Abstract
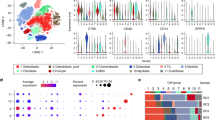
Undifferentiated pleomorphic sarcoma (UPS) is a highly aggressive malignant soft tissue tumor with a poor prognosis; however, the identity and heterogeneity of tumor populations remain elusive. Here, eight major cell clusters were identified through the RNA sequencing of 79,569 individual cells of UPS. UPS originates from mesenchymal stem cells (MSCs) and features undifferentiated subclusters. UPS subclusters were predicted to exist in two bulk RNA datasets, and had a prognostic value in The Cancer Genome Atlas (TCGA) dataset. The functional heterogeneity of malignant UPS cells and the immune microenvironment were characterized. Additionally, the fused cells were innovatively detected by expressing both monocyte/macrophage markers and other subcluster-associated genes. Based on the ligand–receptor interaction analysis, cellular interactions with epidermal growth factor receptor (EGFR) and vascular endothelial growth factor receptor (VEGFR) were abundant. Furthermore, 73% of patients with UPS (48/66) showed positive EGFR expression, which was associated with a poor prognosis. EGFR blockade with cetuximab inhibited tumor growth in a patient-derived xenograft model. Our transcriptomic studies delineate the landscape of UPS intratumor heterogeneity and serve as a foundational resource for further discovery and therapeutic exploration.








Similar content being viewed by others
Data availability
Single-cell sequencing data of human UPS specimens has been deposited at Genome Sequence Archive (GSA) with the identifier: HRA004389. Bulk expression data from complex genetics sarcomas including UPS was downloaded from GSE:21050. TCGA-SARC RNA-seq and survival data were downloaded from the Genomic Data Commons (GDC) (https://portal.gdc.cancer.gov/).
Code availability
Codes used to conduct analyses and generate results of this article is deposited at https://github.com/luyifei111/scUPS.
References
Fletcher C, Bridge JA, Hogendoorn PCW, Mertens F. WHO classification of tumours of soft tissue and bone: WHO classification of tumours, 5. Geneve, Switzerland: World Health Organization; 2013.
Robles-Tenorio A, Solis-Ledesma G. Undifferentiated pleomorphic sarcoma. In: StatPearls. Treasure Island, FL: StatPearls Publishing; 2022.
Canter RJ, Beal S, Borys D, Martinez SR, Bold RJ, Robbins AS. Interaction of histologic subtype and histologic grade in predicting survival for soft-tissue sarcomas. J Am Coll Surg. 2010;210:191–8.e192.
Zhang Y, Wang D, Peng M, Tang L, Ouyang J, Xiong F, et al. Single-cell RNA sequencing in cancer research. J Exp Clin Cancer Res CR. 2021;40:81.
Steele CD, Tarabichi M, Oukrif D, Webster AP, Ye H, Fittall M, et al. Undifferentiated sarcomas develop through distinct evolutionary pathways. Cancer Cell. 2019;35:441–56.e448.
Cancer Genome Atlas Research Network. Electronic address edsc, Cancer Genome Atlas Research N. Comprehensive and integrated genomic characterization of adult soft tissue sarcomas. Cell. 2017;171:950–65.e928.
Gao R, Bai S, Henderson YC, Lin Y, Schalck A, Yan Y, et al. Delineating copy number and clonal substructure in human tumors from single-cell transcriptomes. Nat Biotechnol. 2021;39:599–608.
Noutsias M, Rohde M, Göldner K, Block A, Blunert K, Hemaidan L, et al. Expression of functional T-cell markers and T-cell receptor Vbeta repertoire in endomyocardial biopsies from patients presenting with acute myocarditis and dilated cardiomyopathy. Eur J Heart Fail. 2011;13:611–8.
Ren Q, Ren L, Ren C, Liu X, Dong C, Zhang X. Platelet endothelial cell adhesion molecule-1 (PECAM1) plays a critical role in the maintenance of human vascular endothelial barrier function. Cell Biochem Funct. 2015;33:560–5.
Shen J, Shrestha S, Yen Y-H, Scott MA, Soo C, Ting K, et al. The pericyte antigen RGS5 in perivascular soft tissue tumors. Hum Pathol. 2016;47:121–31.
Maaninka K, Lappalainen J, Kovanen PT. Human mast cells arise from a common circulating progenitor. J Allergy Clin Immunol. 2013;132:463–9.
Cheng S, Li Z, Gao R, Xing B, Gao Y, Yang Y et al. A pan-cancer single-cell transcriptional atlas of tumor infiltrating myeloid cells. Cell. 2021;184:792–809.
McGinnis CS, Murrow LM, Gartner ZJ. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 2019;8:329–37.
Lv F-J, Tuan RS, Cheung KMC, Leung VYL. Concise review: the surface markers and identity of human mesenchymal stem cells. Stem Cells. 2014;32:1408–19.
Wu Y-H, Huang Y-F, Chang T-H, Chen C-C, Wu P-Y, Huang S-C, et al. COL11A1 activates cancer-associated fibroblasts by modulating TGF-β3 through the NF-κB/IGFBP2 axis in ovarian cancer cells. Oncogene. 2021;40:4503–19.
Zhang L-Z, Huang L-Y, Huang A-L, Liu J-X, Yang F. CRIP1 promotes cell migration, invasion and epithelial-mesenchymal transition of cervical cancer by activating the Wnt/β‑catenin signaling pathway. Life Sci. 2018;207:420–7.
Ou L, Fang L, Tang H, Qiao H, Zhang X, Wang Z. Dickkopf Wnt signaling pathway inhibitor 1 regulates the differentiation of mouse embryonic stem cells in vitro and in vivo. Mol Med Rep. 2016;13:720–30.
Traustadóttir GÁ, Lagoni LV, Ankerstjerne LBS, Bisgaard HC, Jensen CH, Andersen DC. The imprinted gene Delta like non-canonical Notch ligand 1 (Dlk1) is conserved in mammals, and serves a growth modulatory role during tissue development and regeneration through Notch dependent and independent mechanisms. Cytokine Growth Factor Rev. 2019;46:17–27.
Pittaway JFH, Lipsos C, Mariniello K, Guasti L. The role of delta-like non-canonical Notch ligand 1 (DLK1) in cancer. Endocr Relat Cancer. 2021;28:R271–287.
Suda T, Yamashita T, Sunagozaka H, Okada H, Nio K, Sakai Y et al. Dickkopf-1 promotes angiogenesis and is a biomarker for hepatic stem cell-like hepatocellular carcinoma. Int J Mol Sci. 2022;23:2801.
Matushansky I, Hernando E, Socci ND, Mills JE, Matos TA, Edgar MA, et al. Derivation of sarcomas from mesenchymal stem cells via inactivation of the Wnt pathway. J Clin Invest. 2007;117:3248–57.
Cao J, Spielmann M, Qiu X, Huang X, Ibrahim DM, Hill AJ, et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature. 2019;566:496–502.
Zhu K, Cai L, Cui C, de Los Toyos JR, Anastassiou D. Single-cell analysis reveals the pan-cancer invasiveness-associated transition of adipose-derived stromal cells into COL11A1-expressing cancer-associated fibroblasts. PLoS Comput Biol. 2021;17:e1009228.
Adachi O, Sugii H, Itoyama T, Fujino S, Kaneko H, Tomokiyo A et al. Decorin promotes osteoblastic differentiation of human periodontal ligament stem cells. Molecules. 2022;27:8224.
Song N-J, Kim S, Jang B-H, Chang S-H, Yun UJ, Park K-M, et al. Small molecule-induced complement factor D (adipsin) promotes lipid accumulation and adipocyte differentiation. PLoS ONE. 2016;11:e0162228.
Gulati GS, Sikandar SS, Wesche DJ, Manjunath A, Bharadwaj A, Berger MJ, et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science. 2020;367:405–11.
Keung EZ, Burgess M, Salazar R, Parra ER, Rodrigues-Canales J, Bolejack V, et al. Correlative analyses of the SARC028 trial reveal an association between sarcoma-associated immune infiltrate and response to pembrolizumab. Clin Cancer Res. 2020;26:1258–66.
Ponzetta A, Carriero R, Carnevale S, Barbagallo M, Molgora M, Perucchini C, et al. Neutrophils driving unconventional T cells mediate resistance against murine sarcomas and selected human tumors. Cell. 2019;178:346–60.e24.
Wisdom AJ, Mowery YM, Hong CS, Himes JE, Nabet BY, Qin X, et al. Single cell analysis reveals distinct immune landscapes in transplant and primary sarcomas that determine response or resistance to immunotherapy. Nat Commun. 2020;11:6410.
Barata JT, Durum SK, Seddon B. Flip the coin: IL-7 and IL-7R in health and disease. Nat Immunol. 2019;20:1584–93.
Tessaro FHG, Ko EY, De Simone M, Piras R, Broz MT, Goodridge HS, et al. Single-cell RNA-seq of a soft-tissue sarcoma model reveals the critical role of tumor-expressed MIF in shaping macrophage heterogeneity. Cell Rep. 2022;39:110977.
Lartigue L, Merle C, Lagarde P, Delespaul L, Lesluyes T, Le Guellec S, et al. Genome remodeling upon mesenchymal tumor cell fusion contributes to tumor progression and metastatic spread. Oncogene. 2020;39:4198–211.
Delespaul L, Gélabert C, Lesluyes T, Le Guellec S, Pérot G, Leroy L, et al. Cell-cell fusion of mesenchymal cells with distinct differentiations triggers genomic and transcriptomic remodelling toward tumour aggressiveness. Sci Rep. 2020;10:21634.
Delespaul L, Merle C, Lesluyes T, Lagarde P, Le Guellec S, Pérot G, et al. Fusion-mediated chromosomal instability promotes aneuploidy patterns that resemble human tumors. Oncogene. 2019;38:6083–94.
Zheng S, Liu Q, Liu T, Yang L, Zhang Q, Shen T, et al. NME4 modulates PD-L1 expression via the STAT3 signaling pathway in squamous cell carcinoma. Biochem Biophys Res Commun. 2020;526:29–34.
Zheng S, Liu T, Liu Q, Yang L, Zhang Q, Han X, et al. Widely targeted metabolomic analyses unveil the metabolic variations after stable knock-down of NME4 in esophageal squamous cell carcinoma cells. Mol Cell Biochem. 2020;471:81–9.
Gast CE, Silk AD, Zarour L, Riegler L, Burkhart JG, Gustafson KT, et al. Cell fusion potentiates tumor heterogeneity and reveals circulating hybrid cells that correlate with stage and survival. Sci Adv. 2018;4:eaat7828.
Kloc M, Subuddhi A, Uosef A, Kubiak JZ, Ghobrial RM. Monocyte-macrophage lineage cell fusion. Int J Mol Sci. 2022;23:6553.
Zhang Y, Du W, Chen Z, Xiang C. Upregulation of PD-L1 by SPP1 mediates macrophage polarization and facilitates immune escape in lung adenocarcinoma. Exp Cell Res. 2017;359:449–57.
Zhang L, Li Z, Skrzypczynska KM, Fang Q, Zhang W, O’Brien SA et al. Single-cell analyses inform mechanisms of myeloid-targeted therapies in colon cancer. Cell. 2020;181:442–59.e29.
Zawada AM, Rogacev KS, Rotter B, Winter P, Marell R-R, Fliser D, et al. SuperSAGE evidence for CD14++CD16+ monocytes as a third monocyte subset. Blood. 2011;118:e50–61.
Wang Z, Dai Z, Zheng L, Xu B, Zhang H, Fan F, et al. Ferroptosis activation scoring model assists in chemotherapeutic agents’ selection and mediates cross-talk with immunocytes in malignant glioblastoma. Front Immunol. 2021;12:747408.
Liu X, Xu J, Li F, Liao Z, Ren Z, Zhu L, et al. Efficacy and safety of the VEGFR2 inhibitor Apatinib for metastatic soft tissue sarcoma: Chinese cohort data from NCT03121846. Biomed Pharmacother. 2020;122:109587.
Kanehisa M, Goto S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 2000;28:27–30.
Osum M, Kalkan R. Cancer stem cells and their therapeutic usage. Adv Exp Med Biol. 2023;1436:69–85.
Sieler M, Weiler J, Dittmar T. Cell-cell fusion and the roads to novel properties of tumor hybrid cells. Cells. 2021;10:1465.
Laberge GS, Duvall E, Haedicke K, Pawelek J. Leukocyte-cancer cell fusion-genesis of a deadly journey. Cells. 2019;8:170.
Ray-Coquard I, Le Cesne A, Whelan JS, Schoffski P, Bui BN, Verweij J, et al. A phase II study of gefitinib for patients with advanced HER-1 expressing synovial sarcoma refractory to doxorubicin-containing regimens. Oncologist. 2008;13:467–73.
Cheng Y, Shen Z, Gao Y, Chen F, Xu H, Mo Q, et al. Phase transition and remodeling complex assembly are important for SS18-SSX oncogenic activity in synovial sarcomas. Nat Commun. 2022;13:2724.
Xie X, Ghadimi MPH, Young ED, Belousov R, Zhu Q-S, Liu J, et al. Combining EGFR and mTOR blockade for the treatment of epithelioid sarcoma. Clin Cancer Res. 2011;17:5901–12.
Dong R, Yang R, Zhan Y, Lai H-D, Ye C-J, Yao X-Y, et al. Single-cell characterization of malignant phenotypes and developmental trajectories of adrenal neuroblastoma. Cancer Cell. 2020;38:716–33.e6.
Tirosh I, Venteicher AS, Hebert C, Escalante LE, Patel AP, Yizhak K, et al. Single-cell RNA-seq supports a developmental hierarchy in human oligodendroglioma. Nature. 2016;539:309–13.
Satija R, Farrell JA, Gennert D, Schier AF, Regev A. Spatial reconstruction of single-cell gene expression data. Nat Biotechnol. 2015;33:495–502.
Wolock SL, Lopez R, Klein AM. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 2019;8:281–91.e9.
Aran D, Looney AP, Liu L, Wu E, Fong V, Hsu A, et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat Immunol. 2019;20:163–72.
Korsunsky I, Millard N, Fan J, Slowikowski K, Zhang F, Wei K, et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat Methods. 2019;16:1289–96.
Bergen V, Lange M, Peidli S, Wolf FA, Theis FJ. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat Biotechnol. 2020;38:1408–14.
Colaprico A, Silva TC, Olsen C, Garofano L, Cava C, Garolini D, et al. TCGAbiolinks: an R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res. 2016;44:e71.
Davis S, Meltzer PS. GEOquery: a bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics. 2007;23:1846–7.
Newman AM, Steen CB, Liu CL, Gentles AJ, Chaudhuri AA, Scherer F, et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat Biotechnol. 2019;37:773–82.
Hänzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 2013;14:7.
Yu G, Wang L-G, Han Y, He Q-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16:284–7.
Fleming SJ, Chaffin MD, Arduini A, Akkad A-D, Banks E, Marioni JC, et al. Unsupervised removal of systematic background noise from droplet-based single-cell experiments using CellBender. Nat Methods. 2023;20:1323–35.
Chuah BY, Putti T, Salto-Tellez M, Charlton A, Iau P, Buhari SA, et al. Serial changes in the expression of breast cancer-related proteins in response to neoadjuvant chemotherapy. Ann Oncol. 2011;22:1748–54.
Tan Z, Gao L, Wang Y, Yin H, Xi Y, Wu X, et al. PRSS contributes to cetuximab resistance in colorectal cancer. Sci Adv. 2020;6:eaax5576.
Acknowledgements
We would like to appreciate BioRender.com, with which some of the icons and elements of diagrams were created. This study was supported by grants from the Genertec Guozhong Healthcare (Grant No. GZKJ-KJXX-QTHT-20220016) (Y.C.), Industry-University Research Innovation Foundation of Science and Technology Development Center of the Ministry of Education (Grant No. 2021JH013) (Y.C.), National Natural Science Foundation of China (Grant No.82373385) (Y.C.).
Author information
Authors and Affiliations
Contributions
Yifei Lu: Methodology, Software, Validation, Formal analysis, Investigation, Data Curation, Writing - Original Draft, Visualization. Deqian Chen: Methodology, Software, Data Curation, Visualization. Bingnan Wang: Validation, Investigation. Wenjun Chai: Methodology, Visualization. Mingxia Yan: Validation, Visualization. Yong Chen: Conceptualization, Funding acquisition. Yong Zhan: Validation, Data Curation. Ran Yang: Data Curation. Enqing Zhou: Data Curation. Shuyang Dai: Data Curation. Yi Li: Data Curation. Rui Dong: Conceptualization, Methodology, Investigation, Resources, Writing - Review & Editing, Supervision, Project administration Biqiang Zheng: Conceptualization, Methodology, Investigation, Resources, Writing - Review & Editing, Supervision, Project administration, Funding acquisition.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lu, Y., Chen, D., Wang, B. et al. Single-cell landscape of undifferentiated pleomorphic sarcoma. Oncogene 43, 1353–1368 (2024). https://doi.org/10.1038/s41388-024-03001-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-024-03001-8
- Springer Nature Limited