Abstract
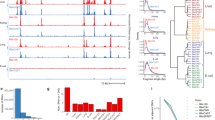
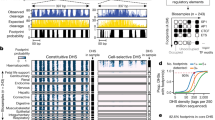
The function of human regulatory regions depends exquisitely on their local genomic environment and on cellular context, complicating experimental analysis of common disease- and trait-associated variants that localize within regulatory DNA. We use allelically resolved genomic DNase I footprinting data encompassing 166 individuals and 114 cell types to identify >60,000 common variants that directly influence transcription factor occupancy and regulatory DNA accessibility in vivo. The unprecedented scale of these data enables systematic analysis of the impact of sequence variation on transcription factor occupancy in vivo. We leverage this analysis to develop accurate models of variation affecting the recognition sites for diverse transcription factors and apply these models to discriminate nearly 500,000 common regulatory variants likely to affect transcription factor occupancy across the human genome. The approach and results provide a new foundation for the analysis and interpretation of noncoding variation in complete human genomes and for systems-level investigation of disease-associated variants.







Similar content being viewed by others
Change history
17 November 2015
In the version of this article initially published online, the Online Methods incorrectly abbreviated mapping quality as MAQ rather than MAPQ. Also in the Online Methods, the procedure for downsampling allele counts for cross–cell type analysis of imbalance was incorrectly written as "we subsampled each site to three cell types and further downsampled the allele counts to mapping quality for the lowest of the three cell types." The sentence should read "we subsampled each site to three cell types and further downsampled to the allele counts to match the lowest of the three cell types." The errors have been corrected for the print, PDF and HTML versions of this article.
References
Gross, D.S. & Garrard, W.T. Nuclease hypersensitive sites in chromatin. Annu. Rev. Biochem. 57, 159–197 (1988).
Thurman, R.E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Maurano, M.T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Degner, J.F. et al. DNase I sensitivity QTLs are a major determinant of human expression variation. Nature 482, 390–394 (2012).
Palmiter, R.D. & Brinster, R.L. Germ-line transformation of mice. Annu. Rev. Genet. 20, 465–499 (1986).
Sanyal, A., Lajoie, B.R., Jain, G. & Dekker, J. The long-range interaction landscape of gene promoters. Nature 489, 109–113 (2012).
Peterson, K.R. & Stamatoyannopoulos, G. Role of gene order in developmental control of human γ- and β-globin gene expression. Mol. Cell. Biol. 13, 4836–4843 (1993).
Thanos, D. & Maniatis, T. Virus induction of human IFN β gene expression requires the assembly of an enhanceosome. Cell 83, 1091–1100 (1995).
Archer, T.K., Lefebvre, P., Wolford, R.G. & Hager, G.L. Transcription factor loading on the MMTV promoter: a bimodal mechanism for promoter activation. Science 255, 1573–1576 (1992).
Mendenhall, E.M. et al. Locus-specific editing of histone modifications at endogenous enhancers. Nat. Biotechnol. 31, 1133–1136 (2013).
Aalfs, J.D. & Kingston, R.E. What does 'chromatin remodeling' mean? Trends Biochem. Sci. 25, 548–555 (2000).
Ronald, J. et al. Simultaneous genotyping, gene-expression measurement, and detection of allele-specific expression with oligonucleotide arrays. Genome Res. 15, 284–291 (2005).
Ni, Y., Hall, A.W., Battenhouse, A. & Iyer, V.R. Simultaneous SNP identification and assessment of allele-specific bias from ChIP-seq data. BMC Genet. 13, 46 (2012).
Knight, J.C., Keating, B.J., Rockett, K.A. & Kwiatkowski, D.P. In vivo characterization of regulatory polymorphisms by allele-specific quantification of RNA polymerase loading. Nat. Genet. 33, 469–475 (2003).
McDaniell, R. et al. Heritable individual-specific and allele-specific chromatin signatures in humans. Science 328, 235–239 (2010).
Kasowski, M. et al. Variation in transcription factor binding among humans. Science 328, 232–235 (2010).
Maurano, M.T., Wang, H., Kutyavin, T. & Stamatoyannopoulos, J.A. Widespread site-dependent buffering of human regulatory polymorphism. PLoS Genet. 8, e1002599 (2012).
Kilpinen, H. et al. Coordinated effects of sequence variation on DNA binding, chromatin structure, and transcription. Science 342, 744–747 (2013).
Reddy, T.E. et al. Effects of sequence variation on differential allelic transcription factor occupancy and gene expression. Genome Res. 22, 860–869 (2012).
McVicker, G. et al. Identification of genetic variants that affect histone modifications in human cells. Science 342, 747–749 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Heap, G.A. et al. Genome-wide analysis of allelic expression imbalance in human primary cells by high-throughput transcriptome resequencing. Hum. Mol. Genet. 19, 122–134 (2010).
Stergachis, A.B. et al. Exonic transcription factor binding directs codon choice and affects protein evolution. Science 342, 1367–1372 (2013).
Zhang, K. et al. Digital RNA allelotyping reveals tissue-specific and allele-specific gene expression in human. Nat. Methods 6, 613–618 (2009).
Henikoff, S. & Shilatifard, A. Histone modification: cause or cog? Trends Genet. 27, 389–396 (2011).
Neph, S. et al. An expansive human regulatory lexicon encoded in transcription factor footprints. Nature 489, 83–90 (2012).
Spivakov, M. et al. Analysis of variation at transcription factor binding sites in Drosophila and humans. Genome Biol. 13, R49 (2012).
Biddie, S.C. et al. Transcription factor AP1 potentiates chromatin accessibility and glucocorticoid receptor binding. Mol. Cell 43, 145–155 (2011).
Sherry, S.T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Badis, G. et al. Diversity and complexity in DNA recognition by transcription factors. Science 324, 1720–1723 (2009).
Zhao, Y. & Stormo, G.D. Quantitative analysis demonstrates most transcription factors require only simple models of specificity. Nat. Biotechnol. 29, 480–483 (2011).
Rohs, R. et al. The role of DNA shape in protein-DNA recognition. Nature 461, 1248–1253 (2009).
Meijsing, S.H. et al. DNA binding site sequence directs glucocorticoid receptor structure and activity. Science 324, 407–410 (2009).
Jolma, A. et al. DNA-binding specificities of human transcription factors. Cell 152, 327–339 (2013).
Lee, J.-H. et al. A robust approach to identifying tissue-specific gene expression regulatory variants using personalized human induced pluripotent stem cells. PLoS Genet. 5, e1000718 (2009).
Ding, J. et al. Gene expression in skin and lymphoblastoid cells: refined statistical method reveals extensive overlap in cis-eQTL signals. Am. J. Hum. Genet. 87, 779–789 (2010).
Price, A.L. et al. Single-tissue and cross-tissue heritability of gene expression via identity-by-descent in related or unrelated individuals. PLoS Genet. 7, e1001317 (2011).
Grundberg, E. et al. Mapping cis- and trans-regulatory effects across multiple tissues in twins. Nat. Genet. 44, 1084–1089 (2012).
Flutre, T., Wen, X., Pritchard, J. & Stephens, M. A statistical framework for joint eQTL analysis in multiple tissues. PLoS Genet. 9, e1003486 (2013).
Veyrieras, J.-B. et al. High-resolution mapping of expression-QTLs yields insight into human gene regulation. PLoS Genet. 4, e1000214 (2008).
John, S. et al. Chromatin accessibility pre-determines glucocorticoid receptor binding patterns. Nat. Genet. 43, 264–268 (2011).
John, S. et al. Genome-scale mapping of DNase I hypersensitivity. Curr. Protoc. Mol. Biol. Chapter 27, Unit 21.27 (2013).
Wang, H. et al. Widespread plasticity in CTCF occupancy linked to DNA methylation. Genome Res. 22, 1680–1688 (2012).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Neph, S. et al. BEDOPS: high-performance genomic feature operations. Bioinformatics 28, 1919–1920 (2012).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Lazarovici, A. et al. Probing DNA shape and methylation state on a genomic scale with DNase I. Proc. Natl. Acad. Sci. USA 110, 6376–6381 (2013).
1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Le Novère, N. MELTING, computing the melting temperature of nucleic acid duplex. Bioinformatics 17, 1226–1227 (2001).
Matys, V. et al. TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Res. 34, D108–D110 (2006).
Portales-Casamar, E. et al. JASPAR 2010: the greatly expanded open-access database of transcription factor binding profiles. Nucleic Acids Res. 38, D105–D110 (2010).
Newburger, D.E. & Bulyk, M.L. UniPROBE: an online database of protein binding microarray data on protein-DNA interactions. Nucleic Acids Res. 37, D77–D82 (2009).
Grant, C.E., Bailey, T.L. & Noble, W.S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Gupta, S., Stamatoyannopoulos, J.A., Bailey, T.L. & Noble, W.S. Quantifying similarity between motifs. Genome Biol. 8, R24 (2007).
Galas, D.J. & Schmitz, A. DNAse footprinting: a simple method for the detection of protein-DNA binding specificity. Nucleic Acids Res. 5, 3157–3170 (1978).
Sing, T., Sander, O., Beerenwinkel, N. & Lengauer, T. ROCR: visualizing classifier performance in R. Bioinformatics 21, 3940–3941 (2005).
Cooper, G.M. et al. Characterization of evolutionary rates and constraints in three Mammalian genomes. Genome Res. 14, 539–548 (2004).
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 15, 1034–1050 (2005).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Gulko, B., Hubisz, M.J., Gronau, I. & Siepel, A. A method for calculating probabilities of fitness consequences for point mutations across the human genome. Nat. Genet. 47, 276–283 (2015).
Lee, D. et al. A method to predict the impact of regulatory variants from DNA sequence. Nat. Genet. 47, 955–961 (2015).
Acknowledgements
This work was supported by US National Institutes of Health grants U54HG004592, U54HG007010, U01ES01156, 1S10RR026770 and 1S10OD017999 to J.A.S. and National Institute of Mental Health fellowship F31MH094073 to M.T.M. J.V. was supported by a National Science Foundation Graduate Research Fellowship under grant DGE-071824.
Author information
Authors and Affiliations
Contributions
M.T.M., E.H. and J.A.S. conceived and designed the experiments. M.T.M. and E.H. analyzed the data. J.V. and M.T.M. performed transcription factor cluster analysis. R.S. provided bioinformatics support. A.S. generated targeted footprinting data. R.K. assisted with data collection. M.T.M. and J.A.S. wrote the manuscript. M.T.M. and J.A.S. jointly supervised research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13 and Supplementary Tables 3–13 and 15–17. (PDF 11308 kb)
Supplementary Table 1. Overview of the DNase I data used in this study.
DNase I mapping of 116 cell types and tissues used in the study, including the shorthand name for the tissue. Signal portion of tags (SPOT) scores are a measure of enrichment and refer to the proportion of reads mapping within a DHS. Read counts include reads mapped uniquely with ≤2 mismatches to an autosomal chromosome; paired-end reads were required to both properly map to the same chromosome. Read counts are in millions. *, FL_E was excluded from the primary analysis and used for independent validation of the predictions in Figure 7. Previously published data sets are labeled by publication (refs. 2,3,24,27,64–67). (TXT 35 kb)
Supplementary Table 2. Overview of the ChIP-seq data used in this study.
ChIP-seq mapping of CTCF and H3K4me3 in 77 cell types and tissues used in the study, Signal portion of tags (SPOT) scores are a measure of enrichment and refer to the proportion of reads mapping within a DHS. Read counts include reads mapped uniquely with ≤2 mismatches to an autosomal chromosome; paired-end reads were required to both properly map to the same chromosome. Read counts are in millions. Previously published data sets are labeled by publication (refs. 2,17,44,68). (TXT 12 kb)
Supplementary Table 14. Clustering of motifs into TF families.
Clustering of motifs from the JASPAR, UniProbe, TRANSFAC and Jolma et al.35 databases. Each TF cluster is listed along with the names of constituent motifs. (TXT 34 kb)
Supplementary Data Set 1. SNPs tested for imbalance in DNA accessibility.
SNPs are listed by their hg19 coordinates. The rsID is used for SNPs in dbSNP 138. SNPs are classified as imbalanced as in Figure 1c. PctRef refers to the proportion of reads mapping to the reference allele (Fig.1d). (TXT 27676 kb)
Supplementary Data Set 2. TF clusters of similar motifs.
Motif weblogos from the JASPAR, UniPROBE and Jolma et al.35 databases grouped by TF cluster. Motifs from TRANSFAC are listed by name without showing a weblogo. (PDF 23365 kb)
Supplementary Data Set 3. SNVs predicted to affect DNA accessibility.
List of SNVs from dbSNP 138 overlapping a TF recognition sequence in a DHS hotspot predicted to affect accessibility with a score greater than 0.10. The file is in extended bed format using hg19 coordinates and includes a header line. Each row contains the SNP coordinates and dbSNP ID, a score scaled as the probability of imbalance, the PWM name and strand, the position of the SNP relative to the PWM match and the two alleles of the SNP. (ZIP 9362 kb)
Rights and permissions
About this article
Cite this article
Maurano, M., Haugen, E., Sandstrom, R. et al. Large-scale identification of sequence variants influencing human transcription factor occupancy in vivo. Nat Genet 47, 1393–1401 (2015). https://doi.org/10.1038/ng.3432
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3432
- Springer Nature America, Inc.
This article is cited by
-
Massively parallel identification of functionally consequential noncoding genetic variants in undiagnosed rare disease patients
Scientific Reports (2022)
-
An effector index to predict target genes at GWAS loci
Human Genetics (2022)
-
JAK inhibitors dampen activation of interferon-stimulated transcription of ACE2 isoforms in human airway epithelial cells
Communications Biology (2021)
-
Tissue context determines the penetrance of regulatory DNA variation
Nature Communications (2021)
-
Genetic perturbation of PU.1 binding and chromatin looping at neutrophil enhancers associates with autoimmune disease
Nature Communications (2021)