Abstract
Selecting sorghum genotypes with higher grain yield and nutritional quality is essential to tackle food insecurity and malnutrition in arid and semi-arid areas. Therefore, this study aimed to determine the genetic diversity, trait association and genotype by yield by trait (GYT) analysis and to select superior sorghum genotypes. One hundred and ten sorghum genotypes were evaluated at three locations in Tigray during the 2018 and 2019 growing seasons using alpha lattice design. Traits such as grain yield, protein content, ash content, starch content, zinc content, iron content, calcium content and magnesium content were profiled. Results showed that wide range and highly significant (p < 0.001) genotype mean performance in each environment as well as combined environments. Several highly performing genotypes were distinguished for each trait studied that could be exploited as breeding parents or direct use. This study further detected highly significant variation (p < 0.001) among the test genotypes for all the traits studied in individual environments and across environments suggesting the presence of sufficient genetic diversity for selection. The high broad-sense heritability (H2 > 0.9) in all individual environments and moderate to high (H2 > 0.0.41 < 0.82) in pooled environments recorded in the present study assured the possibility of effective selection among the genotypes. Besides, strong positive and negative associations were detected between some of the traits in individual and across environments. The significant positive association between traits indicates that both the traits can be improved concurrently through direct selection. Using the GYT analysis, we suggest ten promising sorghum genotypes for direct use or breeding programs in arid and semi-arid areas in general and in Tigray in particular.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Core idea
-
Wide range and higher mean performances of sorghum genotypes were attained in individual and combined environments in Tigray
-
Considerable genetic variations are available among sorghum genotypes in Tigray
-
All the traits studied had higher heritability in individual environments and medium to high heritability in combined environments
-
Significant positive and negative correlations were detected between some of the traits in each individual and combined environments
-
Several superior genotypes of sorghum were selected for all traits studied that could be exploited as breeding parents for sorghum improvement
-
Ten promising sorghum genotypes were selected using the GYT approach for direct use or breeding programs
2 Introduction
Sorghum (Sorghum bicolor [L.] Moench 2n = 2x = 20), a C4 photosynthetic crop, is among the five top cereal crops that feed the world [1]. In Africa, sorghum is the fourth most widely cultivated cereal crop after maize, rice, and wheat and second in area coverage next to maize [1]. Sorghum possesses a wide array of ecological adaptations ranging from the dry lowland areas of the semi-arid tropics to high rainfall and humid regions, as well as in the more temperate regions of the world [2]. It is a staple food for more than half a billion people in Sub-Saharan Africa and Asia [3]. Sorghum is the second cheapest source of energy next to finger millet [4] and its grains are good sources of various minerals, vitamins, carbohydrates and proteins essential for human well-being [5,6,7]. Besides, sorghum is gluten-free [8], which makes it an excellent food crop for people who have gluten intolerance. Due to its high nutritional value, the market interest in sorghum grains is markedly growing worldwide [9].
Sorghum is the most diversified crop having 28 domesticated species categorized into five basic races i.e. bicolor, guinea, caudatum, dura and kafir and ten hybrid races [10]. All races of sorghum are grown across the diverse agroecological zones and farming systems of Ethiopia [11, 12]. Sorghum is the first fully assembled C4 crop using traditional Sanger sequencing [12] and is considered as ‘gold standard’ reference due to its high percentage assembly [13]. The broad genetic base harbored mainly in sorghum landraces is essential for providing the gene pool necessary for the development and deployment of farmers' preferred varieties. Hence, collection and assembly followed by thoughtful quantification of the genetic diversity of such a collection are essential for efficient crop improvement and conservation programs.
Ethiopia is the world’s fourth largest sorghum producer after the United States, Nigeria and Mexico with a total annual production of 4.45 million metric tons [14]. In Ethiopia, sorghum stands fourth in terms of production after maize, wheat and teff, and third in terms of area coverage next to teff and maize [15]. The crop serves as the main stable food for the vast population of the country. It is mainly cultivated in rain-fed and low-input environments, where acute shortage of food and malnutrition prevails. Ethiopian farmers grow diverse forms of sorghum varieties with over 95% of the area allocated for sorghum production covered by landraces [16] Landraces are heterogeneous and adapted to diverse growing conditions and are potential sources of desirable traits for crop improvement [17, 18]. Given its center of origin and diversity [19, 20], and tremendous variation in agro-ecologies, Ethiopia is endowed with extremely rich genetic diversity of sorghum landraces [2, 16, 21, 22], which serves as sources of various important traits. For instance, drought tolerance [16, 23, 24], nutritional quality traits [21, 25, 26], resistance to grain mold [27] and resistant to ergot and green bug [28]. Ethiopia is also among the top nations that have made significant contributions to the sorghum germplasm collections in the world [29]. Moreover, there is still unexplored high genetic variability in Ethiopian sorghum landraces [2, 24].
Selecting genotypes with improved yield and nutritional quality, especially with respect to mineral content would have a significant contribution to intervening the high prevalence of food shortage and malnutrition, especially in arid and semi-arid areas where sorghum is the stable food [30, 31]. Nonetheless, selecting in one trait may result in the reduced level of the other one or more target traits [32], which hinders the progress of crop improvement. To address the challenges of unfavorable association of key traits, Yan and Frégeau-Reid (2018) introduced the genotype by trait by yield (GYT) analysis approach. The GYT approach is a new novel tool to identify superior genotypes based on multiple traits such that grain yield is the main trait that determines the usefulness of the genotypes while the other target traits are valued based on their merit in combining with grain yield [33]. The average tester coordinator (ATC) graph of the GYT biplot meaningfully rank the genotypes based on multiple yield by trait combinations and at the same time shows the strengths and weaknesses of the genotypes [33]. GYT analysis has been widely applied to identify superior genotypes in various crops such as oat [34], wheat [35], cotton [36] and sorghum [37].
Although numerous publications are available regarding the high genetic diversity of sorghum in Ethiopia, most of the studies are concentrated on yield and other agronomic traits with less emphasis on the genetic variability for nutritional quality traits. Genetic variation of local landraces of sorghum in Tigray Region of Ethiopia has not been studied adequately using agronomic and nutritional quality traits. Moreover, there is no research conducted so far to evaluate sorghum genotypes based on the genotype by yield by trait (GYT) approach using grain yield and nutritional quality traits. The objectives of the study were to assess genetic diversity and association in grain yield, starch, protein, ash, zinc (Zn); iron (Fe), calcium (Ca) and magnesium (Mg) contents, and to identify superior genotypes for grain yield and nutritional quality.
3 Materials and methods
3.1 Planting materials and study locations
The study used 108 sorghum landraces collected from Tigray, northern Ethiopia along with two improved varieties (Melkam and Dekeba) obtained from Tigray Agricultural Research Institute (TARI). The field trials were conducted at three locations (Tahtay Adyabo, Tselemti, and Mereb Leke) of Tigray in the 2018 and 2019 main growing seasons. Tigray is located in the northern part of Ethiopia between 12°15ʹN-14°15ʹN latitude and 36°28ʹE-39°59ʹE longitude (Table 1). About 53% of Tigray region is lowlands, 39% mid-altitude and the rest 8% percent is classified as highlands, and the altitude ranges from 500 m above sea level in the northeast to almost 4000 in the southwest [38]. The agro-ecological description of the study sites is provided in Table 1.
3.2 Field evaluation
The descriptions of the genetic materials used in the study are presented in Supplementary Table S1. The experimental layout used was an alpha lattice design with two replications at each experimental site. On each plot, the seeds were sown in a single 5 m long row. The spacing between rows and between plants within a row were 0.75 and 0.25 m, respectively. The land was tilled twice-using traditional oxen-drawn plows, and seedbed preparation, planting and harvesting were done manually. All other agronomic practices were applied as per the recommendations for the area. After harvesting at physiological maturity, panicles were sun-dried for 15 days. Thereafter, panicles from each plot were threshed, winnowed, and seeds packed.
3.3 Data collection
Data on grain yield was collected from each plot and converted to tone per hectare (t ha−1). A composite seed sample of one kg from each accession was tagged and packed in cloth bags. Thereafter, the samples were taken to Mekelle University, Ethiopia, for grain nutritional quality analysis, using Near Infra-Red Spectrophotometers (DA 7250 Perkin) method. The NIRS machine was calibrated and standardized for grain analysis of sorghum by the technicians at Mekelle University. The grain samples were bulked from the two replications and each bulked samples were measured twice. This method is a time- and cost-effective method to analyze grain quality [39]. The traits such as protein, ash and starch content of the sorghum seeds were expressed in terms of percentages, whereas Fe, Zn, Ca and Mg concentrations were expressed in terms of parts per million (ppm).
3.4 Data analysis
The data collected on grain yield and nutritional traits of one hundred and eight sorghum landraces along with the two improved varieties were analyzed using linear mixed models in META-R software [40]. Each location and year were treated as an independent environment. The mean performance, association among the traits, variances and heritability in individual environment (Tahtay Adyabo 2018, Mereb-Leke 2018, Tselemti 2018, Tahtay Adyabo 2019; Mereb-Leke 2019, and Tselemti 2019, hereafter referred as En1, En2, En3, En4, En5 and En6, respectively), and combined across the testing environments were computed using the models implemented in linear mixed effect model in R (lmer) package from package lme4 of R. The models used for this study are described below [41].
The model for the analysis of variance in individual environments is given as
\(Y_{ijkl} = \mu + rep_{i} + block_{k} (rep_{j} ) + gen_{j} + \varepsilon_{ijk}\).
where Yijk = the trait of interest, µ = the mean effect, repi = the effect of the ith replicate, blockj (repi) = the effect of the jth incomplete block within the ith replicate, genk = the effect of the kth genotype, ijk = the error associated with the ith replication, jth incomplete block and the kth genotype.
For the combined analysis, new terms are added to the model for the individual environment as:
where envi and gnvi × genl = the effects of the ith environment and the environment by genotype interaction, respectively.
Broad-sense heritability (H2) in individual and across environments were estimated using the formula:
and
where σ2g, σ2e, σ2ge, nr and nEnvs = the genotype variance, error variance, genotype by environment interaction variance, number of replication and number of environments, respectively.
The least of significance differences (LSD) at 5% of significance was calculated as:
where t = the cumulative Student’s t distribution, 0.05 = the selected α level (5%), dfErr = the degrees of freedom for error in the linear mixed model, and ASED = the average standard error of the differences of the means.
The coefficient of variation was calculated using the formula:
where MSE = the mean squared error.
3.5 Genotype by yield by trait (GYT) analysis
GYT analysis was done following the steps described by Yan and Frégeau-Reid (2018). First, the Genotype by Trait (GT) two-way table was converted to GYT two-way table (Supplementary Table S2). This was done by multiplying grain yield by each trait (grain yield *starch, grain yield*protein, grain yield*ash content, grain yield*zinc, grain yield*iron, grain yield*calcium and grain yield*magnesium, hereafter, referred to as gy*sta, gy*pro, gy*ash, gy*Zn, gy *Fe, gy*Ca and gy*Mg, respectively. Then after, the GYT table was standardized to mean zero and unity variance to minimize biases due to differences using the formula:
where: Pij = the standardized value of genotype i for a trait or yield-trait combination j in the standardized table, Tij = the original value of genotype i for yield-trait combination j in the GY*T table, Tj = the mean across genotypes for yield-trait combination j, and Sj = the standard deviation for yield-trait combination j.
The standardized GYT tables for each trait were used to generate biplots using GGEBiplot GUI package in R-software version 4.0.1 [42]. The biplots were based on singular value decomposition of trait-standardized data (scaled by standard deviation, centered by tester-centered G + E and trait-focused symmetrical singular value partition). For clarity purposes, we use only the numbers of the genotype with the prefix +. For example, LR1, was replaced to + 1; LR80 was replaced to + 80; LR100 was replaced to + 100 and so on for GYT biplot constructions.
The following equation was used to construct the GYT biplot:
where Ϛi1 and Ϛi2 = eigenvalues for PC1 and PC2, respectively, for genotype Г1j; Г2j = eigenvalues for PC1 and PC2, respectively for yield-trait combination (or trait) j, and εij = residual from fitting the PC1 and PC2 for genotype i on yield-trait combination j; \(\lambda_{1}^{\alpha }\) and \(\lambda_{2}^{\alpha }\) = singular values for PC1 and PC2, respectively, and \(\alpha\) = singular value partitioning factor. When α = 1 (i.e., SVP = 1), the biplot is said to be genotype-focused and is suitable for comparing genotypes. When α = 0 (i.e., SVP = 2), the biplot is said to be environment-focused and is suitable for visualizing correlations among environments. The scalar d is chosen such that the length of the longest vector among genotypes equals to that among environments; this is important for generating a functional biplot [43].
4 Results
4.1 Mean performances of the genotypes
The average mean performance of the genotypes for all the traits across the six environments (three locations and two years) is presented in Table 2. The result showed that highly significant variation (p < 0.001) among genotypes in all traits studied. Mean grain yield was 2.7 t ha−1, with a range of 1.4–4.1 t ha−1. The highest yielding genotypes were LR106, LR102, LR25, LR75 and LR23. The average starch, protein, ash, zinc, iron, calcium and magnesium contents were 65.1%, 10.3%, 1.9%, 35.3 ppm, 36.8 ppm, 243 ppm, and 1096.8 ppm, respectively. The highest starch, protein, ash, zinc, iron, calcium and magnesium contents were depicted by LR14 (69.4%), LR65 (11.2%), LR10 (2.3%), LR23 (43.7 ppm), LR38 (47.8 ppm), LR16 (289 ppm) and LR4 (1345.6 ppm), respectively. The minimum values for each of the traits were 1.4 t ha−1, 59.1%, 9.5%, 1.5%, 29.7 ppm, 26.5 ppm, 210.2 ppm and 932 ppm for grain yield, starch, protein, ash, zinc iron, calcium and magnesium, respectively (Table 2).
4.2 Variances and broad-sense heritability
Table 3 demonstrates genotypic variance and heritability for grain yield and nutritional quality traits of the sorghum genotypes in each testing environment. All traits showed significant genotypic variances in all environments. All measured traits showed the highest genotypic variances in En1 except for grain yield, which showed the highest genotype variance in En4. On the contrary, the lowest genotypic variance due to starch, ash, zinc and magnesium content was observed in En3, while the lowest genotype variance due to grain yield, protein and calcium was recorded in En4, En2 and En6, respectively (Table 3).
Combined analyses of variances presented in Table 4 showed highly significant variances (p < 0.001) due to genotypes (σ2g) and genotype by environment interaction (σ2g x e) effects for all studied traits. Similarly, highly significant environmental variances (σ2e) were observed for all traits, except for calcium, which was significant (p < 0.05). The variance due to genotype by environment interaction effect was higher than σ2g and σ2e for all the traits studied except for grain yield in which the variance due to genotype was relatively higher than the σ2g x e and σ2e.
In the present study, all traits were highly heritable (H2 > 0.9) as per the established scale [44] in all individual environments (Table 3). However, a partitioned genotype by environment interaction component reduced the broad-sense heritability across environments (pooled analysis), which ranged from H2 = 0.41 (protein content) to H2 = 0.82 (grain yield) (Table 4).
4.3 Correlation among traits
Pearson’s correlation coefficients among the traits were computed for each environment and across environments using the best linear unbiased prediction (BLUP) mean (Table 5). Some of the traits were significantly (p < 0.001; 0.01; 0.05) correlated while many of them had non-significant correlations (p > 0.05). Protein content had highly significant and positive association with starch (En1, p < 0.01), calcium (En2, p < 0.001 and En3, p < 0.001), magnesium (En3, p < 0.001) contents, and grain yield (En4, p < 0.05; En5, p < 0.05 and En6, p < 0.01). On the other hand, protein content showed a significant and negative association with ash (En1, p < 0.001 and En2, p < 0.05), iron (En1, p < 0.01 and En2, p < 0.001), and zinc (En1, p < 0.01; En2, p < 0.001 and En4, En2, p < 0.05). Ash content showed significant positive association with zinc (En1, En2, p < 0.001; En4, p < 0.001 and En5, p < 0.05) and iron concentrations (En1, p < 0.001; En2, p < 0.001; En4, p < 0.001 and En5, p < 0.05). Similar to protein content, ash content also had a significant and negative association with some traits such as calcium and zinc concentration (En3, p < 0.01). Zinc content exhibited a significant (p < 0.001) and positive correlation with iron in En1, En2, En4 and En5, and significant negative (p < 0.05) associations in En3 and En6. A significant negative (p < 0.05) association was observed between calcium and iron contents in En2 and En6, whereas a significant and positive (p < 0.05) association was found between calcium and grain yield in En4 and En6. Grain yield and magnesium showed a significant positive (p < 0.05) association in En6 only.
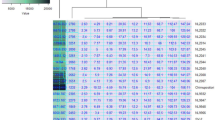
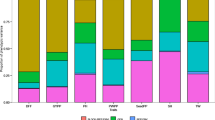
Correlations among the studied traits across the six test environments are presented in Table 6 and Fig. 1. Ash content showed a highly significant and positive (p < 0.001) correlation with zinc and iron contents. Besides, zinc and iron, starch and protein, and zinc and grain yield showed highly significant positive correlations, while calcium and iron had positive significant (p < 0.05) correlation with grain yield and starch, respectively (Table 6). On the other hand, protein had a highly significant (p < 0.001) negative association with iron concentration and significant (p < 0.05) association with ash and zinc contents. The rest traits had no significant association (p > 0.05) (Table 6).
4.4 Genotype by yield by trait (GYT) analysis
In this study, we determine how the grain yield of sorghum is combined with other important nutritional traits using the Pearson correlations coefficient (Table 7) and GYT biplots (Fig. 2) using the combined mean data. Highly significant positive correlations (p < 0.001) were detected among all grain yield-nutritional trait combinations, with correlation coefficients ranging from 0.53 to 0.89. The strongest multi-trait correlation was manifested between grain yield*starch and grain yield*protein (r = 0.89), grain yield*starch and grain yield*calcium (r = 0.87) and grain yield*starch and grain yield * magnesium (r = 0.86). The strong positive correlation between the trait combinations is also indicated by the acute angles between the vectors of the traits on the scatter plot generated from genotype by yield by trait data (Fig. 2A). The first two PCs of the GYT biplot (Fig. 2) explained about 87.5% (PC1 = 78.34%, PC2 = 9.2%) of the total variation. The polygon of which won where/what (Fig. 2B) divided the biplot into six sectors and the yield trait combinations are located in three sectors only. The ATC view of the GYT biplot (Fig. 2C) shows the weakness and strengths of the genotypes and graphically ranks the genotypes based on their overall superiority. Furthermore, top ten promising and five bottom sorghum genotypes were distinguished using the over overall superiority index of the GYT as presented in Table 8 below.
5 Discussions
In the present study, we found highly significantly differences among genotypes for grain yield and nutritional traits across all environments (Table 2). As shown in Table 2, the five top performing genotypes were not the same for all traits such that different genotypes showed better performance for different traits. This suggests that the highest-performing genotypes for a particular trait could serve as a source germplasm for further breeding activities. Genotypes that contain iron and zinc better than the acceptable level, which is > 60 ppm for iron and > 32 ppm for zinc [45] can be used for bio-fortification programs [31] to alleviate acute malnutrition deeply rooted in underdeveloped countries such as Ethiopia. From the present study, LR106, LR102, LR25 and LR16 can be selected for higher grain yield and starch zinc and calcium, LR23, LR45, LR38 and LR12 can be selected for higher zinc and iron content, and LR14, LR27, LR65 and LR57 can be selected for higher protein and starch contents. Nonetheless, selection in one trait may result in the reduction of the level of the other target trait [32], suggesting the need to evaluate the genotypes based on their ability to combine grain yield with other important traits such as nutritional quality traits used in this study as suggested by [33]. Interestingly, all traits studied showed a wide array of variability. The wide range showed in the value of the traits revealed that it is possible to distinguish among the genotypes for effective breeding activities. Similar to the present finding, earlier studies detected wide range of values for grain yield [2, 4], ash [46, 47], starch and protein [21, 48], iron and zinc [25, 49], calcium [25, 50] and magnesium [30, 51], in sorghum genotypes.
The tested genotypes showed highly significant variations (p < 0.001) for all traits studied in all individual environments. Likewise, the combined analysis showed strong significant variance (P < 0.001) for all traits studied due to genotype (σ2g), genotype by environment interaction (δ2 g x e) and environments (σ2e) with the exception that variance due to environment (σ2e) that had significant variation (< 0.05) for calcium content. This indicates that sorghum landraces collected from Tigray harbor high genetic variation useful for the development and deployment of nutritionally enhanced and high-yielding sorghum varieties. This agrees with previous finding [52] that reported enormous variations among sorghum landraces that are evolved in Vavilov centers of crop origin and diversity such as Ethiopia. The large genotype by environment interaction (σ2 g x e) variance relative to the genotypic variance indicates the need to select for trait stability in the target environment. The high level of genetic variation for various quantitative traits identified in the present study could be attributed to the genetic constitution of the genotypes, varied agro-ecologies and environmental conditions that could affect mineral uptake translocation and distribution. Gene flow between wild and cultivated relatives could be also a likely reason for the high genetic diversity in sorghum landraces [53]. The present results support the high genetic diversity of various sorghum traits reported earlier [4, 21, 30, 49].
The genetic variance (σ2 g) component is also helpful to estimate heritability such that high σ2g implies minimal environmental effect on the trait and vice versa. According to the classification made by [44], the magnitude of broad-sense heritability was higher (H2 > 0.9) in each environment for all the traits studied indicating that the minimal environmental effect, and thus any of the traits can be used for selection [4, 54]. The broad-sense heritability of the pooled over traits was medium (H2 > 30 < 60) for protein (H2 = 0.4) and zinc (H2 = 0.5), and high (H2 > 0.6) for the other traits studied. The medium to high broad sense heritability over environments indicates the possibility of effective selection among the genotypes for further breeding activities. The high broad sense heritability recorded in the present study also indicates these traits are controlled by the additive gene action, which could be improved through selection. As in this study, high to medium heritability in sorghum germplasm has been also reported for various agronomic and nutritional traits of sorghum by [4, 31]. As compared to the individual environment, the heritability of the pooled over analysis was reduced to the range from 0.4 (protein content) to 0.82 (grain yield). This is due to the strong effect of genotype by environment interaction (σ2 g x e) effect. This calls for the need to conduct the genotype by environment interaction analysis of the traits to identify high-yielding and stable genotypes.
In the present study, both negative and positive significant associations between some of the traits were detected in individual environments (Table 5) as well as in combined environments (Table 6; Fig. 1). However, the correlations were not in the same trend though out the individual environments. For instance, zinc content had a significant positive correlation with iron in En1, En2, En4 and En5 while significant negative associations of zinc and iron were recorded in En3 and En6. Besides, ash content had a significant positive association with zinc in En1, En4 and En5 whereas; zinc content had a significant negative association with iron concentration in En3. The variation in the association of traits in different environments might be due to variations in the nutrient content of the soil and genotype by environment interaction that may affect mineral adsorption and translocation of crops [55]. In the pooled analysis, significant positive associations were detected in traits such as ash content with zinc, iron and magnesium contents, zinc with iron and grain yield, and calcium with grain yield. On the other hand, protein showed a significant negative association with iron, with ash, iron and zinc contents (Table 6, Fig. 1).
The strong positive associations between traits might be due to common and overlapping quantitative trait loci [56] suggesting that the traits can be improved concurrently through direct selection. In line with the present study, earlier researchers noted a significant positive association between iron and zinc [21, 31], and zinc and protein contents [25, 30, 48]. Whereas negative associations were reported for grain yield with zinc and iron content [31, 57], and calcium content with iron and magnesium content [30]. Growing and selection among large segregating populations could break the negative association of grain yield with other key traits reported in the present study as well as in other earlier research [58, 59].
The GYT analysis is based on the concept that yield is the main objective and the other target traits are evaluated based on their value to combine with grain yield. In the present study, the GYT biplot (Fig. 2) explores about 87.5% of the total variation indicating the suitability of the biplot to graphically display the data for further interpretation. The positive association between the traits as revealed by the acute angle (Fig. 2A) and Pearson correlation coefficient (Table 7) is because the entire yield by trait combinations have yield as a component that is the special feature of genotype by yield by trait (GYT) [33]. The highly significant (p < 0.001) positive associations between the traits identified in the present study (Table 7) indicates that there is possibility of selection among the genotype based on their yield performance and nutritional quality. The finding of the present study agrees with the previous finding 33 (oat), 36 (durum wheat) 37 (cotton) and 61 (sorghum) that reported strong positive association of various traits.
The ‘which-won-where’ view of the GYT biplot (Fig. 2B) was useful to demonstrate the trait profile of the genotypes. The genotypes located in the same sector are closely related, and different genotypes are associated with different traits in each sector. The polygon peak contains the most effective genotypes (vertex genotypes) associated with the trait/s profile in each sector [60]. In this case, genotypes such as LR12, LR25, LR23 and LR 106 were the best genotypes in combining grain yield zinc, ash and iron contents, while LR102, LR103, LR107 were the best genotypes in combining grain yield with protein and calcium, and genotype LR75 and LR28 were best genotypes in combining grain yield with starch and magnesium.
The line perpendicular to the average tester axis (ATA) (Fig. 2C) separates genotypes with below-average means (example: LR56, LR45, LR11, LR92, LR60 and LR57) from those with above-average means (example: LR12, LR23, LR25, LR106, and LR1). Genotypes far from the ideal center are not desirable, whereas genotypes displayed near/at the ideal center are superior genotypes [61]. Therefore, landraces such as LR12, LR23, LR25, LR106, and LR1 are superior genotypes whereas, LR56, LR45, LR57, LR92, LR60 are less performed landraces. Genotypes holding higher mineral content with high-yielding backgrounds can improve the nutritional value of the crop [30]. In the present study, ten promising and five bottom landraces were identified based on the overall superiority index (Table 8) and average tester coordinator (ATC) view of the GYT biplot. These genotypes could contribute significantly to overcoming malnutrition in arid and semi-arid areas including Tigray. As in this study, [33, 35,36,37] also stated the ATC view of the GYT biplot is an instrumental to rank based on overall superiority and to show their overall superiority and their strengths and weaknesses. They can be also sources of parental lines for further breeding activities. Our finding confirms that the Ethiopian sorghum landraces often harbored various important traits explained by earlier researchers [2, 5, 22, 24].
6 Conclusions
Identification and selection of superior sorghum genotypes for grain yield and nutritional traits is essential to tackle the challenges of acute food shortage and malnutrition in arid and semi-arid areas like Tigray region. This study identified sorghum landraces with higher mean performance, large and useful genetic variations for grain yield and nutritional quality traits. Accordingly, various landraces were selected as breeding parents for each of the traits studied. All the traits studied were highly heritable in each environment (H2 > 0.9) and moderate to highly heritable in the pooled environments implying the possibility of effective selection among the genotypes for further breeding activities. The study further found strong significant positive and negative associations between some of the traits in each environment as well as in combined environments. Nonetheless, highly significant positive (p < 0.001) associations were detected in the genotype by yield by trait combinations. The significant positive association between traits suggests that the traits can be improved concurrently through direct selection. Whereas, the strong negative association between traits implies the need for growing and selection among large segregating populations to break the unfavorable association. Using average tester coordination (ATC) view of GYT biplot and overall superiority index (Mean SI) of the GYT analysis, ten promising sorghum landraces were selected for direct use or breeding programs in arid and semi-arid areas in general and in Tigray in particular.
Data availability
All data supporting the findings of this study will be available up on request.
References
FAOSTAT, FAOSTAT statistical database. In: Publisher: FAO (Food and Agriculture Organization of the United Nations), Rome, Italy, 2021. Available from: https.faostat.fao.org. (Accessed 10 June 2023).
Girma G, Nida H, Tirfessa A, Lule D, Bejiga T, Seyoum A, Mekonen M, Nega A, Dessalegn K, Birhanu C. A comprehensive phenotypic and genomic characterization of Ethiopian sorghum germplasm defines core collection and reveals rich genetic potential in adaptive traits. The Plant Genome. 2020;13(3): e20055. https://doi.org/10.1002/tpg2.20055.
Mace ES, Tai S, Gilding EK, Li Y, Prentis PJ, Bian L, Campbell BC, Hu W, Innes DJ, Han X. Whole-genome sequencing reveals untapped genetic potential in Africa’s indigenous cereal crop sorghum. Nat Commun. 2013;4(1):2320. https://doi.org/10.1038/ncomms3320.
Phuke RM, Anuradha K, Radhika K, Jabeen F, Anuradha G, Ramesh T, Hariprasanna K, Mehtre SP, Deshpande SP, Anil G. Genetic variability, genotype× environment interaction, correlation, and GGE biplot analysis for grain iron and zinc concentration and other agronomic traits in RIL population of sorghum (Sorghum bicolor L. Moench). Front Plant Sci. 2017;8:712. https://doi.org/10.3389/fpls.2017.00712.
Shegro A, Labuschagne MT, Shargie NG, Biljon AV. Multivariate analysis of nutritional diversity in sorghum landrace accessions from western Ethiopia. J Biol Sci. 2013;13(2):67–74. https://doi.org/10.3923/jbs.2013.67.74.
Kumar AA, Anuradha K, Ramaiah B, Grando S, Frederick H, Rattunde W, Virk P, Pfeiffer WH. Recent advances in sorghum biofortification research. Plant Breed Rev. 2015; 89–124.
Dykes L. Sorghum phytochemicals and their potential impact on human health. Methods Protoc. 2019. https://doi.org/10.1007/978-1-4939-9039-9_9.
Dicko MH, Gruppen H, Traoré AS, Voragen AG, van Berkel WJ. Sorghum grain as human food in Africa: relevance of content of starch and amylase activities. Afr J Biotechnol. 2006;5(5):384–95.
Angkuratipakorn T, Chung C, Koo CK, Mundo JLM, McClements DJ, Decker EA, Singkhonrat J. Development of food-grade Pickering oil-in-water emulsions: tailoring functionality using mixtures of cellulose nanocrystals and lauric arginate. Food Chem. 2020;327: 127039. https://doi.org/10.1016/j.foodchem.2020.127039.
Harlan JR, de Wet JM. A simplified classification of cultivated sorghum 1. Crop Sci. 1972;12(2):172–6.
Teshome A, Baum B, Fahrig L, Torrance J, Arnason TJ, Lambert J. Sorghum [Sorghum bicolor (L.) Moench] landrace variation and classification in north Shewa and south Welo, Ethiopia. Euphytica. 1997;97:255–63. https://doi.org/10.1023/A:1003074008785.
Ayana A, Bekele E. Geographical patterns of morphological variation in sorghum (Sorghum bicolor (L.) Moench) germplasm from Ethiopia and Eritrea: qualitative characters. Hereditas. 1998;129(3):195–205. https://doi.org/10.1111/j.1601-5223.1998.t01-1-00195.x.
Boyles RE, Brenton ZW, Kresovich S. Genetic and genomic resources of sorghum to connect genotype with phenotype in contrasting environments. Plant J. 2019;97(1):19–39. https://doi.org/10.1111/tpj.14113.
Statistica, Sorghum production worldwide in 2021/2022, by leading country (in 1,000 metric tons) [internet]. Available on: https://www.statista.com/statistics/1134651/global-sorghum-production-by-country/. (Accessed on 12 June 2023).
T.f.d.r.o.E.E.s.s.a.s. survey, Report on area and production of major crops, 2022, p. 136. (Accessed on 12 June 2023).
Adugna A. Analysis of in situ diversity and population structure in Ethiopian cultivated Sorghum bicolor (L.) landraces using phenotypic traits and SSR markers. Springerplus. 2014;3(1):1–14. https://doi.org/10.1186/2193-1801-3-212.
Habyarimana E, Lorenzoni C, Marudelli M, Redaelli R, Amaducci S. A meta-analysis of bioenergy conversion relevant traits in sorghum landraces, lines and hybrids in the Mediterranean region. Ind Crops Prod. 2016;81:100–9.
Ochieng G, Ngugi K, Wamalwa LN, Manyasa E, Muchira N, Nyamongo D, Odeny DA. Novel sources of drought tolerance from landraces and wild sorghum relatives. Crop Sci. 2021;61(1):104–18. https://doi.org/10.1002/csc2.20300.
De Wet J, Harlan J. The origin and domestication of Sorghum bicolor. Econ Bot. 1971;25(2):128–35.
Dillon SL, Shapter FM, Henry RJ, Cordeiro G, Izquierdo L, Lee LS. Domestication to crop improvement: genetic resources for Sorghum and Saccharum (Andropogoneae). Ann Bot. 2007;100(5):975–89. https://doi.org/10.1093/aob/m.
Shegro A, Shargie NG, van Biljon A, Labuschagne MT. Diversity in starch, protein and mineral composition of sorghum landrace accessions from Ethiopia. J Crop Sci Biotechnol. 2012;15:275–80. https://doi.org/10.1007/s12892-012-0008-z.
Mamo W, Enyew M, Mekonnen T, Tesfaye K, Feyissa T. Genetic diversity and population structure of sorghum [Sorghum bicolor (L.) Moench] genotypes in Ethiopia as revealed by microsatellite markers. Heliyon. 2023. https://doi.org/10.1016/j.heliyon.2023.e12830.
Reddy BV, Ramesh S, Reddy PS, Kumar AA, Tropics S-A. Genetic enhancement for drought tolerance in sorghum. Plant Breed Rev. 2009;31:189–222.
Enyew M, Carlsson AS, Geleta M, Tesfaye K, Hammenhag C, Seyoum A, Feyissa T. Novel sources of drought tolerance in sorghum landraces revealed via the analyses of genotype-by-environment interactions. Front Plant Sci. 2022. https://doi.org/10.3389/fpls.2022.1062984.
Gerrano A, Labuschagne M, Van Biljon A, Shargie N. Quantification of mineral composition and total protein content in sorghum [Sorghum bicolor (L.) Moench] genotypes. Cereal Res Commun. 2016;44:272–85. https://doi.org/10.1556/0806.43.2015.046.
Rhodes DH, Hoffmann L, Rooney WL, Herald TJ, Bean S, Boyles R, Brenton ZW, Kresovich S. Genetic architecture of kernel composition in global sorghum germplasm. BMC Genom. 2017;18(1):1–8. https://doi.org/10.1186/s12864-016-3403-x.
Nida H, Girma G, Mekonen M, Lee S, Seyoum A, Dessalegn K, Tadesse T, Ayana G, Senbetay T, Tesso T. Identification of sorghum grain mold resistance loci through genome wide association mapping. J Cereal Sci. 2019;85:295–304. https://doi.org/10.1016/j.jcs.2018.12.016.
Wu Y, Huang Y, Tauer C, Porter DR. Genetic diversity of sorghum accessions resistant to greenbugs as assessed with AFLP markers. Genome. 2006;49(2):143–9. https://doi.org/10.1139/g05-095.
Doggett H. The development of the cultivated sorghums. Crop Evol. 1965; 50–69.
Badigannavar A, Girish G, Ramachandran V, Ganapathi T. Genotypic variation for seed protein and mineral content among post-rainy season-grown sorghum genotypes. Crop J. 2016;4(1):61–7. https://doi.org/10.1016/j.cj.2015.07.0022.
Andiku C, Shimelis H, Shayanowako AI, Gangashetty PI, Manyasa E. Genetic diversity analysis of East African sorghum (Sorghum bicolor [L.] Moench) germplasm collections for agronomic and nutritional quality traits. Heliyon. 2022;8(6):e09690. https://doi.org/10.1016/j.heliyon.2022.e09690.
Yan W, Kang MS, Ma B, Woods S, Cornelius PL. GGE biplot vs. AMMI analysis of genotype-by-environment data. Crop Sci. 2007;47(2):643–53. https://doi.org/10.2135/cropsci2006.06.0374.
Yan W, Frégeau-Reid J. Genotype by yield* trait (GYT) biplot: a novel approach for genotype selection based on multiple traits. Sci Rep. 2018;8(1):1–10. https://doi.org/10.1038/s41598-018-26688-8.
Yan W, Frégeau-Reid J, Mountain N, Kobler J. Genotype and management evaluation based on Genotype by Yield* Trait (GYT) analysis. Crop Breed Genet Genom. 2019. https://doi.org/10.20900/cbgg20190002.
Merrick LF, Glover KD, Yabwalo D, Byamukama E. Use of Genotype by Yield* Trait (GYT) analysis to select hard red spring wheat with elevated performance for agronomic and disease resistance traits. Crop Breed Genet Genom. 2020. https://doi.org/10.20900/cbgg20200009.
Peixoto MA, Evangelista JSPC, Coelho IF, Carvalho LP, Farias FJC, Teodoro PE, Bhering LL. Genotype selection based on multiple traits in cotton crops: the application of genotype by yield* trait biplot. Acta Sci Agron. 2022. https://doi.org/10.4025/actasciagron.v44i1.54136.
Welderufael S, Abay F, Ayana A, Amede T. Genotype by Trait (GT) and Genotype by Yield* Traits (GYT) Analysis of Sorghum Landraces in Tigray, Northern Ethiopia. Crop Breed Genet Genom. 2023. https://doi.org/10.20900/cbgg20230002.
GD Atakilte B, Haile M. Understanding diversity in farming practices in Tigray. 2001. 20 May 2018. Available from: https://www.iied.org/9043iied. Accessed 2023 June 12.
Caporaso N, Whitworth MB, Fisk ID. Near-Infrared spectroscopy and hyperspectral imaging for non-destructive quality assessment of cereal grains. Appl Spectrosc Rev. 2018;53(8):667–87. https://doi.org/10.1080/05704928.2018.1425214.
Alvarado G, López M, Vargas M, Pacheco A, Rodríguez F, Burgueño J, Crossa J. User’s Manual META-R (Multi-Environment Trial Analysis with R), CIMMYT: El Batan, Mexico (2015) 1–34.
Alvarado G, Rodríguez FM, Pacheco A, Burgueño J, Crossa J, Vargas M, Pérez-Rodríguez P, Lopez-Cruz MA. META-R: a software to analyze data from multi-environment plant breeding trials. Crop J. 2020;8(5):745–56. https://doi.org/10.1016/j.cj.2020.03.010.
R.c. Team, R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, (2021). Available at: https://www.R-project.org/.
Yan W. Crop variety trials: data management and analysis. John Wiley & Sons. 2014. https://doi.org/10.1002/9781118688571.
Robinson H. Quantitative genetics in relation to breeding on centennial of Mendelism, Indian J Genet Plant Breed., INDIAN SOC GENET PLANT BREED INDIAN AGRICULTURE RES INST, NEW DELHI DELHI …, 1966, pp. 171.
Chapke R, Tonapi V. Best practices for sorghum cultivation and importance of value-addition, ICAR-Indian Institute of Millets Research, Hyderabad 500 030, India, 2016.
Moharram Y, Youssef A. Sorghum grain and quality of its edible products. J Food Sci. 1995. https://doi.org/10.1016/S0167-4501(06)80156-1.
Chung I-M, Kim E-H, Yeo M-A, Kim S-J, Seo MC, Moon H-I. Antidiabetic effects of three Korean sorghum phenolic extracts in normal and streptozotocin-induced diabetic rats. Int Food Res J. 2011;44(1):127–32. https://doi.org/10.1016/j.foodres.2010.10.051.
Ng’uni D, Geleta M, Hofvander P, Fatih M, Bryngelsson T. Comparative genetic diversity and nutritional quality variation among some important Southern African sorghum accessions [‘Sorghum bicolor’ (L.) Moench]. Aust J Crop Sci. 2012;6(1):56–64.
Makebe A, Shimelis H. diversity analysis for grain nutrient content and agronomic traits among newly bred striga-resistant and Fusarium oxysporum f. sp. strigae (FOS)-compatible sorghum genotypes. Diversity. 2023;15(3):371. https://doi.org/10.3390/d15030371.
Osman A, Abd El-Wahab A, Ahmed MFE, Buschmann M, Visscher C, Hartung CB, Lingens JB. Nutrient composition and in vitro fermentation characteristics of sorghum depending on variety and year of cultivation in Northern Italy. Foods. 2022;11(20):3255. https://doi.org/10.3390/foods11203255.
Ragaee S, Abdel-Aal E-SM, Noaman M. Antioxidant activity and nutrient composition of selected cereals for food use. Food Chem. 2006;98(1):32–8. https://doi.org/10.1016/j.foodchem.2005.04.039.
Hummer KE, Hancock JF. Vavilovian centers of plant diversity: Implications and impacts. Hortic Sci. 2015;50(6):780–3. https://doi.org/10.2135/cropsci2007.08.0441.
Tesso T, Kapran I, Grenier C, Snow A, Sweeney P, Pedersen J, Marx D, Bothma G, Ejeta G. The potential for crop-to-wild gene flow in sorghum in Ethiopia and Niger: a geographic survey. Crop Sci. 2008;48(4):1425–31. https://doi.org/10.2135/cropsci2007.08.0441.
Bashir EM, Ali AM, Ali AM, Melchinger AE, Parzies HK, Haussmann BI. Characterization of Sudanese pearl millet germplasm for agro-morphological traits and grain nutritional values. Plant Genet Res. 2014;12(1):35–47. https://doi.org/10.1017/S1479262113000233.
Sankaran RP, Huguet T, Grusak MA. Identification of QTL affecting seed mineral concentrations and content in the model legume Medicago truncatula. Theor Appl Genet. 2009;119:241–53. https://doi.org/10.1007/s00122-009-1033-2.
Kumar S, Hash CT, Thirunavukkarasu N, Singh G, Rajaram V, Rathore A, Senapathy S, Mahendrakar MD, Yadav RS, Srivastava RK. Mapping quantitative trait loci controlling high iron and zinc content in self and open pollinated grains of pearl millet [Pennisetum glaucum (L.) R. Br.]. Front Plant Sci. 2016;7:1636. https://doi.org/10.3389/fpls.2016.01636.
Ashok Kumar A, Reddy BV, Ramaiah B, Sahrawat KL, Pfeiffer WH. Genetic variability and character association for grain iron and zinc contents in sorghum germplasm accessions and commercial cultivars. Eur J Plant Sci Biotechnol. 2012;6(1):1–5.
Ambekar S, Kamatar M, Ganesamurthy K, Ghorade R, Saxena U, Chand P, Jadav B, Das I, Nageshwararao T, Audilakshmi S. Genetic enhancement of Sorghum (Sorghum bicolor (L) Moench) for grain mould resistance: II. Breeding for grain mould resistance. Crop Prot. 2011;30(7):759–64. https://doi.org/10.1016/j.cropro.2010.06.024.
Diatta C, Tovignan TK, Adoukonou-Sagbadja H, Aidara O, Diao Y, Sarr MP, Ifie BE, Offei SK, Danquah EY, Cisse N. Development of sorghum hybrids for stable yield and resistance to grain mold for the Center and South-East of Senegal. J Crop Prot. 2019;119:197–207. https://doi.org/10.1016/j.cropro.2019.02.001.
Yan W, Tinker NA. Biplot analysis of multi-environment trial data: principles and applications. Can J Plant Sci. 2006;86(3):623–45. https://doi.org/10.4141/P05-169.
Yan W, Rajcan I. Biplot analysis of test sites and trait relations of soybean in Ontario. Crop Sci. 2002;42(1):11–20. https://doi.org/10.2135/cropsci2002.1100.
Funding
This study was supported by Mekelle University (Grant PG/CoDANR/Ph.D./Recurrent /75/2011), International Foundation for Science (Grant IFS C/6182-1).
Author information
Authors and Affiliations
Contributions
SW: planed, excused the field and laboratory experiments, organizes and writes the manuscript; FA, AA and TA, contributed in planning the research, supervised the field and laboratory experiments, contribute in preparing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Welderufael, S., Abay, F., Ayana, A. et al. Genetic diversity, correlation and genotype × yield × trait (GYT) analysis of grain yield and nutritional quality traits in sorghum (Sorghum bicolor [L.] Moench) genotypes in Tigray, northern Ethiopia. Discov Agric 2, 4 (2024). https://doi.org/10.1007/s44279-024-00016-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44279-024-00016-3