Abstract
Growing concern around global warming has led to an increase in research focused on plant responses to increased temperature. In this review, we highlight recent advances in our understanding of plant adaptation to high ambient temperature and heat stress, emphasizing the roles of plant light signaling in these responses. We summarize how high temperatures regulate plant cotyledon expansion and shoot and root elongation and explain how plants use light signaling to combat severe heat stress. Finally, we discuss several future avenues for this research and identify various unresolved questions within this field.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Most people living in Europe, China, and the US have experienced extreme heat waves over the course of this summer (Witze, 2022). Climate researchers predict that long-lasting heat stress will become increasingly common in the future (Chen et al., 2020) and suggest that even a 1 °C increase in the global average temperature will result in a reduction in global crop yields by 3.1% (in soybean) to 7.4% (in maize) (Zhao et al., 2017). Therefore, in addition to the work being done to control global carbon emissions, we must urgently focus on the production of high-temperature resilient plants to solve the food security issue presented by global warming.
Plants, as sessile organisms, are highly sensitive to increasing temperatures. However, they are adaptive as can be seen when using Arabidopsis thaliana as an example. In these plants, increases in temperature from their normal growth (22 °C) to high ambient temperature (28 °C) induce a series of morphological changes (thermomorphogenesis), including hypocotyl elongation, leaf petiole elongation, root elongation, reduced cotyledon expansion, and leaf hyponasty (Casal and Balasubramanian, 2019; Delker et al., 2022; Quint et al., 2016), to reduce temperature stress. Experimental evidence demonstrates that plants use thermomorphogenesis to cool their leaf surface as a means of reducing heat damage (Crawford et al., 2012). However, when temperatures further increase, moving beyond 37 °C, most plants will encounter severe heat stress, which reduces long-term survival because most plants cannot survive long-term exposure to these extreme temperatures (Zhu et al., 2022).
Over the past decade, many well-designed studies have illustrated how plants sense and signal high temperatures to regulate their growth and development. Interestingly, plant photoreceptors and light-signaling components play crucial roles in plant thermomorphogenesis, with several elegant review papers summarizing the interactions between light and temperature (Bian et al., 2022; Ding and Yang, 2022; Jin and Zhu, 2019; Li et al., 2022; Qi et al., 2022). Given this, we have avoided potential content overlap with these reviews by focusing on a discussion of the various interactions between light and temperature and their roles in the regulation of individual plant organ growth. Readers are encouraged to read the other review papers mentioned above for a more nuanced view of the crosstalk between light and high ambient temperature signaling. Please be noted that plants mentioned in this review are Arabidopsis thaliana if not specified. The temperature ranges for thermomorphogenesis are 26–30 °C (aka high ambient temperature) and heat stress temperatures are 37–45 °C.
Light signaling and thermomorphogenesis
Light, the most influential environmental cue in plants, regulates and facilitates photosynthesis and serves as a signal to control plant growth and development at almost all stages of life, ranging from seedling germination to fruit ripening. Plants use various photoreceptors to sense light of various wavelengths, with the best-described systems found in Arabidopsis thaliana. UV RESISTANCE LOCUS8 (UVR8) senses short-wavelength UV-B light (280–320 nm) (Podolec et al., 2021), while blue light (390–500 nm) sensing is supported by several different types of blue light photoreceptors. CRYPTOCHROME1 (CRY1) and CRY2 are two of the most important blue light photoreceptors known to control plant hypocotyl elongation, flowering time, and circadian rhythm (Wang and Lin, 2020). From an evolutionary biology perspective, CRYs are the only photoreceptors found in all organisms, ranging from bacteria to humans. Three F-box proteins, ZEITLUPE (ZTL), light/oxygen/voltage (LOV) KELCH PROTEIN2 (LKP2), and FLAVIN-BINDING KELCH REPEAT-BOX1 (FKF1), are also able to directly sense blue light as photoreceptors and regulate floral transition and clock (Ito et al., 2012), while PHOTOTROPIN1 (PHOT1) and PHOT2 blue light photoreceptors, contribute to the phototropic response (Hart and Gardner, 2021). Red and far-red light (600–750 nm) photoreceptors phytochromes are the firstly identified plant photoreceptors. There are five phytochromes in A. thaliana (phyA-phyE). Unlike other photoreceptors, phytochromes are found in both active and inactive forms that are reversibly switched under red or far-red light irradiation. In these systems, red light absorption activates the phytochromes (Pfr form) and induces their translocation to the nucleus, where they trigger the red light response. However, far-red light treatment inactivates Pfr, returning them to their inactive form and inhibiting their signaling function (Cheng et al., 2021).
Light signal transduction mechanisms have been largely resolved through extensive genetic screening and protein–protein interaction assays. Interestingly, although individual photoreceptors perceive a specific light spectrum, the molecular basis of their downstream signaling is highly conserved. In general, there are two major paradigms of light signaling. The first model for light signaling relies on protein degradation, with E3 ligase CONSTITUTIVE PHOTOMORPHOGENIC1 (COP1) acting as the central negative regulator of these systems. COP1, initially identified using forward genetic screening in the 1990s (Deng et al., 1991, 1992), directly targets many transcription factors for degradation via the 26S proteasome-mediated protein degradation pathway. Transcription factor ELONGATED HYPOCOTYL5 (HY5) was the first reported COP1 substrate known to directly control the expression of light-responsive genes (Osterlund et al., 2000; Zhang et al., 2011). Light-activated photoreceptors repress COP1 activity to stabilize COP1-targeted transcription factors and elicit a light response. However, the interaction between photoreceptor CRY and COP1 is not light-dependent (Wang et al., 2001; Yang et al., 2000); therefore, there was a large gap in our understanding of this pathway until a series of breakthrough revelations were reported in 2011. COP1 E3 ligase activity is stimulated through interaction with the SUPPRESSOR OF PHYA (SPA) proteins (SPA1-SPA4) (Seo et al., 2003). Chentao Lin’s group and Hong-Quan Yang’s group separately reported that CRYs interact with SPA proteins in a blue light-dependent manner, which further curtails COP1-SPA interactions and reduces COP1 activity (Lian et al., 2011; Liu et al., 2011a; Zuo et al., 2011). Subsequent studies have also demonstrated that phytochromes specifically interact with SPA following red light exposure and function to abrogate COP1-SPA interactions (Lu et al., 2015; Sheerin et al., 2015). These studies clearly illustrate how photoreceptors regulate COP1 activity in a light-dependent manner.
The second paradigm involves interaction between photoreceptors and transcription factors. This mechanism is significant because all photoreceptors, except for the membrane-localized phototropin, translocate or are constitutively located within the nucleus upon light irradiation (Chen et al., 2005; Hiltbrunner et al., 2005; Liu et al., 2022; Qian et al., 2016; Yin et al., 2016; Yu et al., 2007). These receptors then mediate their impact via various PHYTOCHROME INTERACTING FACTORs (PIFs), which were the first reported phytochrome-interacting transcription factors (Ni et al., 1998; 1999). CRY1/CRY2 also interact with PIFs (Ma et al., 2016; Pedmale et al., 2016). In A. thaliana, there are seven PIF family members, belonging to the basic helix-loop-helix (bHLH) transcription factor family. These seven PIF proteins play partially redundant roles, although several are also known to perform fairly specific functions under certain conditions (Xu et al., 2015). Most PIF transcription factors are positive regulators of cell elongation that directly bind to auxin biosynthesis-related gene promoters and activate their transcription (Franklin et al., 2011; Sun et al., 2012). Phytochromes interact with PIFs and rapidly trigger PIF protein degradation (Al-Sady et al., 2006; Lorrain et al., 2008; Park et al., 2004; Shen et al., 2008, 2007), while interactions between CRY and PIF result in the inhibition of PIF transcriptional activity (Ma et al., 2016; Pedmale et al., 2016). Photoreceptors are not limited to their interactions with PIF transcription factors but are also known to interact with the majority of transcription factors associated with plant hormone signaling with different regulatory mechanisms. For example, CRY1 and PhyB interact with AUXIN/INDOLE-3-ACETIC ACID-INDUCIBLE (AUX/IAA) proteins (Xu et al., 2018). AUX/IAA proteins are key transcriptional repressors of auxin signaling; these interactions safeguard the stability of AUX/IAA proteins and modulate auxin signaling (Xu et al., 2018). CRY1 and PhyB also interact with AUXIN RESPONSE FACTOR6 (ARF6) and ARF8, regulating their DNA-binding activity and controlling hypocotyl elongation (Mao et al., 2020). UVR8, PhyB, and CRY1 also interact with BRI1-EMS-SUPPRESSOR1 (BES1) to repress hypocotyl elongation via the modulation of brassinosteroid (BR) signaling (Liang et al., 2018; Wang et al., 2018; Wu et al., 2019).
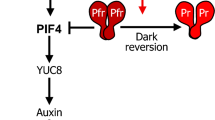
Interestingly, most of these light signaling components also participate in plant responses to high ambient temperature. This is because high ambient temperature stimulates the dark reversion process changing the ratio of active to inactive photoreceptor. PhyB, the first thermosensor identified in A. thaliana, is known to rely on changes in the Pr:Pfr ratio to help the plant sense changes in temperature (Jung et al., 2016; Legris et al., 2016). A recent study revealed that phyB undergoes liquid–liquid phase separation (LLPS) under red light and colocalizes with various signaling components, including PIF4, within the phyB condensates. This suggests that the phyB condensates may serve as a hub for signal incorporation. Furthermore, high-temperature treatments reduce phyB LLPS, which is essential for thermomorphogenesis (Chen et al., 2022). High temperatures also modulate the LLPS of EARLY FLOWERING3 (ELF3), a key component in the plant evening complex, which in turn negatively regulates PIF4 expression and transcriptional activity (Box et al., 2015; Nieto et al., 2015; Nusinow et al., 2011; Raschke et al., 2015). The prion domain (PrD) in ELF3 senses high ambient temperatures and facilitates ELF3 LLPS to inactivate the evening complex (Jung et al., 2020). Phototropins also sense temperature changes, as illustrated in a study of Marchantia polymorpha. In this study, the plants were shown to take advantage of temperature-sensitive photoactivated phototropin activity to regulate chloroplast positions for optimized photosynthesis (Fujii et al., 2017).
PIF4 is a central positive regulator of thermomorphogenesis, with plants without PIF4 (pif4 mutant) known to be largely insensitive to high temperatures at both the phenotype and genome-wide transcriptome levels (Jin et al., 2020; Koini et al., 2009; Kumar et al., 2012). PIF7 interacts with PIF4 and co-regulates gene expression and thermomorphogenesis (Chung et al., 2020; Fiorucci et al., 2020). A recent study argued that PIF7, but not PIF4, is essential for the synergistic effects observed for hypocotyl elongation in response to combining higher ambient temperatures and shade conditions (Burko et al., 2022). Further studies demonstrated that high temperatures enhance PIF7 mRNA translation because of a change in the conformation of the 5′-untranslated region of PIF7 mRNA, which increases translation efficiency (Chung et al., 2020). These hairpin structures also exist in many other high-temperature-responsive mRNAs, including WRKY22 and HsfA2, suggesting that this hairpin structure is a conserved thermo-sensing mechanism (Chung et al., 2020).
In addition, high-temperature-triggered hypocotyl elongation is repressed specifically under blue light irradiation. Blue light-activated CRY1 interacts with PIF4 and inhibits its transcriptional activity, which suppresses thermomorphogenesis (Ma et al., 2016). Another recent study reported that CRY2 accumulation is controlled by temperature, and low ambient temperature (16 °C) conditions increase CRY2 protein degradation via its strong interaction with E3 ligase Light-Response Bric-a-Brack/Tramtrack/Broad (LRB) proteins (LRB1/LRB2/LRB3) (Ma et al., 2021). LRBs directly ubiquitinate CRY2 and stimulate protein turnover via the 26S proteasome (Chen et al., 2021; Ma et al., 2021). These results demonstrate that CRYs participate in plant responses to high ambient temperatures.
COP1-SPA1 also positively regulates thermomorphogenesis (Delker et al., 2014; Nieto et al., 2022; Park et al., 2017) because high temperatures stimulate COP1 nuclear localization and facilitate HY5 degradation (Park et al., 2017). HY5 directly represses PIF4 transcription, and the degradation of HY5 results in the accumulation of PIF4 mRNA (Delker et al., 2014). HY5 also competes with PIF4 to occupy the PIF4 DNA-binding sites, adding another layer of antagonism to these two transcription factors (Bian et al., 2022; Gangappa and Kumar, 2017). In addition to its interactions with COP1, SPA1 also exerts some Ser/Thr kinase activity (Lee et al., 2020; Paik et al., 2019; Wang et al., 2021a). Thus, SPA1 can directly phosphorylate PIF4, with this activation being essential for PIF4 protein stability. This was confirmed by the fact that the absence of all four SPA proteins (spaQ mutants) results in a significant reduction in PIF4 expression. This data also revealed that spaQ plants presented with much shorter hypocotyls under high ambient temperature conditions (Lee et al., 2020).
Taken together, these data reveal that light-signaling components are critical players in the high-temperature response for most plants. In the next sections, we describe the roles of various light signaling components in the regulation of plant growth at high ambient temperatures (Fig. 1) and under high-temperature stress (Fig. 2).
High ambient temperature inhibits plant leaf expansion and promotes cell elongation in both hypocotyl and root. A In the leaves, high temperature activated transcription factor PIF4 interacts with TCP4. The TCP4-PIF4 module binds to KRP1 promoter and induces KRP1 expression to inhibit cell division. BES1 is a positive regulator for cell expansion. PIF4 directly represses BES1 transcription. On another side, high temperature increases COP1 nuclear abundances to directly target BES1 protein for degradation. These mechanisms act together to restrict leaf expansion under high ambient temperature. B PIF4 plays central roles in the high temperature induced hypocotyl elongation. PIF4 directly binds to the promoter of YUCCA8 to induce YUCCA8 expression for boosting auxin biosynthesis. The accumulated auxin further promotes the expression of SAURs. PIF4 is also able to induce SAUR transcription directly. SAUR proteins interact with PP2C.D enzymes and inhibit their enzymatic activities. The inactivation of PP2C.D further releases their repression on H+-ATPase to trigger proton transport into the apoplast. The reduction of apoplastic pH results in cell elongation due to the cell wall loosen and water uptake increase. A lot of co-factors (depicted as INO80-COMPASS, BIC1-BZR1) interact with PIF4 to enhance its transcriptional activities. SPA1 phosphorylates PIF4 to control PIF4 protein stability. The binding of PIF4 with DNA will facilitate other transcription factor (like CDF2) binding to DNA as well. High temperature also stimulates COP1 nucleus localization and enhances HY5 protein turn over. HY5 not only directly represses PIF4 transcription but also interacts with PIF4 protein to abrogate its transcriptional activity. The removal of HY5 results in the activation of PIF4. C High temperature triggers HY5 protein accumulation in roots, which directly controls the expression of brassinosteroid (BR) metabolism genes (CPD, BAS1 and SOB7) and root cell elongation. SPA1 also regulates HY5 protein phosphorylation and protein stability
Regulating cotyledon expansion
Cotyledons are specialized organs designed to provide nutrients for seed germination (Chandler, 2008), and both the initiation and outgrowth of these structures are integrated into the embryonic patterning program. In A. thaliana, the zygote undergoes cell division to form a radially symmetrical embryo, called a globular embryo, and the cotyledon primordia arise as two distinct bumps from the apex of this structure. These embryos are then converted into one embryo with two planes of bilateral symmetry (Bowman and Floyd, 2008; Chandler et al., 2008; Ito et al., 2011). Previous studies have indicated that the symmetrical positioning of the cotyledon primordia and establishment of cotyledon boundaries correlate with changes in auxin distribution, and treatment with exogenous or polar auxin transport inhibitors results in varying defects across the apical patterning of these embryos, including abnormal positioning and cotyledon fusion (Blilou et al., 2005; Friml et al., 2003; Vieten et al., 2005).
Cotyledon expansion is a major agronomic trait for early seedling vigor (Hahm et al., 2020; Li et al., 2012; Neff and Van Volkenburgh, 1994), known to depend exclusively on cell expansion (Gonzalez et al., 2012; Tsukaya et al., 1994). A. thaliana SPATULA (SPT) has been identified as a potent regulator of cotyledon expansion, with spt mutants displaying significantly expanded cotyledons with larger pavement cells than the wild-type plants when grown under red-light conditions (Josse et al., 2011; Penfield et al., 2005). Furthermore, cell expansion in plants is often associated with endoreplication, a cell cycle variant during which several rounds of genome duplication occur without subsequent cell division (De Veylder et al., 2011; Marco D'Ario, 2021; Shu et al., 2018). Exogenous nitrate upregulates the CDK-specific inhibitor gene, LOSS in GIANT CELLS FROM ORGANS (LGO), to increase cotyledon endoreplication and cell size (Moreno et al., 2020). Notably, the bHLH transcription factor CYTOKININ-RESPONSIVE GROWTH REGULATOR (CGK) reportedly promotes cell cycle movement into the S phase, inducing cellular expansion in a ploidy-independent manner in these organs (Park et al., 2021).
High ambient temperatures suppress cotyledon expansion in A. thaliana (Delker et al., 2022; Hahm et al., 2020), with increased temperatures activating PIF4 and thus inducing auxin biosynthesis in the cotyledons (Jung et al., 2016; Legris et al., 2016; Procko et al., 2014). Although auxin moves to the hypocotyl to promote cell elongation, excessive accumulation of auxin in the cotyledons inhibits cell expansion (Bellstaedt et al., 2019; Enders et al., 2017). BRI1-EMS-SUPPRESSOR1 (BES1) is a vital transcription factor known to control brassinosteroid-regulated gene expression and promote hypocotyl and cotyledon growth in A. thaliana (Nolan et al., 2020). A recent study found that pif4 mutants presented with elevated BES1 mRNA levels in cotyledons, indicating that PIF4 likely represses BES1 expression in cotyledon cells. Further analysis also showed that shade or heat exposure increases PIF4 nuclear localization in cotyledon cells increasing the transcriptional repression of BES1 and restricting cotyledon expansion (Costigliolo Rojas et al., 2022). In addition, post-transcriptional evaluations reveal that both shade and heat exposure increase COP1 nuclear localization in cotyledon cells, facilitating increased BES1 protein degradation and thereby reducing cotyledon expansion (Costigliolo Rojas et al., 2022). Therefore, the spatial regulation of COP1 and PIF4 nuclear localization in cotyledon cells shapes cotyledon growth plasticity under high ambient temperatures.
As cotyledons and true leaves have similar organ expansion programs and mature morphologies, they can be considered homologous (Chandler, 2008). Thus, it is not surprising that high ambient temperature also reduces the expansion of the true leaf blade (Ibanez et al., 2017; Jin et al., 2011). These observations were supported by a recent report which showed that PIF4/TCP4 suppresses leaf area by inhibiting cell division under high temperatures (Saini et al., 2022). TCP4 is a CINCINNATA (CIN)-like TEOSINTE BRANCHED1/CYCLOIDEA/PCF (TCP) transcription factor, which inhibits cell division by activating cell cycle inhibitor gene KIP-RELATED PROTEIN1 (KRP1) (Schommer et al., 2014). PIF4 physically interacts with TCP4 to promote the expression of KRP1, which in turn inhibits cell division, reducing leaf size under high temperatures (Saini et al., 2022).
Taken together, these results reveal that high ambient temperature initially increases auxin biosynthesis in cotyledons by activating PIF4, and excessive residual auxin levels in cotyledons inhibit cell expansion. Second, high ambient temperature downregulates nuclear BES1 levels via PIF4-mediated transcriptional regulation and COP1-mediated post-transcriptional control to suppress cotyledon expansion. Third, high ambient temperatures induce the accumulation of PIF4 in leaf cells to reduce cell division and leaf size in a TCP4-dependent manner (Fig. 1A).
Regulating hypocotyl elongation
A. thaliana hypocotyls elongate in response to increased ambient temperature (Gray et al., 1998), and these elongated hypocotyls are the primary characteristic by which we can recognize thermomorphogenesis. Like in the examples above, PIF4 acts as a central regulator for plant hypocotyl elongation under high ambient temperatures, with these increased temperatures inducing PIF4 transcription and stabilizing PIF4 protein for its activation (Koini et al., 2009; Qiu, 2020). These activated proteins then directly associate with the YUCCA8 promoter and induce YUCCA8 expression (Sun et al., 2012). YUCCA8, in turn, encodes a flavin monooxygenase necessary for auxin biosynthesis, which converts indole-3-pyruvic acid (IPA) into indole-3-acetic acid (IAA) (Dai et al., 2013; Mashiguchi et al., 2011; Stepanova et al., 2011; Won et al., 2011). The loss-of-function yucca8 mutants display short hypocotyls under high ambient temperature, which further suggests that the induction of YUCCA8 expression is required for thermomorphogenesis (Sun et al., 2012). Interestingly, YUCCA8 expression is significantly increased in the cotyledons following exposure to high temperatures. Additional experiments demonstrated that hypocotyl elongation is dependent on cotyledon-derived auxins, suggesting that plant cotyledons sense high temperatures and trigger auxin production, which in turn stimulates cell elongation, eventually causing hypocotyl elongation (Bellstaedt et al., 2019). Tissue-specific expression analysis has also shown that epidermal expression of PIF4 is sufficient to induce YUCCA8 expression and promote hypocotyl elongation (Kim et al., 2020).
Thus, the PIF4-YUCCA8 regulatory pathway serves as a hub connecting chromatin remodeling and phytohormone signaling. Recent studies have shown that the INO80 chromatin remodeling complex interacts with PIF4 and mediates H2A.Z eviction, including YUCCA8, at PIF4 binding sites (Xue et al., 2021). Interestingly, PIF4 also interacts with BRASSINAZOLE RESISTANT1 (BZR1), a key component in brassinosteroid signaling (Ibanez et al., 2018; Oh et al., 2012). PIF4-BZR1 co-occupies overlapping DNA regions on a genome-wide scale and synergistically regulates hypocotyl elongation (Ibanez et al., 2018; Oh et al., 2012). In addition, another recent study reported that BLUE-LIGHT INHIBITOR OF CRYPTOCHROME1 (BIC1) interacts with BZR1 and PIF4 to co-activate the transcription of their target genes, such as YUCCA8 (Yang et al., 2021). Interestingly, both BZR1 and BIC1 also positively regulate the expression of PIF4, thereby establishing a positive feedback loop to strengthen PIF4 activity (Ibanez et al., 2018; Yang et al., 2021). Further, the PIF4 protein can bind to its promoter to activate PIF4 transcription, creating a closed positive feedback loop (Zhai et al., 2020). PIF4 binding of the YUCCA8 promoter also changes the chromatin conformation and enhances the binding of the single zinc-finger transcription factor CYCLING DOF FACTOR2 (CDF2) to the YUCCA8 promoter (Gao et al., 2022). PIF4 also directly interacts with CDF2, and these two transcription factors cooperate to control hypocotyl elongation (Gao et al., 2022).
In addition, the auxin-mediated acid growth model is widely accepted for explaining cell elongation (Du et al., 2020). Here, accumulated auxin rapidly induces the expression of the SMALL AUXIN UP RNA (SAUR) genes (Stortenbeker and Bemer, 2019); these genes are also regulated via PIF4 interactions with their promotors, which enable increased expression of SAURs at high ambient temperatures (Franklin et al., 2011). SAUR19, in turn, interacts with the D-clade type-2C protein phosphatase (PP2C.D) and represses its enzymatic activity (Spartz et al., 2014), reducing plasma membrane H+-ATPase activity in these organs (Spartz et al., 2014; Takahashi et al., 2012), which in turn results in increased proton transport into the apoplast. The resulting decrease in apoplast pH loosens the cell wall and increases water uptake and cell elongation (Du et al., 2020). Thus, while the SAUR-PP2C–H+-ATPase signaling cascade remains reasonably straightforward, its modulation can facilitate very nuanced changes in its activity, facilitating complex signal crosstalk in response to changes in temperature. For example, ethylene exposure represses thermomorphogenesis through its key transcription factor, ETHYLENE INSENSITIVE3 (EIN3). Although EIN3 does not directly interact with PIF4, EIN3 does activate the expression of PP2C.D (ARABIDOPSIS PP2C CLADE D7, APD7), repressing H+-ATPase activity (Kim et al., 2021) and facilitating much more complex regulation of these response systems.
Thus taken together, we can conclude that high ambient temperatures initially activate PIF4 within the epidermis, inducing auxin biosynthesis. Auxin is then moved to the hypocotyl, where it acidifies the cell walls, inducing cellular elongation (Fig. 1B).
Regulating root elongation
High ambient temperature not only affects the growth and development of the aerial portions of plants but also their roots (Martins et al., 2017). However, unlike the extensive studies described for hypocotyl elongation, the root response to high ambient temperatures is much less well-defined (CFF et al., 2021). Similar to the promotion of hypocotyl elongation, high ambient temperatures stimulate root elongation under continuous light or long-day conditions (root thermomorphogenesis) (Borniego et al., 2022; Gaillochet et al., 2020; Lee et al., 2021; Martins et al., 2017). A detailed analysis has also demonstrated that high temperatures trigger cell elongation, but not division, to promote root elongation (Martins et al., 2017), and there are several interesting differences between hypocotyl and root thermomorphogenesis.
First, in contrast to the critical role of YUCCA8-dependent auxin biosynthesis in the regulation of hypocotyl elongation, high-temperature treatment does not significantly alter YUCCA8 expression, auxin marker gene expression, or auxin accumulation (Martins et al., 2017) in the roots. Auxin receptor mutants exhibit identical root elongation phenotypes to wild-type plants, suggesting that auxin biosynthesis is not involved in root thermomorphogenesis (Martins et al., 2017). Second, the thermosensors (phyB, ELF3, and PIF7) identified in hypocotyl thermomorphogenesis are not required for root thermomorphogenesis. This finding was exemplified by several studies that showed that even in phyB and elf3 mutants, the plants still produced normal-length roots (Borniego et al., 2022; Gaillochet et al., 2020). High-temperature treatment also did not reduce phyB nuclear body size, further indicating that phyB is not a thermosensor in root thermomorphogenesis. In addition, while high temperatures enhance PIF7 translation during hypocotyl thermomorphogenesis, there is no increase in its accumulation in the roots (Borniego et al., 2022). Therefore, we can assume that none of these three major thermosensors fulfill these roles in the roots. Third, PIF4 and its homologs are dispensable for root thermomorphogenesis, with the root elongation phenotypes in PIF4 loss-of-function mutants (pif4) and pifQ (pif1 pif3 pif4 pif5) quadruple mutants shown to be similar to those of wild-type plants (Gaillochet et al., 2020; Lee et al., 2021). In fact, PIF4 is primarily expressed in the aerial parts of the plant but not in the roots (Lee et al., 2021). Taken together, these results suggest that plant root responses to high ambient temperatures are distinct from that of their hypocotyls.
This was further supported by two independent studies, which reveal that HY5 is necessary for plant root thermomorphogenesis. They showed that hy5 mutants display short root phenotypes under high ambient temperatures (Gaillochet et al., 2020; Lee et al., 2021), and HY5 is predominantly expressed in the roots. These experiments also show that high temperatures induce HY5 transcription in the roots (Lee et al., 2021). In addition, subsequent experiments using cotyledon-specific promoter CAB3 to express the HY5 protein tagged with HA-YFP-HA (DOF-HY5) to restrict HY5 protein movement from shoot to root, results in the hy5 mutant background (pCAB3:DOF-HY5/hy5) not being able to rescue the hy5 short root phenotypes under high ambient temperature (Gaillochet et al., 2020). Transcriptomic analysis revealed that high temperatures elicit distinct differentially expressed gene profiles, and unlike the regulation of growth-related genes in hypocotyls, high-temperature exposure preferentially modulates metabolism-related genes in root samples (Gaillochet et al., 2020; Lee et al., 2021). For example, HY5 directly controls the expression of several genes (CPD, BAS1, and SOB7) involved in brassinosteroid metabolism, allowing this factor to modulate brassinosteroid signaling (Lee et al., 2021) involved in root thermomorphogenesis (Martins et al., 2017) (Fig. 1C).
Furthermore, SPA proteins phosphorylate PIF4, and this phosphorylation is required for PIF4 stability and hypocotyl thermomorphogenesis (Lee et al., 2020). Interestingly, this scenario recapitulates during root thermomorphogenesis, where SPA1 phosphorylates HY5 at serine 36 (S36) (Wang et al., 2021a). Overexpression of phosphor-null HY5 (S36A) in the hy5 mutants resulted in an inability to rescue the hy5 mutant root phenotypes under high ambient temperature, while overexpression of phospho-mimic HY5 (S36D) in the hy5 background mediated clear root elongation (Lee et al., 2021). Further immunoblot studies showed that HY5 phosphorylation was positively correlated with its protein stability in the roots. Thus, it was not surprising that spaQ mutants retain their short root phenotype under high ambient temperatures (Lee et al., 2021). Thus, we can conclude that SPA proteins are necessary for both hypocotyl and root thermomorphogenesis.
Roles in heat stress responses
The previous sections focused on plant responses to high ambient temperatures, such as temperatures that trigger thermomorphogenesis in A. thaliana (26–30 °C). However, when plants grow in temperatures above this limit, they no longer experience thermomorphogenesis but rather enter heat stress. Heat stress has many adverse effects on plant growth and development, including inhibition of germination, growth, reduced fertility, and cell death (heat stress responses) (Ohama et al., 2017). Heat shock transcription factors (Hsf) are rapidly induced in response to heat treatment and are known to function as master regulators of various HEAT SHOCK PROTEINs (HSPs) (Charng et al., 2007; Liu et al., 2011b; Ohama et al., 2017). HSPs are well-known molecular chaperones that facilitate heat-induced refolding of misfolded proteins (Jacob et al., 2017). A recent study revealed a mobile signal underlying the plant heat response, with heat exposure initially triggering nitric oxygen (NO) production at the apex of the inflorescence. This accumulated NO then interacts with glutathione to form S-nitrosoglutathione (GSNO), which then rapidly moves from the shoot to the root and promotes the production of transcription factor GT-1 S-nitrosylation. S-nitrosylated GT-1 then binds to additional NO-responsive elements in the HsfA2 promoter and induces HsfA2 expression, enhancing plant heat tolerance (He et al., 2022). This elegant study not only described the potential heat-sensing organs in plants but also reported a mobile signal for transmitting heat signals from the shoot to the root.
In addition, since PIF4 is a central regulator of thermomorphogenesis, many studies have evaluated its role in the heat stress response. These investigations revealed that similar to high ambient temperatures, heat treatment induces PIF4 transcription and stabilizes the PIF4 protein (Yang et al., 2022). Phenotypic observations (4-day-old etiolated seedlings grown at 22 °C and then placed at 45 °C for 1 h before being recovered at 22 °C for 3 days) illustrate that pif4 mutants are less tolerant of heat treatment, while plants overexpressing PIF4 show enhanced heat tolerance. Further studies have found that PIF4 directly binds to the HsfA2 promoter and induces HsfA2 expression, increasing heat tolerance (Yang et al., 2022). However, another study in adult plants demonstrated that PIF4 acts as a positive regulator of plant senescence at high temperatures (Li et al., 2021). One such study used 3-week-old plants treated at 42 °C for 5 h and then recovered for 3 days for observation and revealed that all of these plants started to produce obvious senescent phenotypes (Li et al., 2021). Additional evaluations revealed clear reductions in chlorophyll content in the wild-type plants following heat stress but significant increases in chlorophyll in pif4 mutants (Li et al., 2021). Consistent with the mutant phenotypes, chlorophyll content was greatly reduced in PIF4 overexpressing lines, and other data revealed that PIF4 directly binds to the promoter of several senescence-related genes (NAC019, SAG113, and IAA29), inducing their transcription and supporting the transition to senescence in response to heat treatment (Li et al., 2021). Although these studies suggest different roles for PIF4, both sets of results further confirm that PIF4-mediated transcriptional regulation is a critical factor in plant heat responses.
Light conditions also affect plant heat-stress responses. For example, comparisons of white light and shade (low-red/far-red ratios) growth conditions reveal a clear increase in heat shock (45 °C for 45 min) tolerance in the latter group (Arico et al., 2019). PhyB mutants consistently demonstrate enhanced heat tolerance, even under white light conditions (Arico et al., 2019; Song et al., 2017), and the expression levels of fatty acid desaturase (FAD) genes decreased in the shade, suggesting that this may contribute to the reduction in total unsaturated fatty acids and increased membrane stability in these plants (Arico et al., 2019).
In addition, the COP1-HY5 regulatory module also participates in the regulation of seed germination under heat stress, where ABA-INSENSITIVE5 (ABI5) acts as the key transcription factor in the abscisic acid (ABA) signaling pathway, which, in turn, regulates seed germination. Heat promotes COP1 movement from the nucleus to the cytoplasm, increasing HY5 stability and promoting its interactions with the ABI5 promoter, inducing its expression and increasing the inhibition of seed germination (Chen et al., 2019).
Taken together, these results show that light signaling components are also critical to the heat stress response in plants (Fig. 2).
Roles of the major light signaling components in plant heat responses. Light activated transcription factor PIF4, which directly induces HsfA2 expression to confer seedling heat tolerance. However, PIF4 also triggers senescence associated gene (NAC019, SAG113 and IAA29) expressions in adult plants to induce plant senescence. Simulated shade enhances plant heat tolerance through the repression of FAD expression to reduce the total unsaturated fatty acids. Heat also inhibits COP1 activity to stabilize HY5, which directly induces ABI5 expression to repress seed germination
Perspectives
Although we have achieved many milestones in understanding the high-temperature signaling response in plants, several unexplored phenomena need to be studied. These include:
-
(1)
Evaluation of the precise temperature sensing mechanisms. This is because plants tend to reduce growth in response to lower temperatures following chilling or freezing stress, albeit to different degrees, but increase their growth in response to higher ambient temperatures (26–30 °C; Phase 1), eventually entering restricted growth when temperatures reach the high-temperature stress threshold (Phase 2). We do not know how a plant measures these temperature changes or makes contrasting decisions to enter either of these two phases. Previous studies have focused only on phase 1 or phase 2, and further studies need to focus on their cross-over and simultaneous induction, with initial investigations focusing on known photoreceptors, including PhyB.
-
(2)
The roles of PIF4 in different developmental stage and temperature range are deserved to be further investigated. As we described, PIF4 positively regulates cell elongation in hypocotyl cells but negatively controls cell expansion in cotyledon cells under high ambient temperatures. Moreover, PIF4 enhances plant heat stress tolerance in seedlings but triggers senescence in adult plants. Therefore, we assume that the function of PIF4 varies according to distinct stages. The recently popular single cell analysis may contribute to a detailed understanding of PIF4 in a more spatial and temporal way.
-
(3)
The mechanisms underlying plant root thermomorphogenesis are not well understood. Although HY5-mediated signaling is required for root elongation at high ambient temperatures, other mechanisms are worthy of further study. This is important because we know that detached roots can elongate under high ambient temperatures, even in the absence of cotyledons and hypocotyls (Bellstaedt et al., 2019), indicating that plant roots can sense temperature without the need for various aerial components of these organisms. In fact, currently reported thermosensors (phyB, ELF3, and PIF7) do not participate in root thermomorphogenesis (Borniego et al., 2022), suggesting that the identification of root thermosensors may be a particularly fruitful avenue of investigation.
-
(4)
The LLPS of thermosensors, phyB and ELF3, suggests that high temperature-induced condensates are a common regulatory mechanism in plants, facilitating their nuanced response to changes in temperature. Since CRY2 also undergoes LLPS in response to blue light and co-condense with m6A writer proteins to regulate N6-methyladenosine modification and circadian rhythm (Wang et al., 2021b), the LLPS status of blue-light-activated CRY2 under high ambient temperatures deserves further exploration.
-
(5)
Finally, although we clearly understand these processes in Arabidopsis, there is an urgent need to evaluate its translational value in other plants to transfer this knowledge to the cultivation of food and fruit crops and increase their high-temperature tolerance.
Availability of data and materials
Not applicable.
Abbreviations
- UVR8:
-
UV RESISTANCE LOCUS8
- CRY1:
-
CRYPTOCHROME1
- ZTL:
-
ZEITLUPE
- LKP2:
-
Light/oxygen/voltage (LOV) KELCH PROTEIN2
- FKF1:
-
FLAVIN-BINDING KELCH REPEAT-BOX1
- PHOT1:
-
PHOTOTROPIN1
- COP1:
-
CONSTITUTIVE PHOTOMORPHOGENIC1
- HY5:
-
ELONGATED HYPOCOTYL5
- SPA:
-
SUPPRESSOR OF PHYA
- PIF4:
-
PHYTOCHROME INTERACTING FACTOR4
- AUX/IAA proteins:
-
AUXIN/INDOLE-3-ACETIC ACID-INDUCIBLE proteins
- ARF6:
-
AUXIN RESPONSE FACTOR6
- ELF3:
-
EARLY FLOWERING3
- LRB1:
-
Light-Response Bric-a-Brack/Tramtrack/Broad protein1
- BES1:
-
BRI1-EMS-SUPPRESSOR1
- BZR1:
-
BRASSINAZOLE RESISTANT1
- TCP4:
-
CINCINNATA (CIN)-like TEOSINTE BRANCHED1/CYCLOIDEA/PCF (TCP) transcription factor4
- KRP1:
-
KIP-RELATED PROTEIN1
- BIC1:
-
BLUE-LIGHT INHIBITOR OF CRYPTOCHROME1
- SAUR:
-
SMALL AUXIN UP RNA
- EIN3:
-
ETHYLENE INSENSITIVE3
- APD7:
-
ARABIDOPSIS PP2C CLADE D7
- HSP:
-
HEAT SHOCK PROTEIN
References
Al-Sady B, Ni W, Kircher S, Schafer E, Quail PH (2006) Photoactivated phytochrome induces rapid PIF3 phosphorylation prior to proteasome-mediated degradation. Mol Cell 23:439–446. https://doi.org/10.1016/j.molcel.2006.06.011
Arico D, Legris M, Castro L, Garcia CF, Laino A, Casal JJ, Mazzella MA (2019) Neighbour signals perceived by phytochrome B increase thermotolerance in Arabidopsis. Plant Cell Environ 42:2554–2566. https://doi.org/10.1111/pce.13575
Bellstaedt J, Trenner J, Lippmann R, Poeschl Y, Zhang X, Friml J, Quint M, Delker C (2019) A Mobile Auxin Signal Connects Temperature Sensing in Cotyledons with Growth Responses in Hypocotyls. Plant Physiol 180:757–766. https://doi.org/10.1104/pp.18.01377
Bian Y, Chu L, Lin H, Qi Y, Fang Z, Xu D (2022) PIFs- and COP1-HY5-mediated temperature signaling in higher plants. Stress Biology 2:35. https://doi.org/10.1007/s44154-022-00059-w
Blilou I, Xu J, Wildwater M, Willemsen V, Paponov I, Friml J, Heidstra R, Aida M, Palme K, Scheres B (2005) The PIN auxin efflux facilitator network controls growth and patterning in Arabidopsis roots. Nature 433:39–44. https://doi.org/10.1038/nature03184
Borniego MB, Costigliolo-Rojas C, Casal JJ (2022) Shoot thermosensors do not fulfil the same function in the root. New Phytol. https://doi.org/10.1111/nph.18332
Bowman JL, Floyd SK (2008) Patterning and polarity in seed plant shoots. Annu Rev Plant Biol 59:67–88. https://doi.org/10.1146/annurev.arplant.57.032905.105356
Box MS, Huang BE, Domijan M, Jaeger KE, Khattak AK, Yoo SJ, Sedivy EL, Jones DM, Hearn TJ, Webb AAR et al (2015) ELF3 controls thermoresponsive growth in Arabidopsis. Curr Biol 25:194–199. https://doi.org/10.1016/j.cub.2014.10.076
Burko Y, Willige BC, Seluzicki A, Novak O, Ljung K, Chory J (2022) PIF7 is a master regulator of thermomorphogenesis in shade. Nat Commun 13:4942. https://doi.org/10.1038/s41467-022-32585-6
Casal JJ, Balasubramanian S (2019) Thermomorphogenesis. Annu Rev Plant Biol 70:321–346. https://doi.org/10.1146/annurev-arplant-050718-095919
CFF, D.E.L., Kleine-Vehn, J., De Smet, I., and Feraru, E. (2021) Getting to the Root of Belowground High Temperature Responses in Plants. J Exp Bot. https://doi.org/10.1093/jxb/erab202
Chandler JW (2008) Cotyledon organogenesis. J Exp Bot 59:2917–2931. https://doi.org/10.1093/jxb/ern167
Chandler J, Nardmann J, Werr W (2008) Plant development revolves around axes. Trends Plant Sci 13:78–84. https://doi.org/10.1016/j.tplants.2007.11.010
Charng YY, Liu HC, Liu NY, Chi WT, Wang CN, Chang SH, Wang TT (2007) A heat-inducible transcription factor, HsfA2, is required for extension of acquired thermotolerance in Arabidopsis. Plant Physiol 143:251–262. https://doi.org/10.1104/pp.106.091322
Chen M, Tao Y, Lim J, Shaw A, Chory J (2005) Regulation of phytochrome B nuclear localization through light-dependent unmasking of nuclear-localization signals. Current Biology : CB 15:637–642. https://doi.org/10.1016/j.cub.2005.02.028
Chen Z, Huang Y, Yang W, Chang G, Li P, Wei J, Yuan X, Huang J, Hu X (2019) The hydrogen sulfide signal enhances seed germination tolerance to high temperatures by retaining nuclear COP1 for HY5 degradation. Plant Sci 285:34–43. https://doi.org/10.1016/j.plantsci.2019.04.024
Chen X, Zhou T, Wu P, Guo Z, Wang M (2020) Emergent constraints on future projections of the western North Pacific Subtropical High. Nat Commun 11:2802. https://doi.org/10.1038/s41467-020-16631-9
Chen Y, Hu X, Liu S, Su T, Huang H, Ren H, Gao Z, Wang X, Lin D, Wohlschlegel JA et al (2021) Regulation of Arabidopsis photoreceptor CRY2 by two distinct E3 ubiquitin ligases. Nat Commun 12:2155. https://doi.org/10.1038/s41467-021-22410-x
Chen D, Lyu M, Kou X, Li J, Yang Z, Gao L, Li Y, Fan LM, Shi H, Zhong S (2022) Integration of light and temperature sensing by liquid-liquid phase separation of phytochrome B. Mol Cell 82(3015–3029):e3016. https://doi.org/10.1016/j.molcel.2022.05.026
Cheng MC, Kathare PK, Paik I, Huq E (2021) Phytochrome Signaling Networks. Annu Rev Plant Biol 72:217–244. https://doi.org/10.1146/annurev-arplant-080620-024221
Chung BYW, Balcerowicz M, Di Antonio M, Jaeger KE, Geng F, Franaszek K, Marriott P, Brierley I, Firth AE, Wigge PA (2020) An RNA thermoswitch regulates daytime growth in Arabidopsis. Nature Plants 6:522–532. https://doi.org/10.1038/s41477-020-0633-3
Costigliolo Rojas C, Bianchimano L, Oh J, Romero Montepaone S, Tarkowska D, Minguet EG, Schon J, Garcia Hourquet M, Flugel T, Blazquez MA et al (2022) Organ-specific COP1 control of BES1 stability adjusts plant growth patterns under shade or warmth. Dev Cell 57(2009–2025):e2006. https://doi.org/10.1016/j.devcel.2022.07.003
Crawford AJ, McLachlan DH, Hetherington AM, Franklin KA (2012) High temperature exposure increases plant cooling capacity. Curr Biol 22:R396-397. https://doi.org/10.1016/j.cub.2012.03.044
Dai X, Mashiguchi K, Chen Q, Kasahara H, Kamiya Y, Ojha S, DuBois J, Ballou D, Zhao Y (2013) The biochemical mechanism of auxin biosynthesis by an arabidopsis YUCCA flavin-containing monooxygenase. J Biol Chem 288:1448–1457. https://doi.org/10.1074/jbc.M112.424077
De Veylder L, Larkin JC, Schnittger A (2011) Molecular control and function of endoreplication in development and physiology. Trends Plant Sci 16:624–634. https://doi.org/10.1016/j.tplants.2011.07.001
Delker C, Sonntag L, James GV, Janitza P, Ibanez C, Ziermann H, Peterson T, Denk K, Mull S, Ziegler J et al (2014) The DET1-COP1-HY5 pathway constitutes a multipurpose signaling module regulating plant photomorphogenesis and thermomorphogenesis. Cell Rep 9:1983–1989. https://doi.org/10.1016/j.celrep.2014.11.043
Delker C, Quint M, Wigge PA (2022) Recent advances in understanding thermomorphogenesis signaling. Curr Opin Plant Biol 68:102231. https://doi.org/10.1016/j.pbi.2022.102231
Deng XW, Caspar T, Quail PH (1991) cop1: a regulatory locus involved in light-controlled development and gene expression in Arabidopsis. Genes Dev 5:1172–1182. https://doi.org/10.1101/gad.5.7.1172
Deng XW, Matsui M, Wei N, Wagner D, Chu AM, Feldmann KA, Quail PH (1992) COP1, an Arabidopsis regulatory gene, encodes a protein with both a zinc-binding motif and a G beta homologous domain. Cell 71:791–801. https://doi.org/10.1016/0092-8674(92)90555-q
Ding Y, Yang S (2022) Surviving and thriving: How plants perceive and respond to temperature stress. Dev Cell 57:947–958. https://doi.org/10.1016/j.devcel.2022.03.010
Du M, Spalding EP, Gray WM (2020) Rapid Auxin-Mediated Cell Expansion. Annu Rev Plant Biol 71:379–402. https://doi.org/10.1146/annurev-arplant-073019-025907
Enders TA, Frick EM, Strader LC (2017) An Arabidopsis kinase cascade influences auxin-responsive cell expansion. Plant J 92:68–81. https://doi.org/10.1111/tpj.13635
Fiorucci AS, Galvao VC, Ince YC, Boccaccini A, Goyal A, Allenbach Petrolati L, Trevisan M, Fankhauser C (2020) PHYTOCHROME INTERACTING FACTOR 7 is important for early responses to elevated temperature in Arabidopsis seedlings. New Phytol 226:50–58. https://doi.org/10.1111/nph.16316
Franklin KA, Lee SH, Patel D, Kumar SV, Spartz AK, Gu C, Ye S, Yu P, Breen G, Cohen JD et al (2011) Phytochrome-interacting factor 4 (PIF4) regulates auxin biosynthesis at high temperature. Proc Natl Acad Sci U S A 108:20231–20235. https://doi.org/10.1073/pnas.1110682108
Friml J, Vieten A, Sauer M, Weijers D, Schwarz H, Hamann T, Offringa R, Jurgens G (2003) Efflux-dependent auxin gradients establish the apical-basal axis of Arabidopsis. Nature 426:147–153. https://doi.org/10.1038/nature02085
Fujii Y, Tanaka H, Konno N, Ogasawara Y, Hamashima N, Tamura S, Hasegawa S, Hayasaki Y, Okajima K, Kodama Y (2017) Phototropin perceives temperature based on the lifetime of its photoactivated state. Proc Natl Acad Sci U S A 114:9206–9211. https://doi.org/10.1073/pnas.1704462114
Gaillochet C, Burko Y, Platre MP, Zhang L, Simura J, Willige BC, Kumar SV, Ljung K, Chory J, Busch W (2020) HY5 and phytochrome activity modulate shoot-to-root coordination during thermomorphogenesis in Arabidopsis. Development 147:dev192625. https://doi.org/10.1242/dev.192625
Gangappa SN, Kumar SV (2017) DET1 and HY5 Control PIF4-Mediated Thermosensory Elongation Growth through Distinct Mechanisms. Cell Rep 18:344–351. https://doi.org/10.1016/j.celrep.2016.12.046
Gao H, Song W, Severing E, Vayssieres A, Huettel B, Franzen R, Richter R, Chai J, Coupland G (2022) PIF4 enhances DNA binding of CDF2 to co-regulate target gene expression and promote Arabidopsis hypocotyl cell elongation. Nat Plants. https://doi.org/10.1038/s41477-022-01213-y
Gonzalez N, Vanhaeren H, Inze D (2012) Leaf size control: complex coordination of cell division and expansion. Trends Plant Sci 17:332–340. https://doi.org/10.1016/j.tplants.2012.02.003
Gray WM, Ostin A, Sandberg G, Romano CP, Estelle M (1998) High temperature promotes auxin-mediated hypocotyl elongation in Arabidopsis. Proc Natl Acad Sci U S A 95:7197–7202. https://doi.org/10.1073/pnas.95.12.7197
Hahm J, Kim K, Qiu Y, Chen M (2020) Increasing ambient temperature progressively disassembles Arabidopsis phytochrome B from individual photobodies with distinct thermostabilities. Nat Commun 11:1660. https://doi.org/10.1038/s41467-020-15526-z
Hart JE, Gardner KH (2021) Lighting the way: Recent insights into the structure and regulation of phototropin blue light receptors. J Biol Chem 296:100594. https://doi.org/10.1016/j.jbc.2021.100594
He NY, Chen LS, Sun AZ, Zhao Y, Yin SN, Guo FQ (2022) A nitric oxide burst at the shoot apex triggers a heat-responsive pathway in Arabidopsis. Nat Plants 8:434–450. https://doi.org/10.1038/s41477-022-01135-9
Hiltbrunner A, Viczian A, Bury E, Tscheuschler A, Kircher S, Toth R, Honsberger A, Nagy F, Fankhauser C, Schafer E (2005) Nuclear accumulation of the phytochrome A photoreceptor requires FHY1. Current Biology : CB 15:2125–2130. https://doi.org/10.1016/j.cub.2005.10.042
Ibanez C, Poeschl Y, Peterson T, Bellstadt J, Denk K, Gogol-Doring A, Quint M, Delker C (2017) Ambient temperature and genotype differentially affect developmental and phenotypic plasticity in Arabidopsis thaliana. BMC Plant Biol 17:114. https://doi.org/10.1186/s12870-017-1068-5
Ibanez C, Delker C, Martinez C, Burstenbinder K, Janitza P, Lippmann R, Ludwig W, Sun H, James GV, Klecker M et al (2018) Brassinosteroids Dominate Hormonal Regulation of Plant Thermomorphogenesis via BZR1. Current Biology : CB 28:303-310 e303. https://doi.org/10.1016/j.cub.2017.11.077
Ito J, Sono T, Tasaka M, Furutani M (2011) MACCHI-BOU 2 is required for early embryo patterning and cotyledon organogenesis in Arabidopsis. Plant Cell Physiol 52:539–552. https://doi.org/10.1093/pcp/pcr013
Ito S, Song YH, Imaizumi T (2012) LOV domain-containing F-box proteins: light-dependent protein degradation modules in Arabidopsis. Mol Plant 5:573–582. https://doi.org/10.1093/mp/sss013
Jacob P, Hirt H, Bendahmane A (2017) The heat-shock protein/chaperone network and multiple stress resistance. Plant Biotechnol J 15:405–414. https://doi.org/10.1111/pbi.12659
Jin H, Zhu Z (2019) Dark, Light, and Temperature: Key Players in Plant Morphogenesis. Plant Physiol 180:1793–1802. https://doi.org/10.1104/pp.19.00331
Jin B, Wang L, Wang J, Jiang KZ, Wang Y, Jiang XX, Ni CY, Wang YL, Teng NJ (2011) The effect of experimental warming on leaf functional traits, leaf structure and leaf biochemistry in Arabidopsis thaliana. BMC Plant Biol 11:35. https://doi.org/10.1186/1471-2229-11-35
Jin H, Lin J, Zhu Z (2020) PIF4 and HOOKLESS1 Impinge on Common Transcriptome and Isoform Regulation in Thermomorphogenesis. Plant Commun 1:100034. https://doi.org/10.1016/j.xplc.2020.100034
Josse EM, Gan Y, Bou-Torrent J, Stewart KL, Gilday AD, Jeffree CE, Vaistij FE, Martinez-Garcia JF, Nagy F, Graham IA, Halliday KJ (2011) A DELLA in disguise: SPATULA restrains the growth of the developing Arabidopsis seedling. Plant Cell 23:1337–1351. https://doi.org/10.1105/tpc.110.082594
Jung JH, Domijan M, Klose C, Biswas S, Ezer D, Gao M, Khattak AK, Box MS, Charoensawan V, Cortijo S et al (2016) Phytochromes function as thermosensors in Arabidopsis. Science 354:886–889. https://doi.org/10.1126/science.aaf6005
Jung JH, Barbosa AD, Hutin S, Kumita JR, Gao M, Derwort D, Silva CS, Lai X, Pierre E, Geng F et al (2020) A prion-like domain in ELF3 functions as a thermosensor in Arabidopsis. Nature 585:256–260. https://doi.org/10.1038/s41586-020-2644-7
Kim S, Hwang G, Kim S, Thi TN, Kim H, Jeong J, Kim J, Kim J, Choi G, Oh E (2020) The epidermis coordinates thermoresponsive growth through the phyB-PIF4-auxin pathway. Nat Commun 11:1053. https://doi.org/10.1038/s41467-020-14905-w
Kim JY, Park YJ, Lee JH, Kim ZH, Park CM (2021) EIN3-Mediated Ethylene Signaling Attenuates Auxin Response during Hypocotyl Thermomorphogenesis. Plant Cell Physiol 62:708–720. https://doi.org/10.1093/pcp/pcab028
Koini MA, Alvey L, Allen T, Tilley CA, Harberd NP, Whitelam GC, Franklin KA (2009) High temperature-mediated adaptations in plant architecture require the bHLH transcription factor PIF4. Current Biology : CB 19:408–413. https://doi.org/10.1016/j.cub.2009.01.046
Kumar SV, Lucyshyn D, Jaeger KE, Alos E, Alvey E, Harberd NP, Wigge PA (2012) Transcription factor PIF4 controls the thermosensory activation of flowering. Nature 484:242–245. https://doi.org/10.1038/nature10928
Lee S, Paik I, Huq E (2020) SPAs promote thermomorphogenesis by regulating the phyB-PIF4 module in Arabidopsis. Development 147:dev189233. https://doi.org/10.1242/dev.189233
Lee S, Wang W, Huq E (2021) Spatial regulation of thermomorphogenesis by HY5 and PIF4 in Arabidopsis. Nat Commun 12:3656. https://doi.org/10.1038/s41467-021-24018-7
Legris M, Klose C, Burgie ES, Rojas CC, Neme M, Hiltbrunner A, Wigge PA, Schafer E, Vierstra RD, Casal JJ (2016) Phytochrome B integrates light and temperature signals in Arabidopsis. Science 354:897–900. https://doi.org/10.1126/science.aaf5656
Li L, Ljung K, Breton G, Schmitz RJ, Pruneda-Paz J, Cowing-Zitron C, Cole BJ, Ivans LJ, Pedmale UV, Jung HS et al (2012) Linking photoreceptor excitation to changes in plant architecture. Genes Dev 26:785–790. https://doi.org/10.1101/gad.187849.112
Li N, Bo C, Zhang Y, Wang L (2021) PHYTOCHROME INTERACTING FACTORS PIF4 and PIF5 promote heat stress induced leaf senescence in Arabidopsis. J Exp Bot 72:4577–4589. https://doi.org/10.1093/jxb/erab158
Li X, Liang T, Liu H (2022) How plants coordinate their development in response to light and temperature signals. Plant Cell 34:955–966. https://doi.org/10.1093/plcell/koab302
Lian HL, He SB, Zhang YC, Zhu DM, Zhang JY, Jia KP, Sun SX, Li L, Yang HQ (2011) Blue-light-dependent interaction of cryptochrome 1 with SPA1 defines a dynamic signaling mechanism. Genes Dev 25:1023–1028. https://doi.org/10.1101/gad.2025111
Liang T, Mei S, Shi C, Yang Y, Peng Y, Ma L, Wang F, Li X, Huang X, Yin Y, Liu H (2018) UVR8 Interacts with BES1 and BIM1 to Regulate Transcription and Photomorphogenesis in Arabidopsis. Dev Cell 44(512–523):e515. https://doi.org/10.1016/j.devcel.2017.12.028
Liu B, Zuo Z, Liu H, Liu X, Lin C (2011a) Arabidopsis cryptochrome 1 interacts with SPA1 to suppress COP1 activity in response to blue light. Genes Dev 25:1029–1034. https://doi.org/10.1101/gad.2025011
Liu HC, Liao HT, Charng YY (2011b) The role of class A1 heat shock factors (HSFA1s) in response to heat and other stresses in Arabidopsis. Plant Cell Environ 34:738–751. https://doi.org/10.1111/j.1365-3040.2011.02278.x
Liu S, Zhang L, Gao L, Chen Z, Bie Y, Zhao Q, Zhang S, Hu X, Liu Q, Wang X, Wang Q (2022) Differential photoregulation of the nuclear and cytoplasmic CRY1 in Arabidopsis. New Phytol 234:1332–1346. https://doi.org/10.1111/nph.18007
Lorrain S, Allen T, Duek PD, Whitelam GC, Fankhauser C (2008) Phytochrome-mediated inhibition of shade avoidance involves degradation of growth-promoting bHLH transcription factors. Plant J 53:312–323. https://doi.org/10.1111/j.1365-313X.2007.03341.x
Lu XD, Zhou CM, Xu PB, Luo Q, Lian HL, Yang HQ (2015) Red-light-dependent interaction of phyB with SPA1 promotes COP1-SPA1 dissociation and photomorphogenic development in Arabidopsis. Mol Plant 8:467–478. https://doi.org/10.1016/j.molp.2014.11.025
Ma D, Li X, Guo Y, Chu J, Fang S, Yan C, Noel JP, Liu H (2016) Cryptochrome 1 interacts with PIF4 to regulate high temperature-mediated hypocotyl elongation in response to blue light. Proc Natl Acad Sci U S A 113:224–229. https://doi.org/10.1073/pnas.1511437113
Ma L, Li X, Zhao Z, Hao Y, Shang R, Zeng D, Liu H (2021) Light-Response Bric-A-Brack/Tramtrack/Broad proteins mediate cryptochrome 2 degradation in response to low ambient temperature. Plant Cell 33:3610–3620. https://doi.org/10.1093/plcell/koab219
Mao Z, He S, Xu F, Wei X, Jiang L, Liu Y, Wang W, Li T, Xu P, Du S et al (2020) Photoexcited CRY1 and phyB interact directly with ARF6 and ARF8 to regulate their DNA-binding activity and auxin-induced hypocotyl elongation in Arabidopsis. New Phytol 225:848–865. https://doi.org/10.1111/nph.16194
Marco D’Ario RT, Schiessl K, Desvoyes B, Gutierrez C, Howard M, Sablowski R (2021) Cell size controlled in plants using DNA content as an internal scale. Science 372:1176–1181. https://doi.org/10.1126/science.abb4348
Martins S, Montiel-Jorda A, Cayrel A, Huguet S, Roux CP, Ljung K, Vert G (2017) Brassinosteroid signaling-dependent root responses to prolonged elevated ambient temperature. Nat Commun 8:309. https://doi.org/10.1038/s41467-017-00355-4
Mashiguchi K, Tanaka K, Sakai T, Sugawara S, Kawaide H, Natsume M, Hanada A, Yaeno T, Shirasu K, Yao H et al (2011) The main auxin biosynthesis pathway in Arabidopsis. Proc Natl Acad Sci U S A 108:18512–18517. https://doi.org/10.1073/pnas.1108434108
Moreno, S., Canales, J., Hong, L.L., Robinson, D., Roeder, A.H.K., and Gutierrez, R.A. (2020). Nitrate Defines Shoot Size through Compensatory Roles for Endoreplication and Cell Division in Arabidopsis thaliana. Curr. Biol. 30, 1988-+. https://doi.org/10.1016/j.cub.2020.03.036.
Neff MM, Van Volkenburgh E (1994) Light-Stimulated Cotyledon Expansion in Arabidopsis Seedlings (The Role of Phytochrome B). Plant Physiol 104:1027–1032. https://doi.org/10.1104/pp.104.3.1027
Ni M, Tepperman JM, Quail PH (1998) PIF3, a phytochrome-interacting factor necessary for normal photoinduced signal transduction, is a novel basic helix-loop-helix protein. Cell 95:657–667. https://doi.org/10.1016/s0092-8674(00)81636-0
Ni M, Tepperman JM, Quail PH (1999) Binding of phytochrome B to its nuclear signalling partner PIF3 is reversibly induced by light. Nature 400:781–784. https://doi.org/10.1038/23500
Nieto C, Lopez-Salmeron V, Daviere JM, Prat S (2015) ELF3-PIF4 interaction regulates plant growth independently of the Evening Complex. Curr Biol 25:187–193. https://doi.org/10.1016/j.cub.2014.10.070
Nieto, C., Catalan, P., Luengo, L.M., Legris, M., Lopez-Salmeron, V., Daviere, J.M., Casal, J.J., Ares, S., and Prat, S. (2022). COP1 dynamics integrate conflicting seasonal light and thermal cues in the control of Arabidopsis elongation. Sci Adv 8, eabp8412. https://doi.org/10.1126/sciadv.abp8412.
Nolan TM, Vukasinovic N, Liu D, Russinova E, Yin Y (2020) Brassinosteroids: Multidimensional Regulators of Plant Growth, Development, and Stress Responses. Plant Cell 32:295–318. https://doi.org/10.1105/tpc.19.00335
Nusinow DA, Helfer A, Hamilton EE, King JJ, Imaizumi T, Schultz TF, Farre EM, Kay SA (2011) The ELF4-ELF3-LUX complex links the circadian clock to diurnal control of hypocotyl growth. Nature 475:398–402. https://doi.org/10.1038/nature10182
Oh E, Zhu JY, Wang ZY (2012) Interaction between BZR1 and PIF4 integrates brassinosteroid and environmental responses. Nat Cell Biol 14:802–809. https://doi.org/10.1038/ncb2545
Ohama N, Sato H, Shinozaki K, Yamaguchi-Shinozaki K (2017) Transcriptional Regulatory Network of Plant Heat Stress Response. Trends Plant Sci 22:53–65. https://doi.org/10.1016/j.tplants.2016.08.015
Osterlund MT, Hardtke CS, Wei N, Deng XW (2000) Targeted destabilization of HY5 during light-regulated development of Arabidopsis. Nature 405:462–466. https://doi.org/10.1038/35013076
Paik I, Chen F, Ngoc Pham V, Zhu L, Kim JI, Huq E (2019) A phyB-PIF1-SPA1 kinase regulatory complex promotes photomorphogenesis in Arabidopsis. Nat Commun 10:4216. https://doi.org/10.1038/s41467-019-12110-y
Park E, Kim J, Lee Y, Shin J, Oh E, Chung WI, Liu JR, Choi G (2004) Degradation of phytochrome interacting factor 3 in phytochrome-mediated light signaling. Plant Cell Physiol 45:968–975. https://doi.org/10.1093/pcp/pch125
Park YJ, Lee HJ, Ha JH, Kim JY, Park CM (2017) COP1 conveys warm temperature information to hypocotyl thermomorphogenesis. New Phytol 215:269–280. https://doi.org/10.1111/nph.14581
Park J, Lee S, Park G, Cho H, Choi D, Umeda M, Choi Y, Hwang D, Hwang I (2021) CYTOKININ-RESPONSIVE GROWTH REGULATOR regulates cell expansion and cytokinin-mediated cell cycle progression. Plant Physiol 186:1734–1746. https://doi.org/10.1093/plphys/kiab180
Pedmale UV, Huang SC, Zander M, Cole BJ, Hetzel J, Ljung K, Reis PA, Sridevi P, Nito K, Nery JR et al (2016) Cryptochromes Interact Directly with PIFs to Control Plant Growth in Limiting Blue Light. Cell 164:233–245. https://doi.org/10.1016/j.cell.2015.12.018
Penfield S, Josse EM, Kannangara R, Gilday AD, Halliday KJ, Graham IA (2005) Cold and light control seed germination through the bHLH transcription factor SPATULA. Curr Biol 15:1998–2006. https://doi.org/10.1016/j.cub.2005.11.010
Podolec R, Demarsy E, Ulm R (2021) Perception and Signaling of Ultraviolet-B Radiation in Plants. Annu Rev Plant Biol 72:793–822. https://doi.org/10.1146/annurev-arplant-050718-095946
Procko C, Crenshaw CM, Ljung K, Noel JP, Chory J (2014) Cotyledon-Generated Auxin Is Required for Shade-Induced Hypocotyl Growth in Brassica rapa. Plant Physiol 165:1285–1301. https://doi.org/10.1104/pp.114.241844
Qi L, Shi Y, Terzaghi W, Yang S, Li J (2022) Integration of light and temperature signaling pathways in plants. J Integr Plant Biol 64:393–411. https://doi.org/10.1111/jipb.13216
Qian C, Mao W, Liu Y, Ren H, Lau OS, Ouyang X, Huang X (2016) Dual-Source Nuclear Monomers of UV-B Light Receptor Direct Photomorphogenesis in Arabidopsis. Mol Plant 9:1671–1674. https://doi.org/10.1016/j.molp.2016.10.005
Qiu Y (2020) Regulation of PIF4-mediated thermosensory growth. Plant Sci 297:110541. https://doi.org/10.1016/j.plantsci.2020.110541
Quint M, Delker C, Franklin KA, Wigge PA, Halliday KJ, van Zanten M (2016) Molecular and genetic control of plant thermomorphogenesis. Nature Plants 2:15190. https://doi.org/10.1038/nplants.2015.190
Raschke A, Ibanez C, Ullrich KK, Anwer MU, Becker S, Glockner A, Trenner J, Denk K, Saal B, Sun X et al (2015) Natural variants of ELF3 affect thermomorphogenesis by transcriptionally modulating PIF4-dependent auxin response genes. BMC Plant Biol 15:197. https://doi.org/10.1186/s12870-015-0566-6
Saini K, Dwivedi A, Ranjan A (2022) High temperature restricts cell division and leaf size by coordination of PIF4 and TCP4 transcription factors. Plant Physiol. https://doi.org/10.1093/plphys/kiac345
Schommer C, Debernardi JM, Bresso EG, Rodriguez RE, Palatnik JF (2014) Repression of cell proliferation by miR319-regulated TCP4. Mol Plant 7:1533–1544. https://doi.org/10.1093/mp/ssu084
Seo HS, Yang JY, Ishikawa M, Bolle C, Ballesteros ML, Chua NH (2003) LAF1 ubiquitination by COP1 controls photomorphogenesis and is stimulated by SPA1. Nature 423:995–999. https://doi.org/10.1038/nature01696
Sheerin DJ, Menon C, zur Oven-Krockhaus, S., Enderle, B., Zhu, L., Johnen, P., Schleifenbaum, F., Stierhof, Y.D., Huq, E., and Hiltbrunner, A. (2015) Light-activated phytochrome A and B interact with members of the SPA family to promote photomorphogenesis in Arabidopsis by reorganizing the COP1/SPA complex. Plant Cell 27:189–201. https://doi.org/10.1105/tpc.114.134775
Shen Y, Khanna R, Carle CM, Quail PH (2007) Phytochrome induces rapid PIF5 phosphorylation and degradation in response to red-light activation. Plant Physiol 145:1043–1051. https://doi.org/10.1104/pp.107.105601
Shen H, Zhu L, Castillon A, Majee M, Downie B, Huq E (2008) Light-induced phosphorylation and degradation of the negative regulator PHYTOCHROME-INTERACTING FACTOR1 from Arabidopsis depend upon its direct physical interactions with photoactivated phytochromes. Plant Cell 20:1586–1602. https://doi.org/10.1105/tpc.108.060020
Shu ZQ, Row S, Deng WM (2018) Endoreplication: The Good, the Bad, and the Ugly. Trends Cell Biol 28:465–474. https://doi.org/10.1016/j.tcb.2018.02.006
Song J, Liu Q, Hu B, Wu W (2017) Photoreceptor PhyB Involved in Arabidopsis Temperature Perception and Heat-Tolerance Formation. Int J Mol Sci 18:1194. https://doi.org/10.3390/ijms18061194
Spartz AK, Ren H, Park MY, Grandt KN, Lee SH, Murphy AS, Sussman MR, Overvoorde PJ, Gray WM (2014) SAUR Inhibition of PP2C-D Phosphatases Activates Plasma Membrane H+-ATPases to Promote Cell Expansion in Arabidopsis. Plant Cell 26:2129–2142. https://doi.org/10.1105/tpc.114.126037
Stepanova AN, Yun J, Robles LM, Novak O, He W, Guo H, Ljung K, Alonso JM (2011) The Arabidopsis YUCCA1 flavin monooxygenase functions in the indole-3-pyruvic acid branch of auxin biosynthesis. Plant Cell 23:3961–3973. https://doi.org/10.1105/tpc.111.088047
Stortenbeker N, Bemer M (2019) The SAUR gene family: the plant’s toolbox for adaptation of growth and development. J Exp Bot 70:17–27. https://doi.org/10.1093/jxb/ery332
Sun J, Qi L, Li Y, Chu J, Li C (2012) PIF4-mediated activation of YUCCA8 expression integrates temperature into the auxin pathway in regulating arabidopsis hypocotyl growth. PLoS Genet 8:e1002594. https://doi.org/10.1371/journal.pgen.1002594
Takahashi K, Hayashi K, Kinoshita T (2012) Auxin activates the plasma membrane H+-ATPase by phosphorylation during hypocotyl elongation in Arabidopsis. Plant Physiol 159:632–641. https://doi.org/10.1104/pp.112.196428
Tsukaya H, Tsuge T, Uchimiya H (1994) The cotyledon: A superior system for studies of leaf development. Planta 195:309–312. https://doi.org/10.1007/BF00199692
Vieten A, Vanneste S, Wisniewska J, Benkova E, Benjamins R, Beeckman T, Luschnig C, Friml J (2005) Functional redundancy of PIN proteins is accompanied by auxindependent cross-regulation of PIN expression. Development 132:4521–4531. https://doi.org/10.1242/dev.02027
Wang Q, Lin C (2020) Mechanisms of Cryptochrome-Mediated Photoresponses in Plants. Annu Rev Plant Biol 71:103–129. https://doi.org/10.1146/annurev-arplant-050718-100300
Wang H, Ma LG, Li JM, Zhao HY, Deng XW (2001) Direct interaction of Arabidopsis cryptochromes with COP1 in light control development. Science 294:154–158. https://doi.org/10.1126/science.1063630
Wang W, Lu X, Li L, Lian H, Mao Z, Xu P, Guo T, Xu F, Du S, Cao X et al (2018) Photoexcited CRYPTOCHROME1 Interacts with Dephosphorylated BES1 to Regulate Brassinosteroid Signaling and Photomorphogenesis in Arabidopsis. Plant Cell 30:1989–2005. https://doi.org/10.1105/tpc.17.00994
Wang W, Paik I, Kim J, Hou X, Sung S, Huq E (2021a) Direct phosphorylation of HY5 by SPA kinases to regulate photomorphogenesis in Arabidopsis. New Phytol 230:2311–2326. https://doi.org/10.1111/nph.17332
Wang X, Jiang B, Gu L, Chen Y, Mora M, Zhu M, Noory E, Wang Q, Lin C (2021b) A photoregulatory mechanism of the circadian clock in Arabidopsis. Nat Plants 7:1397–1408. https://doi.org/10.1038/s41477-021-01002-z
Witze A (2022) Extreme heatwaves: surprising lessons from the record warmth. Nature 608:464–465. https://doi.org/10.1038/d41586-022-02114-y
Won C, Shen X, Mashiguchi K, Zheng Z, Dai X, Cheng Y, Kasahara H, Kamiya Y, Chory J, Zhao Y (2011) Conversion of tryptophan to indole-3-acetic acid by TRYPTOPHAN AMINOTRANSFERASES OF ARABIDOPSIS and YUCCAs in Arabidopsis. Proc Natl Acad Sci U S A 108:18518–18523. https://doi.org/10.1073/pnas.1108436108
Wu J, Wang W, Xu P, Pan J, Zhang T, Li Y, Li G, Yang H, Lian H (2019) phyB Interacts with BES1 to Regulate Brassinosteroid Signaling in Arabidopsis. Plant Cell Physiol 60:353–366. https://doi.org/10.1093/pcp/pcy212
Xu X, Paik I, Zhu L, Huq E (2015) Illuminating Progress in Phytochrome-Mediated Light Signaling Pathways. Trends Plant Sci 20:641–650. https://doi.org/10.1016/j.tplants.2015.06.010
Xu F, He S, Zhang J, Mao Z, Wang W, Li T, Hua J, Du S, Xu P, Li L et al (2018) Photoactivated CRY1 and phyB Interact Directly with AUX/IAA Proteins to Inhibit Auxin Signaling in Arabidopsis. Mol Plant 11:523–541. https://doi.org/10.1016/j.molp.2017.12.003
Xue M, Zhang H, Zhao F, Zhao T, Li H, Jiang D (2021) The INO80 chromatin remodeling complex promotes thermomorphogenesis by connecting H2A.Z eviction and active transcription in Arabidopsis. Mol Plant 14:1799–1813. https://doi.org/10.1016/j.molp.2021.07.001
Yang HQ, Wu YJ, Tang RH, Liu D, Liu Y, Cashmore AR (2000) The C termini of Arabidopsis cryptochromes mediate a constitutive light response. Cell 103:815–827
Yang J, Qu X, Ji L, Li G, Wang C, Wang C, Zhang Y, Zheng L, Li W, Zheng X (2022) PIF4 Promotes Expression of HSFA2 to Enhance Basal Thermotolerance in Arabidopsis. Int J Mol Sci 23:6017. https://doi.org/10.3390/ijms23116017
Yang, Z., Yan, B., Dong, H., He, G., Zhou, Y., and Sun, J. (2021). BIC1 acts as a transcriptional coactivator to promote brassinosteroid signaling and plant growth. EMBO J 40, e104615. https://doi.org/10.15252/embj.2020104615.
Yin R, Skvortsova MY, Loubery S, Ulm R (2016) COP1 is required for UV-B-induced nuclear accumulation of the UVR8 photoreceptor. Proc Natl Acad Sci U S A 113:E4415-4422. https://doi.org/10.1073/pnas.1607074113
Yu X, Klejnot J, Zhao X, Shalitin D, Maymon M, Yang H, Lee J, Liu X, Lopez J, Lin C (2007) Arabidopsis cryptochrome 2 completes its posttranslational life cycle in the nucleus. Plant Cell 19:3146–3156. https://doi.org/10.1105/tpc.107.053017
Zhai H, Xiong L, Li H, Lyu X, Yang G, Zhao T, Liu J, Liu B (2020) Cryptochrome 1 Inhibits Shoot Branching by Repressing the Self-Activated Transciption Loop of PIF4 in Arabidopsis. Plant Commun 1:100042. https://doi.org/10.1016/j.xplc.2020.100042
Zhang H, He H, Wang X, Wang X, Yang X, Li L, Deng XW (2011) Genome-wide mapping of the HY5-mediated gene networks in Arabidopsis that involve both transcriptional and post-transcriptional regulation. Plant J 65:346–358. https://doi.org/10.1111/j.1365-313X.2010.04426.x
Zhao C, Liu B, Piao S, Wang X, Lobell DB, Huang Y, Huang M, Yao Y, Bassu S, Ciais P et al (2017) Temperature increase reduces global yields of major crops in four independent estimates. Proc Natl Acad Sci U S A 114:9326–9331. https://doi.org/10.1073/pnas.1701762114
Zhu T, van Zanten M, De Smet I (2022) Wandering between hot and cold: temperature dose-dependent responses. Trends Plant Sci. https://doi.org/10.1016/j.tplants.2022.06.001
Zuo Z, Liu H, Liu B, Liu X, Lin C (2011) Blue light-dependent interaction of CRY2 with SPA1 regulates COP1 activity and floral initiation in Arabidopsis. Current Biology : CB 21:841–847. https://doi.org/10.1016/j.cub.2011.03.048
Acknowledgements
We would like to celebrate the 120th anniversary of the founding of the Nanjing Normal University and apologize to any colleagues whose work could not be included due to space constraints.
Funding
This work was supported by the National Natural Science Foundation of China (31970256) and the Qing Lan Project from the Jiangsu Department of Education.
Author information
Authors and Affiliations
Contributions
Q.W. and Z.Z. both wrote the manuscript and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no conflicts of interest to declare.
Additional information
Handling editor: Shu-Hua Yang.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, Q., Zhu, Z. Light signaling-mediated growth plasticity in Arabidopsis grown under high-temperature conditions. Stress Biology 2, 53 (2022). https://doi.org/10.1007/s44154-022-00075-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44154-022-00075-w