Abstract
Defects in the PEX5 gene impair the import of peroxisomal matrix proteins, leading to nonfunctional peroxisomes and other associated pathological defects such as Zellweger syndrome. Although PEX5 regulates autophagy process in a stress condition, the mechanisms controlling autophagy by PEX5 under nutrient deprivation are largely unknown. Herein, we show a novel function of PEX5 in the regulation of autophagy via Transcription Factor EB (TFEB). Under serum-starved conditions, when PEX5 is depleted, the mammalian target of rapamycin (mTORC1) inhibitor TSC2 is downregulated, which results in increased phosphorylation of the mTORC1 substrates, including 70S6K, S6K, and 4E-BP-1. mTORC1 activation further suppresses the nuclear localization of TFEB, as indicated by decreased mRNA levels of TFEB, LIPA, and LAMP1. Interestingly, peroxisomal mRNA and protein levels are also reduced by TFEB inactivation, indicating that TFEB might control peroxisome biogenesis at a transcriptional level. Conversely, pharmacological inhibition of mTOR resulting from PEX5 depletion during nutrient starvation activates TFEB by promoting nuclear localization of the protein. In addition, mTORC1 inhibition recovers the damaged-peroxisome biogenesis. These data suggest that PEX5 may be a critical regulator of lysosomal gene expression and autophagy through the mTOR-TFEB-autophagy axis under nutrient deprivation.
Similar content being viewed by others
Introduction
Peroxisome is an essential cellular organelle for performing various metabolic activities, including oxidation of very long and 3-methy branched chain fatty acids, synthesis of bile acid and plasmalogen, and degradation of hydrogen peroxide. Mutations in either a peroxisomal enzyme or genes involved in peroxisome biogenesis cause inheritable genetic disorders, such as Zellweger syndrome, infantile Refsum disease, and neonatal adrenoleukodystropy (ALD)1,2,3. In addition, it has also been suggested that peroxisomes play an important role in regulating various cellular signaling pathways. Peroxisomal Mitochondrial antiviral-signaling protein (MAVS) functions as an antiviral signaling platform to induce the interferon-independent signaling pathways4. Reactive oxygen species (ROS)-mediated signaling has been proposed to regulate autophagy at the peroxisomal membrane through the localization of tuberous sclerosis complex (TSC)5.
Peroxisomes are originally derived from the endoplasmic reticulum (ER) and uptake peroxisomal matrix proteins containing the peroxisome target sequence (PTS) from the cytosol6. During import of matrix proteins into the peroxisome, several Pex proteins participate in the process of peroxisome assembly. Especially, the PTS1 receptor protein, PEX5, plays an important role in peroxisome biogenesis7,8. It has been shown that inactivation of PEX5 results in abnormal peroxisome structure and function, similar to the features of Zellweger syndrome9,10. Zellweger syndrome, also known as the cerebrohepatorenal syndrome, causes multi-organ defects during development and leads to early death within a year of birth. The symptoms include hypotonia, central nervous system abnormalities, severe mental retardation, hepatic cirrhosis, renal cyst, and skeletal malformations2,11. It also involves accumulation of very long chain fatty acid (VLCFA), pristanic acid, phytanic acid, and bile acid intermediates and lack of ether phospholipids due to inability to assemble functional peroxisomes in the patients1,12. In addition to peroxisome biogenesis, it has been proposed that PEX5 is subjected to ubiquitination and is involved in the autophagic degradation of peroxisome, a process known as pexophagy13,14. Ubiquitinated PEX5 is recognized by the autophagic adaptor, SQSTM1/p62, which triggers pexophagy under stress conditions.
It has been well established that nutrient deprivation stimulates autophagy, a catabolic pathway by which cytoplasmic contents and subcellular organelles are delivered to the lysosome for degradation15. Cytoplasmic targets are sequestered within autophagosomes, which fuse with lysosomes to form autolysosomes for degradation, thereby maintaining cellular homeostasis. Autophagy is a tightly regulated process from the very first step of formation of a pre-initiation complex to the final step of degradation and recycling, through coordination of various regulatory components16,17. Among them, the mammalian target of rapamycin complex 1 (mTORC1) plays a critical role as a negative regulator of autophagy during nutrient depletion. The mTOR signaling pathway coordinates energy, growth signals, and nutrient abundance with cell growth and division, and responds to numerous stresses. Dysregulation of the mTORC1 signaling pathway leads to cancer, metabolic diseases, and diabetes18,19. Under nutrient-rich conditions, mTORC1 is activated and directly phosphorylates and inhibits proteins related to the autophagy process. Under nutrient-starved conditions, mTOR1 is inactivated and dissociates from the ULK complex, leading to induction of autophagy. Recently, Transcription Factor EB (TFEB), a member of the basic helix-loop-helix-zipper family of transcription factors, is known to control the expression of genes that regulate lysosomal biogenesis and autophagy20,21. mTOR-mediated phosphorylation regulates the activity and subcellular localization of TFEB under specific conditions, such as starvation or lysosomal dysfunction22.
Previous studies have suggested TSC interacts with PEX5 and localizes to the peroxisome, which in turn regulates mTOR activity and autophagy5. In this study, we provide evidence that PEX5 deficiency prevents autophagy by regulating mTOR signaling and TFEB activity during starvation. Furthermore, inhibition of mTOR activity restores the autophagic pathway that is impaired by PEX5 depletion. In addition, our data suggest that PEX5 function is associated with the lysosome, which is an important cellular organelle involved in the process of autophagy. These findings strongly indicate that peroxisome is a cellular signaling platform for autophagy under starved conditions.
Materials and methods
Antibodies
Anti-PEX5 was purchased from GeneTex (Irvine, CA, USA). Anti-catalase, anti-LAMP-1, anti-PCNA, anti-cathepsin B, and anti-cathepsin D were bought from Santa Cruz Biotechnology (Santa Cruz, CA, USA). Anti-LAMP-2 and anti-LC3 were obtained from Abcam (Cambridge, MA, USA). Anti-TFEB was purchased from MyBioSource (San Diego, CA, USA). Anti-TFEB, anti-p-p70S6K, anti-p70S6K, anti-p-S6R, anti-S6R, anti-p-4E-BP-1, anti-4E-BP-1, and anti-TSC2 were obtained from Cell Signaling Technology (Danvers, MA, USA). Anti-PMP70 was bought from Thermo Fisher Scientific (Waltham, MA, USA). Anti-ACOX1 was purchased from Proteintech (Chicago, IL, USA). Anti-DBP (HSD17B4) was bought from OriGene Technologies (Rockville, MD, USA). Anti- SQSTM1/p62 was obtained from Sigma-Aldrich (St. Louis, MO, USA).
Cell culture and drug treatments
HepG2 (human hepatocyte carcinoma cell line) and HeLa (human epithelial carcinoma cell line) cells were originally purchased from the American Type Culture Collection (ATCC, Manassa, VA, USA). HepG2 and HeLa cells were grown in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% heat-inactivated fetal bovine serum, 2 mM l-glutamine, 100 IU/mL penicillin, and 10 μg/mL streptomycin. All cells were incubated in a humidified 5% CO2 incubator at 37 °C. Cells were treated with the mTORC1 inhibitor, rapamycin (Sigma-Aldrich), for the indicated time. For starvation experiments, cells were washed two times with phosphate buffered saline (PBS, pH 7.4), one time with serum-free media, and further incubated for 24 h at 37 °C in serum-free media.
Immunofluorescence
Cells growing on coverslips were fixed in 4% paraformaldehyde (Sigma–Aldrich) at room temperature (RT) for 10 min, rinsed in PBS, incubated in 0.1% Triton X-100, and then rinsed in PBS. After incubation with 3% bovine serum albumin, cells were incubated with primary antibody (1:500 dilution) at 4 °C. Next, the plates were carefully washed with PBS and stained with Alexa Flour 488- or Alexa Fluor 568-conjugated goat anti-mouse or anti-rabbit secondary antibodies (Invitrogen/Life Technologies, Grand island, NY, USA) for 1 h at RT. The samples were then washed three times with PBS and incubated with 10 μM 4′,6-diamidino-2-phenylindole (DAPI) in PBS at RT for 30 min. After mounting the coverslip, the cells were observed under an Olympus FluoView FV1000 confocal laser scanning microscope.
siRNA and transient transfection
Cells were cultured in 60 mm culture plates until they reached ~50–60% confluence. They were then transfected with 100 nM control siRNA, PEX5 siRNA (#1 sense-CGUAUCCUGGGAUCUCUCU, antisense-AGAGAGAUCCCAGGAUACG, #2 sense-GAAUUCAUCUCUGAAGUUA, antisense-UAACUUCAGAGAGAUGAAUUC-3, Bioneer, Daejeon, Korea), or TFEB siRNA (sc38509, Santa Cruz Biotechnology) for 24 h in serum-free medium, using Lipofectamine RNAiMax transfection reagent (Invitrogen/Life Technologies), and then exposed to serum starvation for 24 h. The effect of gene silencing was determined by Q-PCR and western blot analysis. For plasmid transfection, cells were transiently transfected with GFP-LC323, pEGFP-N1, pEGFP-N1-TFEB (Addgene, # 38119), or TFEB-S142A plasmids by using Lipofectamine 3000 (Invitrogen/Life Technologies), according to the manufacturer’s instructions.
Mutagenesis
The human pEGFP-N1-TFEB mutant was generated using QuickChange lightning site-directed mutagenesis kit (Stratagene, Santa Clara, CA, USA), according to the manufacturer’s instructions. The primer sets were as follows: hTFEB S142A; forward: GGCAACGCTCCCAATGCCCCCATGGCCATGCTGCACA, reverse: TGTGCAGCATGGCCATGGGGGCATTGGGAGCACTGTTGCC.
Quantitative RT-PCR (Q-PCR)
Total RNA was extracted using TRIzol Reagent (Invitrogen/Life Technologies), and RT-PCR was performed using the reverse transcription kit (Roche, Indianapolis, IN, USA), according to the manufacturer’s instructions. Briefly, 1 µg of RNA was used to construct the template cDNA for Q-PCR with LightCycler using FastStart DNA Master SYBR Green I (Roche, Indianapolis). The primer sets were as follows: hPEX5, 5′-GTGCTCTACCTGGCT-3′ and 5′-CTGAAGTGTTCTGCTCCTAC-3′; hPMP70, 5′-GCAGACAGTCCACTCAGTCTTC-3′ and 5′-CCATTGAAAACCGAAACAAAA-3′; hCatalase, 5′-CTCAGGTGCGGGCATTCT-3′ and 5′-CAATGTTCTCACACAGACGTT TCC-3′; hACOX-1, 5′-CATTCACCGTGTTCATATGCTTTC-3′ and 5′-ATGCCTTCAGGTGTATGTGATCTG-3′; hDBP, 5′- ACCTGGTCTCTCAAGCAGGA-3′ and 5′-CTAACGCTCCTCTTT CTGCAA-3′; hNUDT7, 5′-CACCGTCCGGCTAGAGAA-3′ and 5′-GGGTCACGCTTACCTCCA G-3′; hLAMP-1, 5′-GTGGGTCCAGGCTTTCAA-3′ and 5′-AGCATGCTGTTCTCGTCCA-3′; hTFEB, 5′-CGGCAGTGCCTGGTACAT-3′ and 5′-CTGCATGCGCAACCCTAT-3′; hLIPA, 5′-CCCTGAGAGCTGGCACTG-3′ and 5′-AAGAACCGCATTTTCATTACTG-3′; and hPPARα, 5′-GTGCCAGCAGATTCAGTGTC-3′ and 5′-GCCGAGCTCCAAAGCTACTC-3′; hPGC-1α, 5′-CTAGACAGTCTCCCTGTGGATGAA-3′ and 5′-TGTCAGTGGTCACGTCTCCAT-3′; and internal control 36B4, 5′-CACTGGTCTAGGACCCGAGAAG-3′ and 5′-GGTGCCTCTGGAGATTTTCG-3′.
Western blot analysis
Cells were lysed using Pro-Prep protein extraction solution (Intron Biotechnology, Seoul, Korea). The samples were centrifuged at 16,000 × g for 20 min at 4 °C, and the supernatant was collected to determine protein concentration via the Bradford method. Aliquots of 30 µg of protein were subjected to 10% sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis for 2 h at 100 V. The separated proteins were transferred from the SDS-polyacrylamide gels to Hybond-P+ polyvinylidene difluoride membranes (Amersham Bioscience, Buckinghamshire, UK). The membrane was blocked with 5% nonfat milk in Tris-buffered saline containing 0.05% Tween 20 for 2 h at RT and then incubated with primary antibodies as indicated. Signals from the bound antibodies were detected by the horseradish peroxidase-conjugated secondary antibodies and an ECL western blotting detection reagent (Bionote, Gyeonggi-do, Korea).
Preparation of cytosolic and nuclear fractions
Cells were washed twice with ice-cold PBS, lysed with three packed cell volumes of Buffer A (10 mM HEPES [pH 7.9], 10 mM KCl, 0.1 mM EDTA, 1 mM DTT, and protease/phosphatase inhibitors cocktail), and incubated on ice for 30 min. Cell lysates were centrifuged at 16,000 × g at 4 °C for 10 min. Supernatants were collected and nuclear pellets were resuspended in three packed cell volumes of Buffer B (20 mM HEPES [pH 7.9], 0.4 M NaCl, 1 mM EDTA, 1 mM DTT, 10% glycerol (vol/vol), and protease/phosphatase inhibitors cocktail) and incubated for 30 min on ice. After centrifugation at 16,000 × g at 4 °C for 10 min, the supernatant was aliquoted and stored at –80 °C until further use for western blot analysis24.
LysoTracker staining
Cells cultured to 60–70% confluence were subjected to different stimuli under reduced serum (2%) DMEM medium, followed by incubation with 50 nM LysoTracker Red DND-99 (Life Technologies, Grand Island, NY) for 1 h. Cells were then washed thoroughly with filtered PBS, mounted on glass slides, and viewed under Olympus FluoView FV1000 confocal laser scanning microscope.
Statistical analysis
All data are presented as the mean ± s.d. from at least three independent experiments. The P-value was determined by Student’s t-test. A P-value below 0.05 was considered statistically significant.
Results
PEX5 depletion reduces autophagy during starvation
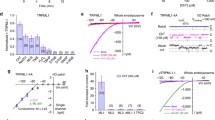
To examine the role of PEX5 in autophagy, HepG2 cells were transiently transfected with two independent PEX5 small interfering RNA (siRNA), PEX5 siRNA #1 and PEX5 siRNA #2, to rule out the off-target effect. Compared to treatment with control siRNA, treatment with two independent PEX5 siRNAs reduced the mRNA and protein expression levels of PEX5, as determined by real-time PCR (Q-PCR) and immunoblot analysis (Supplementary Figure 1). Next, the conversion of LC3-I to LC3-II, which is a marker for autophagosome formation, was measured. Under normal growth conditions, treatment with control or two PEX5 siRNAs did not affect LC3-II levels (Fig. 1a). LC3-II expression was increased in control siRNA-treated HepG2 cells grown in serum-starved medium, but treatment with two PEX5 siRNAs prevented the starvation-induced LC3-II conversion (Fig. 1a). To confirm whether the reduced LC3-II levels resulting from PEX5 depletion were caused by inhibition of autophagic flux, we measured the level of SQSTM1/p62, which is a scaffold protein that can be degraded through autophagy25. Our result revealed that SQSTM1/p62 degradation was suppressed by PEX5 knockdown under serum-free conditions (Fig. 1a). Because mTOR1 activation is well known as a critical negative regulator of the autophagy pathway, we measured the phosphorylated states of the mTOR1 substrates, which are targets for mTOR1 kinase. Starvation suppressed the phosphorylation of S6K, 70S6K, and 4E-BP-1 in cells transfected with control siRNA (Fig. 1b). In contrast, PEX5 depletion by both siRNA increased the phosphorylation of these mTOR1 substrates in starved conditions. Furthermore, to examine the autophagosome formation, GFP-LC3 puncta were measured by immunofluorescence with PEX5 siRNA #1 due to better effect than PEX5 siRNA #2. Serum starvation induced an increase in the numbers of GFP-LC3 puncta in HepG2 cells transfected with control siRNA. However, knockdown with PEX5 siRNA markedly abolished the formation of GFP-LC3 puncta in serum-starved HepG2 cells (Fig. 1c). These results indicated that PEX5 depletion prevents the activation of autophagy during starvation. To further examine whether the effect of PEX5 depletion on autophagy take places only in HepG2 cells, we performed the experiments with HeLa cells. As shown in Supplementary Figure 2a, both of PEX5 siRNAs knockdown prevent the starvation-induced autophagy by measuring LC3II and p62 protein levels in HeLa cells. Also, PEX5 knockdown increased the phosphorylation of these mTORC1 substrates in starved Hela cells in Supplementary Figure 2b. These results indicated that the effects of PEX5 knockdown on autophagy and mTOR signaling are a general regulation. A recent report showed that PEX5 interacts with TSC2 and recruits TCS1 to the peroxisomal membrane5. Since TSC2 is known to repress mTOR1 signaling by regulating Rheb GTPase26, we examined whether PEX5 affects TSC2 expression. The amount of TSC2 protein expressed during starvation was decreased in PEX5 siRNA-transfected HepG2 cells (Fig. 1d) compared to that in control siRNA-transfected cells. Serum starvation increased the expression of TSC2 mRNA in cells transfected with control siRNA. However, PEX5 depletion with siRNA significantly decreased the starvation-induced expression of TSC2 mRNA, which in turn upregulated mTOR activity (Fig. 1e). Interestingly, TSC2 protein levels were not increased as much as mRNA increases upon starvation as compared to normal condition. It is reported that E3 ubiquitin ligase, FBW5–DDB1–CUL4–ROC1 regulates both TSC2 protein stability and TSC complex turnover27. Also it is suggested that TSC1 interacts with TSC2 to stabilize TSC2 by excluding HERC1 ubiquitin ligase from the TSC2 complex28. Therefore, TSC2 protein level may not be relevant to mRNA level due to post-translational regulation. These results suggest that PEX5 depletion during starvation negatively regulates autophagy through the activation of mTOR signaling.
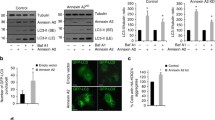
a HepG2 cells were treated with CTL siRNA (100 nM) and PEX5 siRNA #1 or #2 (100 nM) for 24 h. Then, cells were incubated in the presence or absence of serum. After 24 h, cells were harvested and immunoblotted with antibodies against PEX5, LC3I/II, p62, and β-actin. b Immunoblot analysis of p-70S6K, p-S6R, and p-4E-BP-1, and total 70S6K, S6R, and 4E-BP-1. c Cells were transfected with GFP-LC3 plasmid. After 6 h of transfection, cells were treated with CTL and PEX5 siRNAs. Visualization and quantification of GFP-LC3 puncta were performed by fluorescence microscopy after 48 h of incubation. Representative images from three independent determinations are shown (*P < 0.05). Scale bar: 10 μm. d Immunoblot analysis of TSC2 and PEX5 protein levels. e Transcript levels of TSC2 were measured by Q-PCR (**P < 0.01)
PEX5 regulates the expression of peroxisomal genes during starvation
Deficiency of PEX5 in a model for Zellweger syndrome, results in a reduction in peroxisome numbers in the hepatocytes10,29. To examine whether the peroxisome number was affected by PEX5 depletion in serum starvation, we performed immunofluorescence for peroxisome-specific markers, including PMP70 and catalase. During starvation, transfection with PEX5 siRNA markedly reduced the number of peroxisomes compared to transfection with control siRNA (Fig. 2a). We also examined the mRNA expression of peroxisomal genes, including PMP70, catalase, DBP, ACOX1, and NUDT7. Their expression was decreased by PEX5 depletion in serum starvation (Fig. 2b). Since Peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PGC1α) and peroxisome proliferator-activated receptor α (PPARα) are also involved in peroxisome biogenesis, we measured their expression levels30. The mRNA expression of PGC1α and PPARα was significantly decreased by PEX5 depletion in serum starvation (Fig. 2c). Taken together, our data confirm that PEX5 plays a pivotal role in controlling peroxisome biogenesis. Furthermore, dysfunction of peroxisome biogenesis caused by PEX5 depletion is regulated by the modulation of transcriptional activation of genes related to peroxisomes in starvation conditions.
a Cells were treated with CTL and PEX5 siRNAs (100 nM) for 24 h. Then, cells were incubated in the presence or absence of serum. After 24 h, cells were stained with antibodies against PMP70 and catalase, and analyzed by fluorescence microscopy. Scale bar: 10 μm. b Cells were treated with CTL and PEX5 siRNAs for 24 h. Then, cells were incubated in the presence or absence of serum. After 24 h of incubation, RNA was extracted and Q-PCR analysis of peroxisomal genes was performed. Data are presented as mean ± s.d. from three independent experiments (**P < 0.01, *P < 0.05). c Transcript levels of PGC-1α and PPARα were measured by Q-PCR (*P < 0.05)
PEX5 inhibits TFEB nuclear translocation in response to serum starvation
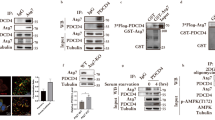
The transcription factor, TFEB, activates genes for lysosomal biogenesis and controls autophagy20,31. In addition, activation of TFEB is tightly regulated by cellular localization. In normal conditions, TFEB, as an inactive form, is mainly localized to the cytosol. However, nutrient deprivation leads to nuclear translocation of TFEB, where it induces transcriptional activation22. Moreover, TFEB has been suggested to regulate the activity and expression of PGC1α and PPARα, thereby controlling lipid metabolism in the liver32. To examine whether nuclear translocation of TFEB is affected by PEX5 depletion, we investigated the subcellular localization of TFEB in the presence or absence of serum by immunofluorescence. Following transfection with either control or PEX5 siRNA, TFEB was evenly scattered throughout the cytosol of cells cultured in normal media containing 10% serum. Starvation of cells transfected with control siRNA resulted in TFEB translocation to the nucleus, whereas its translocation was markedly suppressed by PEX5 knockdown (Fig. 3a). To confirm the location of the TFEB protein, cells were fractionated into nuclear and cytosolic fractions and then subjected to western blot analysis. TFEB localized mainly to the cytosol under normal conditions (Fig. 3b). However, under conditions of starvation, it translocated to the nucleus, as shown by a strong positive anti-TFEB band, which was absent in PEX5 siRNA-transfected cells (Fig. 3b). Consistent with the effect of PEX5 depletion on TFEB in HepG 2 cells, both of PEX5 siRNAs knockdown prevent the nuclear localization of TFEB during starvation in HeLa cells (Supplementary Figure 2c). To investigate the role of TFEB in autophagy, the conversion of LC3-I to LC3-II and the degradation of p62/SQSTM1 were measured (Fig. 3c). Transfection with both control and PEX5 siRNAs did not affect LC3-II conversion and p62/SQSTM1 degradation under normal conditions. Indicators of autophagy, including LC3-II conversion and SQSTM1/p62 degradation, were observed in control siRNA-treated cells during starvation, but not in PEX5 siRNA-treated cells. Next, to examine the link between TFEB activity and peroxisome biogenesis, the expression levels of peroxisomal proteins were measured after knockdown with TFEB siRNA. Compared with control siRNA treatment, TFEB depletion decreased the expression of peroxisomal proteins, including PMP70, DBP, and catalase, only during starvation (Fig. 3d). TFEB depletion using siRNA decreased the mRNA expression levels of peroxisomal genes, including PMP70, ACOX1, catalase, and NUDT7, during starvation, which indicates that TFEB plays an important role in peroxisome biogenesis through transcriptional regulation (Fig. 3e). These results suggest that PEX5 requires TFEB to regulate peroxisome biogenesis during starvation.
a Cells were treated with CTL and PEX5 siRNAs (100 nM) for 24 h. Then, cells were incubated in the presence or absence of serum. After 24 h, cells were stained with an antibody against TFEB and analyzed by fluorescence microscopy. Scale bar: 10 μm. b After HepG2 cells were treated as indicated, cells were subjected to nuclear and cytosol fractionation and immunoblotted with antibodies against TFEB, PCNA, and β-actin. PCNA and β-actin were used as nuclear and cytosolic markers, respectively. c, d Cells were transfected with CTL or TFEB siRNA (100 nM) for 24 h, and then incubated in the presence or absence of serum for 24 h. Protein expression of TFEB, LC3 and SQSTM1/p62 (autophagy proteins), and PMP70, DBP, and catalase (peroxisomal proteins) were measured by immunoblotting. e Q-PCR analysis of peroxisomal genes was performed (**P < 0.01, *P < 0.05)
PEX5 depletion impairs lysosomal biogenesis and function
Since PEX5 depletion resulted in decreased TFEB nuclear translocation, we hypothesized that knockdown of PEX5 might affect lysosomal biogenesis during serum starvation. A marked increase in fluorescent particles stained with LysoTracker was observed in control cells during serum starvation, but not in the PEX5 siRNA-transfected cells, indicating that lysosomal acidification required for the maturation process was largely impaired (Fig. 4a). In addition, we performed flow cytometry analysis to measure the fluorescence intensity of LysoTracker. The results showed that the red fluorescence intensity of LysoTracker was significantly decreased by PEX5 siRNA during starvation compared to that by control siRNA (Fig. 4b). To confirm the transcriptional activation of TFEB, we measured the mRNA expression of target genes, such as LIPA and LAMP-1, which are often associated with the lysosome. mRNA levels of TFEB, LIPA, and LAMP-1 were significantly decreased by treatment with PEX5 siRNA compared to treatment with control siRNA under starvation conditions (Fig. 4c). We also verified whether the expression levels of lysosomal proteins were changed by PEX5 depletion in serum starvation. Both LAMP-1 and LAMP-2 levels were found to be reduced (Fig. 4d). We further investigated the activation and maturation of the lysosomal hydrolase cathepsins B and D, which were decreased by PEX5 depletion during serum starvation (Fig. 4e). These results suggest that PEX5 depletion greatly impaired lysosomal biogenesis and functional maturation by regulating TFEB activation during serum starvation.
Cells were transfected with CTL siRNA (100 nM) and PEX5 siRNA (100 nM) for 24 h, and then incubated in the presence or absence of serum for 24 h. After incubation, cells were stained with LysoTracker Red DND-99 (50 nM) for 3 h and analyzed by confocal microscopy (a) or fluorescence-activated cell sorting and quantified (b). Data are presented as mean ± s.d. from three independent experiments (*P < 0.05). c Transcript levels of TFEB target genes were measured by Q-PCR. Values are expressed as fold increase compared to the starvation control group. Data were presented as mean ± s.d. from three independent experiments (**P < 0.01, *P < 0.05). d, e Protein expression of LAMP-1/LAMP-2 (lysosomal proteins) and cathepsins B/D (lysosomal proteases) were measured by immunoblotting
PEX5 regulates TFEB localization and autophagy via mTORC1
Nuclear localization and transcriptional activation of TFEB are dependent on mTORC1-mediated phosphorylation. In normal conditions, TFEB is phosphorylated by mTORC1 and localized to the cytosol, whereas in serum starvation it is dephosphorylated and further translocated to the nucleus to transcribe target genes21,32. To determine whether PEX5 depletion during starvation regulates TFEB activity through mTORC1, TFEB localization was examined in the presence or absence of the mTOR inhibitor, rapamycin. Consistent with previous observations, upon starvation, most of the TFEB protein was found in the nucleus in cells transfected with control siRNA, and this was dramatically suppressed in PEX5 siRNA-transfected cells. Furthermore, the decreased nuclear localization of TFEB by PEX5 depletion was significantly recovered in serum-starved cells in the presence of rapamycin (Fig. 5a). Next, we investigated the subcellular localization of TFEB by western blot analysis (Fig. 5b). Nuclear translocation of TFEB in control cells was inhibited by treatment with PEX5 siRNA upon serum starvation. However, treatment with rapamycin resulted in the appearance of an anti-TFEB reactive signal in the nuclear fraction along with faded signals in the cytosolic fraction, in PEX5-depleted and serum-starved cells. We also tested whether rapamycin restores autophagy, which is inhibited by PEX5 depletion during starvation. Consistently, depletion of PEX5 with siRNA clearly inhibited the activation of autophagy evidenced by both a faint signal of LC3-II and a recovered strong signal of SQSTM1/p62in cells treated with dimethyl sulfoxide (DMSO) during serum starvation (Fig. 5c). Inhibitory effects of PEX5 depletion on autophagy caused by serum starvation were reversed by treatment with rapamycin, as shown by the increased signal intensities of LC3-II and decreased signals of SQSTM1/p62, recovered to levels similar to those of control cells. Moreover, treatment with rapamycin resulted in decreased phosphorylation of 70S6K, S6K, and 4EBP-1 proteins in cells transfected with PEX5 siRNA (Fig. 5d). These results suggest that PEX5 depletion downregulated TFEB activity via the mTORC1 signaling pathway.
Cells transfected with CTL siRNA (100 nM) and PEX5 siRNA (100 nM) for 24 h were pretreated with DMSO or rapamycin (1 μM) for 6 h, and then incubated in the absence of serum for 24 h. (a) After HepG2 cells were treated as indicated, cells were stained with an antibody against TFEB and analyzed by fluorescence microscopy. Scale bar: 10 μm. Data are presented as mean ± SD from three independent experiments (**P < 0.01). (b) After HepG2 cells were treated as indicated, cells were subjected to nuclear and cytosol fractionation and immunoblotted with antibodies against TFEB, PCNA, and β-actin. PCNA and β-actin were used as nuclear and cytosolic markers, respectively. (c) Immunoblot analysis of LC3 and SQSTM1/p62. (d) Immunoblot analysis of PEX5, p-70S6K, p-S6R, and p-4E-BP-1
Rapamycin restores the functions of peroxisomes and lysosomes in PEX5-depleted cells during serum starvation
Since PEX5 depletion suppressed the mRNA expression of peroxisomal genes, we evaluated the effect of rapamycin on the expression levels of peroxisomal genes and proteins in PEX5-depleted cells during starvation. Treatment with rapamycin increased mRNA expression of peroxisome-related genes, including PMP70, ACOX1, catalase, NUDT7, PGC1α, and PPARα in PEX5-depleted and serum-starved cells compared to control cells in DMSO (Fig. 6a). Consistently, the expression levels of proteins including, PMP70, ACOX1, and catalase in PEX5-depleted and serum-starved cells were recovered by rapamycin treatment as compared to control cells in DMSO (Fig. 6b). These data suggest that rapamycin restored peroxisome biogenesis by regulating mTORC1 activity during PEX5 depletion in starvation. To further confirm TFEB activation, we measured the mRNA expression of target genes. mRNA levels of TFEB, LIPA, and LAMP-1 were decreased by PEX5 depletion, but increased by treatment with rapamycin (Fig. 6c). Further, mTORC1 inhibition by rapamycin markedly increased the expression of lysosomal proteins, including LAMP-1, LAMP-2, cathepsin A, and cathepsin B (Fig. 6d). It was previously suggested that phosphorylation of TFEB by mTORC1 inhibited the nuclear translocation of TFEB. To test whether constitutive activation of TFEB was able to overcome the effect of PEX5 depletion on both autophagy and peroxisome biogenesis, ectopic expression of wild-type (WT) TFEB or constitutively active TFEB(S142A) was individually induced in HepG2 cells. The expression of constitutively active TFEB(S142A) recovered the protein levels of LC3-II, LAMP-1, and PMP70 in PEX5-depleted cells compared with WT-TFEB-overexpressing cells during starvation (Fig. 6e). These results suggest that PEX5 depletion downregulated TFEB activity via the mTORC1 signaling pathway.
Cells transfected with siRNA for 24 h were pretreated with DMSO or rapamycin (1 μM) for 6 h, and then incubated in the absence of serum for 24 h. a Transcript levels of peroxisome-related genes were measured by Q-PCR. Values are expressed as fold increase compared to the starvation control group. Data are presented as mean ± s.d. from three independent experiments (**P < 0.01, *P < 0.05). b Immunoblot analysis of PMP70, ACOX1, DBP, catalase, and PEX5. c Q-PCR analysis of TFEB-regulated genes. Data are presented as mean ± s.d. from three independent experiments (**P < 0.01, *P < 0.05). d Protein expression of LAMP-1/LAMP-2 (lysosomal proteins) and cathepsins B/D (lysosomal proteases) were measured by immunoblotting. e Cells were transfected with empty, GFP-WT-TFEB, and GFP-TFEB-S142A plasmids. After 6 h of transfection, cells were treated with CTL and PEX5 siRNAs. After incubation in the absence of serum for 24 h, cells were subjected to immunoblot analysis with antibodies against PEX5, LC3, LAMP1, PMP70, TFEB, and β-actin
Discussion
The present data suggest that there is a relationship between PEX5 and autophagy mediated through the regulation of TSC2 expression and mTORC1 signaling. It has been shown that TSC1-TSC2 complex inhibits Rheb GTPase activity, suppressing the mTORC1 pathway, a negative modulator of autophagy28,33. Since TSC1 interacts with TSC2 and prevents its ubiquitin-dependent degradation34, it is considered that TSC2 protein stability is critical for regulating mTORC1 signaling. Moreover, recent studies have demonstrated that PEX5 recruits TSC2 to the peroxisomes through interaction between PEX5 and PTS1 in TSC2, indicating that their localization to peroxisomes plays an important role in repressing mTORC1 signaling in response to ROS5. Based on our data, PEX5 depletion repressed the transcriptional expression of TSC2 during starvation, which in turn was unable to inhibit mTOR activation, thereby blocking the autophagy process even in serum-depleted conditions. Although the detailed mechanism needs to be defined, our results suggested that TSC2 mRNA expression could be regulated to control its function, and that PEX5 might affect the transcriptional activity of TSC2 to modulate mTORC1 signaling during starvation.
The number of peroxisomes is tightly regulated by biogenesis and pexophagy to meet the metabolic needs of the cell35,36. Even though, mRNA of PMP70 was increased by PEX5 knockdown in normal condition, its protein level was not changed in Fig. 2. It is because the number of peroxisomes might be enough to supply the metabolic need in normal condition. Therefore, we considered that the number of peroxisomes are not induced despite of increased the level of PMP70 mRNA. On the other hands, PEX5 depletion suppressed peroxisome biogenesis in serum starvation condition. Our data have tried to demonstrate that PEX5 depletion was not able to suppress mTOR signaling during starvation, which prevents TFEB nuclear localization. Then, TFEB target genes including PPAR alpha and PGC1 alpha were not activated, which leads to down-regulate the peroxisome biogenesis. Based on our data, PEX5 depletion blocks peroxisome biogenesis through regulating TFEB during starvation.
In addition to response to lack of nutrient supply, autophagy is also required for modulating metabolic disturbances caused by genetic problems or variety of stresses, including oxidative stress, infection, and hypoxia. Autophagy dysfunction is suggested to contribute to the development and progression of numerous human diseases, including cancer, neurodegenerative disorders, aging, and metabolic diseases. According to previous findings, regulation of the autophagy process can exert therapeutic effects on several disorders37,38,39,40,41. One of the major peroxisomal disorders, X-linked ALD (X-ALD), is associated with the accumulation of VLCFA caused by mutations in ABCD1, a gene encoding the peroxisomal transporter protein42,43,44. Recent studies have suggested that abnormal accumulation of VLCFA attenuates autophagy, and restoration of autophagy by pharmacological treatment alleviates the symptoms of X-ALD caused by ABCD1 deficiency in mouse models45. Since PEX5 is involved in peroxisome biogenesis and its deficiency induces VLCFA accumulation, we hypothesized that PEX5 might play a critical role in regulation of autophagy. According to our results, PEX5-depleted cells showed increased mTORC1 activity and suppressed autophagy at the same time. It is suggested that mTORC1 activity might not be suppressed during starvation owing to the lack of TSC2 protein. Furthermore, PEX5 depletion also inhibited TFEB nuclear translocation in serum-starved cells as compared to control cells. Consistently, the transcription of genes for autophagy was significantly suppressed in PEX5-depleted cells under starvation. Notably, inhibition of mTORC1 restored TFEB activity, lysosomal biogenesis, and autophagy in PEX5-depleted cells under starvation. These results suggest that PEX5 may be a critical regulator of the mTOR-TFEB-autophagy axis during nutrient deprivation. Interestingly, our data suggested that TFEB might be involved in peroxisome biogenesis by regulating the transcription of peroxisomal genes during starvation. Because both PEX5 and TFEB depletion reduced the mRNA expression of peroxisomal genes, TFEB can be considered as a novel regulator of peroxisome biogenesis (Fig. 7).
Schematic representation of regulation of mTOR-TFEB by PEX5 depletion during starvation. In serum starvation, TSC2 suppresses mTOR activity, which promotes the nuclear localization of TFEB, which in turn regulates lysosomal gene expression, peroxisomes biogenesis, and autophagy. Lack of PEX5 reduces TSC2 activity, and thereby, is unable to prevent mTOR activity and activate TFEB in starvation conditions. Rapamycin treatment restores the dysregulation of mTOR-TFEB by PEX5 depletion
Despite regulating peroxisome biogenesis by transporting matrix proteins into peroxisomes, PEX5 induces pexophagy under stress conditions. In response to ROS stress or amino-acid starvation, PEX5 becomes ubiquitinated by E3 ligase located in the peroxisome and recruits the autophagy adaptor proteins, SQSTM1/p62 and NBR1, which target peroxisomes to autophagosomes for pexophagy13,36. We also observed that the number of peroxisomes was reduced by PEX5 knockdown during starvation. However, this reduction may not be due to pexophagy because PEX5 depletion by RNAi is not sufficient to control pexophagy through PEX5 ubiquitination. Therefore, it is possible that the decreased peroxisome numbers are resulted from suppression of peroxisome biogenesis through TFEB inactivation. On the other hand, PEX5 depletion in the liver led to abnormal mitochondrial accumulation10,29. It has been known that the damaged mitochondria are subjected to degradation by mitophagy46,47. From our results, it can be speculated that the presence of damaged mitochondria in PEX5 knockout liver is due to the suppressed autophagy, which in turn inhibits mitophagy. It is possible that elimination of abnormal mitochondria by induction of autophagy might alleviate the symptoms of Zellweger syndrome. Taken together, our study suggested that PEX5 is required for the regulation of autophagy via TFEB, and induction of autophagy could be an attractive therapeutic option for patients with Zellweger syndrome.
References
Fidaleo, M. Peroxisomes and peroxisomal disorders: the main facts. Exp. Toxicol. Pathol. 62, 615–625 (2010).
Steinberg, S. J. et al. Peroxisome biogenesis disorders. Biochim. Biophys. Acta 1763, 1733–1748 (2006).
Van Veldhoven, P. P. Biochemistry and genetics of inherited disorders of peroxisomal fatty acid metabolism. J. Lipid Res. 51, 2863–2895 (2010).
Dixit, E. et al. Peroxisomes are.ignaling platforms for antiviral innate immunity. Cell 141, 668–681 (2010).
Zhang, J. et al. A tuberous sclerosis complex signalling node at the peroxisome regulates mTORC1 and autophagy in response to ROS. Nat. Cell Biol. 15, 1186–1196 (2013).
Ma, C., Agrawal, G. & Subramani, S. Peroxisome assembly: matrix and membrane protein biogenesis. J. Cell Biol. 193, 7–16 (2011).
Francisco, T. et al. A cargo-centered perspective on the PEX5 receptor-mediated peroxisomal protein import pathway. J. Biol. Chem. 288, 29151–29159 (2010). 288.
Lee, J. N. et al. Proteasome inhibitors induce auditory hair cell death through peroxisome dysfunction. Biochem. Biophys. Res. Commun. 456, 269–274 (2015).
Baes, M. et al. A mouse model for Zellweger syndrome. Nat. Genet. 17, 49–57 (1997).
Baumgart, E. et al. Mitochondrial alterations caused by defective peroxisomal biogenesis in a mouse model for Zellweger syndrome (PEX5 knockout mouse). Am. J. Pathol. 159, 1477–1494 (2001).
Goldfischer, S. et al. Peroxisomal and mitochondrial defects in the cerebro-hepato-renal syndrome. Science 182, 62–64 (1973).
Wanders, R. J. A. Metabolic and molecular basis of peroxisomal disorders. Am. J. Hum. Genet. 126, 355–375 (2004).
Zhang, J. et al. ATM functions at the peroxisome to induce pexophagy in response to ROS. Nat. Cell Biol. 17, 1259–1269 (2015).
Nordgren, M. et al. Export-deficient monoubiquitinated PEX5 triggers peroxisome removal in SV40 large T antigen-transformed mouse embryonic fibroblasts. Autophagy 11, 1326–1340 (2015).
Pyo, J. O., Nah, J. & Jung, Y. K. Molecules and their functions in autophagy. Exp. Mol. Med. 44, 73–80 (2012).
Green, D. R. & Levine, B. To be or not to be? How selective autophagy and cell death govern cell fate. Cell 157, 65–75 (2014).
Kim, K. H. & Lee, M. S. Autophagy-a key player in cellular and body metabolism. Nat. Rev. Endocrinol. 10, 322–337 (2014).
Sengupta, S., Peterson, T. R. & Sabatini, D. M. Regulation of the mTOR complex 1 pathway by nutrients, growth factors, and stress. Mol. Cell 40, 310–322 (2010).
Zoncu, R., Efeyan, A. & Sabatini, D. M. mTOR: from growth signal integration to cancer, diabetes and ageing. Nat. Rev. Mol. Cell Biol. 12, 21–35 (2011).
Settembre, C. et al. TFEB links autophagy to lysosomal biogenesis. Science 17, 1429–1433 (2011).
Settembre, C. et al. A lysosome-to-nucleus signalling mechanism senses and regulates the lysosome via mTOR and TFEB. EMBO J. 31, 1095–1108 (2012).
Napolitano, G. & Ballabio, A. TFEB at a glance. J. Cell Sci. 129, 2475–2481 (2016).
Jin, A. et al. 2,2′-dipyridyl induces pexophagy. Biochem. Biophys. Res. Commun. 469, 941–947 (2016).
Lee, J. N. et al. 3-Aminotriazole protects from CoCl2-induced ototoxicity by inhibiting the generation of reactive oxygen species and proinflammatory cytokines in mice. Arch. Toxicol. 90, 781–791 (2016).
Moscat, J., Diaz-Meco, M. T. & Wooten, M. W. Of the atypical PKCs, Par-4 andp62: recent understandings of the biology and pathology of a PB1-dominated complex. Cell Death Differ. 16, 1426–1437 (2009).
Inoki, K., Li, Y., Xu, T. & Guan, K. L. Rheb GTPase is a direct target of TSC2 GAP activity and regulates mTOR signaling. Genes Dev. 17, 1829–1834 (2003).
Hu, J. et al. WD40 protein FBW5 promotes ubiquitination of tumor suppressor TSC2 by DDB1-CUL4-ROC1 ligase. Genes Dev. 22, 866–871 (2008).
Chong-Kopera, H. et al. TSC1 stabilizes TSC2 by inhibiting the interaction between TSC2 and the HERC1 ubiquitin ligase. J. Biol. Chem. 281, 8313–8316 (2006).
Dirkx, R. et al. Absence of peroxisomes in mouse hepatocytes causes mitochondrial and ER abnormalities. Hepatology 41, 868–878 (2005).
Bagattin, A., Hugendubler, L. & Mueller, E. Transcriptional coactivator PGC-1alpha promotes peroxisomal remodeling and biogenesis. Proc. Natl Acad. Sci. USA 107, 20376–20381 (2010).
Sardiello, M. et al. A gene network regulating lysosomal biogenesis and function. Science 325, 473–477 (2009).
Settembre, C. et al. TFEB controls cellular lipid metabolism through a starvation-induced autoregulatory loop. Nat. Cell Biol. 15, 647–658 (2013).
Yoon, M. S. & Choi, C. S. The role of amino acid-induced mammalian target of rapamycin complex 1(mTORC1) signaling in insulin resistance. Exp. Mol. Med. 48, e201 (2016).
Huang, J. & Manning, B. D. The TSC1-TSC2 complex: a molecular switchboard controlling cell growth. Biochem. J. 412, 179–190 (2008).
Nordgren, M., Wang, B., Apanasets, O. & Fransen, M. Peroxisome degradation in mammals: mechanisms of action, recent advances, and perspectives. Front Physiol. 4, 145 (2013).
Sargent, G. et al. PEX2 is the E3 ubiquitin ligase required for pexophagy during starvation. J. Cell Biol. 214, 677–690 (2016).
Chakrabarti, L., Eng, J., Ivanov, N., Garden, G. A. & La Spada, A. R. Autophagy activation and enhanced mitophagy characterize the Purkinje cells of pcd mice prior to neuronal death. Mol. Brain 2, 24 (2009).
Martinet, W., Verheye, S. & De Meyer, G. R. Everolimus-induced mTOR inhibition selectively depletes macrophages in atherosclerotic plaques by autophagy. Autophagy 3, 241–244 (2007).
Ciechanover, A. & Kwon, Y. T. Degradation of misfolded proteins in neurodegenerative diseases: therapeutic targets and strategies. Exp. Mol. Med. 47, e147 (2015).
Sharma, S., Mells, J. E., Fu, P. P., Saxena, N. K. & Anania, F. A. GLP-1 analogs reduce hepatocyte steatosis and improve survival by enhancing the unfolded protein response and promoting macroautophagy. PLoS. ONE. 6, e25269 (2011).
Ravikumar, B. et al. Inhibition of mTOR induces autophagy and reduces toxicity of polyglutamine expansions in fly and mouse models of Huntington disease. Nat. Genet. 36, 585–595 (2004).
Mosser, J. et al. Putative X-linked adrenoleukodystrophy gene shares unexpected homology with ABC transporters. Nature 361, 726–730 (1993).
Moser, H. W. et al. Adrenoleukodystrophy: increased plasma content of saturated very long chain fatty acids. Neurology 31, 1241–1249 (1981).
Berger, J., Forss-Petter, S. & Eichler, F. S. Pathophysiology of X-linked adrenoleukodystrophy. Biochimie 98, 135–142 (2014).
Launay, N. et al. Autophagy induction halts axonal degeneration in a mouse model of X-adrenoleukodystrophy. Acta Neuropathol. 129, 399–415 (2015).
Novak, I. et al. Nix is a selective autophagy receptor for mitochondrial clearance. EMBO Rep. 11, 45–51 (2010).
Youle, R. J. & Narendra, D. P. Mechanisms of mitophagy. Nat. Rev. Mol. Cell Biol. 12, 9–14 (2011).
Acknowledgements
This work was supported by the National Research Foundation of Korea (NRF) under grants funded by the Korean government (No. 2014M3A9D8034463, 2015R1A2A1A15054674, 2011-0030130, and 2017R1A2B2008840) and by GIST Research Institute(GRI) grant funded by GIST in 2017.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims inpublished maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, and provide a link to the Creative Commons license. You do not have permission under this license to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Eun, S.Y., Lee, J.N., Nam, IK. et al. PEX5 regulates autophagy via the mTORC1-TFEB axis during starvation. Exp Mol Med 50, 1–12 (2018). https://doi.org/10.1038/s12276-017-0007-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s12276-017-0007-8
- Springer Nature Limited
This article is cited by
-
TFEB activation triggers pexophagy for functional adaptation during oxidative stress under calcium deficient-conditions
Cell Communication and Signaling (2024)
-
Loss of pex5 sensitizes zebrafish to fasting due to deregulated mitochondria, mTOR, and autophagy
Cellular and Molecular Life Sciences (2023)
-
Autophagy-Dependent Increased ADAM10 Mature Protein Induced by TFEB Overexpression Is Mediated Through PPARα
Molecular Neurobiology (2021)
-
Activation of TFEB ameliorates dedifferentiation of arterial smooth muscle cells and neointima formation in mice with high-fat diet
Cell Death & Disease (2019)
-
Pexophagy in yeast and mammals: an update on mysteries
Histochemistry and Cell Biology (2018)