Abstract
Background:
Mutations in the KRAS gene can be detected in about 70–90% of pancreatic cancer (PC) cases. Whether these mutations have a prognostic or predictive value remains elusive. Furthermore, the clinical relevance of the extended RAS (KRAS+NRAS) mutational status is unclear in PC.
Methods:
We prospectively defined a PC patient population who received erlotinib-free chemotherapy regimens. A statistically significant difference between KRAS wild-type and KRAS mutated tumours in at least 160 patients in this population would support the assumption of a rather prognostic role of KRAS.
Results:
One hundred and seventy-eight tumour samples were collected from prospective clinical studies and successfully analysed for the extended RAS status: 37 tumours were KRAS wild-type (21%), whereas 141 (79%) carried a KRAS mutation; 132 of these mutations were found in KRAS exon 2 (74%), whereas only 9 mutations (5%) were detected in KRAS exon 3. Within KRAS exon 4 and NRAS exons 2–4, no mutations were apparent. There was no significant difference in overall survival for KRAS wild-type vs mutant patients (9.9 vs 8.3 months, P=0.70).
Conclusions:
Together with the results of the AIO-PK-0104-trial, the present analysis supports the notion that KRAS mutation status is rather predictive than prognostic in advanced PC.
Similar content being viewed by others
Main
Pancreatic cancer (PC) is increasingly becoming a leading cause of death from gastrointestinal malignancies (Rahib et al, 2014). During the past years, novel chemotherapy regimens such as FOLFIRINOX or gemcitabine plus nab-paclitaxel have improved therapeutic options for patients with advanced disease (Conroy et al, 2011; Von Hoff et al, 2013). Despite significant efforts by many translational research groups, there are still no established biomarkers that could guide individualised treatment decisions in PC (Kruger et al, 2014).
Thus the current situation in PC represents a clear contrast to other solid tumours such as colorectal, breast or lung cancer, where a molecular workup of tumour specimens before the initiation of targeted therapy has become a standard procedure already some years ago (Amado et al, 2008; Rosell et al, 2012). The only targeted therapy which has been approved for PC in 2007 is erlotinib, an oral tyrosine kinase inhibitor acting at the intracellular adenoside triphosphate-binding site of the epidermal growth factor receptor (EGFR) (Moore et al, 2007). Up to now, there is, however, no predictive biomarker regarding the EGFR or its downstream targets for the use of erlotinib in PC.
In a large randomised phase III trial conducted by the ‘Arbeitsgemeinschaft Internistische Onkologie’ (AIO-PK0104), where erlotinib was used as first-line treatment in combination with either gemcitabine or capecitabine, Kirsten rat sarcoma oncogene (KRAS) mutations in exon 2 were successfully determined in 173 patients. A KRAS exon 2 wild-type status was shown to be associated with an improved overall survival (OS) of 7.9 months compared with 5.7 months in patients with a KRAS mutation (HR 1.68, P=0.005) (Boeck et al, 2013; Heinemann et al, 2013). All patients treated within AIO-PK0104 received erlotinib, thus the potential predictive vs prognostic role of KRAS remained elusive. A recent meta-analysis that pooled data from 17 studies on KRAS reported a potential prognostic role of KRAS in PC (Tao et al, 2016). However, as this meta-analysis included data from patients in different clinical settings (e.g., resected patients and advanced disease) and treatments (no chemotherapy, chemotherapy, EGFR-containing regimens), it still remains unclear how to interpret the relation between KRAS and OS in PC.
In the current study, the authors therefore aimed to collect archival tumour samples from patients with advanced PC treated with erlotinib-free chemotherapy regimens in order to investigate whether the effect of KRAS on OS is prognostic or predictive for the efficacy of erlotinib. Most of the already existing reports evaluating the relevance of KRAS mutations in PC performed KRAS analyses in exon 2 only (or partly in exon 3). To our knowledge, no reports are so far available on the role of NRAS mutation in PC. It is known from colorectal cancer that tumours with mutations in the extended RAS genes (KRAS exons 2–4 and NRAS exons 2–4) do not respond to treatment with EGFR-acting agents (Douillard et al, 2013). We therefore decided to perform an extended RAS analysis also in this translational multicentre study.
Methods
Patient population
For the current pooled prospective analysis, patients were eligible if they had histologically proven locally advanced or metastatic adenocarcinoma of the pancreas with available formalin-fixed paraffin-embedded (FFPE) archival tumour samples and were treated with at least one dose of palliative first-line chemotherapy without EGFR-acting agents, especially erlotinib. Previous chemotherapy or chemoradiotherapy treatment, with the exception of adjuvant therapy, was not allowed. Neuroendocrine tumours of the pancreas were excluded. For a better comparability of survival data, patients receiving the more effective combination regimens FOLFIRINOX or gemcitabine+nab-paclitaxel were excluded.
Three large randomised multicentre phase II/III trials, conducted by our group in the pre-erlotinib era, were screened for eligible patients: Heinemann et al (2006) (phase III, gemcitabine+cisplatin vs gemcitabine), Boeck et al (2008) (phase II, capecitabine+oxaliplatin vs capecitabine+gemcitabine vs gemcitabine+oxaliplatin) and Heinemann et al (2013) (AIO-PK0104 trial preamendment: capecitabine followed by second-line gemcitabine vs gemcitabine followed by second-line capecitabine in a crossover design). Additional patients were identified from a prospectively maintained tumour registry of the outpatient clinic at the Ludwig-Maximilian-University of Munich and the Technical University of Munich. This project was approved by the ethics committees of both universities (project numbers 554-11 and 401-15, respectively).
Statistical analysis
For the current translational analysis, a preplanned sample size calculation was performed based on the translational results of the AIO-PK0104 trial (Heinemann et al, 2013): in 176 patients treated within the AIO-PK0104 trial, patients with KRAS wild-type tumours treated with an erlotinib-containing regimen showed a superior survival compared with patients with KRAS mutant tumours (HR 0.60, P=0.005). To detect the same difference (with a power of 80% by a two-sided log-rank test with a significance level of 5%) in a patient population treated without EGFR-targeting agents, at least 160 patients were needed, estimating a rate of censored observations for OS of 10% and a rate of KRAS wild-type vs mutant tumours of 30% and 70%, respectively. A statistically significant difference between KRAS wild-type and KRAS mutant tumours regarding OS in this patient population not receiving erlotinib thus would support the assumption of a prognostic role for KRAS in advanced PC.
The IBM SPSS Version 23 software package (IBM Corporation, Armonk, NY, USA) was used for statistical analysis. For qualitative data and quantitative data that were dichotomised at a clinically relevant cutoff, the χ2 test was applied. Survival was estimated by the Kaplan–Meier-method. Comparison of survival was conducted by the log-rank test. Hazard ratios (HR) were obtained by Cox Regression. A P-value of <0.05 was regarded as statistically significant. OS was recorded as the time interval between first application of palliative chemotherapy and death from any cause.
KRAS/NRAS mutation analysis
Archival FFPE tumour tissue (gained by routine diagnostic procedures at the initial diagnosis or confirmation of recurrence of PC) was obtained from the participating study centres/pathologists for extended RAS analysis. Cytological specimens were not included. The majority (90%) of KRAS/NRAS mutation analyses were performed centrally at the University of Munich, Institute of Pathology. RAS mutations in exon 2 (codons 12, 13), exon 3 (codons 59, 61) and exon 4 (codons 117, 146) and NRAS mutations in exon 2 (codons 12, 13), exon 3 (codons 59, 61) and exon 4 (codons 117, 146) were investigated by established routine pyrosequencing using the PyroMark Q24 and the PyroMark Q24 2.0.6. software (Qiagen, Hilden, Germany). DNA was extracted with the help of the QIAamp DNA FFPE Tissue Kit on a QIAcube device (Qiagen). PCR primers were purchased from Biomers.net GmbH (Ulm, Germany). The primer sequences are listed in Supplementary Materials and Methods (see Supplementary Table S1). Ten percent (n=18) of the analyses of KRAS exons 2–3 were performed at the Institute of Pathology at the Technical University of Munich by high-resolution melting assay (HRMA) and Sanger sequencing, as described previously (Schlitter et al, 2014, 2015).
Results
Patients and treatments
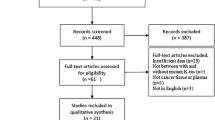
Evaluable tumour material was successfully collected from 178 patients (100 males, 56%; 78 females, 44%). In total, 122 (60%) of the analysed patients were treated in one of the three large prospective, controlled clinical trials: 59 (33%) within the trial by Heinemann et al (2006) (phase III, gemcitabine+cisplatin vs gemcitabine), 45 (25%) within the trial by Boeck et al (2008) (phase II, capecitabine+oxaliplatin vs capecitabine+gemcitabine vs gemcitabine+oxaliplatin), and 18 (10%) in the AIO-PK0104 study (preamendment) (Heinemann et al, 2013). The remaining patients were treated at the outpatient clinics at the Ludwig-Maximilian-University of Munich (n=38, 21%) or at the Technical University of Munich (n=18, 10%). Median age was 63 years (range 36–88 years). All patients were diagnosed with a pancreatic ductal adenocarcinoma, 158 (89%) had metastatic tumours and 20 patients (11%) had locally advanced disease. Most primary tumours (60%) were located in the pancreatic head, 22% in the pancreatic body and 18% in the pancreatic tail. The liver was the most frequent site of metastasis (72%), followed by the lung (15%). All included patients received at least one dose of palliative chemotherapy. Most patients received gemcitabine-based treatments (70%), thereof 35% gemcitabine alone, 24% gemcitabine+cisplatin and 10% gemcitabine+oxaliplatin. Fluoropyrimidine-based regimens were applied in 18% of patients: 8% received capecitabine alone, 8% capecitabine+oxaliplatin and 2% infusional 5-fluorouracil/leucovorin+oxaliplatin (FOLFOX). Combinations of gemcitabine and capecitabine were given in 12%.
Tumour RAS status
Considering KRAS exon 2 mutations (codons 12 and 13), 132 (74%) of the analysed tumours were mutated (see Table 1). KRAS exon 3 (codon 61) mutations were found in 9 cases (5%). Within the 160 tumours evaluated for KRAS exon 4 and in all exons of NRAS, only wild-type status was detected. Overall, in KRAS exons 2–3, a mutation rate of 79% was determined. The most common specific mutation in exon 2 was G12D (49%), followed by G12V (34%) and G12R (14%).
There were no differences in baseline patient characteristics, such as gender, age, KPS, tumour stage (locally advanced vs metastatic) and treatment (gemcitabine-based, fluoropyrimidine-based or gemcitabine+fluoropyrimidine combinations) for KRAS wild-type vs mutant tumours when all KRAS mutations were combined (exons 2 and 3) or for exon 3 only (see Table 2). Splitting the patients into subgroups by the median pretreatment CA 19-9 value (586 U ml−1) revealed a higher proportion of patients with KRAS mutant tumours in exons 2 and 3 in the CA 19-9 high subgroup (87% vs 71%, P=0.016). The origin of the tumour specimen (metastasis vs primary tumour) had no impact on the rate of RAS mutations (P=0.326).
Survival analyses according to tumour RAS status
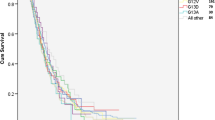
Considering KRAS exon 2 mutations, there was no difference in survival between wild-type (9.56 months; 95% CI: 4.83–14.29) and mutant tumours (8.28 months; 95% CI: 7.35–9.20; HR: 1.13, 95% CI: 0.79–1.61, P=0.51). Adding the 9 cases with mutations in KRAS exon 3 did not change these results: 9.89 months (95% CI: 5.24–14.54) vs 8.28 months (95% CI: 7.32–9.24; HR 1.08, 95% CI: 0.73–1.59, P=0.70; see Table 3 and Figure 1). The 9 patients with KRAS mutations in exon 3 had a median OS of 5.75 months (95% CI: 2.87–8.63). Considering the specific KRAS mutations in codon 12, patients with a G12R mutation had the longest median OS (11.93 months), followed by G12D (8.28 months) and G12V (7.92 months); differences in OS between those subgroups were statistically not significant (P=0.86; see Table 3).
Survival analyses according to tumour RAS status in specific subgroups
As expected, patients with locally advanced disease had a longer median OS (13.63 months) compared with patients with metastatic disease (8.28 months; HR 0.54, 95% CI 0.32–0.92, P=0.022). In locally advanced tumours, KRAS wild-type patients (n=5) had a median OS of 22.18 months, whereas KRAS mutant patients (n=15) had a median OS of 13.63 months (HR 0.55, 95% CI 0.15–1.95, P=0.34). In metastatic patients (n=158), OS for KRAS wild type (n=32) was estimated with 9.10 months vs 8.15 months for patients with a KRAS mutant tumour (HR 0.96, 95% CI 0.64–1.45, P=0.86).
CA 19-9 levels (that were determined locally at the participating centres) at baseline were available from 149 patients; the median CA 19-9 value was 586 U ml−1 (range 0.60–740 000). Twenty-three patients (15%) had a CA 19-9 value below the upper limit of normal of 37 U ml−1 with an OS of 13.18 months, 59 patients (40%) had a CA 19-9 level between 38 and 1000 U ml−1 with a survival of 9.23 months and 67 patients (45%) had CA 19-9 values >1000 U ml−1 with a median OS of 7.60 months. The differences in survival between the subgroups were not statistically significant (P=0.24). Survival according to CA 19-9 and RAS status was as follows: in the subgroup with CA 19-9 values ⩽37 U ml−1, OS for KRAS wild type (n=5) was 20.30 months vs 9.63 months for 18 KRAS mutant patients (HR 0.82, 95% CI 0.26–2.57, P=0.73). Considering the second group with CA 19-9 levels between 38 and 1000 U ml−1, KRAS wild-type patients (n=17) had a median OS of 16.14 months compared with 42 KRAS mutant patients with an OS of 8.58 months. This difference again was statistically not significant (HR 0.98, 95% CI 0.54–1.77, P=0.934). In the largest group with CA 19-9 values >1000 U ml−1, 10 patients were KRAS wild type and 57 were KRAS mutant. There was no difference in median OS between both subgroups: 7.49 months vs 7.92 months (HR 0.85, 95% CI 0.40–1.79, P=0.67).
In 124 patients (70%) who received gemcitabine or gemcitabine-based regimens, median OS was 8.31 months; the 33 patients (19%) who were treated with a fluoropyrimidine-based regimen (capecitabine, capecitabine+oxaliplatin or FOLFOX) had a median OS of 8.81 months. In 21 patients (11%), a combination of gemcitabine+capecitabine was applied resulting in a median OS of 10.97 months. The differences between the groups were not significant (P=0.30). Within the gemcitabine subgroup, OS for KRAS wild-type patients (n=31) was 9.89 months compared with 8.31 months for 93 KRAS mutant patients (HR 0.95, 95% CI 0.62–1.46, P=0.80). In the subgroup receiving fluoropyrimidine-containing chemotherapy, only two patients were KRAS wild type (OS 20.33 months), whereas 31 patients had a KRAS mutant tumour (OS 7.92 months; HR 0.33, 95% CI 0.04–2.45, P=0.25). Considering patients who received gemcitabine and capecitabine, 4 patients were classified as KRAS wild type (OS 6.57 months) and 17 patients had a tumour harbouring a KRAS mutation (OS 12.52 months; HR 2.89, 95% CI 0.86–9.75, P=0.074).
Discussion
Based on extended RAS testing (KRAS and NRAS) in advanced PC, we observed an overall mutation rate of 79%. Seventy-four percent of the mutations were detected in KRAS exon 2. The number of KRAS mutations in PC reported by other investigators varies widely between 47% and 95% (see Table 4). However, these reports also differ substantially in several patient characteristics (e.g., inclusion of resected patients, palliative and adjuvant treatment settings, Asian vs Caucasian patients) and the extent of RAS testing. KRAS mutations are described as nearly ubiquitous in PC (Waddell et al, 2015). Differences in the mutation rate may also be related to technical aspects. However, the methods used for the current analysis (pyrosequencing, Sanger sequencing and HRMA) are highly sensitive with a limit of detection (share of tumour DNA in the whole DNA extracted) of 10%, 15–20% and 10%, respectively (Ogino et al, 2005; Tsiatis et al, 2010).
Screening for additional mutated loci revealed an incidence of 5% in exon 3 (codons 59 and 61), while no mutations were found in KRAS exon 4 and in NRAS exons 2–4. Previous reports indicated a mutation frequency of 1–3% in KRAS exon 3 (codon 61) in PC (Lee et al, 2007; Kim et al, 2011; Shin et al, 2013; Wang et al, 2015). Compared with colorectal cancer, where other RAS-mutations amount to approximately 17% of mutations in KRAS exon 2 wild-type patients (Douillard et al, 2013), these more rare mutations seem to have a secondary role in PC. To our knowledge, the current report is the first to also investigate NRAS mutations in PC, and interestingly, these mutations do not appear to have a substantial role in pancreatic ductal adenocarcinoma.
Considering the frequency of specific KRAS mutations within exon 2 (49% G12D, 34% G12V and 14% G12R), our results are well comparable with data recently reported by Bournet et al (2016) (49.5%, 36% and 14.5%) or Ogura et al (2013) (42.9%, 39.7% and 7.0%), respectively. We could also observe a trend for different outcomes of the specific subgroups, which were – however – not statistically significant and differed substantially from the recently described data by Bournet et al (2016). In their analysis, patients with a G12D mutation had a significantly worse median OS (6 months) than KRAS wild-type patients (9 months), G12V (9 months) or G12R mutations (14 months). In our population, we observed the worst survival for the G12V mutation (7.9 months), followed by G12D (8.3 months) and G12R (11.9 months). Based on the limited patient numbers and the post hoc nature of the analysis, it is not possible to derive clear prognostic information from any of the subgroups. Interestingly, we could observe a higher rate of KRAS mutations in patients with CA 19-9 levels above the median value of 586 U ml−1 (87% vs 71%, P=0.017). Ogura et al (2013) also reported a higher – however, statistically not significantly different – CA 19-9 expression in KRAS-mutated tumours. The potential association between higher CA 19-9 values and KRAS mutant tumours might reflect an unfavourable tumour biology. However, this issue is controversial as Bournet et al (2016) did not observe this effect using the upper limit of normal of 37 U ml−1 as a cutoff value for CA 19-9. In our subgroup analyses, there was no difference according to KRAS status if patients were allocated to groups with CA 19-9 values <37 U ml−1, 38–1000 U ml−1 or >1000 U ml−1.
When the RAS mutational status was correlated with OS, there were no significant differences between wild-type and mutant tumours. This applied for the analysis of exon 2 as well as for the extended RAS analysis, taking into account the additional nine mutations found in KRAS exon 3. These results are in line with data recently reported by Bournet et al (2016) In 162 PC patients treated with palliative chemotherapy without the use of anti-EGFR agents, no OS difference between RAS wild-type and mutant tumours was observed (10.0 months vs 9.5 months; HR 1.02, P=0.88). Based on the observation that the RAS mutation status has no influence on outcome in PC patients receiving anti-EGFR-free treatment and given the survival benefit observed in erlotinib-treated patients with KRAS wild-type tumours, it might be concluded that a KRAS wild-type status may rather be predictive for the efficacy of erlotinib. Considering the current literature on KRAS in PC (summarised within Table 4), the available reports are controversial. However, most of the work so far consists of retrospective analyses missing a clear prospective design and sample size calculation to further elucidate this question.
There are only two randomised studies with data on KRAS status with different treatment arms for gemcitabine alone and gemcitabine and erlotinib (da Cunha Santos et al, 2010; Wang et al, 2015). Wang et al (2015) performed an analysis on the effect of KRAS on all patients only, finding no effect on efficacy; however, the authors did not report the outcome for the different treatment arms. This study furthermore had a low rate of KRAS wild-type patients (5.7%, 5 out of 88 patients), thus the power to detect a predictive role of KRAS in this randomised trial was too low and was consequently not further followed by the authors. In a small molecular subset analysis (n=117 out of 569) from the pivotal PA.3 study, no (statistically significant) survival difference was observed for KRAS wild-type patients between the two treatment arms (gemcitabine+erlotinib vs gemcitabine+placebo): 6.1 vs 4.5 months, HR 0.66, 95% CI 0.28–1.7, P=0.34 (da Cunha Santos et al, 2010). There furthermore was no significant difference for KRAS wild-type vs mutant tumours in the gemcitabine+placebo arm (4.46 vs 7.36 months, HR 0.68, 95% CI 0.33–12.42, P=0.30). This analysis is, however, limited by its retrospective nature and the rather small patient number.
Three other groups chose a similar approach as ours analysing patients with erlotinib-free regimens in palliative first-line treatment: Bournet et al (2016), Ogura et al (2013), and Lee et al (2007). Of note, the findings of Bournet et al (2016) as well as our results are in contrast to the Asian reports: Ogura et al (2013) performed KRAS testing within exon 2 in 242 patients and found a significant advantage in OS for KRAS wild-type patients (479 vs 255 days, P=0.03). However, their analysis included also patients with best supportive care only and accepted also radiation therapy as treatment option, thus the possibility to compare these results directly to those of Bournet et al (2016 and ours is limited. The second analysis by Lee et al (2007) showed an improved OS for patients with KRAS wild-type tumours (13.4 vs 9.1 months, P=0.03), however, was already carried out in 2007, consisted of only 66 Asian patients and had a rather low proportion of KRAS-mutated tumours (64%).
Other reports on KRAS in patients with resectable PC are hardly comparable to our analysis as they cover a completely different therapeutic situation. Furthermore, the studies by Kwon et al (2015), Sinn et al (2014), Shin et al (2013) and Oliveira-Cunha et al (2012) do not deliver information on systemic treatment (especially with EGFR-directed agents; for details, see Table 4). Also the latest meta-analysis reported by Tao et al (2016) is not appropriate for this issue as patients in different clinical settings (surgical treatment, no treatment, chemotherapy treatment, etc.) were mixed up. Their subgroup analysis with chemotherapy-only patients furthermore did not distinguish between anti-EGFR-free and anti-EGFR-containing treatment regimens.
Conclusion
In conclusion, the potential prognostic or predictive role of KRAS mutations in PC currently remains a matter of debate. However, the current study – at least to our knowledge – represents the largest analysis in a homogeneous patient population with an upfront statistical design in this setting, showing no statistically significant impact of KRAS mutation status on OS in an erlotinib-free patient population. Considering the differences in the outcome observed by the AIO-PK0104 study in patients receiving erlotinib (Boeck et al, 2013), one might support the hypothesis for a predictive role of KRAS for the efficacy of erlotinib (or other anti-EGFR-targeting agents). A main limitation of the current data set is based on the fact that we included patients from three different trials and two tumour registries. However, the conclusion that we were not able to find evidence (also within different subgroup analyses) for a prognostic role of RAS in the analysed patient population is – at least in our opinion – robust. The subsequent ‘cross-trial’ comparison to AIO-PK0104 should of course be regarded with caution, as the role of RAS as a prognostic vs predictive biomarker in advanced PC could only be defined within a solely prospective, randomised biomarker trial.
Change history
23 May 2017
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Amado RG, Wolf M, Peeters M, Van Cutsem E, Siena S, Freeman DJ, Juan T, Sikorski R, Suggs S, Radinsky R, Patterson SD, Chang DD (2008) Wild-type KRAS is required for panitumumab efficacy in patients with metastatic colorectal cancer. J Clin Oncol 26 (10): 1626–1634.
Boeck S, Hoehler T, Seipelt G, Mahlberg R, Wein A, Hochhaus A, Boeck HP, Schmid B, Kettner E, Stauch M, Lordick F, Ko Y, Geissler M, Schoppmeyer K, Kojouharoff G, Golf A, Neugebauer S, Heinemann V (2008) Capecitabine plus oxaliplatin (CapOx) versus capecitabine plus gemcitabine (CapGem) versus gemcitabine plus oxaliplatin (mGemOx): final results of a multicenter randomized phase II trial in advanced pancreatic cancer. Ann Oncol 19 (2): 340–347.
Boeck S, Jung A, Laubender RP, Neumann J, Egg R, Goritschan C, Vehling-Kaiser U, Winkelmann C, Fischer von Weikersthal L, Clemens MR, Gauler TC, Marten A, Klein S, Kojouharoff G, Barner M, Geissler M, Greten TF, Mansmann U, Kirchner T, Heinemann V (2013) EGFR pathway biomarkers in erlotinib-treated patients with advanced pancreatic cancer: translational results from the randomised, crossover phase 3 trial AIO-PK0104. Br J Cancer 108 (2): 469–476.
Bournet B, Muscari F, Buscail C, Assenat E, Barthet M, Hammel P, Selves J, Guimbaud R, Cordelier P, Buscail L (2016) KRAS G12D mutation subtype is a prognostic factor for advanced pancreatic adenocarcinoma. Clin Transl Gastroenterol 7: e157.
Conroy T, Desseigne F, Ychou M, Bouche O, Guimbaud R, Becouarn Y, Adenis A, Raoul JL, Gourgou-Bourgade S, de la Fouchardiere C, Bennouna J, Bachet JB, Khemissa-Akouz F, Pere-Verge D, Delbaldo C, Assenat E, Chauffert B, Michel P, Montoto-Grillot C, Ducreux M Groupe Tumeurs Digestives of Unicancer, PRODIGE Intergroup (2011) FOLFIRINOX versus gemcitabine for metastatic pancreatic cancer. N Engl J Med 364 (19): 1817–1825.
da Cunha Santos G, Dhani N, Tu D, Chin K, Ludkovski O, Kamel-Reid S, Squire J, Parulekar W, Moore MJ, Tsao MS (2010) Molecular predictors of outcome in a phase 3 study of gemcitabine and erlotinib therapy in patients with advanced pancreatic cancer: National Cancer Institute of Canada Clinical Trials Group Study PA.3. Cancer 116 (24): 5599–5607.
Douillard JY, Oliner KS, Siena S, Tabernero J, Burkes R, Barugel M, Humblet Y, Bodoky G, Cunningham D, Jassem J, Rivera F, Kocakova I, Ruff P, Blasinska-Morawiec M, Smakal M, Canon JL, Rother M, Williams R, Rong A, Wiezorek J, Sidhu R, Patterson SD (2013) Panitumumab-FOLFOX4 treatment and RAS mutations in colorectal cancer. N Engl J Med 369 (11): 1023–1034.
Heinemann V, Quietzsch D, Gieseler F, Gonnermann M, Schonekas H, Rost A, Neuhaus H, Haag C, Clemens M, Heinrich B, Vehling-Kaiser U, Fuchs M, Fleckenstein D, Gesierich W, Uthgenannt D, Einsele H, Holstege A, Hinke A, Schalhorn A, Wilkowski R (2006) Randomized phase III trial of gemcitabine plus cisplatin compared with gemcitabine alone in advanced pancreatic cancer. J Clin Oncol 24 (24): 3946–3952.
Heinemann V, Vehling-Kaiser U, Waldschmidt D, Kettner E, Marten A, Winkelmann C, Klein S, Kojouharoff G, Gauler TC, von Weikersthal LF, Clemens MR, Geissler M, Greten TF, Hegewisch-Becker S, Rubanov O, Baake G, Hohler T, Ko YD, Jung A, Neugebauer S, Boeck S (2013) Gemcitabine plus erlotinib followed by capecitabine versus capecitabine plus erlotinib followed by gemcitabine in advanced pancreatic cancer: final results of a randomised phase 3 trial of the ‘Arbeitsgemeinschaft Internistische Onkologie’ (AIO-PK0104). Gut 62 (5): 751–759.
Kim ST, Lim do H, Jang KT, Lim T, Lee J, Choi YL, Jang HL, Yi JH, Baek KK, Park SH, Park YS, Lim HY, Kang WK, Park JO (2011) Impact of KRAS mutations on clinical outcomes in pancreatic cancer patients treated with first-line gemcitabine-based chemotherapy. Mol Cancer Ther 10 (10): 1993–1999.
Kruger S, Haas M, Ormanns S, Bachmann S, Siveke JT, Kirchner T, Heinemann V, Boeck S (2014) Translational research in pancreatic ductal adenocarcinoma: current evidence and future concepts. World J Gastroenterol 20 (31): 10769–10777.
Kwon MJ, Jeon JY, Park HR, Nam ES, Cho SJ, Shin HS, Kwon JH, Kim JS, Han B, Kim DH, Choi YL (2015) Low frequency of KRAS mutation in pancreatic ductal adenocarcinomas in Korean patients and its prognostic value. Pancreas 44 (3): 484–492.
Lee J, Jang KT, Ki CS, Lim T, Park YS, Lim HY, Choi DW, Kang WK, Park K, Park JO (2007) Impact of epidermal growth factor receptor (EGFR) kinase mutations, EGFR gene amplifications, and KRAS mutations on survival of pancreatic adenocarcinoma. Cancer 109 (8): 1561–1569.
Moore MJ, Goldstein D, Hamm J, Figer A, Hecht JR, Gallinger S, Au HJ, Murawa P, Walde D, Wolff RA, Campos D, Lim R, Ding K, Clark G, Voskoglou-Nomikos T, Ptasynski M, Parulekar W National Cancer Institute of Canada Clinical Trials Group (2007) Erlotinib plus gemcitabine compared with gemcitabine alone in patients with advanced pancreatic cancer: a phase III trial of the National Cancer Institute of Canada Clinical Trials Group. J Clin Oncol 25 (15): 1960–1966.
Ogino S, Kawasaki T, Brahmandam M, Yan L, Cantor M, Namgyal C, Mino-Kenudson M, Lauwers GY, Loda M, Fuchs CS (2005) Sensitive sequencing method for KRAS mutation detection by pyrosequencing. J Mol Diagn 7 (3): 413–421.
Ogura T, Yamao K, Hara K, Mizuno N, Hijioka S, Imaoka H, Sawaki A, Niwa Y, Tajika M, Kondo S, Tanaka T, Shimizu Y, Bhatia V, Higuchi K, Hosoda W, Yatabe Y (2013) Prognostic value of K-ras mutation status and subtypes in endoscopic ultrasound-guided fine-needle aspiration specimens from patients with unresectable pancreatic cancer. J Gastroenterol 48 (5): 640–646.
Oliveira-Cunha M, Hadfield KD, Siriwardena AK, Newman W (2012) EGFR and KRAS mutational analysis and their correlation to survival in pancreatic and periampullary cancer. Pancreas 41 (3): 428–434.
Rahib L, Smith BD, Aizenberg R, Rosenzweig AB, Fleshman JM, Matrisian LM (2014) Projecting cancer incidence and deaths to 2030: the unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res 74 (11): 2913–2921.
Rosell R, Carcereny E, Gervais R, Vergnenegre A, Massuti B, Felip E, Palmero R, Garcia-Gomez R, Pallares C, Sanchez JM, Porta R, Cobo M, Garrido P, Longo F, Moran T, Insa A, De Marinis F, Corre R, Bover I, Illiano A, Dansin E, de Castro J, Milella M, Reguart N, Altavilla G, Jimenez U, Provencio M, Moreno MA, Terrasa J, Munoz-Langa J, Valdivia J, Isla D, Domine M, Molinier O, Mazieres J, Baize N, Garcia-Campelo R, Robinet G, Rodriguez-Abreu D, Lopez-Vivanco G, Gebbia V, Ferrera-Delgado L, Bombaron P, Bernabe R, Bearz A, Artal A, Cortesi E, Rolfo C, Sanchez-Ronco M, Drozdowskyj A, Queralt C, de Aguirre I, Ramirez JL, Sanchez JJ, Molina MA, Taron M, Paz-Ares L Spanish Lung Cancer Group in collaboration with Groupe Francais de Pneumo-Cancérologie, Associazione Italiana Oncologia Toracica (2012) Erlotinib versus standard chemotherapy as first-line treatment for European patients with advanced EGFR mutation-positive non-small-cell lung cancer (EURTAC): a multicentre, open-label, randomised phase 3 trial. Lancet Oncol 13 (3): 239–246.
Schlitter AM, Born D, Bettstetter M, Specht K, Kim-Fuchs C, Riener MO, Jeliazkova P, Sipos B, Siveke JT, Terris B, Zen Y, Schuster T, Hofler H, Perren A, Kloppel G, Esposito I (2014) Intraductal papillary neoplasms of the bile duct: stepwise progression to carcinoma involves common molecular pathways. Mod Pathol 27 (1): 73–86.
Schlitter AM, Jang KT, Kloppel G, Saka B, Hong SM, Choi H, Offerhaus GJ, Hruban RH, Zen Y, Konukiewitz B, Regel I, Allgauer M, Balci S, Basturk O, Reid MD, Esposito I, Adsay V (2015) Intraductal tubulopapillary neoplasms of the bile ducts: clinicopathologic, immunohistochemical, and molecular analysis of 20 cases. Mod Pathol 28 (9): 1249–1264.
Shin SH, Kim SC, Hong SM, Kim YH, Song KB, Park KM, Lee YJ (2013) Genetic alterations of K-ras, p53, c-erbB-2, and DPC4 in pancreatic ductal adenocarcinoma and their correlation with patient survival. Pancreas 42 (2): 216–222.
Sinn BV, Striefler JK, Rudl MA, Lehmann A, Bahra M, Denkert C, Sinn M, Stieler J, Klauschen F, Budczies J, Weichert W, Stenzinger A, Kamphues C, Dietel M, Riess H (2014) KRAS mutations in codon 12 or 13 are associated with worse prognosis in pancreatic ductal adenocarcinoma. Pancreas 43 (4): 578–583.
Tao LY, Zhang LF, Xiu DR, Yuan CH, Ma ZL, Jiang B (2016) Prognostic significance of K-ras mutations in pancreatic cancer: a meta-analysis. World J Surg Oncol 14 (1): 146.
Tsiatis AC, Norris-Kirby A, Rich RG, Hafez MJ, Gocke CD, Eshleman JR, Murphy KM (2010) Comparison of Sanger sequencing, pyrosequencing, and melting curve analysis for the detection of KRAS mutations: diagnostic and clinical implications. J Mol Diagn 12 (4): 425–432.
Von Hoff DD, Ervin T, Arena FP, Chiorean EG, Infante J, Moore M, Seay T, Tjulandin SA, Ma WW, Saleh MN, Harris M, Reni M, Dowden S, Laheru D, Bahary N, Ramanathan RK, Tabernero J, Hidalgo M, Goldstein D, Van Cutsem E, Wei X, Iglesias J, Renschler MF (2013) Increased survival in pancreatic cancer with nab-paclitaxel plus gemcitabine. N Engl J Med 369 (18): 1691–1703.
Waddell N, Pajic M, Patch AM, Chang DK, Kassahn KS, Bailey P, Johns AL, Miller D, Nones K, Quek K, Quinn MC, Robertson AJ, Fadlullah MZ, Bruxner TJ, Christ AN, Harliwong I, Idrisoglu S, Manning S, Nourse C, Nourbakhsh E, Wani S, Wilson PJ, Markham E, Cloonan N, Anderson MJ, Fink JL, Holmes O, Kazakoff SH, Leonard C, Newell F, Poudel B, Song S, Taylor D, Waddell N, Wood S, Xu Q, Wu J, Pinese M, Cowley MJ, Lee HC, Jones MD, Nagrial AM, Humphris J, Chantrill LA, Chin V, Steinmann AM, Mawson A, Humphrey ES, Colvin EK, Chou A, Scarlett CJ, Pinho AV, Giry-Laterriere M, Rooman I, Samra JS, Kench JG, Pettitt JA, Merrett ND, Toon C, Epari K, Nguyen NQ, Barbour A, Zeps N, Jamieson NB, Graham JS, Niclou SP, Bjerkvig R, Grutzmann R, Aust D, Hruban RH, Maitra A, Iacobuzio-Donahue CA, Wolfgang CL, Morgan RA, Lawlor RT, Corbo V, Bassi C, Falconi M, Zamboni G, Tortora G, Tempero MA . Australian Pancreatic Cancer Genome Initiative Gill AJ, Eshleman JR, Pilarsky C, Scarpa A, Musgrove EA, Pearson JV, Biankin AV, Grimmond SM (2015) Whole genomes redefine the mutational landscape of pancreatic cancer. Nature 518 (7540): 495–501.
Wang JP, Wu CY, Yeh YC, Shyr YM, Wu YY, Kuo CY, Hung YP, Chen MH, Lee WP, Luo JC, Chao Y, Li CP (2015) Erlotinib is effective in pancreatic cancer with epidermal growth factor receptor mutations: a randomized, open-label, prospective trial. Oncotarget 6 (20): 18162–18173.
Acknowledgements
The authors would like to thank all patients and their families, nurses, study coordinators and investigators for their active participation in the clinical trials that contributed data to this translational study. This work is part of the doctoral thesis of Sibylle Baechmann and Anna Remold.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
No direct payment was received for the current work by a company or any third parties. The follwing authors declare a potential conflict of interest: Michael Haas: Honoraria for scientific presentations: Celgene, Research funding: Roche, Boehringer, Travel accommodation expenses: Boehringer, Celgene. Andreas Jung: Honoraria for scientific presentations: Amgen, AstraZeneca, Merck, Qiagen, Roche, Travel accommodation expenses: Amgen, AstraZeneca, Merck, Qiagen, Roche. Stefan Boeck: Honoraria for scientific presentations: Roche, Celgene, Consulting or Advisory Board: Celgene, Baxalta, Research funding: Roche, Celgene, Clovis Oncology, Travel accommodation expenses: Celgene, Roche. Volker Heinemann: Honoraria for scientific presentations: Merck, Roche, Celgene, Consulting or Advisory Board: Merck, Roche Celgene, Boehringer, AMGEN, Research funding: Merck, AMGEN, Roche Celgene, Boehringer, Travel accommodation expenses: Merck, Roche. All other authors declare that they have no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Haas, M., Ormanns, S., Baechmann, S. et al. Extended RAS analysis and correlation with overall survival in advanced pancreatic cancer. Br J Cancer 116, 1462–1469 (2017). https://doi.org/10.1038/bjc.2017.115
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2017.115
- Springer Nature Limited
Keywords
This article is cited by
-
TPX2 expression as a negative predictor of gemcitabine efficacy in pancreatic cancer
British Journal of Cancer (2023)
-
Oncogenic KRAS blockade therapy: renewed enthusiasm and persistent challenges
Molecular Cancer (2021)
-
Bacterial lipopolysaccharide as negative predictor of gemcitabine efficacy in advanced pancreatic cancer – translational results from the AIO-PK0104 Phase 3 study
British Journal of Cancer (2020)
-
POLE gene hotspot mutations in advanced pancreatic cancer
Journal of Cancer Research and Clinical Oncology (2018)