Abstract
During the final stage of cell division, cytokinesis, the Aurora-B-dependent abscission checkpoint (NoCut) delays membrane abscission to avoid DNA damage and aneuploidy in cells with chromosome segregation defects. This arrest depends on Aurora-B-mediated phosphorylation of CHMP4C, a component of the endosomal sorting complex required for transport (ESCRT) machinery that mediates abscission, but the mechanism remains unknown. Here we describe ANCHR (Abscission/NoCut Checkpoint Regulator; ZFYVE19) as a key regulator of the abscission checkpoint, functioning through the most downstream component of the ESCRT machinery, the ATPase VPS4. In concert with CHMP4C, ANCHR associates with VPS4 at the midbody ring following DNA segregation defects to control abscission timing and prevent multinucleation in an Aurora-B-dependent manner. This association prevents VPS4 relocalization to the abscission zone and is relieved following inactivation of Aurora B to allow abscission. We propose that the abscission checkpoint is mediated by ANCHR and CHMP4C through retention of VPS4 at the midbody ring.







Similar content being viewed by others
References
Guttinger, S., Laurell, E. & Kutay, U. Orchestrating nuclear envelope disassembly and reassembly during mitosis. Nat. Rev. Mol. Cell Biol. 10, 178–191 (2009).
Agromayor, M. & Martin-Serrano, J. Knowing when to cut and run: mechanisms that control cytokinetic abscission. Trends Cell Biol. 23, 433–441 (2013).
Fededa, J. P. & Gerlich, D. W. Molecular control of animal cell cytokinesis. Nat. Cell Biol. 14, 440–447 (2012).
Green, R. A., Paluch, E. & Oegema, K. Cytokinesis in animal cells. Annu. Rev. Cell Dev. Biol. 28, 29–58 (2012).
Guizetti, J. & Gerlich, D. W. Cytokinetic abscission in animal cells. Semin. Cell Dev. Biol. 21, 909–916 (2010).
Schiel, J. A., Childs, C. & Prekeris, R. Endocytic transport and cytokinesis: from regulation of the cytoskeleton to midbody inheritance. Trends Cell Biol. 23, 319–327 (2013).
Schiel, J. A. & Prekeris, R. Membrane dynamics during cytokinesis. Curr. Opin. Cell Biol. 25, 92–98 (2013).
Steigemann, P. & Gerlich, D. W. Cytokinetic abscission: cellular dynamics at the midbody. Trends Cell Biol. 19, 606–616 (2009).
Carlton, J. G. & Martin-Serrano, J. Parallels between cytokinesis and retroviral budding: a role for the ESCRT machinery. Science 316, 1908–1912 (2007).
Morita, E. et al. Human ESCRT and ALIX proteins interact with proteins of the midbody and function in cytokinesis. EMBO J. 26, 4215–4227 (2007).
Guizetti, J. et al. Cortical constriction during abscission involves helices of ESCRT-III-dependent filaments. Science 331, 1616–1620 (2011).
Elia, N., Sougrat, R., Spurlin, T. A., Hurley, J. H. & Lippincott-Schwartz, J. Dynamics of endosomal sorting complex required for transport (ESCRT) machinery during cytokinesis and its role in abscission. Proc. Natl Acad. Sci. USA 108, 4846–4851 (2011).
Guizetti, J. & Gerlich, D. W. ESCRT-III polymers in membrane neck constriction. Trends Cell Biol. 22, 133–140 (2012).
Hurley, J. H. & Hanson, P. I. Membrane budding and scission by the ESCRT machinery: it’s all in the neck. Nat. Rev. Mol. Cell Biol. 11, 556–566 (2010).
McCullough, J., Colf, L. A. & Sundquist, W. I. Membrane fission reactions of the mammalian ESCRT pathway. Annu. Rev. Biochem. 82, 663–692 (2013).
Schiel, J. A. et al. FIP3-endosome-dependent formation of the secondary ingression mediates ESCRT-III recruitment during cytokinesis. Nat. Cell Biol. 14, 1068–1078 (2012).
Janssen, A. & Medema, R. H. Genetic instability: tipping the balance. Oncogene 32, 4459–4470 (2013).
Mendoza, M. & Barral, Y. Co-ordination of cytokinesis with chromosome segregation. Biochem. Soc. Trans. 36, 387–390 (2008).
Petronczki, M. & Uhlmann, F. Cell biology. ESCRTing DNA at the cleavage site during cytokinesis. Science 336, 166–167 (2012).
Colnaghi, R., Carpenter, G., Volker, M. & O’Driscoll, M. The consequences of structural genomic alterations in humans: genomic disorders, genomic instability and cancer. Semin. Cell Dev. Biol. 22, 875–885 (2011).
Gascoigne, K. E. & Cheeseman, I. M. Induced dicentric chromosome formation promotes genomic rearrangements and tumorigenesis. Chromosome. Res. 21, 407–418 (2013).
Orthwein, A. et al. Mitosis inhibits DNA double-strand break repair to guard against telomere fusions. Science 344, 189–193 (2014).
Shimizu, N., Shingaki, K., Kaneko-Sasaguri, Y., Hashizume, T. & Kanda, T. When, where and how the bridge breaks: anaphase bridge breakage plays a crucial role in gene amplification and HSR generation. Exp. Cell Res. 302, 233–243 (2005).
Burrell, R. A. et al. Replication stress links structural and numerical cancer chromosomal instability. Nature 494, 492–496 (2013).
Chan, K. L. & Hickson, I. D. New insights into the formation and resolution of ultra-fine anaphase bridges. Semin. Cell Dev. Biol. 22, 906–912 (2011).
Dykhuizen, E. C. et al. BAF complexes facilitate decatenation of DNA by topoisomerase IIalpha. Nature 497, 624–627 (2013).
Germann, S. M. et al. TopBP1/Dpb11 binds DNA anaphase bridges to prevent genome instability. J. Cell Biol. 204, 45–59 (2014).
Wang, L. H., Mayer, B., Stemmann, O. & Nigg, E. A. Centromere DNA decatenation depends on cohesin removal and is required for mammalian cell division. J. Cell Sci. 123, 806–813 (2010).
Caldwell, C. M., Green, R. A. & Kaplan, K. B. APC mutations lead to cytokinetic failures in vitro and tetraploid genotypes in Min mice. J. Cell Biol. 178, 1109–1120 (2007).
Steigemann, P. et al. Aurora B-mediated abscission checkpoint protects against tetraploidization. Cell 136, 473–484 (2009).
Burrell, R. A., McGranahan, N., Bartek, J. & Swanton, C. The causes and consequences of genetic heterogeneity in cancer evolution. Nature 501, 338–345 (2013).
Ganem, N. J. & Pellman, D. Linking abnormal mitosis to the acquisition of DNA damage. J. Cell Biol. 199, 871–881 (2012).
Hoffelder, D. R. et al. Resolution of anaphase bridges in cancer cells. Chromosoma 112, 389–397 (2004).
Janssen, A., van der Burg, M., Szuhai, K., Kops, G. J. & Medema, R. H. Chromosome segregation errors as a cause of DNA damage and structural chromosome aberrations. Science 333, 1895–1898 (2011).
Norden, C. et al. The NoCut pathway links completion of cytokinesis to spindle midzone function to prevent chromosome breakage. Cell 125, 85–98 (2006).
Thompson, S. L. & Compton, D. A. Examining the link between chromosomal instability and aneuploidy in human cells. J. Cell Biol. 180, 665–672 (2008).
Davoli, T. & de, L. T. The causes and consequences of polyploidy in normal development and cancer. Annu. Rev. Cell Dev. Biol. 27, 585–610 (2011).
Fujiwara, T. et al. Cytokinesis failure generating tetraploids promotes tumorigenesis in p53-null cells. Nature 437, 1043–1047 (2005).
Gordon, D. J., Resio, B. & Pellman, D. Causes and consequences of aneuploidy in cancer. Nat. Rev. Genet. 13, 189–203 (2012).
Holland, A. J. & Cleveland, D. W. Losing balance: the origin and impact of aneuploidy in cancer. EMBO Rep. 13, 501–514 (2012).
Oromendia, A. B., Dodgson, S. E. & Amon, A. Aneuploidy causes proteotoxic stress in yeast. Genes Dev. 26, 2696–2708 (2012).
Siegel, J. J. & Amon, A. New insights into the troubles of aneuploidy. Annu. Rev. Cell Dev. Biol. 28, 189–214 (2012).
Storchova, Z. & Pellman, D. From polyploidy to aneuploidy, genome instability and cancer. Nat. Rev. Mol. Cell Biol. 5, 45–54 (2004).
Norden, C. et al. The NoCut pathway links completion of cytokinesis to spindle midzone function to prevent chromosome breakage. Cell 125, 85–98 (2006).
Mendoza, M. et al. A mechanism for chromosome segregation sensing by the NoCut checkpoint. Nat. Cell Biol. 11, 477–483 (2009).
Carlton, J. G., Caballe, A., Agromayor, M., Kloc, M. & Martin-Serrano, J. ESCRT-III governs the Aurora B-mediated abscission checkpoint through CHMP4C. Science 336, 220–225 (2012).
Capalbo, L. et al. The chromosomal passenger complex controls the function of endosomal sorting complex required for transport-III Snf7 proteins during cytokinesis. Open Biol. 2, 120070 (2012).
Chen, C. T., Hehnly, H. & Doxsey, S. J. Orchestrating vesicle transport, ESCRTs and kinase surveillance during abscission. Nat. Rev. Mol. Cell Biol. 13, 483–488 (2012).
Mackay, D. R., Makise, M. & Ullman, K. S. Defects in nuclear pore assembly lead to activation of an Aurora B-mediated abscission checkpoint. J. Cell Biol. 191, 923–931 (2010).
Capalbo, L. et al. The chromosomal passenger complex controls the function of endosomal sorting complex required for transport-III Snf7 proteins during cytokinesis. Open. Biol. 2, 120070 (2012).
Sagona, A. P. et al. A tumor-associated mutation of FYVE-CENT prevents its interaction with Beclin 1 and interferes with cytokinesis. PLoS One 6, e17086 (2011).
Thoresen, S. B., Pedersen, N. M., Liestol, K. & Stenmark, H. A phosphatidylinositol 3-kinase class III sub-complex containing VPS15, VPS34, Beclin 1, UVRAG and BIF-1 regulates cytokinesis and degradative endocytic traffic. Exp. Cell Res. 316, 3368–3378 (2010).
Doxsey, S. J. Molecular links between centrosome and midbody. Mol. Cell 20, 170–172 (2005).
Doxsey, S., McCollum, D. & Theurkauf, W. Centrosomes in cellular regulation. Annu. Rev. Cell Dev. Biol. 21, 411–434 (2005).
Caballe, A. & Martin-Serrano, J. ESCRT machinery and cytokinesis: the road to daughter cell separation. Traffic 12, 1318–1326 (2011).
Carmena, M., Ruchaud, S. & Earnshaw, W. C. Making the Auroras glow: regulation of Aurora A and B kinase function by interacting proteins. Curr. Opin. Cell Biol. 21, 796–805 (2009).
Kettenbach, A. N. et al. Quantitative phosphoproteomics identifies substrates and functional modules of Aurora and Polo-like kinase activities in mitotic cells. Sci. Signal. 4, rs5 (2011).
Von Schwedler, U. K. et al. The protein network of HIV budding. Cell 114, 701–713 (2003).
Bajorek, M. et al. Biochemical analyses of human IST1 and its function in cytokinesis. Mol. Biol. Cell 20, 1360–1373 (2009).
Lee, S. et al. MITD1 is recruited to midbodies by ESCRT-III and participates in cytokinesis. Mol. Biol. Cell 23, 4347–4361 (2012).
Morita, E. et al. Human ESCRT-III and VPS4 proteins are required for centrosome and spindle maintenance. Proc. Natl Acad. Sci. USA 107, 12889–12894 (2010).
Chinwalla, V. et al. A t(11;15) fuses MLL to two different genes, AF15q14 and a novel gene MPFYVE on chromosome 15. Oncogene 22, 1400–1410 (2003).
Marina, O. et al. Serologic markers of effective tumor immunity against chronic lymphocytic leukemia include nonmutated B-cell antigens. Cancer Res. 70, 1344–1355 (2010).
Mackay, D. R., Elgort, S. W. & Ullman, K. S. The nucleoporin Nup153 has separable roles in both early mitotic progression and the resolution of mitosis. Mol. Biol. Cell 20, 1652–1660 (2009).
Campeau, E. et al. A versatile viral system for expression and depletion of proteins in mammalian cells. PLoS One 4, e6529 (2009).
Dull, T. et al. A third-generation lentivirus vector with a conditional packaging system. J. Virol. 72, 8463–8471 (1998).
Acknowledgements
A. Engen and H.P. Bjønnes are acknowledged for expert handling of cell cultures, and T. Høiby and C. Herrmann for invaluable technical assistance. O. Mjaavatten at the Proteomics Unit at the University of Bergen (PROBE) and G. de Souza at the Proteomics Core Facility Unit at Oslo University Hospital are acknowledged for performing mass spectrometry analyses of GFP–ANCHR and GFP–VPS4 immunoprecipitates. The confocal microscopy core facility of Oslo University Hospital is acknowledged for providing access to microscopes. We also thank D. Gerlich at the Institute of Molecular Biotechnology (IMBA), Vienna, for supplying plasmids for the generation of GFP–α-tubulin+mCherry–H2B stable cell lines, and J. Martin-Serrano, King’s College London, UK, for advice on NoCut activation. S.B.T. and M.V. are PhD students of the South-Eastern Norway Regional Health Authority. C.R. is a senior researcher and C.C. a postdoctoral fellow of the Norwegian Cancer Society. H.S. was supported by an Advanced Grant from the European Research Council. This work was partly supported by the Research Council of Norway through its Centres of Excellence funding scheme, project number 179571.
Author information
Authors and Affiliations
Contributions
S.B.T. generated plasmid constructs, and performed confocal, high-content and live imaging, biochemical work, cell transfections, image processing, data analysis, statistical analyses and preparation of figures. C.C. generated plasmid constructs and stable cell lines, and performed live microscopy, photoconversion experiments, image processing, in vitro kinase assays and data analysis. C.R. carried out and analysed the siRNA screen targeting PtdIns(3)P-binding proteins, and performed cell transfections for epistasis studies. M.V. performed cell transfections, biochemical work and image analysis. K.O.S. generated plasmid constructs and stable cell lines, and performed photoconversion experiments and image processing. K.L. did the statistical analysis of the siRNA screen. J.S.A. performed mass spectrometry analysis of Aurora-B-phosphorylated ANCHR. H.S. coordinated the study and oversaw experiments. S.B.T., C.C. and H.S. wrote the paper. All authors discussed the results and assisted in revising the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 The FYVE domain protein ANCHR (ZFYVE19) regulates abscission and prevents cleavage furrow regression in the presence of lagging chromatin.
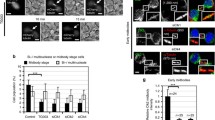
(a) HeLa cells were transfected with smart-pool siRNAs targeting the PtdIns(3)P-binding FYVE- and PX- domain families of proteins, and after 72 or 96 h fixed and stained for α-tubulin, Aurora B and DNA (Hoechst). Images collected on a high-content ScanR microscope were then scored manually to quantify multinuclear cells. The values obtained are ranked in ascending order according to the frequency of multinuclear cells. Each dot represents the average of four individual observations. Red dots represent PtdIns(3)P-binding proteins, while green and blue dots represent positive and negative controls, respectively. Dashed box highlights ANCHR (ZFYVE19). (b,c) Montage showing sequential images of asynchronous HeLa cells expressing GFP-α-tubulin and mCherry-H2B treated with siRNAs through cytokinesis, with a time interval of 5 min. T = time, T = 0 defined as the point of complete cleavage furrow ingression. Scale bar, 10 μm. (b) Arrows indicate microtubule disassembly. (c) shows furrow regression in ANCHR knock-down cells in the presence of lagging anaphase chromatin (indicated by arrows).
Supplementary Figure 2 ANCHR (ZFYVE19) binds PtdIns(3)P.
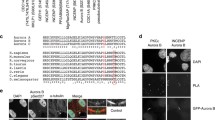
(a) Immunoblot showing GST alone or GST-ANCHR incubated with PtdIns(3)P-liposomes. (b) Representative confocal images showing the localization of Myc-ANCHR1 − 133, but not Myc-ANCHR1 − 133R101A to PtdIns(3)P-rich early endosomes. The cells were co-stained with antibodies against the early endosome marker EEA1 and β-Actin. Scale bar, 20 μm. (c) Representative confocal images showing the localization of GFP-ANCHR1 − 133 and GFP-ANCHR R101A to the midbody. The cells were co-stained with antibodies against α-tubulin and DNA (Hoechst). Scale bar, 10 μm.
Supplementary Figure 3 ANCHR interacts with VPS4 at overexpressed and moderate, non-arresting levels.
(a) Coomassie-stained gel of GFP-trap lysates used for mass spectromeric analysis of proteins interacting with overexpressed GFP or GFP-ANCHR. Black arrows indicate GFP and GFP-ANCHR. Red arrows indicate bands specifically detected in the GFP-ANCHR pull down which were excised and sent for mass spec analysis. (b) Western blot analysis of lysates from stable transgenic HeLa lines used for mass spec analysis presented in Supplementary Table 2, probed with anti-ANCHR (left panel) or anti-VPS4 (right panel) antibodies to determine the expression level of transgene compared to endogenous allele. Quantification indicated that the eGFP-ANCHR transgene was overexpressed 6 fold compared to endogenous, whereas eGFP-VPS4 levels were equal to endogenous. (c) GFP-trap immunoprecipitates (IP) from HeLa cells stably expressing GFP-ANCHR analysed by western blotting using antibodies against GFP and VPS4.
Supplementary Figure 4 Localization of ANCHR and VPS4 during interphase and early mitosis.
Localization of ANCHR and VPS4 during interphase and early mitosis. (a) Representative confocal images showing HeLa cells stained with antibodies against endogenous ANCHR and VPS4, and DNA (Hoechst). Scale bar, 10 μm. (b–d) Assessment of VPS4 antibody specificity. HeLa cells were treated with either non-targeting siRNA or VPS4A siRNA for 72 h and stained with antibodies against endogenous ANCHR and VPS4, and DNA (Hoechst). (b) Representative confocal images showing late cytokinetic cells containing chromatin bridges. Circles highlight the midbody ring. Scale bar, 10 μm. (c) Quantification of VPS4 intensities on midbody rings. (D) Western blot from whole cell lysates blotted for VPS4 and β-actin. Arrows indicate VPS4B (top), VPS4A (middle) and an unspecific band (bottom).
Supplementary Figure 5 ANCHR interacts with VPS4 at the midbody.
(a) Representative confocal images showing GFP-ANCHR and endogenous VPS4 at the midbody in a normally segregating cell. The cells were co-stained with Hoechst (DNA). Scale bar, 10 μm. (b) Representative confocal images from HeLa cells stably expressing GFP-VPS4A and mCherry-CEP55 showing their colocalization at the Flemming body (midbody ring). The cells were co-stained with antibodies against α-tubulin and DNA (Hoechst). Scale bar, 10 μm (c) Montage showing sequential stills of a HeLa cell expressing GFP-VPS4 and mCherry-CEP55, imaged at a time interval of 5 min. GFP-VPS4 co-localizes with the mCherry-CEP55-positive Flemming body (midbody ring) before abscission. Arrow indicates severed cytokinetic bridge. Scale bar, 2 μm. (d) The structure of ANCHR showing the defined domains, including the MIM-A and MIM-B. Alignment of the ANCHR MIM-A and MIM-B with other type-1 MIMs from different ESCRTs.
Supplementary Figure 6 The midbody is intact in ANCHR-depleted cells.
(a) Representative confocal images showing ANCHR, VPS4 and RacGAP1 at the midbody in untransfected (left panel) or ANCHR-depleted (right panel) cells. The cells were co-stained with Hoechst (DNA). Scale bar, 10 μm. (b) Quantification of RagGAP1 mean intensity on midbody ring (a.u. = arbitrary units) ± s.d. n = 15.
Supplementary Figure 7 VPS4 interaction with CHMP4C is reduced on Aurora B inhibition.
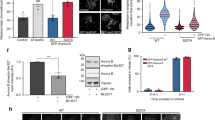
Quantification of VPS4 from GFP-trap blots illustrated in Fig. 5f. Values are averaged from three independent experiments.
Supplementary Figure 8 In vitro and in vivo phosphorylation of ANCHR.
(a) ANCHR can be phosphorylated by Aurora B at S22 in vitro. In vitro kinase assays using purified recombinant GST or GST-ANCHR, incubated with active Aurora B kinase in the presence of 33P. Incorporation of 33P was visualized by autoradiography, protein loading was assessed by coomassie staining. Mass spectrometry analysis (Supplementary Table 3) of in vitro phosphorylated GST-ANCHR using the non-radioactive phosphorus isotope identified S22 as the dominant phosphorylation site. Subsequent in vitro kinase assays using mutated GST-ANCHR S22A abolished phosphorylation by Aurora B, indicating that S22 is the sole Aurora B target in ANCHR in vitro. (b) ANCHR is not a prominent Aurora B target in vivo. HeLa cells transfected with the indicated constructs were labelled with 33P phosphate overnight, and treated with 2 μm AZD1152 for 2.5 h where indicated. Following lysis, GFP-fusions were immunoprecipitated using GFP antibodies. Incorporation of 33P was evaluated by autoradiography.
Supplementary Figure 9
Uncropped western blots used in this study.
Supplementary information
Supplementary Information
Supplementary Information (PDF 962 kb)
Supplementary Table 1
Supplementary Information (XLS 27 kb)
Supplementary Table 2
Supplementary Information (XLS 39 kb)
Supplementary Table 3
Supplementary Information (XLS 26 kb)
Supplementary Table 4
Supplementary Information (XLSX 10 kb)
Supplementary Table 5
Supplementary Information (XLSX 9 kb)
Localization of eGFP-Vps4 to the midbody ring before abscission.
HeLa cells stably expressing near-endogenous levels of eGFP-VPS4 and the midbody marker mCherry-CEP55 were imaged as the cell proceeded through abscission. Note that eGFP-VPS4 colocalizes with mCherry-CEP55 before abscission (t = 12 min). (MOV 429 kb)
Rights and permissions
About this article
Cite this article
Thoresen, S., Campsteijn, C., Vietri, M. et al. ANCHR mediates Aurora-B-dependent abscission checkpoint control through retention of VPS4. Nat Cell Biol 16, 547–557 (2014). https://doi.org/10.1038/ncb2959
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2959
- Springer Nature Limited
This article is cited by
-
The many functions of ESCRTs
Nature Reviews Molecular Cell Biology (2020)
-
The EGFR-ZNF263 signaling axis silences SIX3 in glioblastoma epigenetically
Oncogene (2020)
-
Understanding the birth of rupture-prone and irreparable micronuclei
Chromosoma (2020)
-
ESCRT-III is necessary for the integrity of the nuclear envelope in micronuclei but is aberrant at ruptured micronuclear envelopes generating damage
Oncogenesis (2019)
-
Building bridges between chromosomes: novel insights into the abscission checkpoint
Cellular and Molecular Life Sciences (2019)