Abstract
The Severe Acute Respiratory Syndrome (SARS)-CoV-2-induced Coronavirus Disease 2019 (COVID-19) pandemic has caused an overwhelming influence on public health because of its high morbidity and mortality. Critical-illness cases often manifest as acute respiratory distress syndrome (ARDS). Previous evidence has suggested platelets and thrombotic events as key mediators of SARS-CoV-2-associated ARDS. However, how the balance of platelet regeneration from the hematopoietic system is changed in ARDS remains elusive. Here, we reported a more severe inflammation condition and hyperactivity of platelets in COVID-19 ARDS patients compared with those infected but without ARDS. Analysis of peripheral blood revealed an increased proportion of hematopoietic stem cells (HSCs), common myeloid progenitors (CMPs), megakaryocyte-erythrocyte progenitors (MEPs), and megakaryocyte progenitors (MkPs) in ARDS patients, suggesting a megakaryocytic-differentiation tendency. Finally, we found altered gene expression pattern in HSCs in COVID-19 ARDS patients. Surprisingly, genes representing platelet-primed HSCs were downregulated, indicating these cells are being stimulated to differentiate. Taken together, our findings shed light on the response of the hematopoietic system to replenish platelets that were excessively consumed in COVID-19 ARDS, providing a mechanism for disease progression and further therapeutic development.
Graphical Abstract

Highlights
• Hyperactivity of platelets and mild decrease in plaletlet number are observed in COVID-19 patients with ARDS.
• A more active state of the hematopoietic system is established to replenish platelet pool in COVID-19 ARDS patients.
• The influence of SARS-CoV-2 on the hematopoietic system is also manifested in altered gene expression pattern in HSC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Coronavirus Disease 2019 (COVID-19) pandemic, caused by the Severe Acute Respiratory Syndrome-CoV-2 (SARS-CoV-2) virus, has spread globally and resulted in over 670 million infections and 6.8 million deaths to date. Despite widespread vaccination efforts, localized outbreaks continue to occur due to the virus’ evolution and variation. These outbreaks often lead to severe multi-organ dysfunction, particularly in the lung, with elderly individuals being at a higher risk [1, 2]. A significant number of hospitalized COVID-19 patients (15–30%) develop Acute Respiratory Distress Syndrome (ARDS), characterized by bilateral chest opacities on X-ray and hypoxemia [3, 4]. COVID-19-induced ARDS is related to a high mortality rate which increases with severity [5, 6]. Abundant research has suggested a high relevance of thrombosis in pulmonary capillaries [3] and a conversion of the immune system toward an inflammatory state [7] in these COVID-19 patients diagnosed with ARDS. However, how the hematopoietic system is changed during the development of COVID-19-related ARDS remains poorly understood.
Thombopoiesis is the production of platelets through a multistep process involving Hematopoietic Stem Cells (HSCs), multipotent progenitors (MPPs), Common Myeloid Progenitors (CMPs), Megakaryocyte-Erythrocyte Progenitors (MEPs), Megakaryocyte Progenitors (MkPs) and megakaryocytes (MKs) [8]. Platelets play a crucial role in thrombosis and hemostasis by being recruited to the vascular injury sites, activated, and coagulation [9, 10]. Besides, their interaction with various leukocytes and the release of pro-inflammatory cytokines upon activation indicated a high correlation between platelets and inflammation [11, 12]. Recent studies identified Vwf+ platelet-primed HSCs in mice with higher expression levels of platelet-related genes such as Vwf, Clu, Fhl1, Gpr64, and Sdpr [13]. Under an acute inflammation state, HSC-like megakaryocyte-committed progenitors are stimulated to mature, express more MK-related proteins and differentiate, contributing to platelet recovery [14]. Abundant evidence demonstrated the hyperactivity of platelets after SARS-CoV-2 infection [15,16,17,18]. Especially, elevated D-dimer and abnormal fibrinogen concentrations of platelets in COVID-19-induced ARDS patients are related to high mortality rates [19, 20]. In addition, platelets from COVID-19 ARDS patients are primed for aggregation [21], interacting with leukocytes to form immunothrombosis [22]. These data indicate a key role of platelets in the development of ARDS. Therefore, we aimed to explore how the hematopoietic system responds to SARS-CoV-2-induced ARDS and its impacts on platelet production from HSCs during acute inflammation.
Considering the rapid transmission, high infectious capacity, and high fatality rate of SARS-CoV-2, China activated the first-level public health emergency response to contain the outbreak shortly after its occurance, and was able to achieve remarkable epidemic prevention and control. And because of this, most of the research teams in China didn’t have access to acquire enough COVID-19 samples. Therefore, studies of the mechanism and progress of SARS-CoV-2-related ARDS at a cellular and molecular level were limited and left behind in China. As the virulence of the strain diminished, the gradual liberalization of prevention and control was realized last December. Meanwhile, the policy temporarily resulted in a peak of SARS-CoV-2 infection, with a prevalence of ARDS. Therefore, we were able to obtain samples for this research, which we think is fundamental to a deeper understanding and novel therapeutics for the disease, especially when it develops with ARDS.
In this study, we examined peripheral blood samples from critically ill COVID-19-related ARDS patients. Platelet activation and coagulation function tests revealed hyperactivity of platelets in these ARDS patients, suggesting a higher thrombus burden despite a lower platelet count. Then, we focused on the response of the hematopoietic system to severe thrombopenia in the inflammatory process of ARDS development. Multiparameter flow cytometry analysis revealed a differentiation tendency towards myeloid lineage and platelets. RNA-seq analysis indicated the elevated ability of the hematopoietic system to restore the platelet pool. Taken together, these findings elucidated an emergency mechanism of the hematopoietic system to relieve life-threatening thrombopenia when ARDS occurred in COVID-19 patients.
Results
COVID-19 ICU-ARDS patients possessed more severe infection and inflammation conditions than non-ICU patients
To assess the essential blood function under COVID-19, blood routines were collected from 70 patients, 22 of whom were in the Intensive Care Unit (ICU) and diagnosed with ARDS (Table 1). Notably, the ratio of lymphocytes and monocytes was out of the normal range in COVID-19 patients whether diagnosed with ARDS or not, indicating that COVID-19 infection led to unusual blood cell composition. Further analysis showed the lymphocyte and monocyte levels in ICU-ARDS patients were significantly lower than those in non-ICU patients. Interestingly, ICU-ARDS patients also exhibited an abnormal increase in neutrophils, indicating that more severe infection and inflammation occurred in ICU-ARDS patients, and consistent with this, Giemsa staining showed a large amount of neutrophils in ICU-ARDS patient samples (Supplementary Fig. 1). Besides, more ICU-ARDS patients (27.27%) were identified with a low level of platelets compared with non-ICU patients (8.33%), suggesting that severe COVID-19 infection which resulted in ARDS reduced the number of platelets in peripheral blood.
As severe inflammation is usually accompanied by systematic cytokine storm, we also collected inflammation-related cytokine data to evaluate the inflammatory level in ICU-ARDS patients (Fig. 1A). Although most ICU-ARDS patients possess a normal level of Interleukin (IL)-1β, IL-8, IL-10 and TNF-α in peripheral blood, the IL-6 level was significantly increased in ICU-ARDS patients, confirming a systemic inflammation in the blood system. In summary, certain values of blood routines and cytokines in COVID-19 patients were significantly abnormal. ICU-ARDS patients had a more serious COVID-19 infection and inflammation accompanied by mild platelet reduction compared with non-ICU patients.
The altered levels of cytokines and hyperactivity of platelets in the peripheral blood under the acute inflammation condition induced by COVID-19 ARDS. A Numbers of individuals whose pro-inflammatory cytokines were within (Normal) or elevated beyond (Hyper) the reference ranges in 20 COVID-19 ARDS patients. B The expression of CD62P on the surface of platelets in whole blood was measured both before and after activation by 10 μM ADP. H, non-infected group (n=6); V, COVID-19 ARDS group (n=27). C-E Levels of Brain-Derived Neurotrophic Factor (BDNF) (C) Epidermal Growth Factor (EGF) (D) and MMP3 (E) in Platelet-Rich Plasma (PRP) prepared from the blood of COVID-19 ARDS patients (n = 7) and uninfected participants (control, n = 6) were measured by ELISA. *P < .05; ****P < .0001; ns, not significant
Elevated platelet activation ability and thrombus burden in ICU-ARDS patients
In order to evaluate the function of the platelets, we treated the whole blood samples with ADP to activate the platelets and analyzed the level of CD62P+ in CD41+ cells (Fig. 1B). It is noteworthy that ICU-ARDS samples were already more active before ADP treatment and could be activated to a higher level than non-COVID-19 samples, demonstrating that platelets affected by severe COVID-19 infection exhibited a ready-to-activate state. Given that IL-6 level was significantly increased in ICU-ARDS patients, the systemic inflammation may contribute to the hyperactivity of platelets in peripheral blood. Besides, we also tested the coagulation function in part of ICU-ARDS patients (n = 21) (Table 2). D-dimer, an index that is highly related to the prognosis and severity of COVID-19 [23, 24], was significantly increased in ICU-ARDS patients, revealing an increased burden of thrombus and more severe infection of COVID-19. Besides, the level of fibrinogen and prothrombin time in ICU-ARDS patients also increased, which further clarified ICU-ARDS patients had an abnormal function of blood coagulation. Taken together, ICU-ARDS patients exhibited hyperactivity of platelets and an elevated thrombus burden caused by severe infection of COVID-19, which also indicated that the reduction of platelets in ICU-ARDS patients was at least partially due to the consumption mediated by platelet activation and thrombus formation.
BDNF and EGF level decreased in PRP from peripheral blood of ICU-ARDS patients
To evaluate the cytokine level in platelets, we prepared platelet-rich plasma (PRP) from the peripheral blood of ICU-ARDS and non-COVID-19 samples and measured the level of several cytokines (Fig. 1C-E). Serum BDNF usually decreased after being infected by COVID-19 [25], similarly, the BDNF level in ICU-ARDS PRP significantly decreased. Besides, the EGF level in ICU-ARDS PRP was also lower than that in non-COVID-19 samples. However, the MMP3 level, a serum endopeptidase elevated in acute lung injury and severe COVID-19 infection, was not significantly different between PRP from ICU-ARDS and non-COVID-19 peripheral blood. These indicated that severe COVID-19 infection affected the level of certain growth factors released by platelets, possibly due to reduced platelet cell number in these patients.
ICU-ARDS patients exhibited abnormal components of the hematopoietic compartment and increased megakaryocytic lineage differentiation
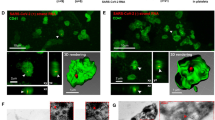
HSC is capable of differentiating into common lymphoid progenitor (CLP) and CMP cells which further produced several essential blood cells including platelets and play an indispensable role in maintaining hematopoiesis homeostasis. To evaluate the level of various hematopoietic lineage cells in ICU-ARDS patients, we combined multiple cell surface markers to distinguish HSC/MPP, CLP, CMP, granulocyte-monocyte progenitor (GMP), MEP, MkP and erythroid progenitor (EP) cells in peripheral blood samples. FACS analysis showed an increased level of lineage-negative cells in ICU-ARDS patients which indicated the population of immature cells were increased under ARDS (Fig. 2A-B). An elevated level of lineage-negative cells in ICU-ARDS patients also intimated that certain stem/progenitor cells were activated to a proliferating state. Indeed, the level of HSC/MPP cells significantly increased in ICU-ARDS patients’ samples compared with non-COVID-19 samples, indicating the HSC compartment rapidly turned into activating state triggered by severe COVID-19 infection (Fig. 2C). Severe COVID-19 was usually marked by emergency myelopoiesis and our FACS analysis revealed that CMP cells in ICU-ARDS samples significantly increased while there was no obvious difference in the number of CLP cells (Fig. 2C).
The hyperactive state of the hematopoietic compartment in COVID-19 ARDS patients. A A representative example of the FACS plot of peripheral blood showed changes in the hematopoietic stem/progenitor cell (HSPC) population in the non-infected group (top panel) and COVID-19 ARDS patients (bottom panel). B Statistical analysis of the percentage of Lineage-negative cells (Lin−) showed a profound increase in COVID-19 ARDS patients (n = 26) compared with non-infected participants (n = 10). C-G Flow cytometry analysis of HSPCs at different hierarchies in peripheral blood from ICU-ARDS patients (n = 26) and non-infected participants (n = 10), including HSCs/MPPs, CLPs, CMPs, GMPs, MEPs, MkPs, and EPs. *P < .05; **P < .01; ns, not significant
Since CMP cells could differentiate into GMP or MEP cells, measurement of GMP and MEP cells could further determine the specific direction of differentiation in the myeloid compartment. Notably, MEP cells in ICU-ARDS samples were significantly higher than that in non-COVID-19 samples, while there was no significant difference in the number of GMP cells (Fig. 2D-E), demonstrating a specific direction of differentiation towards megakaryocytic and erythroid lineages. Although there was no significant change in MkP cells between ICU-ARDS and non-COVID-19 samples, unusually high levels of MkP cells were identified in part of ICU-ARDS samples and the mean level of MkP cells was relatively higher than that of the non-COVID-19 samples (Fig. 2F). No significant differences in EP cells were observed between ICU-ARDS and non-COVID-19 samples (Fig. 2G). These results further illustrated that cells in the myeloid compartment, especially MEP and MkP cells, exhibited elevated proliferative ability after being affected by severe COVID-19 infection. Such emergent megakaryocytic lineage differentiation was possibly a self-response to elevated consumption of platelets in ICU-ARDS patients due to hyperactivation-induced thrombus formation. Overall, we discovered anomalous myelopoiesis and an increased tendency of megakaryopoiesis in the peripheral blood of ICU-ARDS patients.
ICU-ARDS patients showed an increased level of megakaryocytes and granulocytes in peripheral blood
To further clarify the specific changes in the blood system, we used FACS to measure different mature blood cells in ICU-ARDS and non-COVID-19 peripheral blood samples (Fig. 3). Severe COVID-19 was usually associated with lymphopenia and T cell number reduction and indeed we found that the number of T cells in ICU-ARDS samples was significantly lower than that in non-COVID-19 samples (9.64% vs 14.87%). There was no significant difference in the level of B cells (1.53% vs 2.13%) or natural killer (NK) cells (6.02% vs 6.15%) (Fig. 3A-B). However, the mean levels of myeloid cells (78.29% vs 74.40%) and megakaryocytes (7.77% vs 7.31%) were mildly elevated in ICU-ARDS patients. These results further elucidate that systemic myelopoiesis occurred in ICU-ARDS patients’ peripheral blood. An elevated level of megakaryocytes was a signal of improved platelet regeneration which contributes to replenishing excessively depleted platelets due to ARDS-induced thrombus formation.
The alternation in the components of mature blood cell populations in the COVID-19 ARDS group. A A representative example of the FACS plot of peripheral blood after erythrocyte lysis showed changes in the proportion of mature blood cells in the non-infected group (top panel) and COVID-19 ARDS patients (bottom panel). B Summary of flow cytometry analysis for the levels of the different hematopoietic cells including T, B, NK, myeloid cells, and MKs in the non-infected group (Control, n=10) and COVID-19 ARDS patients (n=26). *P < .05; ns, not significant
RNA-seq revealed hyper-proliferation and myeloid differentiation ability of HSC/MPP cells in ICU-ARDS patients
To interpret the internal changes in HSC/MPP compartment under COVID-19-induced ARDS, we sorted Lin−CD34+CD38−CD45RA− cells and employed RNA-sequencing to evaluate the transcriptome response. RNA-seq identified 395 differentially expressed genes (DEGs), 18 of which are highly related to platelet function (Fig. 4A). Notably, platelet adhesion-related gene MMP8 and activation-related gene OLR1 were significantly elevated in ICU-ARDS samples which further proved the existence of increased thrombus burden. Besides, LCN2, a gene expressed in platelet and could promote the proliferation of CD34+ cells, was highly expressed in ICU-ARDS samples, demonstrating that early in HSC/MPP cells there is a tendency of active platelet regeneration. However, genes representing platelet-biased or MK-biased HSCs were downregulated in ICU-ARDs samples, including CLU, SDPR, and CD9, suggesting a highly active state of platelet regeneration from these HSC populations under the COVID-19 ARDS inflammation state to replenish the consumed platelets. Consistent with this, the proportion of CD41+ HSCs/MPPs was reduced in the systemic inflammation state (Supplementary Fig. 2A), while the average CD41 expression level on the surface of CD41-positive HSCs/MPPs was significantly increased (Supplementary Fig. 2B), indicating higher MK protein synthesis. Heatmap analysis revealed that all ICU-ARDS groups were significantly clustered and distinguished from the non-COVID-19 group (Fig. 4B). Besides, we sorted out the pathways related to hematopoiesis and inflammation in the gene ontology (GO) database and summarized the corresponding gene sets to further look for potential associations. There were 50 hematopoietic-cell-lineage-related genes and 60 inflammatory-response-related genes overlapped with DEGs and 26 DEGs were highly related to both hematopoiesis and inflammation, illustrating the connection between systemic inflammation and hematopoietic disorder under ARDS induced by severe COVID-19 infection (Fig. 4C). Furthermore, GO analysis illustrated that several platelet- and myeloid-related pathways were enriched in ICU-ARDS groups, demonstrating the trend of myelopoiesis and platelet production in the HSC/MPP compartment (Fig. 4D). Interestingly, HSC/MPP cells affected by severe COVID-19 infection exhibited more sensitivity to platelet-derived growth factors (GO:0036119 and GO0036120). Given that platelets in ICU-ARDS patients were in a ready-to-activate state and platelet activation directly led to the release of platelet derived growth factor (PDGF), PDGF produced by hyperactivated platelet may contribute to the elevated proliferation ability of HSC/MPP cells in ARDS patients. Furthermore, gene-set enrichment analysis (GSEA) based on GO gene sets revealed stem-cell-development- and immune-related pathways were significantly enriched, further indicating the abnormal activity of HSCs/MPPs in ICU-ARDS patients (Fig. 4E-G). However, antigen processing and presentation and type I interferon signaling pathway were inhibited in ICU-ARDS samples, demonstrating the immunosuppression under ARDS induced by COVID-19 which was in accordance with the systemic T cell reduction (Fig. 3B). In summary, RNA-seq analysis illustrated that HSC/MPP cells in ICU-ARDS patients exhibited elevated proliferation ability and incline of megakaryocytic/platelet lineage differentiation as a response to ARDS caused by severe COVID-19 infection.
COVID-19 ARDS patients showed altered gene expression pattern of HSCs/MPPs at the transcriptional level. RNA-seq was conducted on RNA extracted from HSCs/MPPs isolated from the peripheral blood of uninfected participants (n = 7) and COVID-19 ARDS patients (n = 9). A Volcano plot representing differentially expressed genes (DEGs) between non-infected and COVID-19 ARDS groups. Red dots, significantly upregulated genes; Blue dots, significantly downregulated genes. B Unsupervised hierarchical clustering of the control and patient groups. C Venn diagram representing DEGs involved in the hematopoietic cell lineage and inflammatory response. D GO analysis showed the pathways associated with inflammation response, platelet activation, and myeloid-lineage differentiation enriched significantly in patients. E–G GSEA of antigen processing and presentation (E) stem cell development (F) and type I interferon signaling pathway (G) in COVID-19 ARDS-HSCs/MPPs and control-HSCs/MPPs
Discussion
Since the COVID-19 pandemic swept across the world in 2019, many efforts have been made to uncover hidden mechanisms and therapies. Here, we explored the response cascade of the hematopoietic system to the perturbation of platelet count and activity during the process of COVID-19-related ARDS development at the stem/progenitor-cell hierarchy and transcriptome level. It is noteworthy that we uncovered a myeloid/MK-differentiation tendency, evidenced by larger pools of HSCs/MPPs, CMPs, MEPs, and MkPs (Fig. 1). In line with our findings, previous studies identified emergency myelopoiesis in severe COVID-19 [26,27,28]. The analysis of RNA-seq data for HSCs/MPPs in the peripheral blood shows hundreds of DEGs in COVID-19 patients developing ARDS (Fig. 4). Further investigations are necessary to validate the functions of these genes on coronavirus-related ARDS development.
HSCs, as a type of adult stem cells, are essential for constructing the blood and immune system. In addition to the widely accepted lymphoid and myeloid-biased HSCs, evidence shows that there are other types of lineage-biased HSCs. Vwf + or CD9+ HSCs have been identified to have higher MK lineage-generation potential [13, 29]. We found that genes representing MK or platelet biased-HSCs including CLU, SDPR, and CD9 were downregulated in the ARDS group. Interestingly, the proportion of CD41+ HSCs/MPPs was reduced in the systemic inflammation state, but the average CD41 expression level on the surface of CD41-positive HSCs/MPPs was significantly increased. Considering all evidence, we postulated that platelet or MK-primed HSCs were driven to differentiate towards the megakaryocytic lineage to replenish circulating platelets under this pathological condition to cope with thrombocytopenia.
COVID-related cytokine storm is linked with illness severity and poor prognosis, characterized by the burst of pro-inflammatory cytokines or chemokines, such as IL-1, IL-6, IL-8, IL-12, and tumor necrosis factor (TNF)α [30]. Circulating IL-1β, IL-6, and IL-8 are associated with platelet hyperactivity and hypercoagulability, a hallmark of systemic inflammation [31]. These three cytokines have corresponding receptors on the platelets, promoting thrombosis [31]. IL-6-receptor blockers have been tested in critically ill COVID-19 patients, contributing to improved clinical outcomes through interdicting IL-6/JAK/STAT3 signaling [30, 32]. Besides, a study on thrombocytopenia pointed out a negative correlation of platelet count with serum IL-1β and IL-8 [33]. Our findings are consistent with these previous studies, referring to the hyperactivity of platelets (Fig. 1B), while platelet number were reduced (Table 1). Excessively high levels of pro-inflammatory cytokines in the serum (Fig. 1A) were also detected in ARDS patients induced by SARS-CoV-2, as we expected. Meanwhile, we evaluated the levels of some cytokines in PRP. BDNF and EGF, rich in platelets, were found to be declined in PRP from ARDS patients, possibly due to the reduced platelet count.
In summary, we highlight an active state of platelet regeneration under the inflammation condition related to SARS-CoV-2-induced ARDS and a myeloid/MK-differentiation bias. Previously, strict policies against the COVID-19 epidemic have curbed the sustained transmission of SARS-CoV-2 in China, but on the other hand, chances to dissect pathological mechanisms were limited to a certain extent. With the realization of heard immunity recently, the number of SARS-CoV-2-induced ARDS patients has significantly decreased, which further limited our access to obtain a sufficient number of clinical samples for validation of our results and more investigations beyond our findings presented here. Future studies could benefit from collaborations with multiple research centers. Despite these limitations, our findings provide new insight into the response of the hematopoietic system to SARS-CoV-2, which is fundamental for deepening our understanding and developing new therapeutics for coronavirus infection.
Methods
Patients in this study
In this cohort study, we enrolled 70 COVID-19 patients, including 48 patients with mild lung infections (< 30% area) and 22 severe ARDS patients and we investigated blood routines and levels of cytokines from these patients.
The whole blood, from 18 non-COVID-19 samples and 40 severe ARDS patients with COVID-19 hospitalized at Shanghai General Hospital, is collected into anticoagulant citrate dextrose-anticoagulated (ACD-A) treated tube. A total of 58 whole blood samples were used for FACS analysis, evaluation of platelet activation, ELISA, RNA-seq, and Giemsa staining. All in vitro experiments were conducted in the laboratory of HemaCell Biotechnology Inc. We used N-95 respirators to prevent the transmission of COVID-19. Shanghai General Hospital Health Regulatory Agency and the ethics committee approved the study.
Flow cytometry analysis and sorting
The human hematopoietic stem and progenitor populations include HSC/MPP, CLP, CMP, GMP, MEP, MkP, and EP, and the cell surface marker cocktail for each population was defined as follows: HSC/MPP, Lin-CD34+CD38-; HSC, Lin-CD34+CD38-CD90+CD45RA-; MPP, Lin-CD34+CD38-CD90-CD45RA-; CMP, Lin-CD34+CD38+CD123+CD45RA-; CLP, Lin-CD34+CD38+CD127+; GMP, Lin-CD34+CD38+CD123+CD45RA+; MEP, Lin-CD34+CD38+CD123-CD45RA-; MkP, Lin-CD34+CD38+CD123-CD45RA-CD41+; EP, Lin-CD34+CD38+CD123-CD45RA-CD71+CD105+.
The cell surface markers used to define the mature blood populations are: T cell, CD3+; B cell, CD19+; NK cell, CD56+; Myeloid cell, CD36+; MK cell, CD41+.
The antibodies used in this study are: Lin FITC, CD45RA BV510, CD34 PE/Cy7, CD38 PE/Cy5, CD90 APC, CD123 AF700, CD127 PE, CD41 PerCP/Cy5.5, CD71 APC/Cy7, CD105 BV421, CD3 PE, CD11b AF780, CD19 FITC, CD56 APC, live/dead dyes: PI and DAPI (BioLegend).
Flow cytometry analysis was performed on LSRFortessa X-20 (BD Biosciences). Flow cytometry sorting for HSCs/MPPs (refer to Supplementary Fig. 3 for gating strategy) was performed on FACS Aria (BD Biosciences). All flow cytometry data were analyzed with FlowJo (Tree Star).
RNA-sequencing
The cells were isolated from whole blood samples after red blood cell (RBC) lysis buffer (Invitrogen) treatment. HSCs/MPPs sorted from non-COVID-19 individuals and COVID-19 ICU-ARDS patients were used to generate Smart-Seq2 libraries at Suzhou Geekgene Technology Co., Ltd, and subject to RNA-sequencing analysis. Differentially expressed genes (DEGs) are marked in red (p value ≤ 0.05 and fold change of gene expression ≥ 1.2) and in blue (p value ≤ 0.05 and fold change of gene expression ≤ -1.2) in Volcano plot. The DEG set, together with the hematopoietic cell lineage gene set and inflammatory response gene set, constituted the Venn diagram. The Gene Ontology (GO) analysis, KEGG pathway analysis, and Gene Set Enrichment Analysis (GSEA) were performed with the Geekgene Bioinformatics Service Portal/Platform (http://bioinfo.geekgene.com.cn/).
Platelet activation assay
Platelet activation was analyzed by incubating 10 μM ADP (Sigma-Aldrich) for 30 min. The expression of CD62P and CD41 was analyzed with flow cytometry. The antibodies used in this study are CD41 PE and CD62P APC (BioLegend).
Enzyme-Linked Immunosorbent Assay (ELISA)
We collected the supernatant by centrifuging the whole blood. Platelet-rich plasma (PRP) was obtained by supernatant frozen in liquid nitrogen and thawed at 37 °C, which was repeated three times. PRP concentrations of BDNF, EGF, and MMP3 were evaluated using ELISA kits (ExCell Bio, Taicang) following the manufacturer’s instructions. PRP was diluted with phosphate-buffered saline (PBS). The dilution ratios are 1:25 (BDNF, MMP3) and 1:2 (EGF).
Giemsa staining
The Giemsa Staining Kit (Abcam, Shanghai) was used for cell morphology analysis.
Statistical analysis
Data are presented as the mean plus or minus standard deviation. The significance of differences between groups was determined by a two-tailed unpaired t-test. P-values < 0.05 were considered statistically significant. Statistical analysis was performed with GraphPad Prism 8.
Availability of data and materials
The data supporting the findings of this study are available within the paper and its Supplementary Information files.
References
Verity R, Okell LC, Dorigatti I, Winskill P, Whittaker C, Imai N, Cuomo-Dannenburg G, Thompson H, Walker PGT, Fu H, et al. Estimates of the severity of coronavirus disease 2019: a model-based analysis. Lancet Infect Dis. 2020;20(6):669–77. https://doi.org/10.1016/S1473-3099(20)30243-7.
Osuchowski MF, Winkler MS, Skirecki T, Cajander S, Shankar-Hari M, Lachmann G, Monneret G, Venet F, Bauer M, Brunkhorst FM, et al. The COVID-19 puzzle: deciphering pathophysiology and phenotypes of a new disease entity. Lancet Respir Med. 2021;9(6):622–42. https://doi.org/10.1016/S2213-2600(21)00218-6.
Attaway AH, Scheraga RG, Bhimraj A, Biehl M, Hatipoglu U. Severe covid-19 pneumonia: pathogenesis and clinical management. BMJ. 2021;372:n436.
Meyer NJ, Gattinoni L, Calfee CS. Acute respiratory distress syndrome. Lancet. 2021;398(10300):622–37. https://doi.org/10.1016/S0140-6736(21)00439-6.
Network C-IGobotR, the C-ICUI: Clinical characteristics and day-90 outcomes of 4244 critically ill adults with COVID-19: a prospective cohort study. Intensive Care Med 2021, 47(1):60–73. https://doi.org/10.1007/s00134-020-06294-x
Mazeraud A, Jamme M, Mancusi RL, Latroche C, Megarbane B, Siami S, Zarka J, Moneger G, Santoli F, Argaud L, et al. Intravenous immunoglobulins in patients with COVID-19-associated moderate-to-severe acute respiratory distress syndrome (ICAR): multicentre, double-blind, placebo-controlled, phase 3 trial. Lancet Respir Med. 2022;10(2):158–66. https://doi.org/10.1016/S2213-2600(21)00440-9.
Monsel A, Hauw-Berlemont C, Mebarki M, Heming N, Mayaux J, Nguekap Tchoumba O, Diehl JL, Demoule A, Annane D, Marois C, et al. Treatment of COVID-19-associated ARDS with mesenchymal stromal cells: a multicenter randomized double-blind trial. Crit Care. 2022;26(1):48. https://doi.org/10.1186/s13054-022-03930-4.
Seita J, Weissman IL. Hematopoietic stem cell: self-renewal versus differentiation. Wiley Interdiscip Rev Syst Biol Med. 2010;2(6):640–53. https://doi.org/10.1002/wsbm.86.
van der Meijden PEJ, Heemskerk JWM. Platelet biology and functions: new concepts and clinical perspectives. Nat Rev Cardiol. 2019;16(3):166–79. https://doi.org/10.1038/s41569-018-0110-0.
Engelmann B, Massberg S. Thrombosis as an intravascular effector of innate immunity. Nat Rev Immunol. 2013;13(1):34–45. https://doi.org/10.1038/nri3345.
Franco AT, Corken A, Ware J. Platelets at the interface of thrombosis, inflammation, and cancer. Blood. 2015;126(5):582–8. https://doi.org/10.1182/blood-2014-08-531582.
Koupenova M, Clancy L, Corkrey HA, Freedman JE. Circulating platelets as mediators of immunity, inflammation, and thrombosis. Circ Res. 2018;122(2):337–51. https://doi.org/10.1161/CIRCRESAHA.117.310795.
Sanjuan-Pla A, Macaulay IC, Jensen CT, Woll PS, Luis TC, Mead A, Moore S, Carella C, Matsuoka S, Bouriez Jones T, et al. Platelet-biased stem cells reside at the apex of the haematopoietic stem-cell hierarchy. Nature. 2013;502(7470):232–6. https://doi.org/10.1038/nature12495.
Haas S, Hansson J, Klimmeck D, Loeffler D, Velten L, Uckelmann H, Wurzer S, Prendergast AM, Schnell A, Hexel K, et al. Inflammation-induced emergency megakaryopoiesis driven by hematopoietic stem cell-like megakaryocyte progenitors. Cell Stem Cell. 2015;17(4):422–34. https://doi.org/10.1016/j.stem.2015.07.007.
Martins-Goncalves R, Campos MM, Palhinha L, Azevedo-Quintanilha IG, Abud Mendes M, Ramos Temerozo J, Toledo-Mendes J, Rosado-de-Castro PH, Bozza FA, Souza Rodrigues R, et al. Persisting platelet activation and hyperactivity in COVID-19 survivors. Circ Res. 2022;131(11):944–7. https://doi.org/10.1161/CIRCRESAHA.122.321659.
Manne BK, Denorme F, Middleton EA, Portier I, Rowley JW, Stubben C, Petrey AC, Tolley ND, Guo L, Cody M, et al. Platelet gene expression and function in patients with COVID-19. Blood. 2020;136(11):1317–29. https://doi.org/10.1182/blood.2020007214.
Hottz ED, Azevedo-Quintanilha IG, Palhinha L, Teixeira L, Barreto EA, Pao CRR, Righy C, Franco S, Souza TML, Kurtz P, et al. Platelet activation and platelet-monocyte aggregate formation trigger tissue factor expression in patients with severe COVID-19. Blood. 2020;136(11):1330–41. https://doi.org/10.1182/blood.2020007252.
Zhang S, Liu Y, Wang X, Yang L, Li H, Wang Y, Liu M, Zhao X, Xie Y, Yang Y, et al. SARS-CoV-2 binds platelet ACE2 to enhance thrombosis in COVID-19. J Hematol Oncol. 2020;13(1):120. https://doi.org/10.1186/s13045-020-00954-7.
Grasselli G, Tonetti T, Protti A, Langer T, Girardis M, Bellani G, Laffey J, Carrafiello G, Carsana L, Rizzuto C, et al. Pathophysiology of COVID-19-associated acute respiratory distress syndrome: a multicentre prospective observational study. Lancet Respir Med. 2020;8(12):1201–8. https://doi.org/10.1016/S2213-2600(20)30370-2.
Long W, Yang J, Li Z, Li J, Chen S, Chen D, Wang S, Li Q, Hu D, Huang J, et al. Abnormal fibrinogen level as a prognostic indicator in coronavirus disease patients: a retrospective cohort study. Front Med (Lausanne). 2021;8:687220.
Zaid Y, Guessous F, Puhm F, Elhamdani W, Chentoufi L, Morris AC, Cheikh A, Jalali F, Boilard E, Flamand L. Platelet reactivity to thrombin differs between patients with COVID-19 and those with ARDS unrelated to COVID-19. Blood Adv. 2021;5(3):635–9. https://doi.org/10.1182/bloodadvances.2020003513.
Middleton EA, He XY, Denorme F, Campbell RA, Ng D, Salvatore SP, Mostyka M, Baxter-Stoltzfus A, Borczuk AC, Loda M, et al. Neutrophil extracellular traps contribute to immunothrombosis in COVID-19 acute respiratory distress syndrome. Blood. 2020;136(10):1169–79. https://doi.org/10.1182/blood.2020007008.
Li Y, Zhao K, Wei H, Chen W, Wang W, Jia L, Liu Q, Zhang J, Shan T, Peng Z, et al. Dynamic relationship between D-dimer and COVID-19 severity. Br J Haematol. 2020;190(1):e24–7. https://doi.org/10.1111/bjh.16811.
Yu HH, Qin C, Chen M, Wang W, Tian DS. D-dimer level is associated with the severity of COVID-19. Thromb Res. 2020;195:219–25. https://doi.org/10.1016/j.thromres.2020.07.047.
Petrella C, Zingaropoli MA, Ceci FM, Pasculli P, Latronico T, Liuzzi GM, Ciardi MR, Angeloni A, Ettorre E, Menghi M et al: COVID-19 affects serum brain-derived neurotrophic factor and neurofilament light chain in aged men: implications for morbidity and mortality. Cells 2023, 12(4).
Schulte-Schrepping J, Reusch N, Paclik D, Bassler K, Schlickeiser S, Zhang B, Kramer B, Krammer T, Brumhard S, Bonaguro L, et al. Severe COVID-19 Is Marked by a Dysregulated Myeloid Cell Compartment. Cell. 2020;182(6):1419-1440 e1423. https://doi.org/10.1016/j.cell.2020.08.001.
Silvin A, Chapuis N, Dunsmore G, Goubet AG, Dubuisson A, Derosa L, Almire C, Henon C, Kosmider O, Droin N, et al. Elevated calprotectin and abnormal myeloid cell subsets discriminate severe from mild COVID-19. Cell. 2020;182(6):1401-1418 e1418.
Qin G, Liu S, Yang L, Yu W, Zhang Y. Myeloid cells in COVID-19 microenvironment. Signal Transduct Target Ther. 2021;6(1):372. https://doi.org/10.1038/s41392-021-00792-0.
Liu Y, Zuo X, Chen P, Hu X, Sheng Z, Liu A, Liu Q, Leng S, Zhang X, Li X, et al. Deciphering transcriptome alterations in bone marrow hematopoiesis at single-cell resolution in immune thrombocytopenia. Signal Transduct Target Ther. 2022;7(1):347. https://doi.org/10.1038/s41392-022-01167-9.
Yang L, Xie X, Tu Z, Fu J, Xu D, Zhou Y. The signal pathways and treatment of cytokine storm in COVID-19. Signal Transduct Target Ther. 2021;6(1):255. https://doi.org/10.1038/s41392-021-00679-0.
Bester J, Pretorius E. Effects of IL-1beta, IL-6 and IL-8 on erythrocytes, platelets and clot viscoelasticity. Sci Rep. 2016;6:32188. https://doi.org/10.1038/srep32188.
Investigators R-C, Gordon AC, Mouncey PR, Al-Beidh F, Rowan KM, Nichol AD, Arabi YM, Annane D, Beane A, van Bentum-Puijk W, et al. Interleukin-6 Receptor antagonists in critically Ill patients with Covid-19. N Engl J Med. 2021;384(16):1491–502. https://doi.org/10.1056/NEJMoa2100433.
Zhang S, Qu J, Wang L, Li M, Xu D, Zhao Y, Zhang F, Zeng X. Activation of toll-like receptor 7 signaling pathway in primary sjogren’s syndrome-associated thrombocytopenia. Front Immunol. 2021;12:637659.
Acknowledgements
The authors thank members of the Zhu lab and of HemaCell Biotechnology Inc. for technical support and for discussion during preparation of this manuscript.
Funding
This work was supported by National Natural Science Foundation of China (82100140, 82070335 and 81570293), the STI2030 project (STI2030-SQ2021AAA010745), the 111 project (B21024), and the National Key R&D Program (2020YFC2004703).
Author information
Authors and Affiliations
Contributions
GW and FZ designed the study; GW and JH enrolled patients in the study; LW, HL, JL, HY, CW, XL, KX and GW performed laboratory work and analyzed the data; LW, HL, JL, HY and FZ wrote the manuscript; all authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
All COVID-19 patients were identified under study protocols approved by the Institutional Review Board (IRB) of Shanghai General Hospital (IRB number 2023–141). Non-infected group were enrolled under the same IRB protocol. All study participants or their legal authorized representatives gave written informed consent for study enrollment in accordance with Shanghai General Hospital.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1:
Supplementary Figure 1. The Giemsa staining for the blood smear of theperipheral blood from COVID-19 ARDS patients. Neutrophils, indicated by redcircles. Scale bar, 20 μm. Supplementary Figure 2. The characterization of CD41 expression in HSCs/MPPs. (A) The decreased percentage of CD41-positive cells in HSC/MPP of COVID-19 ARDS patients. (B) The average expression intensity of CD41 on the surface of CD41+ HSCs/MPPs. Supplementary Figure 3. The gating strategyto sort out Lin-CD34+CD38-CD45RA- HSCs/MPPs for RNA-seq.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, L., Liu, H., Liu, J. et al. Hyperactivity of platelets and increased megakaryopoiesis in COVID-19 patients with acute respiratory distress syndrome. Med-X 1, 8 (2023). https://doi.org/10.1007/s44258-023-00009-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44258-023-00009-9