Abstract
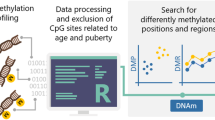
The general psychopathology factor (GPF) has been proposed as a way to capture variance shared between psychiatric symptoms. Despite a growing body of evidence showing both genetic and environmental influences on GPF, the biological mechanisms underlying these influences remain unclear. In the current study, we conducted epigenome-wide meta-analyses to identify both probe- and region-level associations of DNA methylation (DNAm) with school-age general psychopathology in six cohorts from the Pregnancy And Childhood Epigenetics (PACE) Consortium. DNAm was examined both at birth (cord blood; prospective analysis) and during school-age (peripheral whole blood; cross-sectional analysis) in total samples of N = 2178 and N = 2190, respectively. At school-age, we identified one probe (cg11945228) located in the Bromodomain-containing protein 2 gene (BRD2) that negatively associated with GPF (p = 8.58 × 10–8). We also identified a significant differentially methylated region (DMR) at school-age (p = 1.63 × 10–8), implicating the SHC Adaptor Protein 4 (SHC4) gene and the EP300-interacting inhibitor of differentiation 1 (EID1) gene that have been previously implicated in multiple types of psychiatric disorders in adulthood, including obsessive compulsive disorder, schizophrenia, and major depressive disorder. In contrast, no prospective associations were identified with DNAm at birth. Taken together, results of this study revealed some evidence of an association between DNAm at school-age and GPF. Future research with larger samples is needed to further assess DNAm variation associated with GPF.


Similar content being viewed by others
Data availability
Site-level meta-analytical results will be made publicly available (Supplementary data file) upon acceptance for publication. For access to cohort-level data, requests can be sent directly to individual studies.
Code availability
Analytical codes can be requested from authors.
References
Angold A, Costello EJ, Erkanli A. Comorbidity. J Child Psychol Psychiatry. 1999;40:57–87.
Kessler RC, Chiu WT, Demler O, Merikangas KR, Walters EE. Prevalence, severity, and comorbidity of 12-month DSM-IV disorders in the National Comorbidity Survey Replication. Arch Gen Psychiatry. 2005;62:617–27.
Cuffe SP, Visser SN, Holbrook JR, Danielson ML, Geryk LL, Wolraich ML, et al. ADHD and Psychiatric Comorbidity: Functional Outcomes in a School-Based Sample of Children. J Atten Disord. 2020;24:1345–54.
Roy-Byrne PP, Stang P, Wittchen HU, Ustun B, Walters EE, Kessler RC. Lifetime panic-depression comorbidity in the National Comorbidity Survey. Association with symptoms, impairment, course and help-seeking. Br J Psychiatry. J Ment Sci. 2000;176:229–35.
Lahey BB, Rathouz PJ, Keenan K, Stepp SD, Loeber R, Hipwell AE. Criterion validity of the general factor of psychopathology in a prospective study of girls. J Child Psychol Psychiatry. 2015;56:415–22.
Rijlaarsdam J, Cecil CAM, Buil JM, van Lier PAC, Barker ED. Exposure to Bullying and General Psychopathology: A Prospective, Longitudinal Study. Res Child Adolesc Psychopathol. 2021;49:727–36.
Neumann A, Pappa I, Lahey BB, Verhulst FC, Medina-Gomez C, Jaddoe VW, et al. Single Nucleotide Polymorphism Heritability of a General Psychopathology Factor in Children. J Am Acad Child Adolesc Psychiatry. 2016;55:1038–.e4.
Caspi A, Moffitt TE. All for One and One for All: Mental Disorders in One Dimension. Am J Psychiatry. 2018;175:831–44.
Pettersson E, Lahey BB, Larsson H, Lichtenstein P. Criterion Validity and Utility of the General Factor of Psychopathology in Childhood: Predictive Associations With Independently Measured Severe Adverse Mental Health Outcomes in Adolescence. J Am Acad Child Adolesc Psychiatry. 2018;57:372–83.
Sallis H, Szekely E, Neumann A, Jolicoeur-Martineau A, van IJzendoorn M, Hillegers M, et al. General psychopathology, internalising and externalising in children and functional outcomes in late adolescence. J Child Psychol Psychiatry. 2019;60:1183–90.
Brikell I, Larsson H, Lu Y, Pettersson E, Chen Q, Kuja-Halkola R, et al. The contribution of common genetic risk variants for ADHD to a general factor of childhood psychopathology. Mol Psychiatry. 2020;25:1809–21.
Riglin L, Thapar AK, Leppert B, Martin J, Richards A, Anney R, et al. Using Genetics to Examine a General Liability to Childhood Psychopathology. Behav Genet. 2020;50:213–20.
Brodbeck J, Fassbinder E, Schweiger U, Fehr A, Späth C, Klein JP. Differential associations between patterns of child maltreatment and comorbidity in adult depressed patients. J Affect Disord. 2018;230:34–41.
Caspi A, Houts RM, Belsky DW, Goldman-Mellor SJ, Harrington H, Israel S, et al. The p Factor: One General Psychopathology Factor in the Structure of Psychiatric Disorders? Clin Psychol Sci J Assoc. Psychol Sci. 2014;2:119–37.
Campbell M, Jahanshad N, Mufford M, Choi KW, Lee P, Ramesar R, et al. Overlap in genetic risk for cross-disorder vulnerability to mental disorders and genetic risk for altered subcortical brain volumes. J Affect Disord. 2021;282:740–56.
Cross-Disorder Group of the Psychiatric Genomics Consortium. Electronic address: plee0@mgh.harvard.edu, Cross-Disorder Group of the Psychiatric Genomics Consortium. Genomic Relationships, Novel Loci, and Pleiotropic Mechanisms across Eight Psychiatric Disorders. Cell. 2019;179:1469–.e11.
Teschendorff AE, Relton CL. Statistical and integrative system-level analysis of DNA methylation data. Nat Rev Genet. 2018;19:129–47.
Meaney MJ. Epigenetics and the biological definition of gene x environment interactions. Child Dev. 2010;81:41–79.
Cecil CAM, Walton E, Jaffee SR, O’Connor T, Maughan B, Relton CL, et al. Neonatal DNA methylation and early-onset conduct problems: A genome-wide, prospective study. Dev Psychopathol. 2018;30:383–97.
Neumann A, Walton E, Alemany S, Cecil C, González JR, Jima DD, et al. Association between DNA methylation and ADHD symptoms from birth to school age: a prospective meta-analysis. Transl Psychiatry. 2020;10:398.
Hannon E, Dempster E, Viana J, Burrage J, Smith AR, Macdonald R, et al. An integrated genetic-epigenetic analysis of schizophrenia: evidence for co-localization of genetic associations and differential DNA methylation. Genome Biol. 2016;17:176.
Zhu Y, Strachan E, Fowler E, Bacus T, Roy-Byrne P, Zhao J. Genome-wide profiling of DNA methylome and transcriptome in peripheral blood monocytes for major depression: A Monozygotic Discordant Twin Study. Transl Psychiatry. 2019;9:215.
Rijlaarsdam J, Barker ED, Caserini C, Koopman-Verhoeff ME, Mulder RH, Felix JF, et al. Genome-wide DNA methylation patterns associated with general psychopathology in children. J Psychiatr Res. 2021;140:214–20.
Zhou W, Laird PW, Shen H. Comprehensive characterization, annotation and innovative use of Infinium DNA methylation BeadChip probes. Nucl Acids Res. 2017;45:e22.
Solomon O, MacIsaac J, Quach H, Tindula G, Kobor MS, Huen K, et al. Comparison of DNA methylation measured by Illumina 450K and EPIC BeadChips in blood of newborns and 14-year-old children. Epigenetics. 2018;13:655–64.
Patalay P, Fonagy P, Deighton J, Belsky J, Vostanis P, Wolpert M. A general psychopathology factor in early adolescence. Br J Psychiatry J Ment Sci. 2015;207:15–22.
Gervin K, Salas LA, Bakulski KM, van Zelm MC, Koestler DC, Wiencke JK, et al. Systematic evaluation and validation of reference and library selection methods for deconvolution of cord blood DNA methylation data. Clin Epigenetics. 2019;11:125.
Houseman EA, Accomando WP, Koestler DC, Christensen BC, Marsit CJ, Nelson HH, et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinforma. 2012;13:86.
Salas LA, Koestler DC, Butler RA, Hansen HM, Wiencke JK, Kelsey KT, et al. An optimized library for reference-based deconvolution of whole-blood biospecimens assayed using the Illumina HumanMethylationEPIC BeadArray. Genome Biol. 2018;19:64.
Rosseel Y. lavaan: An R Package for Structural Equation Modeling. J Stat Softw. 2012;48:1–36.
Willer CJ, Li Y, Abecasis GR. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinforma Oxf Engl. 2010;26:2190–1.
Min JL, Hemani G, Davey Smith G, Relton C, Suderman M. Meffil: efficient normalization and analysis of very large DNA methylation datasets. Bioinforma Oxf Engl. 2018;34:3983–9.
Suderman M, Staley JR, French R, Arathimos R, Simpkin A, Tilling K. dmrff: identifying differentially methylated regions efficiently with power and control. bioRxiv. 2018. https://www.biorxiv.org/content/biorxiv/early/2018/12/31/508556.full.pdf:508556.
Battram T, Yousefi P, Crawford G, Prince C, Sheikhali Babaei M, Sharp G, et al. The EWAS Catalog: a database of epigenome-wide association studies. Wellcome Open Res. 2022;7:41.
Li M, Zou D, Li Z, Gao R, Sang J, Zhang Y, et al. EWAS Atlas: a curated knowledgebase of epigenome-wide association studies. Nucleic Acids Res. 2019;47:D983–D988.
Hannon E, Knox O, Sugden K, Burrage J, Wong CCY, Belsky DW, et al. Characterizing genetic and environmental influences on variable DNA methylation using monozygotic and dizygotic twins. PLoS Genet. 2018;14:e1007544.
Hannon E, Lunnon K, Schalkwyk L, Mill J. Interindividual methylomic variation across blood, cortex, and cerebellum: implications for epigenetic studies of neurological and neuropsychiatric phenotypes. Epigenetics 2015;10:1024–32.
Edgar RD, Jones MJ, Meaney MJ, Turecki G, Kobor MS. BECon: a tool for interpreting DNA methylation findings from blood in the context of brain. Transl Psychiatry. 2017;7:e1187.
Braun P, Han S, Nagahama Y, Gaul L, Heinzman J, Hing B, et al. 28 - IMAGE-CpG: DEVELOPMENT OF A WEB-BASED SEARCH TOOL FOR GENOME-WIDE DNA METHYLATION CORRELATION BETWEEN LIVE HUMAN BRAIN AND PERIPHERAL TISSUES WITHIN INDIVIDUALS. Eur Neuropsychopharmacol. 2019;29:S796.
Phipson B, Maksimovic J, Oshlack A. missMethyl: an R package for analyzing data from Illumina’s HumanMethylation450 platform. Bioinforma Oxf Engl. 2016;32:286–8.
isglobal-brge/EASIER: EwAS: quality control, meta-analysIs and EnRichment version 0.1.2.8 from GitHub. https://rdrr.io/github/isglobal-brge/EASIER/. Accessed May 2022.
Lawrence M, Huber W, Pagès H, Aboyoun P, Carlson M, Gentleman R, et al. Software for computing and annotating genomic ranges. PLoS Comput Biol. 2013;9:e1003118.
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature. 2014;511:421–7.
Nagel M, Jansen PR, Stringer S, Watanabe K, de Leeuw CA, Bryois J, et al. Meta-analysis of genome-wide association studies for neuroticism in 449,484 individuals identifies novel genetic loci and pathways. Nat Genet. 2018;50:920–7.
Demontis D, Walters RK, Martin J, Mattheisen M, Als TD, Agerbo E, et al. Discovery of the first genome-wide significant risk loci for attention deficit/hyperactivity disorder. Nat Genet. 2019;51:63–75.
Levey DF, Gelernter J, Polimanti R, Zhou H, Cheng Z, Aslan M, et al. Reproducible Genetic Risk Loci for Anxiety: Results From ∼200,000 Participants in the Million Veteran Program. Am J Psychiatry. 2020;177:223–32.
Aragam N, Wang K-S, Pan Y. Genome-wide association analysis of gender differences in major depressive disorder in the Netherlands NESDA and NTR population-based samples. J Affect Disord. 2011;133:516–21.
Howard DM, Hall LS, Hafferty JD, Zeng Y, Adams MJ, Clarke T-K, et al. Genome-wide haplotype-based association analysis of major depressive disorder in Generation Scotland and UK Biobank. Transl Psychiatry. 2017;7:1263.
Budde M, Friedrichs S, Alliey-Rodriguez N, Ament S, Badner JA, Berrettini WH, et al. Efficient region-based test strategy uncovers genetic risk factors for functional outcome in bipolar disorder. Eur Neuropsychopharmacol J Eur Coll Neuropsychopharmacol. 2019;29:156–70.
Blokland GAM, Grove J, Chen C-Y, Cotsapas C, Tobet S, Handa R, et al. Sex-Dependent Shared and Nonshared Genetic Architecture Across Mood and Psychotic Disorders. Biol Psychiatry. 2022;91:102–17.
Boraska V, Davis OSP, Cherkas LF, Helder SG, Harris J, Krug I, et al. Genome-wide association analysis of eating disorder-related symptoms, behaviors, and personality traits. Am J Med Genet Part B Neuropsychiatr Genet Publ Int Soc Psychiatr Genet. 2012;159B:803–11.
Goes FS, McGrath J, Avramopoulos D, Wolyniec P, Pirooznia M, Ruczinski I, et al. Genome-wide association study of schizophrenia in Ashkenazi Jews. Am J Med Genet Part B Neuropsychiatr Genet Publ Int Soc Psychiatr Genet. 2015;168:649–59.
Wang N, Wu R, Tang D, Kang R. The BET family in immunity and disease. Signal Transduct Target Ther. 2021;6:23.
Gyuris A, Donovan DJ, Seymour KA, Lovasco LA, Smilowitz NR, Halperin ALP, et al. The chromatin-targeting protein Brd2 is required for neural tube closure and embryogenesis. Biochim Biophys Acta. 2009;1789:413–21.
Garcia-Gutierrez P, Juarez-Vicente F, Wolgemuth DJ, Garcia-Dominguez M. Pleiotrophin antagonizes Brd2 during neuronal differentiation. J Cell Sci. 2014;127:2554–64.
DeMars KM, Yang C, Candelario-Jalil E. Neuroprotective effects of targeting BET proteins for degradation with dBET1 in aged mice subjected to ischemic stroke. Neurochem Int. 2019;127:94–102.
Pathak S, Miller J, Morris EC, Stewart WCL, Greenberg DA. DNA methylation of the BRD2 promoter is associated with juvenile myoclonic epilepsy in Caucasians. Epilepsia. 2018;59:1011–9.
Wockner LF, Noble EP, Lawford BR, Young RM, Morris CP, Whitehall VLJ, et al. Genome-wide DNA methylation analysis of human brain tissue from schizophrenia patients. Transl Psychiatry. 2014;4:e339.
McKinney B, Ding Y, Lewis DA, Sweet RA. DNA methylation as a putative mechanism for reduced dendritic spine density in the superior temporal gyrus of subjects with schizophrenia. Transl Psychiatry. 2017;7:e1032.
Skariah G, Seimetz J, Norsworthy M, Lannom MC, Kenny PJ, Elrakhawy M, et al. Mov10 suppresses retroelements and regulates neuronal development and function in the developing brain. BMC Biol. 2017;15:54.
Vissers LELM, Gilissen C, Veltman JA. Genetic studies in intellectual disability and related disorders. Nat Rev Genet. 2016;17:9–18.
Hanson E, Bernier R, Porche K, Jackson FI, Goin-Kochel RP, Snyder LG, et al. The cognitive and behavioral phenotype of the 16p11.2 deletion in a clinically ascertained population. Biol Psychiatry. 2015;77:785–93.
Zufferey F, Sherr EH, Beckmann ND, Hanson E, Maillard AM, Hippolyte L, et al. A 600 kb deletion syndrome at 16p11.2 leads to energy imbalance and neuropsychiatric disorders. J Med Genet. 2012;49:660–8.
Weiss LA, Shen Y, Korn JM, Arking DE, Miller DT, Fossdal R, et al. Association between microdeletion and microduplication at 16p11.2 and autism. N. Engl J Med. 2008;358:667–75.
Chang H, Li L, Li M, Xiao X. Rare and common variants at 16p11.2 are associated with schizophrenia. Schizophr Res. 2017;184:105–8.
You Y, Li W, Gong Y, Yin B, Qiang B, Yuan J, et al. ShcD interacts with TrkB via its PTB and SH2 domains and regulates BDNF-induced MAPK activation. BMB Rep. 2010;43:485–90.
Liu R, Lei JX, Luo C, Lan X, Chi L, Deng P, et al. Increased EID1 nuclear translocation impairs synaptic plasticity and memory function associated with pathogenesis of Alzheimer’s disease. Neurobiol Dis. 2012;45:902–12.
Sullivan PF, de Geus EJC, Willemsen G, James MR, Smit JH, Zandbelt T, et al. Genome-wide association for major depressive disorder: a possible role for the presynaptic protein piccolo. Mol Psychiatry. 2009;14:359–75.
Mulder RH, Neumann A, Cecil CAM, Walton E, Houtepen LC, Simpkin AJ, et al. Epigenome-wide change and variation in DNA methylation in childhood: trajectories from birth to late adolescence. Hum Mol Genet. 2021;30:119–34.
Singham T, Viding E, Schoeler T, Arseneault L, Ronald A, Cecil CM, et al. Concurrent and Longitudinal Contribution of Exposure to Bullying in Childhood to Mental Health: The Role of Vulnerability and Resilience. JAMA Psychiatry. 2017;74:1112–9.
Bale TL, Epperson CN. Sex as a Biological Variable: Who, What, When, Why, and How. Neuropsychopharmacol Publ Am Coll Neuropsychopharmacol. 2017;42:386–96.
Author information
Authors and Affiliations
Contributions
JR: analysis plan and study design, Generation R analysis, quality control of data and meta-analyses, interpretation of results, manuscript drafting; MCT: Helix and INMA analyses, quality control of data, shadow meta-analysis, functional and genomic enrichment analyses, results interpretation, manuscript drafting, revision; LS: contributed to analysis plan and study design, ALSPAC analysis, interpretation of results, and manuscript review; SaA: DCHS analysis, and manuscript review; AM: GLAKU analysis and manuscript review; AN: Generation R coordination, manuscript review; JF: study design, Generation R coordination, critical revision of the manuscript; JS: INMA coordination, provided funding, manuscript review; KBG: MoBa Helix funding, MoBa coordination; RG: KANC-Helix funding, KANC coordination; JW: BiB-Helix funding, BiB coordination; MK: Rhea-Helix data acquisition; HJZ: DCHS coordination, and manuscript review; DJS: DCHS coordination, and manuscript review; KH: GLAKU coordination, manuscript review; KR: GLAKU coordination, manuscript review; JL: GLAKU coordination, manuscript review; AH: DCHS analysis, and manuscript review; DC: contributed to analysis plan and review of drafts and manuscript; SiA: analysis plan and study design, INMA analysis, manuscript review; CC: analysis plan and study design, Generation R coordination, funding, manuscript drafting and review.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
All studies acquired approval from local ethics committees and informed consent was obtained for all participants. Full details are listed in the methods supplement.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rijlaarsdam, J., Cosin-Tomas, M., Schellhas, L. et al. DNA methylation and general psychopathology in childhood: an epigenome-wide meta-analysis from the PACE consortium. Mol Psychiatry 28, 1128–1136 (2023). https://doi.org/10.1038/s41380-022-01871-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-022-01871-6
- Springer Nature Limited