Abstract
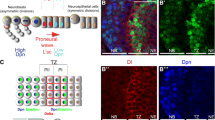
Brain areas each generate specific neuron subtypes during development. However, underlying regional variations in neurogenesis strategies and regulatory mechanisms remain poorly understood. In Drosophila, neurons in four optic lobe ganglia originate from two neuroepithelia, the outer (OPC) and inner (IPC) proliferation centers. Using genetic manipulations, we found that one IPC neuroepithelial domain progressively transformed into migratory progenitors that matured into neural stem cells (neuroblasts) in a second domain. Progenitors emerged by an epithelial-mesenchymal transition–like mechanism that required the Snail-family member Escargot and, in subdomains, Decapentaplegic signaling. The proneural factors Lethal of scute and Asense differentially controlled progenitor supply and maturation into neuroblasts. These switched expression from Asense to a third proneural protein, Atonal. Dichaete and Tailless mediated this transition, which was essential for generating two neuron populations at defined positions. We propose that this neurogenesis mode is central for setting up a new proliferative zone to facilitate spatio-temporal matching of neurogenesis and connectivity across ganglia.








Similar content being viewed by others
Change history
23 December 2014
In the version of this article initially published, there were misworded sentences in the abstract and introduction and formatting errors in the genotypes in the Online Methods. The abstract referred to “neural stem cells and neuroblasts” where it should have read “neural stem cells (neuroblasts).” The fourth paragraph began, “The IPC produces a larval neuron population known as the distal cells, whose neurites in the adult extend into either the medulla and lobula or the medulla and lamina, lobula plate neurons, whose neurites connect the lobula plate with the medulla or lobula, and lobula neurons.” The corrected sentence reads, “The IPC produces three populations: first, a larval neuron population known as the distal cells, whose neurites in the adult extend into either the medulla and lobula or the medulla and lamina; second, lobula plate neurons, whose neurites connect the lobula plate with the medulla or lobula; and third, lobula neurons.” In the Online Methods, first paragraph, the following should have been superscripted: x in brkx47 in all three instances, strII in tkvstrII in item (5) of the second numbered list, 1 in “dac1 from F. Pignoni”, IR KK100642 in UAS-fas3IR KK100642, and IR KK104691 in UAS-l′scIR KK104691. The following should not have been subscripted: sc in UAS-l′scIR TRiP.JF02399. The errors have been corrected in the HTML and PDF versions of the article.
References
Lui, J.H., Hansen, D.V. & Kriegstein, A.R. Development and evolution of the human neocortex. Cell 146, 18–36 (2011).
Florio, M. & Huttner, W.B. Neural progenitors, neurogenesis and the evolution of the neocortex. Development 141, 2182–2194 (2014).
Brand, A.H. & Livesey, F.J. Neural stem cell biology in vertebrates and invertebrates: more alike than different? Neuron 70, 719–729 (2011).
Homem, C.C. & Knoblich, J.A. Drosophila neuroblasts: a model for stem cell biology. Development 139, 4297–4310 (2012).
Ayala, R., Shu, T. & Tsai, L.H. Trekking across the brain: the journey of neuronal migration. Cell 128, 29–43 (2007).
Hofbauer, A. & Campos-Ortega, J.A. Proliferation and and early differentiation of the optic lobes in Drosophila melanogaster. Rouxs Arch. Dev. Biol. 198, 264–274 (1990).
Fischbach, K.F. & Dittrich, A.P.M. The optic lobe of Drosophila melanogaster. I. A Golgi analysis of wild-type structure. Cell Tissue Res. 258, 441–475 (1989).
Green, P., Hartenstein, A.Y. & Hartenstein, V. The embryonic development of the Drosophila visual system. Cell Tissue Res. 273, 583–598 (1993).
Egger, B., Boone, J.Q., Stevens, N.R., Brand, A.H. & Doe, C.Q. Regulation of spindle orientation and neural stem cell fate in the Drosophila optic lobe. Neural Dev. 2, 1 (2007).
Yasugi, T., Umetsu, D., Murakami, S., Sato, M. & Tabata, T. Drosophila optic lobe neuroblasts triggered by a wave of proneural gene expression that is negatively regulated by JAK/STAT. Development 135, 1471–1480 (2008).
Egger, B., Gold, K.S. & Brand, A.H. Notch regulates the switch from symmetric to asymmetric neural stem cell division in the Drosophila optic lobe. Development 137, 2981–2987 (2010).
Apitz, H. & Salecker, I. A challenge of numbers and diversity: neurogenesis in the Drosophila optic lobe. J. Neurogenet. 28, 233–249 (2014).
Li, X. et al. Temporal patterning of Drosophila medulla neuroblasts controls neural fates. Nature 498, 456–462 (2013).
Suzuki, T., Kaido, M., Takayama, R. & Sato, M. A temporal mechanism that produces neuronal diversity in the Drosophila visual center. Dev. Biol. 380, 12–24 (2013).
Huang, Z. & Kunes, S. Hedgehog, transmitted along retinal axons, triggers neurogenesis in the developing visual centers of the Drosophila brain. Cell 86, 411–422 (1996).
Huang, Z., Shilo, B.Z. & Kunes, S. A retinal axon fascicle uses spitz, an EGF receptor ligand, to construct a synaptic cartridge in the brain of Drosophila. Cell 95, 693–703 (1998).
Meinertzhagen, I.A. & Hanson, T.E. The development of the optic lobe. in The Development of Drosophila melanogaster (eds. Bate, M. & Martinez-Arias, A.) 1363–1491 (Cold Spring Harbor Laboratory Press, 1993).
Oliva, C. et al. Proper connectivity of Drosophila motion detector neurons requires Atonal function in progenitor cells. Neural Dev. 9, 4 (2014).
Maisak, M.S. et al. A directional tuning map of Drosophila elementary motion detectors. Nature 500, 212–216 (2013).
Tuthill, J.C., Nern, A., Holtz, S.L., Rubin, G.M. & Reiser, M.B. Contributions of the 12 neuron classes in the fly lamina to motion vision. Neuron 79, 128–140 (2013).
Harzsch, S. & Walossek, D. Neurogenesis in the developing visual system of the branchiopod crustacean Triops longicaudatus (LeConte, 1846): corresponding patterns of compound-eye formation in Crustacea and Insecta? Dev. Genes Evol. 211, 37–43 (2001).
Strausfeld, N.J. The evolution of crustacean and insect optic lobes and the origins of chiasmata. Arthropod Struct. Dev. 34, 235–256 (2005).
Lee, T. & Luo, L. Mosaic analysis with a repressible cell marker for studies of gene function in neuronal morphogenesis. Neuron 22, 451–461 (1999).
Lanzuolo, C. & Orlando, V. Memories from the polycomb group proteins. Annu. Rev. Genet. 46, 561–589 (2012).
Thiery, J.P., Acloque, H., Huang, R.Y. & Nieto, M.A. Epithelial-mesenchymal transitions in development and disease. Cell 139, 871–890 (2009).
Whiteley, M., Noguchi, P.D., Sensabaugh, S.M., Odenwald, W.F. & Kassis, J.A. The Drosophila gene escargot encodes a zinc finger motif found in snail-related genes. Mech. Dev. 36, 117–127 (1992).
Burstyn-Cohen, T. & Kalcheim, C. Association between the cell cycle and neural crest delamination through specific regulation of G1/S transition. Dev. Cell 3, 383–395 (2002).
Heldin, C.H., Landstrom, M. & Moustakas, A. Mechanism of TGF-beta signaling to growth arrest, apoptosis, and epithelial-mesenchymal transition. Curr. Opin. Cell Biol. 21, 166–176 (2009).
Affolter, M. & Basler, K. The Decapentaplegic morphogen gradient: from pattern formation to growth regulation. Nat. Rev. Genet. 8, 663–674 (2007).
Clandinin, T.R. & Feldheim, D.A. Making a visual map: mechanisms and molecules. Curr. Opin. Neurobiol. 19, 174–180 (2009).
Brand, M., Jarman, A.P., Jan, L.Y. & Jan, Y.N. asense is a Drosophila neural precursor gene and is capable of initiating sense organ formation. Development 119, 1–17 (1993).
Hassan, B.A. et al. Atonal regulates neurite arborization but does not act as a proneural gene in the Drosophila brain. Neuron 25, 549–561 (2000).
Younossi-Hartenstein, A., Nassif, C., Green, P. & Hartenstein, V. Early neurogenesis of the Drosophila brain. J. Comp. Neurol. 370, 313–329 (1996).
González, F., Romani, S., Cubas, P., Modolell, J. & Campuzano, S. Molecular analysis of the asense gene, a member of the achaete-scute complex of Drosophila melanogaster, and its novel role in optic lobe development. EMBO J. 8, 3553–3562 (1989).
Bello, B.C., Izergina, N., Caussinus, E. & Reichert, H. Amplification of neural stem cell proliferation by intermediate progenitor cells in Drosophila brain development. Neural Dev. 3, 5 (2008).
Boone, J.Q. & Doe, C.Q. Identification of Drosophila type II neuroblast lineages containing transit amplifying ganglion mother cells. Dev. Neurobiol. 68, 1185–1195 (2008).
Bowman, S.K. et al. The tumor suppressors Brat and Numb regulate transit-amplifying neuroblast lineages in Drosophila. Dev. Cell 14, 535–546 (2008).
Hay, E.D. The mesenchymal cell, its role in the embryo, and the remarkable signaling mechanisms that create it. Dev. Dyn. 233, 706–720 (2005).
Itoh, Y. et al. Scratch regulates neuronal migration onset via an epithelial-mesenchymal transition-like mechanism. Nat. Neurosci. 16, 416–425 (2013).
Theveneau, E. & Mayor, R. Neural crest delamination and migration: from epithelium-to-mesenchyme transition to collective cell migration. Dev. Biol. 366, 34–54 (2012).
Li, G., Fang, L., Fernandez, G. & Pleasure, S.J. The ventral hippocampus is the embryonic origin for adult neural stem cells in the dentate gyrus. Neuron 78, 658–672 (2013).
Machold, R., Klein, C. & Fishell, G. Genes expressed in Atoh1 neuronal lineages arising from the r1/isthmus rhombic lip. Gene Expr. Patterns 11, 349–359 (2011).
Lois, C., Garcia-Verdugo, J.M. & Alvarez-Buylla, A. Chain migration of neuronal precursors. Science 271, 978–981 (1996).
Doetsch, F. & Alvarez-Buylla, A. Network of tangential pathways for neuronal migration in adult mammalian brain. Proc. Natl. Acad. Sci. USA 93, 14895–14900 (1996).
Shen, S.P., Aleksic, J. & Russell, S. Identifying targets of the Sox domain protein Dichaete in the Drosophila CNS via targeted expression of dominant negative proteins. BMC Dev. Biol. 13, 1 (2013).
Wallace, K., Liu, T.H. & Vaessin, H. The pan-neural bHLH proteins DEADPAN and ASENSE regulate mitotic activity and cdk inhibitor dacapo expression in the Drosophila larval optic lobes. Genesis 26, 77–85 (2000).
Castro, D.S. et al. A novel function of the proneural factor Ascl1 in progenitor proliferation identified by genome-wide characterization of its targets. Genes Dev. 25, 930–945 (2011).
Bertrand, N., Castro, D.S. & Guillemot, F. Proneural genes and the specification of neural cell types. Nat. Rev. Neurosci. 3, 517–530 (2002).
Kohwi, M. & Doe, C.Q. Temporal fate specification and neural progenitor competence during development. Nat. Rev. Neurosci. 14, 823–838 (2013).
Shimozaki, K. et al. SRY-box-containing gene 2 regulation of nuclear receptor tailless (Tlx) transcription in adult neural stem cells. J. Biol. Chem. 287, 5969–5978 (2012).
Samakovlis, C. et al. Genetic control of epithelial tube fusion during Drosophila tracheal development. Development 122, 3531–3536 (1996).
Bourbon, H.M. et al. A P-insertion screen identifying novel X-linked essential genes in Drosophila. Mech. Dev. 110, 71–83 (2002).
Campbell, G. & Tomlinson, A. Transducing the Dpp morphogen gradient in the wing of Drosophila: regulation of Dpp targets by brinker. Cell 96, 553–562 (1999).
Tsai, S.F. et al. Gypsy retrotransposon as a tool for the in vivo analysis of the regulatory region of the optomotor-blind gene in Drosophila. Proc. Natl. Acad. Sci. USA 94, 3837–3841 (1997).
Chotard, C., Leung, W. & Salecker, I. glial cells missing and gcm2 cell autonomously regulate both glial and neuronal development in the visual system of Drosophila. Neuron 48, 237–251 (2005).
Bazigou, E. et al. Anterograde Jelly belly and Alk receptor tyrosine kinase signaling mediates retinal axon targeting in Drosophila. Cell 128, 961–975 (2007).
Nellen, D., Affolter, M. & Basler, K. Receptor serine/threonine kinases implicated in the control of Drosophila body pattern by decapentaplegic. Cell 78, 225–237 (1994).
Bellaïche, Y., Gho, M., Kaltschmidt, J.A., Brand, A.H. & Schweisguth, F. Frizzled regulates localization of cell-fate determinants and mitotic spindle rotation during asymmetric cell division. Nat. Cell Biol. 3, 50–57 (2001).
Jarman, A.P., Brand, M., Jan, L.Y. & Jan, Y.N. The regulation and function of the helix-loop-helix gene, asense, in Drosophila neural precursors. Development 119, 19–29 (1993).
Jarman, A.P., Grell, E.H., Ackerman, L., Jan, L.Y. & Jan, Y.N. Atonal is the proneural gene for Drosophila photoreceptors. Nature 369, 398–400 (1994).
Bier, E., Vaessin, H., Younger-Shepherd, S., Jan, L.Y. & Jan, Y.N. deadpan, an essential pan-neural gene in Drosophila, encodes a helix-loop-helix protein similar to the hairy gene product. Genes Dev. 6, 2137–2151 (1992).
Russell, S.R., Sanchez-Soriano, N., Wright, C.R. & Ashburner, M. The Dichaete gene of Drosophila melanogaster encodes a SOX-domain protein required for embryonic segmentation. Development 122, 3669–3676 (1996).
Patel, N.H., Snow, P.M. & Goodman, C.S. Characterization and cloning of fasciclin III: a glycoprotein expressed on a subset of neurons and axon pathways in Drosophila. Cell 48, 975–988 (1987).
Tayler, T.D., Robichaux, M.B. & Garrity, P.A. Compartmentalization of visual centers in the Drosophila brain requires Slit and Robo proteins. Development 131, 5935–5945 (2004).
Hayden, M.A., Akong, K. & Peifer, M. Novel roles for APC family members and Wingless/Wnt signaling during Drosophila brain development. Dev. Biol. 305, 358–376 (2007).
Kraut, R. & Campos-Ortega, J.A. inscuteable, a neural precursor gene of Drosophila, encodes a candidate for a cytoskeleton adaptor protein. Dev. Biol. 174, 65–81 (1996).
Ohshiro, T., Yagami, T., Zhang, C. & Matsuzaki, F. Role of cortical tumour-suppressor proteins in asymmetric division of Drosophila neuroblast. Nature 408, 593–596 (2000).
Wodarz, A., Ramrath, A., Grimm, A. & Knust, E. Drosophila atypical protein kinase C associates with Bazooka and controls polarity of epithelia and neuroblasts. J. Cell Biol. 150, 1361–1374 (2000).
Spana, E.P. & Doe, C.Q. The prospero transcription factor is asymmetrically localized to the cell cortex during neuroblast mitosis in Drosophila. Development 121, 3187–3195 (1995).
Kosman, D., Small, S. & Reinitz, J. Rapid preparation of a panel of polyclonal antibodies to Drosophila segmentation proteins. Dev. Genes Evol. 208, 290–294 (1998).
Cardona, A. et al. An integrated micro- and macroarchitectural analysis of the Drosophila brain by computer-assisted serial section electron microscopy. PLoS Biol. 8, e1000502 (2010).
Acknowledgements
We thank A. Baena-Lopez (MRC-NIMR), A. Carmena (CSIC University of Alicante), A. Gould (MRC-NIMR), Y.N. Jan (University of California, San Francisco), H. Jäckle (Max Planck Institute), A. Jarman (University of Edinburgh), G.X.S. Jefferis (MRC-LMB), J. Mueller (Max Planck Institute), J. Skeath (Washington University), R. Sousa-Nunes (King's College), E. Piddini (Gurdon institute), F. Pignoni (SUNY Upstate Medical University), J.P. Vincent (MRC-NIMR), U. Walldorf (University of Homburg), the Bloomington Drosophila Stock Center (US National Institutes of Health P40OD018537), the Drosophila Genomics Resource Center, the Vienna Drosophila RNAi Center and the Developmental Studies Hybridoma Bank for fly strains and antibodies. We thank A. Alifandi for help with EdU feeding experiments, A. Bailey and C. Desplan for helpful discussions, and A. Baena-Lopez, F. Guillemot, E. Ober, J.P. Vincent, B. Richier and N. Shimosako for critical reading of the manuscript. This work was supported by the Medical Research Council (U117581332).
Author information
Authors and Affiliations
Contributions
H.A. and I.S. conceived the project, H.A. carried out the experiments, and H.A. and I.S. analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Expression of escargot (esg) and genetic approach for achieving IPC-specific knockdown.
(a) esgMH766-Gal4 UAS-cd8GFP (green) and esg-lacZB7-2-22 (red) show similar expression patterns. Both transgenes label cells at the p-IPC neuroepithelial margins (arrowheads), as well as migratory progenitors within cell streams (arrows) of late third instar larvae (3L). (b,c) fasciclin 3 (fas3)NP1233-Gal4 drives expression of UAS-cd8GFP (green) in the IPC and its progeny, while ey3.5-Gal80 blocks Gal4 activity in R-cells (arrows, b). This approach efficiently drives expression of UAS-RNAi transgenes, as revealed by the knockdown of fas3 in p-IPC neuroepithlial cells and their progeny (asterisks, red, c). (d) In a EdU pulse-chase experiment, brains of mid third instar larvae (mid 3L) were assessed 4 hours after 2.5 hours EdU feeding. At this stage, p-IPC neuroepithelial cells extensively incorporate EdU (red) and thus are proliferative. dc, distal cells; ln, lamina neurons; lopn, lobula plate neurons; mn, medulla neurons. For genotypes and sample numbers, see Supplementary Table 2. Scale bars represent 50 μm.
Supplementary Figure 2 Validation of lethal of scute (l’sc) RNAi-mediated knockdown and characterization of phenotypes in the developing IPC.
(a) In control animals, maintained at 18°C, L’sc is expressed in neuroepithelial cells of the OPC and p-IPC (red, arrows). (b,c) Experimental animals were grown at 29°C during the third instar larval stage. IPC-specific expression of two different l’sc RNAi transgenes - l’scIR JF02399 and l’scIR KK104691 - using fasciclin 3 (fas3)NP1233-Gal4 results in efficient knockdown of L’sc expression in the p-IPC (asterisks) without affecting expression in the OPC (arrows). (d-g) IPC-specific expression of l’scIR KK104691 results in identical phenotypes as l’scIR JF02399 (cf. Fig. 7). Compared to controls, Deadpan-positive (Dpn+) neuroblasts (red, arrows, d,e) and Dachshund-positive (Dac+) progeny (red, arrows, f,g), including lobula plate neurons (lopn), are reduced. (h-k) Knockdown of l’sc does not affect esg-lacZ expression (red, h,i), p-IPC morphology (arrow, h,i), and Dichaete expression in cell streams (red, arrows, j,k). Although the d-IPC is reduced in size, Dichaete continues to be expressed (j,k). For genotypes and sample numbers, see Supplementary Table 2. Scale bars represent 50 μm.
Supplementary Figure 3 Validation of Dichaete RNAi-mediated knockdown in the IPC.
(a) In controls, Dichaete (red) is expressed in cell streams (arrow), the d-IPC and medulla neurons (mn). (b,c) IPC-specific expression of two different RNAi transgenes - DichaeteIR KK107194 and DichaeteIR GD49549 – using fasciclin 3 (fas3)NP1233-Gal4 results in efficient knockdown of Dichaete expression in progenitors in cell streams and neuroblasts in the d-IPC (asterisks), without affecting expression in medulla neurons. (d-f) Asense (Ase, red) is not expressed in cell streams (arrows) in control animals (d) and upon Dichaete knockdown (e,f). Both Dichaete RNAi transgenes show identical phenotypes in impairing d-IPC neurogenesis. Discs large (Dlg) immunolabeling is shown in blue. For genotypes and sample numbers, see Supplementary Table 2. Scale bars represent 50 μm.
Supplementary Figure 4 Knockdown of tailless (tll) affects early p-IPC development.
(a,b) Compared to controls (a), IPC-specific expression of a tll RNAi transgene using fasciclin 3 (fas3)NP1233-Gal4 strongly impairs early p-IPC neuroepithelial development, visualized by aPKC labeling at the third instar larval stage (red, b). The p-IPC is absent (asterisk) and the d-IPC is reduced (arrow). (c,d) neuralized (neur)-Gal4 mediated tll RNAi transgene expression results in knockdown of Tll (red) in the d-IPC (asterisk, d) without affecting expression in the lamina (La) and p-IPC (arrows). Low levels of Tll expression remain in the d-IPC. Discs large (Dlg) immunolabeling is shown in blue. For genotypes and sample numbers, see Supplementary Table 2. Scale bars represent 50 μm.
Supplementary Figure 5 Model illustrating two neuroblast competence stages in the d-IPC.
Migratory progenitors mature into neuroblasts (Nb) in the d-IPC, where they transition through two competence stages that are defined by the sequential expression of Asense (Ase) and Atonal (Ato)/Dachshund (Dac). These give rise to Twin of Eyeless-positive (Toy+) distal cells (dc) and Dachshund-positive (Dac+) lobula plate neurons (lopn), respectively. Dichaete (D) acts upstream of tailless (tll) to promote the transition between the two neuroblast stages. Genetic manipulations indicate that Dichaete is required to induce tll. tll is required to suppress Dichaete and ase, and to induce ato and dac.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Tables 1 and 2 (PDF 4643 kb)
3D model of proliferation zones and cell streams in the optic lobe.
To generate the model of a third instar larval optic lobe, the following areas were reconstructed using escargot (esg)-Gal4MH766, UAS-cd8GFP, Asense and E-cadherin as markers: Opc neuroepithelium (purple), lamina (light grey), medulla neuroblasts and ganglion mother cells (Gmc) (light purple), p-Ipc neuroepithelium (light green), progenitor cell streams (yellow green), d-Ipc neuroblasts and Gmcs (green), as well as s-Ipc derived lobula clusters (lo1 and lo2) including neuroblasts, Gmcs, postmitotic neurons and extending neurite tracts (grey). For labels see Fig. 1l. (MOV 10741 kb)
Source data
Rights and permissions
About this article
Cite this article
Apitz, H., Salecker, I. A region-specific neurogenesis mode requires migratory progenitors in the Drosophila visual system. Nat Neurosci 18, 46–55 (2015). https://doi.org/10.1038/nn.3896
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.3896
- Springer Nature America, Inc.
This article is cited by
-
Regulatory modules mediating the complex neural expression patterns of the homeobrain gene during Drosophila brain development
Hereditas (2022)
-
Complementary brains
Nature Ecology & Evolution (2022)
-
Neuronal diversity and convergence in a visual system developmental atlas
Nature (2021)
-
Spatio-temporal relays control layer identity of direction-selective neuron subtypes in Drosophila
Nature Communications (2018)
-
The transcription factor SoxD controls neuronal guidance in the Drosophila visual system
Scientific Reports (2018)