGKE (Beijing, China). Human NR1H3 expression plasmid GV-NR1H3 was synthesized by Genechen Co. (Shanghai, China). The negative control is referring to with a control siRNA sequence. The THP-1 cells were treated with siRNA or expression plasmid after PMA stimulation.
Co-culture experiments
Co-culture experiments were carried out using a transwell system (3.0 μm pore size, Transparent (PET) Membrane; Corning Life Sciences, Lowell, MA, USA). For experiments assessing THP-1 biology, 3.5 ×104 ZR751 cells were seeded in the upper chamber with PMA-stimulated THP-1 in the bottom for 24 h. PMA-stimulated THP-1 cells cultured alone served as controls.
To obtain adipose tissue homogenate, 50 mg of pericancerous adipose tissue was transferred to a pre-cooled 1 ml RPMI 1640 (10% FBS, 1% PS) on ice. The tissues were then homogenized evenly using a sample homogenate solution. Following centrifugation at 3000 rpm for 10 min at 4 °C, the water liquid layer in the middle was retained. PMA-stimulated THP-1 were cultured with 30% conditioned medium (CM) containing adipose tissue homogenate for 24 h. There isn’t any PMA in the conditioned media given to the epithelial cell culture. Samples were obtained from fresh surgical resections.
RNA sequencing and analysis
Total RNAs were isolated from the THP-1 cells with or without NR1H3 knockdown using TRIzol LS reagent (Thermo Fisher Scientific, USA). The quality and quantity of the total RNA were assessed using Qubit 3.0 (Thermo Fisher Scientific, USA) and Agilent 2100 Bioanalyzer (Agilent Technologies, USA), respectively. The mRNA library was constructed according to the protocols of TruSeq Stranded Total RNA Library Prep Kit v2 (Illumina, USA) by Genesky Biotechnologies Inc., Shanghai (China).
The raw reads of mRNA sequencing were evaluated for quality using FastQC. The clean reads of mRNA were aligned to the human reference genome hg19 using HISAT2, and StringTie was used for transcript assembly and quantification.
RT-qPCR
Total RNAs were isolated from tissues or cultured cells using TRIzol LS reagent (Thermo Fisher Scientific, USA) and reverse-transcribed into cDNA using All-In-One 5X RT MasterMix (abm, Vancouver, Canada). Then qPCR was carried out using BlasTaqTM 2X qPCR MasterMix (abm, Vancouver, Canada). The primers was synthesized by Sangon Biotech (Shanghai, China). Sense and reverse primers were: GAPDH, Forward 5’-GGAGCGAGATCCCTCCAAAAT-3’, Reverse 5’-GGCTGTTGTCATACTTCTCATGG-3’; NR1H3, Forward 5’-G CCTTCAGAACCCACAGAGATCC-3’, Reverse 5’-ACGCTGCATAGCTCGTTCC-3’; FABP4, Forward 5’-ACTGGGCCAGGAATTTGACG-3’, Reverse 5’- CTCGTGGAAGTGACGCCTT-3’.
Chromatin immunoprecipitation and quantitative PCR analyses
Chromatin immunoprecipitation (ChIP) and quantitative PCR (qPCR) analyses were performed as described by Hyperactive pG-MNase CUT&RUN Assay Kit for PCR/qPCR (HD101-01, Nanjing, China). Calculate the circulating threshold of immunoprecipitated DNA for various samples using qPCR. Antibodies used for ChIP: normal rabbit IgG (ABcolonal, AC005), anti-H3K4me3 (ABcolonal, A22146) and anti-NR1H3 (Abcam, ab41902). Primers used for ChIP PCR analyses: FABP4, Forward 5’-ACTGGGCCAGGAATTTGACG-3’, Reverse 5’- CTCGTGGAAGTGACGCCTT-3’; SREBP1, Forward 5’-ACTTCTGGAGGCATCGCAAGCA-3’, Reverse 5’-AGGTTCCAGAGGAGGCTACAAG-3’.
Western blot
Protein lysates were obtained from THP-1 cells transfected with siRNA. The lysates were separated by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and transferred on polyvinylidene difluoride (PVDF) membranes. After being blocked with 5% fat-free dry milk, the membranes were incubated with using anti-β-actin (Gene Tex, Irvine, CA, USA), anti-NR1H3 (Abcam, Shanghai, China), anti-FABP4 (Abcam, Shanghai, China), anti-SREBP1 (ABcolonal, Wuhan, China).
ELISA
Culture supernatant from from THP-1 cells transfected with siRNA were obtained via centrifugation (10 min, 4 °C, 3000 RPM). We used enzyme-linked immunosorbent assay (ELISA) kits for FABP4 (BOSTER, Wuhan, China). All plates were analyzed using the SpectraMax i3x multimode reader (Molecular Devices, Shanghai, China) at 450-nm wavelengths, according to the manufacturers’ recommendations.
CCK-8 cell proliferation assay
To assess cell proliferation, MCF7, ZR-751, MDA-MB-231 and SKBR3 cell lines were cultured with 30% conditioned medium (CM) with or without siNR1H3 from PMA-stimulated THP-1 for 24 h. Then, 5 × 103 cells/well were seeded in a 96-well plate and were cultured with 30% CM every 24 h. Cell proliferation was measured at OD450 nm using the CCK-8 kit (Dojindo CO. Ltd., Kumamoto, Japan) every 24 h. There isn’t any siRNA or PMA in the conditioned media given to the epithelial cell culture.
Transwell assay
Briefly, MCF7 and ZR751 cell lines were cultured with 30% CM with or without siNR1H3 from PMA-stimulated THP-1 for 24 h. Then, 3.5 × 104 MCF7 or 5 × 104 ZR751 in 200 μL of serum-free medium were added to the upper chambers of the Transwell plate, and the lower chambers were filled with 600 μL of medium containing 20% FBS as a chemoattractant. After 24 h, the migrated cells were fixed in 4% paraformaldehyde and stained with 0.1% crystal violet, and the non-migratory cells on the upper surface of the chambers were removed by a cotton swab. Then, at least four random fields were selected and observed by microscopy (Olympus, Japan). There isn’t any siRNA or PMA in the conditioned media given to the epithelial cell culture.
Tumor challenges
To monitor the progression of tumors in lean and obese mice, we followed the following experimental procedure. Six-week-old female C57BL/6 mice were randomly divided into two groups, with 8 mice in each group, and fed either a low-fat diet (LFD) or a high-fat diet (HFD) for a duration of 13 weeks. Subsequently, 5*105/100ul (PBS) E0771 cells were orthotopically implanted into the mammary fat pad of the lean and obese mice, respectively. Tumor growth was measured using calipers, and the tumor volume was calculated using the formula 0.5 * (large diameter) * (small diameter)2 twice a week. The mice were then continuously fed with their respective diets, either LFD or HFD, for an additional 21 days. After the designated period, tumor tissues were collected for frozen sections, and immunofluorescence analysis was performed. For the immunofluorescence staining, the following antibodies were utilized: anti-DAP12 antibody (1:200, Abcam, ab283679) and anti-FABP4 antibody (1:200, Santa Cruz, sc-261529). The researchers were informed of the grouping during the experiment and when evaluating the results.
Statistical analysis
All data processing was performed using R 3.6.1 software. The statistical methods and threshold for corresponding analysis were explicitly described in the results or figure legends or Materials and Methods. All statistical results with a p-value < 0.05 were considered to indicate statistical significance.
Data availability
Single-cell RNA-sequencing data have been deposited in the GSA database under accession code HRA004141. All other relevant data are within the paper and Supplementary files.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin. 2021;71:209–49.
Pan XF, Wang L, Pan A. Epidemiology and determinants of obesity in China. lancet Diabetes Endocrinol. 2021;9:373–92.
Whitlock G, Lewington S, Sherliker P, Clarke R, Emberson J, Halsey J, et al. Body-mass index and cause-specific mortality in 900 000 adults: collaborative analyses of 57 prospective studies. Lancet. 2009;373:1083–96.
Bluher M. Obesity: global epidemiology and pathogenesis. Nat Rev Endocrinol. 2019;15:288–98.
Marengo A, Rosso C, Bugianesi E. Liver Cancer: Connections with Obesity, Fatty Liver, and Cirrhosis. Annu Rev Med. 2016;67:103–17.
Bardou M, Barkun AN, Martel M. Obesity and colorectal cancer. Gut. 2013;62:933–47.
Neuhouser ML, Aragaki AK, Prentice RL, Manson JE, Chlebowski R, Carty CL, et al. Overweight, Obesity, and Postmenopausal Invasive Breast Cancer Risk: A Secondary Analysis of the Women’s Health Initiative Randomized Clinical Trials. JAMA Oncol. 2015;1:611–21.
Bhaskaran K, Douglas I, Forbes H, dos-Santos-Silva I, Leon DA, Smeeth L. Body-mass index and risk of 22 specific cancers: a population-based cohort study of 5.24 million UK adults. Lancet. 2014;384:755–65.
Renehan AG, Tyson M, Egger M, Heller RF, Zwahlen M. Body-mass index and incidence of cancer: a systematic review and meta-analysis of prospective observational studies. Lancet. 2008;371:569–78.
Michels KB, Terry KL, Willett WC. Longitudinal study on the role of body size in premenopausal breast cancer. Arch Intern Med. 2006;166:2395–402.
Chan DSM, Vieira AR, Aune D, Bandera EV, Greenwood DC, McTiernan A, et al. Body mass index and survival in women with breast cancer-systematic literature review and meta-analysis of 82 follow-up studies. Ann Oncol. 2014;25:1901–14.
Jiralerspong S, Goodwin PJ. Obesity and Breast Cancer Prognosis: Evidence, Challenges, and Opportunities. J Clin Oncol Oncol. 2016;34:4203–16.
Hao J, Zhang Y, Yan X, Yan F, Sun Y, Zeng J, et al. Circulating Adipose Fatty Acid Binding Protein Is a New Link Underlying Obesity-Associated Breast/Mammary Tumor Development. Cell Metab. 2018;28:689–705 e685.
Zhang C, Yue C, Herrmann A, Song J, Egelston C, Wang T, et al. STAT3 Activation-Induced Fatty Acid Oxidation in CD8(+) T Effector Cells Is Critical for Obesity-Promoted Breast Tumor Growth. Cell Metab. 2020;31:148–161 e145.
Qian J, Olbrecht S, Boeckx B, Vos H, Laoui D, Etlioglu E, et al. A pan-cancer blueprint of the heterogeneous tumor microenvironment revealed by single-cell profiling. Cell Res. 2020;30:745–62.
Zheng L, Qin S, Si W, Wang A, Xing B, Gao R, et al. Pan-cancer single-cell landscape of tumor-infiltrating T cells. Science. 2021;374:abe6474.
Azizi E, Carr AJ, Plitas G, Cornish AE, Konopacki C, Prabhakaran S, et al. Single-Cell Map of Diverse Immune Phenotypes in the Breast Tumor Microenvironment. Cell. 2018;174:1293–1308 e1236.
Nguyen QH, Pervolarakis N, Blake K, Ma D, Davis RT, James N, et al. Profiling human breast epithelial cells using single cell RNA sequencing identifies cell diversity. Nat Commun. 2018;9:2028.
Geldhof V, de Rooij L, Sokol L, Amersfoort J, De Schepper M, Rohlenova K, et al. Single cell atlas identifies lipid-processing and immunomodulatory endothelial cells in healthy and malignant breast. Nat Commun. 2022;13:5511.
Lee SB, Kim HK, Choi Y, Ju YW, Lee HB, Han W, et al. Dynamic and subtype-specific interactions between tumour burden and prognosis in breast cancer. Sci Rep. 2020;10:15445.
Bassez A, Vos H, Van Dyck L, Floris G, Arijs I, Desmedt C, et al. A single-cell map of intratumoral changes during anti-PD1 treatment of patients with breast cancer. Nat Med. 2021;27:820–32.
Wu SZ, Al-Eryani G, Roden DL, Junankar S, Harvey K, Andersson A, et al. A single-cell and spatially resolved atlas of human breast cancers. Nat Genet. 2021;53:1334–47.
Zeng J, Sauter ER, Li B. FABP4: A New Player in Obesity-Associated Breast Cancer. Trends Mol Med. 2020;26:437–40.
Gyamfi J, Yeo JH, Kwon D, Min BS, Cha YJ, Koo JS, et al. Interaction between CD36 and FABP4 modulates adipocyte-induced fatty acid import and metabolism in breast cancer. NPJ Breast Cancer. 2021;7:129.
Guillen NAG, Gallardo JA, Diaz G, de la Rosa M, Hernandez JV. IH et al. The nuclear receptor LXRalpha controls the functional specialization of splenic macrophages. Nat Immunol. 2013;14:831–9.
Pourcet B, Feig JE, Vengrenyuk Y, Hobbs AJ, Kepka-Lenhart D, Garabedian MJ, et al. LXRalpha regulates macrophage arginase 1 through PU.1 and interferon regulatory factor 8. Circ Res. 2011;109:492–501.
Wu SZ, Roden DL, Wang C, Holliday H, Harvey K, Cazet AS, et al. Stromal cell diversity associated with immune evasion in human triple-negative breast cancer. EMBO J. 2020;39:e104063.
Elyada E, Bolisetty M, Laise P, Flynn WF, Courtois ET, Burkhart RA, et al. Cross-Species Single-Cell Analysis of Pancreatic Ductal Adenocarcinoma Reveals Antigen-Presenting Cancer-Associated Fibroblasts. Cancer Discov. 2019;9:1102–23.
Hornburg M, Desbois M, Lu S, Guan Y, Lo AA, Kaufman S, et al. Single-cell dissection of cellular components and interactions shaping the tumor immune phenotypes in ovarian cancer. Cancer Cell. 2021;39:928–44 e926.
Bhardwaj P, Au CC, Benito-Martin A, Ladumor H, Oshchepkova S, Moges R, et al. Estrogens and breast cancer: Mechanisms involved in obesity-related development, growth and progression. J Steroid Biochem Mol Biol. 2019;189:161–70.
Cereijo R, Gavalda-Navarro A, Cairo M, Quesada-Lopez T, Villarroya J, Moron-Ros S, et al. CXCL14, a Brown Adipokine that Mediates Brown-Fat-to-Macrophage Communication in Thermogenic Adaptation. Cell Metab. 2018;28:750–763 e756.
Takahashi M, Takahashi Y, Takahashi K, Zolotaryov FN, Hong KS, Iida K, et al. CXCL14 enhances insulin-dependent glucose uptake in adipocytes and is related to high-fat diet-induced obesity. Biochem Biophys Res Commun. 2007;364:1037–42.
Tanegashima K, Okamoto S, Nakayama Y, Taya C, Shitara H, Ishii R, et al. CXCL14 deficiency in mice attenuates obesity and inhibits feeding behavior in a novel environment. PloS One. 2010;5:e10321.
Glessner JT, Bradfield JP, Wang K, Takahashi N, Zhang H, Sleiman PM, et al. A genome-wide study reveals copy number variants exclusive to childhood obesity cases. Am J Hum Genet. 2010;87:661–6.
Wheeler E, Huang N, Bochukova EG, Keogh JM, Lindsay S, Garg S, et al. Genome-wide SNP and CNV analysis identifies common and low-frequency variants associated with severe early-onset obesity. Nat Genet. 2013;45:513–7.
Hutchinson SA, Websdale A, Cioccoloni G, Roberg-Larsen H, Lianto P, Kim B, et al. Liver x receptor alpha drives chemoresistance in response to side-chain hydroxycholesterols in triple negative breast cancer. Oncogene. 2021;40:2872–83.
Bian X, Liu R, Meng Y, Xing D, Xu D, Lu Z. Lipid metabolism and cancer. J Exp Med. 2021;218:e20201606.
Galbraith L, Leung HY, Ahmad I. Lipid pathway deregulation in advanced prostate cancer. Pharm Res. 2018;131:177–84.
Zeng H, Qin H, Liao M, Zheng E, Luo X, Xiao A, et al. CD36 promotes de novo lipogenesis in hepatocytes through INSIG2-dependent SREBP1 processing. Mol Metab. 2022;57:101428.
Zhou C, Qian W, Li J, Ma J, Chen X, Jiang Z, et al. High glucose microenvironment accelerates tumor growth via SREBP1-autophagy axis in pancreatic cancer. J Exp Clin Cancer Res. 2019;38:302.
Wang B, Tontonoz P. Liver X receptors in lipid signalling and membrane homeostasis. Nat Rev Endocrinol. 2018;14:452–63.
Nelson ER, Wardell SE, Jasper JS, Park S, Suchindran S, Howe MK, et al. 27-Hydroxycholesterol links hypercholesterolemia and breast cancer pathophysiology. Science. 2013;342:1094–8.
McDonnell DP, Park S, Goulet MT, Jasper J, Wardell SE, Chang CY, et al. Obesity, cholesterol metabolism, and breast cancer pathogenesis. Cancer Res. 2014;74:4976–82.
Furuhashi M, Tuncman G, Gorgun CZ, Makowski L, Atsumi G, Vaillancourt E, et al. Treatment of diabetes and atherosclerosis by inhibiting fatty-acid-binding protein aP2. Nature. 2007;447:959–65.
Hao J, Yan F, Zhang Y, Triplett A, Schultz DA, Sun Y, et al. Expression of Adipocyte/Macrophage Fatty Acid-Binding Protein in Tumor-Associated Macrophages Promotes Breast Cancer Progression. Cancer Res. 2018;78:2343–55.
Aibar S, Gonzalez-Blas CB, Moerman T, Huynh-Thu VA, Imrichova H, Hulselmans G, et al. SCENIC: single-cell regulatory network inference and clustering. Nat Methods. 2017;14:1083–6.
Van de Sande B, Flerin C, Davie K, De Waegeneer M, Hulselmans G, Aibar S, et al. A scalable SCENIC workflow for single-cell gene regulatory network analysis. Nat Protoc. 2020;15:2247–76.
Patel AP, Tirosh I, Trombetta JJ, Shalek AK, Gillespie SM, Wakimoto H, et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science. 2014;344:1396–401.
Acknowledgements
We thank the Hangzhou Lianchuan Biotechnology Co., Ltd. for its guidance on single cell sequencing, and the Genesky Biotechnologies Inc., Shanghai (China) for its guidance on data analysis. We thank the National Science Foundation of China (Grant No. 82473021), the Natural Science Foundation of Shandong Province (Grant No. ZR2022MH084), the Qingdao Science and Technology Plan Project (Grant No. 23-2-8-smjk-13-nsh) and the Flexible Talent guidance Project of Qilu Hospital (Qingdao) of Shandong University (Grant No. QDKY2021RX02) for funding this research.
Author information
Authors and Affiliations
Contributions
GZ, XZ, KD, and HG conceptualized and designed the experiments. GZ, LM, KD, SS, and HG performed the scRNA-seq experiments and data analyses. LM, XZ, and TJ performed tissue sample collection, immunofluorescence staining, and data analysis. XZ, and TJ performed cell-related experiments. GZ, and ZL performed clinical data analysis. GZ, and HG wrorte the manuscript, and all authors edited and proofed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
All clinical samples and animal procedures were in compliance with the relevant guidelines and regulationss and approved by the Animal Care and Use Committee, Qilu Hospital (Qingdao), Shandong University (KYLL-KS-2020054).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhao, G., Zhang, X., Meng, L. et al. Single-cell RNA-sequencing reveals a unique landscape of the tumor microenvironment in obesity-associated breast cancer. Oncogene (2024). https://doi.org/10.1038/s41388-024-03161-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41388-024-03161-7
- Springer Nature Limited