Abstract
Coastal ecosystems are an important region for biogeochemical cycling, are a hotspot of anthropogenic disturbance and play a crucial role in global carbon cycling through the metabolic activities of bacterioplankton. Bacterioplankton can be broadly classified into two lifestyles: free-living (FL) and particle-attached (PA). However, how coastal bacterioplankton the community structure, co-occurrence networks and carbon metabolic functions with different lifestyles are differentiated is still largely unknown. Understanding these processes is necessary to better determine the contributions of coastal bacterioplankton to carbon cycling. Here, the characteristics of community structure and carbon metabolism function of bacterioplankton with two lifestyles in the coastal areas of Guangdong Province were investigated using amplicon sequencing, metagenomic, and metatranscriptomic techniques. The results show that the main bacterioplankton responsible for carbon metabolism were the Pseudomonadota, Bacteroidota, and Actinomycetota. The microbial community structure, carbon metabolic function, and environmental preferences differ between different lifestyles. FL and PA bacteria exhibited higher carbon fixation and degradation potentials, respectively. A range of environmental factors, such as dissolved oxygen, pH, and temperature, were associated with the community structure and carbon metabolic functions of the bacterioplankton. Human activities, such as nutrient discharge, may affect the distribution of functional genes and enhance the carbon degradation functions of bacterioplankton. In conclusion, this study increased the understanding of the role of microorganisms in regulating carbon export in coastal ecosystems with intense human activity.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The ocean, as the largest carbon reservoir on earth, plays a crucial role in the carbon cycle and can buffer the impacts of climate change (Dithugoe Choaro et al. 2023). Primary production in surface seawater and the fixed carbon export to the deep sea are considered primary contributors to natural carbon sequestration (Schlitzer 2002; Siegenthaler and Sarmiento 1993). Bacterioplankton are a key component of marine ecosystems, actively participating in the biological (BCP) and microbial carbon pumps (MCP) through carbon fixation and degradation, and affecting the absorption and release of CO2 (Jiao et al. 2010). For instance, picocyanobacteria, represented by Prochlorococcus and Synechococcus, are the most abundant prokaryotic photoautotrophs on Earth. Through photosynthesis, they convert CO2 into organic carbon, contributing approximately 25% of the net marine primary productivity (Flombaum et al. 2013). Ideally, carbon fixed by these phytoplankton in surface water sinks and is buried as sediment, contributing to carbon storage for atmospheric CO2. However, the fixed carbon may subsequently be affected by microbial carbon degradation processes which can lead to remineralization and release during the sinking process. As a result, only a small portion of the fixed carbon reaches the bottom of the ocean (Giering et al. 2014; Martin et al. 1987), affecting the efficiency of carbon export (Azam et al. 1991; Fenchel 2008).
Bacterioplankton can be classified into two lifestyles: free-living (FL) and particle-attached (PA). In recent years, a number of studies have compared the differences between FL and PA microorganisms in terms of community structure, physiological characteristics, and metabolic functions (Crump et al. 1999; DeLong et al. 1993; Ganesh et al. 2014; Li et al. 2018). PA bacteria often exhibit higher diversity, being primarily composed of complex biopolymers/carbohydrates degrading taxa. They contain more genes associated with nutrient cycling, such as carbon degradation, nitrogen fixation, and polyphosphate degradation (Jain et al. 2019; Liu et al. 2020; Smith et al. 2013). However, the functional characteristics and differences in specific carbon metabolism processes between the two bacterial communities remain unclear, resulting in the differing influence of these two communities on carbon export process, especially in the highly human-disturbed coastal areas, being undefined. Coastal regions are one of the most important sites for the cycling and transformation of essential elements such as carbon and nitrogen (Djurhuus et al. 2020). Investigating the influence of bacterioplankton and human activities on carbon export process in this habitat may lead to a better understanding of the carbon sequestration function of marine ecosystems.
In recent years, rapid industrialization and urbanization have resulted in an influx of a large quantity of inorganic and organic particulates into coastal areas of Guangdong, leading to severe water eutrophication. The ecosystem and biogeochemical cycles in the coastal area of Guangdong Province are being strongly disturbed by pollutants (Duan et al. 2022; Strokal et al. 2015). Therefore, it is essential to identify environmental effects on microorganisms in this region and the ecological functions performed by them. To deepen the understanding of microbial communities in the coastal areas of Guangdong, water samples from 22 sites along the coastline were collected, and high-throughput amplicon sequencing, metagenomic, and metatranscriptomic techniques were used to determine the diversity, community structure, network properties, community assembly processes, environmental drivers, and carbon metabolism functions between the two different lifestyles. This study aims to enhance the knowledge of the ecological functions of bacterioplankton communities by investigating and analyzing different bacterial lifestyles, providing scientific data and a theoretical reference for the study of carbon cycling functions in coastal Guangdong.
Materials and methods
Study area and sampling strategy
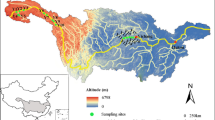
A field survey was conducted in June 2019, at the interface between the Pearl River Estuary and the shallow continental shelf of the South China Sea. A total of 22 sampling sites were set up on the coastline of Guangdong Province. These sites covered coastal regions, including Jiangmen, Zhuhai, Shenzhen, Huizhou, Shanwei, Jieyang, and Shantou, that are significantly impacted by human activities (Fig. 1A, Table S1). Seawater samples were collected at a depth of 1 m. Environmental parameters of the seawater were measured in situ using a Conductivity Temperature Depth (CTD, Sea-Bird Scientific) profiler. These included temperature, dissolved oxygen (DO), Chlorophyll a (Chl a), salinity, and pH. Immediately after collecting, each of the water samples was filtered through a 3 μm filter (Isopore™, 142 mm, Millipore) and a 0.2 μm filter (Supor-200, 142 mm, Pall Life Sciences) to collect PA and FL microbial communities, respectively. The 44 membrane filters (2 from each sampling site) were then transferred to sterile plastic bags and frozen in liquid nitrogen before being stored in a –80 °C freezer in the laboratory. The filtered water samples were stored in sterile centrifuge tubes at 4 °C for further chemical analysis.
Overview of microbial community in Guangdong coastal zone. A Geographical location of sampling sites in Guangdong coastal zone. B Alpha diversity of different lifestyles. The free-living (FL) and particle-attached (PA) are denoted as orange and purple correspondingly. Abbreviation of alpha diversity indices: ISI: Inverse Simpson Index; SE: Shannon Entropy; SR: Species Richness. C Principal Coordinates Analysis (PCoA) plot, including all samples of the two lifestyles. D Cluster trees based on Bray–Curtis dissimilarity among all samples aligned with the relative abundance of different phyla. Top 10 abundant phyla were showed and the rest were incorporated and exhibited as “Others”. Redundancy Analysis (RDA) results of FL E and PA F microbial community
Biochemical properties, DNA and RNA extraction, and sequencing
Dissolved organic carbon (DOC) analysis was performed using a TOC analyzer (TOC-vcph; Shimadzu Corp., Tokyo, Japan) following standard methods. The concentrations of nitrate (NO3−), nitrite (NO2−), ammonium (NH4+), dissolved nitrogen (DN), and soluble reactive phosphorus (SRP) were determined using a fully automated intelligent chemical analyzer (SmartChem; Westoco Scientific Instruments Inc., Brookfield, CT, USA).
Total DNA and RNA were extracted from each of the 44 filter samples (22 sites × 2 lifestyles) using RNeasy PowerSoil Total RNA Kit (QIAGEN, Germany) according to the manufacturer’s protocol. The DNA and RNA were separately eluted from the capture column by RNeasy PowerSoil DNA Elution Kit (QIAGEN) and RNeasy PowerSoil Total RNA Kit (QIAGEN). DNA and RNA samples were then sent to the Guangdong Magigene Biotechnology Company for amplicon, metagenomic and metatranscriptomic sequencing. The V4 and V5 hypervariable regions of the bacterial 16S rRNA gene were amplified using forward primer 515F (5′-GTGYCAGCMGCCGCGGTAA-3′) and reverse primer 926R (5′-CCGYCAATTYMTTTRAGTTT-3′) (Parada et al. 2016). Amplicon sequencing was performed on the Illumina NovaSeq 6000 PE250 platform. The metagenomic and metatranscriptomic sequencing were performed on the Illumina Hiseq PE150 platform.
Microbial community analysis
The amplicon sequencing data were further analyzed following the USEARCH pipeline (Edgar 2010); after merging paired-end reads, primer sequences were removed using CUTADAPT v2.4 (Martin 2011). Subsequently, sequences with a maximum expected error greater than 1.0 or a length less than 350 bp were filtered out. The remaining high-quality reads were dereplicated using the unoise3 algorithm, clustering sequences into amplicon sequence variants (ASVs) at a 100% similarity threshold. Taxonomic annotation of ASVs was performed using the Naïve Bayes classifier, based on the SILVA 132 database (Quast et al. 2012) on the QIIME2 platform (qiime2-2022.11) (Bolyen et al. 2019). ASVs that were not annotated as bacteria or archaea were removed (Kaehler et al. 2019). The reads, with primers removed, were then mapped back to ASV representative sequences to generate an ASV table. To correct for the impact of sequencing depth, the ASV table was rarefied to the lowest number of reads across all samples (46,403), resulting in a final dataset of 5,262,136 high-quality sequences. ASVs along with their frequency distribution and taxonomic annotation information were used for downstream statistical and diversity analysis.
Alpha diversity indices were used to measure the diversity within individual samples, reflecting microbial community abundance and diversity. Bray–Curtis distance was calculated to assess differences between samples, followed by Principal Coordinate Analysis (PCoA) for visualization. The similarity analysis (ANOSIM) in the vegan package was used to determine if the differences between groups were significantly different from the differences within group.
Redundancy Analysis (RDA) was employed to establish links between changes in microbial community composition and the corresponding environmental variables. Before RDA, Variance Inflation Factors (VIFs) were calculated for each environmental variable to identify collinearity. Variables with the highest VIFs were removed until all variables had a VIF below 10 (Zhang et al. 2021). Next, the Bioenv function from the vegan package was used to further select combinations of environmental factors that exhibited strong correlations with community differences. The first two canonical axes of the RDA results were visualized to show sites and explanatory variables. Additionally, the Hierarchical Partitioning (HP) method was used to assess the contribution of environmental factors to changes in community structure. This step utilized the rdacca.hp function from the rdacca.hp package (Lai et al. 2022).
Co-occurrence network analysis
The ASV table was taxonomically annotated to obtain abundance charts of bacterial communities at different taxonomic levels. Subsequently, to enhance the robustness of network construction, rare ASVs, i.e., those present in less than one-third of the samples, were filtered out. SparCC software was then employed to infer correlations between species within the two microbial community types, constructing a microbial species interaction network (Friedman and Alm 2012). Based on the constructed network, several topologic characteristics were calculated. To determine the ecological roles of each node, two topological properties were considered: within-module degree (Zi) and among-module connectivity (Pi), indicating how ‘well connected’ a node is to nodes within their own module or from other modules, respectively (Guimerà and Amaral 2005). Specifically, based on their Zi and Pi values, all nodes were assigned to one of four roles: module hubs (Zi > 0.25, Pi ≤ 0.62), connectors (Zi ≤ 0.25, Pi > 0.62), network hubs (Zi > 0.25, Pi > 0.62), and peripherals (Zi ≤ 0.25, Pi < 0.62) (Olesen et al. 2007). Nodes other than peripheral nodes in each network were defined as potential ecologically important key species (Zhang et al. 2021). The network was visualized using Gephi (0.10.1) (Bastian et al. 2009).
Quantification of microbial community assembly processes
In this study, the relative importance of different community assembly processes was quantified using the null-model approach described by Stegen et al. (2013). The null-model approach distinguishes between selection processes and neutral processes by calculating standardized phylogenetic turnover (βNTI) between communities. The βNTI of FL and PA were calculated, respectively, using the iCAMP package in R. βNTI less than -2 or greater than 2 indicate homogeneous selection or heterogeneous selection, respectively. |βNTI|< 2 suggests neutral processes. Subsequently, the Bray–Curtis-based Raup–Crick metric (RCbray) was used to further categorize specific neutral processes, with RCbray < -0.95 inferring homogeneous dispersal, RCbray > 0.95 representing dispersal limitation, and |RCbray|≤ 0.95 indicating drift (Osburn et al. 2021; Stegen et al. 2013, 2015).
Identification and calculation of carbon metabolism genes
The raw metagenomic reads were trimmed using fastp (Chen et al. 2018) (v0.20.0, parameters: -n 0 -l 30 –cut_front –cut_right –cut_window_size 4 –cut_mean_quality 30). High-quality metagenomic reads for each sample were then assembled using MEGAHIT v1.2.9 (Li et al. 2016) with parameters '–k-min 27 –k-max 127 –k-step 20 –min-contig-len 500’. Open-reading frames (ORFs) were predicted from all contigs using Prodigal v2.6.3 (Hyatt et al. 2010) with parameters ‘-m -p meta’.
A reference database related to carbon metabolism-related genes was developed using amino acid (aa) sequences manually downloaded from the Nr database, according to previous studies (Shi et al. 2019; Tu et al. 2014), which is mainly consisted of seven common carbon sources in natural carbon degradation processes (Table S2) and six microbial carbon fixation processes (Table S3). Then, the ORFs were searched against the reference database using DIAMOND v0.9 (Buchfink et al. 2015) with parameters ‘-k 1 –evalue 0.00001 –id 50 –query-cover 80’. To construct a non-redundant gene catalog (467,177 ORFs), the results after annotation were clustered at a 95% similarity level (Li et al. 2014; Sunagawa et al. 2015) using CD-HIT-EST v4.8.1 (Li and Godzik 2006) with parameters ‘-c 0.95 -n 10 -G 0 -aS 0.9 -g 1 -d 0 -T 15 -M 0’ (Bahram et al. 2018). The abundance of each ORF across samples was calculated by mapping metagenomic high-quality reads to the gene catalog using BBMap (Bushnell 2014) with parameters ‘minid = 0.95’. In this study, Fragments Per Kilobase of exon model per Million mapped fragments (FPKM) was used to measure the relative abundance of mapped genes related to carbon degradation and carbon fixation.
The quality of raw metatranscriptomic reads was also controlled by fastp (Chen et al. 2018) (v0.20.0, parameters: -n 0 -l 30 –cut_front –cut_right –cut_window_size 4 –cut_mean_quality 30). Before further analysis, RiboDetector was used to clear ribosomal RNA. Then, the transcript abundance of each ORF was calculated by mapping high-quality metatranscriptomic reads to the gene catalog using BBMap (Bushnell 2014) with parameters ‘minid = 0.95’. Similarly, the relative transcript abundance of mapped genes related to carbon degradation and fixation was also shown by FPKM.
Taxonomy assignment of functional genes
To determine the taxonomy of the carbon metabolism genes, they were searched against the GTDB r214 protein database (https://gtdb.ecogenomic.org/) by mmseqs easy-taxonomy (Mirdita et al. 2021) with default parameters. The contribution of different phyla to each carbon metabolism-related gene was calculated as the sum of the abundances of sequences that were assigned to that phylum.
Genome binning, curation, dereplication, and annotation
Metagenome-Assembled Genomes (MAGs) were recovered from each sample using previously described methods (Luo et al. 2022; Wang et al. 2022). Briefly, scaffolds of each sample were binned by MetaBAT2, based on the coverage variation of contigs across samples and tetranucleotide frequencies with default parameters (Kang et al. 2019). The qualities of MAGs were evaluated using CheckM v1.0.12 (Parks et al. 2015), and then, the potential contaminations of each MAG were identified and removed by RefineM v0.0.25 (Parks et al. 2017) and manual curation. MAGs with high-to-medium-quality (completeness > 60%, contamination < 5%) from all samples were combined and dereplicated using dRep v2.6.2 (Olm et al. 2017) with default settings. The taxonomy of each MAG was determined by GTDB-Tk v1.2.0 (Chaumeil et al. 2019). ORFs in MAGs were predicted using Prodigal v2.6.3 (Hyatt et al. 2010) with default parameters. Then, the ORFs were searched against the reference database by DIAMOND v0.9 (Buchfink et al. 2015) with parameters as described in Sect. “Identification and calculation of carbon metabolism genes” to construct a gene catalog (7865 ORFs). Gene abundance and transcript abundance of carbon metabolism genes in each MAG were quantified as described in Sect. “Identification and calculation of carbon metabolism genes”. The abundance of each MAG in every sample was estimated based on the FPKM values of its constituent scaffolds.
Statistical analysis
All statistical analyses were performed using R software (version 4.3.1). The significance of differences in α-diversity, genomic abundance, and transcriptomic abundance of carbon metabolism genes between FL and PA were determined using Wilcoxon test. The relationship between relative abundance of carbon metabolism genes and environmental traits was analyzed based on Pearson's correlation method. The spatial distributions of abundance of carbon metabolism genes were interpolated across the PRE using the IDW method in GSTAT package (Gräler et al. 2016).
Results
Structure, environmental drivers, and assembly process of bacterioplankton community with different lifestyles
Amplicon data from 39 filter samples were successfully obtained. There was no significant difference in α-diversity between FL and PA communities (P > 0.05, Fig. 1B). The result of principal coordinate analysis (PCoA) indicates significant differences (ANOSIM, R = 0.4681, P < 0.05) in community structure between the FL and PA groups (Fig. 1C).
A total of 19 phyla and 60 classes were detected. Pseudomonadota and Bacteroidota were the most common phyla in each community (ranging from 64.8% to 99.0%). The FL community was mainly composed of Bacteroidota, while the PA community was mainly composed of Pseudomonadota (Fig. 1D). Additionally, the microbial composition in each sample was clustered based on their Bray–Curtis distances and, except for a few samples, FL and PA clustered separately (Fig. 1D).
To identify the environmental variables influencing the community structure of FL and PA, RDA was performed separately for FL and PA. After filtering based on VIF and BioEnv analysis, nine and six environmental variables were retained for RDA analysis in FL and PA groups, respectively. In the FL group, temperature, DO, and Chl a were identified as the primary factors shaping microbial community structure (Fig. 1E), with temperature exerting the most significant impact (6.9%, P < 0.05). In the PA group, NH4+, temperature, and NO3− were the main environmental drivers (Fig. 1F), with NH4+ being the most important driver (7.3%, P < 0.05).
Stochastic processes were the primary assembly mechanisms for the two bacterial communities (Fig. S1A). The role of dispersal limitation was important for bacterial community assembly in both the FL (51.9%) and PA (51.6%) community (Fig. S1B). Additionally, heterogeneous selection processes constituted a significant proportion in FL (36.7%) and PA (34.6%), indicating microbial adaptability to diverse environmental conditions in the Guangdong coastal zone (Fig. S1B).
Co-occurrence networks with different lifestyles
Microbial interaction networks for FL and PA were constructed using the SparCC method (Fig. 2A, B). These networks exhibited a range of topological properties. Compared to FL, the PA network had a higher number of nodes and connections, indicating a more complex network structure (Table 1). Additionally, the PA network exhibited higher values for average degree (AD), graph density (GD), and clustering coefficient (CC), suggesting that the bacterial community structure in PA was more tightly interconnected than in FL.
The molecular ecological networks of the free-living (FL) and particle-attached (PA) microbial communities. A FL network. B PA network. The color of the nodes indicates different modules and the size is proportional to log2(ASV numbers). C The different ecological role of all nodes, partitioned at Zi = 2.5 and Pi = 0.62. The nodes of free-living (FL) and particle-attached (PA) are denoted as orange and purple correspondingly. D The taxonomy of connectors, module hubs and network hubs at phylum level
To identify potential keystone taxa in each network, each node was classified into four ecological roles based on their within-module degree (Zi) and among-module connectivity (Pi) (see methods, Fig. 2C). The PA community had nine keystone nodes, most of which served as module hubs critical for maintaining the stability of module structures. These keystone taxa belonged to Pseudomonadota, Bacteroidota, and Planctomycetota (Fig. 2D). In contrast, the FL community had 32 keystone nodes, which was significantly more than PA, comprising 12 module hubs, 18 connector nodes, and 2 network hubs. These nodes played essential ecological roles in maintaining the stability and functionality of the network. Similarly, they were primarily composed of bacteria from Pseudomonadota and Bacteroidota (Fig. 2D).
Carbon metabolism functions of bacteria with different lifestyles
Metagenomic data from 38 samples and metatranscriptomic data from 21 samples were successfully obtained (Table S4). The gene abundance and transcriptional abundance in carbon degradation (CD) and carbon fixation (CF) processes were first compared between FL and PA. The results showed that PA had a higher abundance of carbon degradation genes, while FL had a higher abundance of carbon fixation genes (Fig. S2). This suggests that in this study, FL and PA may dominate different carbon cycling processes in coastal ecosystems.
To investigate whether there were differences in specific carbon metabolism processes among bacterioplankton with different lifestyles, the composition (Fig. 3A–H) and abundance (Fig. 3I–L, Fig. S2) of genes involved in carbon degradation (Table S2) and carbon fixation (Table S3) in FL and PA were analyzed. Seven common carbon sources in natural carbon degradation processes and six microbial carbon fixation processes are listed in Fig. 3. A PCoA analysis at both the gene and metabolic pathway levels was conducted and the results showed that there were significant differences in carbon degradation and carbon fixation functions between FL and PA (ANOSIM, P < 0.05), suggesting that FL and PA exhibit distinct preferences for specific functional genes and metabolic pathways when performing carbon cycling. This phenomenon is further illustrated in the barplot in Fig. 3, where genes with significant differences in abundance in the genomes (Fig. 3I–J) and the transcriptome (Fig. 3K–L) between FL and PA are shown. These differences highlight the distinct carbon metabolism characteristics of the two kinds of bacteria with different lifestyles. For example, in terms of the degradation of aromatic hydrocarbons, FL had a higher variety and abundance of dioxygenase and monooxygenase genes (such as nbac, xylE, carAa, and camdcab), which played crucial roles in the cleavage of aromatic rings. PA, however, seemed to have the advantage in the side-chain modification process of aromatic hydrocarbons (such as mdld, nfsa, and bend). In terms of the six carbon fixation processes, there was a higher abundance of functional genes in FL, indicating that FL may have higher carbon fixation potential than PA in coastal surface seawater.
Differences in carbon degradation and carbon fixation functions between the free-living (FL) and particle-attached (PA) microbial communities. Principal coordinates analysis (PCoA) of carbon degradation (CD) based on genes (A, E) or pathways (C, G) and PCoA of carbon fixation (CF) based on genes (B, F) or pathways (D, H). The circular dots represent metagenomic data (A ~ D) and the triangular dots represent metatranscriptomic data (E ~ H). Fragments Per Kilobase Million (FPKM), after taking natural logarithm, were used to represent the abundance of carbon metabolism genes in metagenomes (I, J) and metatranscriptomes (K, L) of the sampling points. Wilcoxon rank-sum test of each gene was performed to examine whether the difference between FL and PA was significant. Only genes with significant differences were shown. Gene directed toward the arrow showed FL (or PA) means that there was more of that gene in FL (or PA). Abbreviation in carbon fixation process: rTCA cycle: reductive tricarboxylic acid cycle; WL: reductive acetyl-CoA pathway; 3HP bicycle: 3-hydroxypropionate bicycle; 3HP/4HB: 3-hydroxypropionate/4-hydroxybutyrate cycle; DC/4HB: dicarboxylate/4-hydroxybutyrate cycle
Spatial distribution of carbon metabolism genes and their response to environmental factors
The genes contributing to the same carbon metabolism processes were combined and the distribution characteristics of carbon metabolism genes at different sampling sites are summarized (Fig. 4). Overall, both the carbon degradation and carbon fixation gene abundances were relatively higher in Pearl River Estuary, suggesting that estuaries may be hotspots for carbon cycling in coastal regions.
Additionally, the response of bacterioplankton in carbon metabolism functions to the physicochemical properties in coastal environment was explored, and the correlation between carbon metabolic gene abundance and environmental factors was investigated (Fig. 5). The results showed that six environmental factors can affect the carbon metabolic function of bacterioplankton, including pH, Chl a, salinity, DO (Fig. 5B, E, G), as well as nutrient-related factors such as NO2− and SRP (Fig. 5C, D, F). Notably, as with community structure, the abundance of carbon metabolism genes in FL was associated with more environmental factors, suggesting that FL communities may be more sensitive to environmental changes.
Correlation analysis between relative abundance of carbon metabolism genes and environmental traits. A Pearson correlation heatmap between abundance of carbon degradation genes (CDGs) and carbon fixation genes (CFGs) in FL and PA. The significance levels were denoted as “*”, 0.01 < P ≤ 0.05; “**”, 0.001 < P ≤ 0.01; “***”, P ≤ 0.001. B The relationship between CDGs abundance and salinity, pH or Chl a in FL bacteria. C The relationship between CDGs abundance and SRP or NO2− in FL bacteria. D The relationship between CFGs abundance and NO2− in FL bacteria. E The relationship between CDGs abundance and Chl a in PA bacteria. F The relationship between CDGs abundance and SRP in PA bacteria. G The relationship between CFGs abundance and pH or DO in PA bacteria. PPMCC: Pearson product–moment correlation coefficient
Relative contributions of community members to different processes
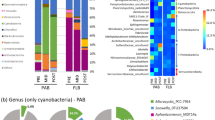
To further explain the relationship between community members and functional capacities, the taxonomy of each functional gene was annotated. The results indicate that more genes were annotated as Pseudomonadota (32.6%), Actinomycetota (11.3%), and Bacteroidota (10.7%). Relative contributions of different phyla to certain carbon metabolism process were assessed based on the relative abundance of genes or transcripts from each phylum (Fig. 6A–C). Overall, in the PA community, Pseudomonadota had a notably higher gene abundance (Fig. 6A, B) and transcriptional activity (Fig. 6C). In contrast, in the FL community, the relative contribution of Pseudomonadota was comparatively lower, with a wider variety of phyla playing a significant role.
Relative contributions of different phyla to different carbon metabolism processes in Guangdong coastal zone. The relative contributions of different phyla to the gene abundances (A, B) and transcriptional abundance (C) of carbon metabolism-related genes. Average abundance of carbon degradation (D) and carbon fixation (E) related genes in MAGs from different phyla, expressed as a percentage of the total abundance across all phyla. (F) Transcriptional abundance of carbon metabolism-related genes in MAGs of different phyla with different lifestyles during each carbon cycle process. The length of the segments is proportional to ln (FPKM + 1). Wilcoxon rank-sum test was performed to examine whether the difference between FL and PA group is significant. The significance levels are denoted as “*”, 0.01 < P ≤ 0.05; “**”, 0.001 < P ≤ 0.01; “***”, P ≤ 0.001
MAGs were recovered to identify the key taxa involved in carbon cycling functions within the coastal habitats of Guangdong. A total of 2793 prokaryotic genomes were assembled from 38 metagenomes. The MAGs were then refined and dereplicated into 332 representative genomes (Table S5). Most of the obtained MAGs belonged to Pseudomonadota (114/332), followed by Bacteroidota (110/332). Average abundance of carbon degradation (Fig. 6D) and carbon fixation (Fig. 6E) related genes, as well as transcriptional abundance (Fig. 6F) in MAGs from different phyla were calculated. Similarly, Pseudomonadota and Bacteroidota exhibited higher levels of carbon metabolism. The distinct characteristics shown by many taxa in the two lifestyles were further compared. Among Pseudomonadota, which demonstrated a significant role, the choice of lifestyle did not significantly affect its carbon metabolic capability. This may suggest that Pseudomonadota exhibited a wide and strong adaptability to various lifestyles. In contrast, in Actinomycetota, Bacillota, Bacteroidota, Planctomycetota, and Verrucomicrobiota, FL and PA displayed more inconsistencies, indicating that these phyla may have different selective preference for lifestyle.
Discussion
This study showed that within the coastal regions of Guangdong, the primary bacterioplankton responsible for carbon metabolism were Pseudomonadota, Bacteroidota, and Actinomycetota. The selection of lifestyle significantly influenced the structure, ecological function, and environmental preferences of bacterioplankton communities, subsequently impacting the carbon cycling mechanisms within these coastal habitats. Overall, FL demonstrated a greater capacity for carbon fixation, while PA had a greater capacity for carbon degradation. When it comes to specific pathways of carbon metabolism, FL and PA communities performed better in different pathways, seeming to be able to complement each other. The distribution of their functional genes was influenced by a range of physicochemical factors and human activities.
Distribution patterns of carbon metabolic functions in Guangdong coastal zone
In the coastal habitats of Guangdong, the primary phyla were Pseudomonadota, Bacteroidota, Actinomycetota, Bacillota, and Cyanobacteriota, which is consistent with studies from other relevant aquatic environments (Harrison et al. 2008). Additionally, as with the amplicon results, the annotation of functional genes and MAGs indicates that a broad and strong carbon cycle functional spectrum was distributed within Pseudomonadota, Bacteroidota, and Actinomycetota (Fig. 6). Similar results have been found in studies from the Southern Ocean, where Pseudomonadota and Actinomycetota had a greater number of genes involved in the degradation of carbohydrates, such as chitin and cellulose (Dithugoe Choaro et al. 2023). Notably, some low-abundance groups also played important roles in certain carbon metabolism processes in this study. For instance, Myxococcota, which was less abundant, lacked the degradation activities for hemicellulose and pectin, but possessed a relatively strong ability for aromatics degradation and Calvin cycle. Similarly, some studies have also emphasized the significant roles of Myxococcota in photosynthesis (Li et al. 2023).
In this study, the Pearl River Estuary showed greater vitality in carbon metabolism, with most of the carbon metabolic genes highly accumulated within the regions (Fig. 4). Some of carbon metabolic genes, however, were distributed with different patterns. For example, it was found that the abundance of chitin degradation genes was highest near Shanwei, Jieyang, and Shantou, which are distant from the estuary. This may have resulted from aquaculture activities, because these three coastal cities are known for mariculture, including chitin-rich crustaceans such as prawn (e.g., Litopenaeus vannamei) and crabs (e.g., Scylla serrata), which may contribute to the accumulation of chitin in the region and eventually led to the enrichment of genes related to chitin degradation.
Community structure and functional characteristics of different lifestyles
In this study, particles had significant impact on the composition, co-occurrence patterns and carbon metabolism capability of bacterial communities. In the PA community, Pseudomonadota had a higher relative abundance compared to the FL community. Additionally, at the class level, Gammaproteobacteria was the most abundant taxon within the PA group (Fig. S3). This may result from the process of microbial colonization on particle surfaces, as previous research has suggested that Gammaproteobacteria are pioneering organisms in marine biofilm formation (Lee et al. 2008).
The co-occurrence network formed by PA bacteria was denser, with more distinct modules and more complex correlations within modules than FL bacteria. Similarly, many studies have shown that PA bacteria often have a more complex co-occurrence network structure compared to FL bacteria (Xu et al. 2018; Zhang et al. 2016). This suggests that particles serve as colonization points that make interactions such as cooperation and competition among bacteria more pronounced. Cooperation can help different PA bacteria collectively degrade the particles they colonized, while the limited space on particle surfaces may lead to strong competition for ecological niches among PA bacteria (Clauset et al. 2004; Xu et al. 2021). Despite the lower complexity of the FL network, the number of potential keystone species in the FL network exceeded that of the PA network, whose metabolic functions may be important in ecological services (Banerjee et al. 2018; Rafrafi et al. 2013). For example, two network hubs in the FL network, Acinetobacter and Flavobacterium were identified, both of which can degrade a range of organic pollutants, such as petroleum and polycyclic aromatic hydrocarbons (PAHs), suggesting the potential importance of FL communities in pollution remediation in the coastal areas of Guangdong (Chen et al. 2014; Liu et al. 2019). The key taxon found in both of the microbial networks was Rhodobacteraceae, which is mainly comprised of aerobic photo- and chemoheterotrophs but also purple non-sulfur bacteria, performing photosynthesis in anaerobic environments. It is deeply involved in sulfur and carbon biogeochemical cycling and symbiosis with micro- and macroorganisms in marine ecosystems (Dogs et al. 2017) and makes a significant contribution to biogeochemical processes.
The characteristics of planktonic bacterial carbon metabolic potentials in the surface water of the Guangdong coastal region were further investigated from a molecular aspect. First, carbon degradation and carbon fixation potential of FL and PA were broadly investigated. The results show that different lifestyle selection affected the carbon metabolic function of the bacterioplankton (Fig. 3A–H, with higher abundance of carbon degradation genes in PA and higher abundance of carbon fixation genes in FL, respectively (Fig. S2). This is consistent with the results of previous studies, in which PA were found to be more active in the degradation of particulate matter, while FL tended to be oligotrophic autotrophs (Herndl and Reinthaler 2013; Karner and Herndl 1992; Smith et al. 2013). In this study, the abundance of Cyanobacteriota in FL was higher, and Synechococcus and Cyanobium were the two key groups in the FL network; this further demonstrates the dominant position of the FL group in carbon fixation processes. The carbon metabolic genes that differed between FL and PA are further listed in Fig. 3I–L. These suggest that they may dominate different specific carbon metabolism processes and complement each other. For example, the degradation of cellulose requires the joint action of cellobiase and endoglucanase, and the transcription levels of genes encoding these two enzymes were more abundant in FL or PA, respectively (Fig. 3K).
Environmental preference of different lifestyles
The microbial community structures of these two lifestyles were influenced by a range of environmental factors. Overall, temperature had a substantial impact on both the FL and PA community structures. Additionally, DO and NH4+ were major factors affecting the FL and PA communities, respectively (Fig. 1E–F). Similar results have been observed in the previous studies (Ma et al. 2023). Temperature can regulate bacterial metabolic capabilities by influencing enzyme activity, ultimately affecting bacterial community distribution and function. High DO is essential for aerobic metabolism in aquatic bacteria, with FL being more dependent on DO than PA (Aldunate et al. 2018; Mohiuddin et al. 2019). Studies have shown that organic particles in eutrophic water can absorb NH4+ (Lehmann et al. 2004), and the NH4+ adsorbed on particles affects the nitrogen cycle of PA, significantly influencing their community structure.
There was a negative correlation of pH and salinity with carbon degradation gene (CDG) abundance in FL groups (Fig. 5B). Similarly, pH exhibited a negative correlation with CFG abundance in PA groups (Fig. 5G). This might be because higher pH and salinity inhibited enzyme activity. It is worth noting that NO2− and SRP, which can serve as N and P nutrients, were positively correlated with CDG abundance (Fig. 5C, F), seemingly promoting bacterial carbon degradation. Similar results have been observed in other habitats, including soil (Li et al. 2021a, 2021b), mangroves (Li et al. 2024), and estuaries (Yuan et al. 2010), where the input of N and P tended to stimulate microbial mineralization, which is detrimental to carbon storage. This suggests that achieving an increase in carbon sequestration requires a stronger control of nutrient inputs to the ocean (Jiao et al. 2011). Similarly, the concentration of Chl a can serve as an indicator of water eutrophication, which may be one of reasons why its effect on the abundance of CDGs was consistent with that of nutrients (Fig. 5B, E). Additionally, DO exhibited a negative correlation with CFG abundance in PA (Fig. 5G); this may be due to the fact that many processes among the six carbon fixation pathways are dependent on anaerobic conditions, such as the dicarboxylate/4-hydroxybutyrate cycle (DC/4HB), which is a strictly anaerobic carbon fixation process found in archaea (Huber et al. 2008).
Conclusion
Utilizing amplicon, metagenomic and metatranscriptomic techniques, it was found that within the coastal regions of Guangdong, the primary bacterioplankton responsible for carbon metabolism were Pseudomonadota, Bacteroidota, and Actinomycetota. Different selection of lifestyle mode affected the structure, function, and environmental preference of bacterioplankton communities. FL and PA communities had stronger carbon fixation and carbon degradation potential, respectively, and thus made different contributions and had different influences on carbon storage processes in coastal ecosystems. Similar to previous studies, this study shows that nutrient input enhances the carbon degradation function of bacterioplankton, suggesting that nutrient input from land needs to be reduced. This would contribute to the realization of ocean negative carbon emissions (ONCE).
Data availability
All sequencing data from Guangdong coastal zone are available at National Center for Biotechnology Information (https://www.ncbi.nlm.nih.gov/) under the accession number PRJNA1062668 (amplicon), PRJNA1063027 (metagenome), and PRJNA1063025 (metatranscriptome).
References
Aldunate M, De la Iglesia R, Bertagnolli AD, Ulloa O (2018) Oxygen modulates bacterial community composition in the coastal upwelling waters off central Chile. Deep Sea Res 2 Top Stud Oceanogr 156:68–79
Azam F, Smith DC, Hollibaugh JT (1991) The role of the microbial loop in Antarctic pelagic ecosystems. Polar Res 10:239–244
Bahram M, Hildebrand F, Forslund SK, Anderson JL, Soudzilovskaia NA, Bodegom PM, Bengtsson-Palme J, Anslan S, Coelho LP, Harend H, Huerta-Cepas J, Medema MH, Maltz MR, Mundra S, Olsson PA, Pent M, Põlme S, Sunagawa S, Ryberg M, Tedersoo L et al (2018) Structure and function of the global topsoil microbiome. Nature 560:233–237
Banerjee S, Schlaeppi K, van der Heijden MGA (2018) Keystone taxa as drivers of microbiome structure and functioning. Nat Rev Microbiol 16:567–576
Bastian M, Heymann S, Jacomy M (2009) Gephi: An open source software for exploring and manipulating networks. Proc Int AAAI Conf Web Soc Media 3:361–362
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, Alexander H, Alm EJ, Arumugam M, Asnicar F, Bai Y, Bisanz JE, Bittinger K, Brejnrod A, Brislawn CJ, Brown CT, Callahan BJ, Caraballo-Rodríguez AM, Chase J, Cope EK et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857
Buchfink B, Xie C, Huson DH (2015) Fast and sensitive protein alignment using DIAMOND. Nat Meth 12:59–60
B Bushnell (2014) BBMap: a fast, accurate, splice-aware aligner. Lawrence Berkeley National Laboratory (LBNL)
Chaumeil PA, Mussig AJ, Hugenholtz P, Parks DH (2019) GTDB-Tk: a toolkit to classify genomes with the genome taxonomy database. Bioinformatics 36:1925–1927
Chen Y, Li C, Zhou Z, Wen J, You X, Mao Y, Lu C, Huo G, Jia X (2014) Enhanced biodegradation of alkane hydrocarbons and crude oil by mixed strains and bacterial community analysis. Appl Biochem Biotechnol 172:3433–3447
Chen S, Zhou Y, Chen Y, Gu J (2018) fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34:i884–i890
Clauset A, Newman MEJ, Moore C (2004) Finding community structure in very large networks. Phys Rev E: Stat, Nonlinear, Soft Matter Phys 70:066111
Crump BC, Armbrust EV, Baross JA (1999) Phylogenetic analysis of particle-attached and free-living bacterial communities in the Columbia river, its estuary, and the adjacent coastal ocean. Appl Environ Microbiol 65:3192–3204
DeLong EF, Franks DG, Alldredge AL (1993) Phylogenetic diversity of aggregate-attached vs. free-living marine bacterial assemblages. Limnol Oceanogr 38:924–934
Dithugoe Choaro D, Bezuidt Oliver KI, Cavan Emma L, Froneman William P, Thomalla Sandy J, Makhalanyane Thulani P (2023) Bacteria and archaea regulate particulate organic matter export in suspended and sinking marine particle fractions. mSphere 8:e00420–e00422
Djurhuus A, Closek CJ, Kelly RP, Pitz KJ, Michisaki RP, Starks HA, Walz KR, Andruszkiewicz EA, Olesin E, Hubbard K, Montes E, Otis D, Muller-Karger FE, Chavez FP, Boehm AB, Breitbart M (2020) Environmental DNA reveals seasonal shifts and potential interactions in a marine community. Nat Commun 11:254
Dogs M, Wemheuer B, Wolter L, Bergen N, Daniel R, Simon M, Brinkhoff T (2017) Rhodobacteraceae on the marine brown alga Fucus spiralis are abundant and show physiological adaptation to an epiphytic lifestyle. Syst Appl Microbiol 40:370–382
Duan L, Li J-L, Yin L-Z, Luo X-Q, Ahmad M, Fang B-Z, Li S-H, Deng Q-Q, Wang P, Li W-J (2022) Habitat-dependent prokaryotic microbial community, potential keystone species, and network complexity in a subtropical estuary. Environ Res 212:113376
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461
Fenchel T (2008) The microbial loop – 25 years later. J Exp Mar Biol Ecol 366:99–103
Flombaum P, Gallegos JL, Gordillo RA, Rincón J, Zabala LL, Jiao N, Karl DM, Li WKW, Lomas MW, Veneziano D, Vera CS, Vrugt JA, Martiny AC (2013) Present and future global distributions of the marine Cyanobacteria Prochlorococcus and Synechococcus. Proc Natl Acad Sci USA 110:9824–9829
Friedman J, Alm EJ (2012) Inferring correlation networks from genomic survey data. PLoS Comput Biol 8:e1002687
Ganesh S, Parris DJ, DeLong EF, Stewart FJ (2014) Metagenomic analysis of size-fractionated picoplankton in a marine oxygen minimum zone. ISME J 8:187–211
Giering SLC, Sanders R, Lampitt RS, Anderson TR, Tamburini C, Boutrif M, Zubkov MV, Marsay CM, Henson SA, Saw K, Cook K, Mayor DJ (2014) Reconciliation of the carbon budget in the ocean’s twilight zone. Nature 507:480–483
Gräler B, Pebesma EJ, Heuvelink GBM (2016) Spatio-Temporal Interpolation Using Gstat R J 8:204
Guimerà R, Amaral LAN (2005) Cartography of complex networks: modules and universal roles. J Stat Mech 2005:P02001
Harrison PJ, Yin K, Lee JHW, Gan J, Liu H (2008) Physical–biological coupling in the Pearl River Estuary. Cont Shelf Res 28:1405–1415
Herndl GJ, Reinthaler T (2013) Microbial control of the dark end of the biological pump. Nat Geosci 6:718–724
Huber H, Gallenberger M, Jahn U, Eylert E, Berg IA, Kockelkorn D, Eisenreich W, Fuchs G (2008) A dicarboxylate/4-hydroxybutyrate autotrophic carbon assimilation cycle in the hyperthermophilic Archaeum Ignicoccus hospitalis. Proc Natl Acad Sci USA 105:7851–7856
Hyatt D, Chen G-L, LoCascio PF, Land ML, Larimer FW, Hauser LJ (2010) Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11:119
Jain A, Krishnan KP, Singh A, Thomas FA, Begum N, Tiwari M, Bhaskar VP, Gopinath A (2019) Biochemical composition of particles shape particle-attached bacterial community structure in a high Arctic fjord. Ecol Indic 102:581–592
Jiao N, Herndl GJ, Hansell DA, Benner R, Kattner G, Wilhelm SW, Kirchman DL, Weinbauer MG, Luo T, Chen F, Azam F (2010) Microbial production of recalcitrant dissolved organic matter: long-term carbon storage in the global ocean. Nat Rev Microbiol 8:593–599
Jiao N, Tang K, Cai H, Mao Y (2011) Increasing the microbial carbon sink in the sea by reducing chemical fertilization on the land. Nat Rev Microbiol 9:75–75
Kaehler BD, Bokulich NA, McDonald D, Knight R, Caporaso JG, Huttley GA (2019) Species abundance information improves sequence taxonomy classification accuracy. Nat Commun 10:4643
Kang DD, Li F, Kirton E, Thomas A, Egan R, An H, Wang Z (2019) MetaBAT 2: an adaptive binning algorithm for robust and efficient genome reconstruction from metagenome assemblies. PeerJ 7:e7359
Karner M, Herndl GJ (1992) Extracellular enzymatic activity and secondary production in free-living and marine-snow-associated bacteria. Mar Biol 113:341–347
Lai J, Zou Y, Zhang J, Peres-Neto PR (2022) Generalizing hierarchical and variation partitioning in multiple regression and canonical analyses using the rdacca.hp R package. Methods Ecol Evol 13:782–788
Lee J-W, Nam J-H, Kim Y-H, Lee K-H, Lee D-H (2008) Bacterial communities in the initial stage of marine biofilm formation on artificial surfaces. J Microbiol 46:174–182
Lehmann MF, Bernasconi SM, McKenzie JA, Barbieri A, Simona M, Veronesi M (2004) Seasonal variation of the δC and δN of particulate and dissolved carbon and nitrogen in Lake Lugano: constraints on biogeochemical cycling in a eutrophic lake. Limnol Oceanogr 49:415–429
Li W, Godzik A (2006) Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22:1658–1659
Li J, Jia H, Cai X, Zhong H, Feng Q, Sunagawa S, Arumugam M, Kultima JR, Prifti E, Nielsen T, Juncker AS, Manichanh C, Chen B, Zhang W, Levenez F, Wang J, Xu X, Xiao L, Liang S, Zhang D et al (2014) An integrated catalog of reference genes in the human gut microbiome. Nat Biotechnol 32:834–841
Li D, Luo R, Liu C-M, Leung C-M, Ting H-F, Sadakane K, Yamashita H, Lam T-W (2016) MEGAHIT v1.0: a fast and scalable metagenome assembler driven by advanced methodologies and community practices. Methods 102:3–11
Li J-L, Salam N, Wang P-D, Chen L-X, Jiao J-Y, Li X, Xian W-D, Han M-X, Fang B-Z, Mou X-Z, Li W-J (2018) Discordance between resident and active bacterioplankton in free-living and particle-associated communities in estuary ecosystem. Microb Ecol 76:637–647
Li JH, Cheng BH, Zhang R, Li WJ, Shi XM, Han YW, Ye LF, Ostle NJ, Bardgett RD (2021a) Nitrogen and phosphorus additions accelerate decomposition of slow carbon pool and lower total soil organic carbon pool in alpine meadows. Land Degrad Dev 32:1761–1772
Li JH, Zhang R, Cheng BH, Ye LF, Li WJ, Shi XM (2021b) Effects of nitrogen and phosphorus additions on decomposition and accumulation of soil organic carbon in alpine meadows on the Tibetan Plateau. Land Degrad Dev 32:1467–1477
Li L, Huang D, Hu Y, Rudling NM, Canniffe DP, Wang F, Wang Y (2023) Globally distributed Myxococcota with photosynthesis gene clusters illuminate the origin and evolution of a potentially chimeric lifestyle. Nat Commun 14:6450
Li X, Cheng X, Cheng K, Cai Z, Feng S, Zhou J (2024) The influence of tide-brought nutrients on microbial carbon metabolic profiles of mangrove sediments. Sci Total Environ 906:167732
Liu X, Liu M, Chen X, Yang Y, Hou L, Wu S, Zhu P (2019) Indigenous PAH degraders along the gradient of the Yangtze Estuary of China: relationships with pollutants and their bioremediation implications. Mar Pollut Bull 142:419–427
Liu Y, Lin Q, Feng J, Yang F, Du H, Hu Z, Wang H (2020) Differences in metabolic potential between particle-associated and free-living bacteria along Pearl River Estuary. Sci Total Environ 728:138856
Luo X-Q, Wang P, Li J-L, Ahmad M, Duan L, Yin L-Z, Deng Q-Q, Fang B-Z, Li S-H, Li W-J (2022) Viral community-wide auxiliary metabolic genes differ by lifestyles, habitats, and hosts. Microbiome 10:190
Ma Y, Li P, Zhong H, He M, Wang B, Mou X, Wu L (2023) The ecological differentiation of particle-attached and free-living bacterial communities in a seasonal flooding lake—the Poyang Lake. Microb Ecol 86:795–809
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. Embnet J 17:3
Martin JH, Knauer GA, Karl DM, Broenkow WW (1987) VERTEX: carbon cycling in the northeast Pacific. Deep-Sea Res, Part A 34:267–285
Mirdita M, Steinegger M, Breitwieser F, Söding J, Levy Karin E (2021) Fast and sensitive taxonomic assignment to metagenomic contigs. Bioinformatics 37:3029–3031
Mohiuddin MM, Botts SR, Paschos A, Schellhorn HE (2019) Temporal and spatial changes in bacterial diversity in mixed use watersheds of the Great Lakes region. J Gt Lakes Res 45:109–118
Olesen JM, Bascompte J, Dupont YL, Jordano P (2007) The modularity of pollination networks. Proc Natl Acad Sci USA 104:19891–19896
Olm MR, Brown CT, Brooks B, Banfield JF (2017) dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication. ISME J 11:2864–2868
Osburn ED, Aylward FO, Barrett JE (2021) Historical land use has long-term effects on microbial community assembly processes in forest soils. ISME Commun 1:48
Parada AE, Needham DM, Fuhrman JA (2016) Every base matters: assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples. Environ Microbiol 18:1403–1414
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW (2015) CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res 25:1043–1055
Parks DH, Rinke C, Chuvochina M, Chaumeil P-A, Woodcroft BJ, Evans PN, Hugenholtz P, Tyson GW (2017) Recovery of nearly 8,000 metagenome-assembled genomes substantially expands the tree of life. Nat Microbiol 2:1533–1542
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2012) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596
Rafrafi Y, Trably E, Hamelin J, Latrille E, Meynial-Salles I, Benomar S, Giudici-Orticoni M-T, Steyer J-P (2013) Sub-dominant bacteria as keystone species in microbial communities producing bio-hydrogen. Int J Hydrogen Energy 38:4975–4985
Schlitzer R (2002) Carbon export fluxes in the Southern Ocean: results from inverse modeling and comparison with satellite-based estimates. Deep Sea Res 2 Top Stud Oceanogr 49:1623–1644
Shi Z, Yin H, Van Nostrand JD, Voordeckers JW, Tu Q, Deng Y, Yuan M, Zhou A, Zhang P, Xiao N, Ning D, He Z, Wu L, Zhou J (2019) Functional gene array-based ultrasensitive and quantitative detection of microbial populations in complex communities. mSystems 4:e00296-19
Siegenthaler U, Sarmiento JL (1993) Atmospheric carbon dioxide and the ocean. Nature 365:119–125
Smith M, Zeigler Allen L, Allen A, Herfort L, Simon H (2013) Contrasting genomic properties of free-living and particle-attached microbial assemblages within a coastal ecosystem. Front Microbiol 4:120
Stegen JC, Lin X, Fredrickson JK, Chen X, Kennedy DW, Murray CJ, Rockhold ML, Konopka A (2013) Quantifying community assembly processes and identifying features that impose them. ISME J 7:2069–2079
Stegen JC, Lin X, Fredrickson JK, Konopka AE (2015) Estimating and mapping ecological processes influencing microbial community assembly. Front Microbiol 6:370
Strokal M, Kroeze C, Li L, Luan S, Wang H, Yang S, Zhang Y (2015) Increasing dissolved nitrogen and phosphorus export by the Pearl River (Zhujiang): a modeling approach at the sub-basin scale to assess effective nutrient management. Biogeochemistry 125:221–242
Sunagawa S, Coelho LP, Chaffron S, Kultima JR, Labadie K, Salazar G, Djahanschiri B, Zeller G, Mende DR, Alberti A, Cornejo-Castillo FM, Costea PI, Cruaud C, d’Ovidio F, Engelen S, Ferrera I, Gasol JM, Guidi L, Hildebrand F, Kokoszka F et al (2015) Structure and function of the global ocean microbiome. Science 348:1261359
Tu Q, Yu H, He Z, Deng Y, Wu L, Van Nostrand JD, Zhou A, Voordeckers J, Lee Y-J, Qin Y, Hemme CL, Shi Z, Xue K, Yuan T, Wang A, Zhou J (2014) GeoChip 4: a functional gene-array-based high-throughput environmental technology for microbial community analysis. Mol Ecol Resour 14:914–928
Wang P, Li J-L, Luo X-Q, Ahmad M, Duan L, Yin L-Z, Fang B-Z, Li S-H, Yang Y, Jiang L, Li W-J (2022) Biogeographical distributions of nitrogen-cycling functional genes in a subtropical estuary. Funct Ecol 36:187–201
Xu H, Zhao D, Huang R, Cao X, Zeng J, Yu Z, Hooker KV, Hambright KD, Wu QL (2018) Contrasting network features between free-living and particle-attached bacterial communities in Taihu Lake. Microb Ecol 76:303–313
Xu H, Zeng J, Li C, Zhao D, He R, Wu QL (2021) Distinct microhabitats affect the relative balance of ecological processes shaping the spatial distribution of bacterial communities in lakeshore habitats. Freshw Biol 66:1475–1489
Yuan X, Yin K, Harrison PJ, Cai W, He L, Xu J (2010) Bacterial production and respiration in subtropical Hong Kong waters: influence of the Pearl River discharge and sewage effluent. Aquat Microb Ecol 58:167–179
Zhang Y, Xiao W, Jiao N (2016) Linking biochemical properties of particles to particle-attached and free-living bacterial community structure along the particle density gradient from freshwater to open ocean. J Geophys Res: Biogeosci 121:2261–2274
Zhang L, Delgado-Baquerizo M, Shi Y, Liu X, Yang Y, Chu H (2021) Co-existing water and sediment bacteria are driven by contrasting environmental factors across glacier-fed aquatic systems. Water Res 198:117139
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 32200090), the Key-Area Research and Development Program of Guangdong Province (No. 2022B0202110001), Guangdong Basic and Applied Basic Research Foundation, China (Nos. 2023A1515012270, 2022A1515010756), the Science and Technology Program by Department of Natural Resources of Guangdong Province (GDNRC [2023]41), and Key Research Program Project of Guangzhou Science and Technology Bureau (No. 2024B03J1276).
Author information
Authors and Affiliations
Contributions
W.J.L., J.L.L., P.D.W., J.X.W., and Z.Q.P. conceived this study; J.L.L., W.D.X., W.C.Y., and Y.Y. collected the samples; Q.Q.D., X.Q.L., and Z.W.Y. performed the DNA extraction and physicochemical measurement; Z.Q.P. and P.D.W. conducted bioinformatic analysis. Z.Q.P. wrote the first draft manuscript; P.D.W., J.L.L., X.Z.M., and Z.Q.P. revised the manuscript with contributions and inputs from all authors. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Animal and human rights statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Edited by Chengchao Chen.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Peng, Z., Wang, P., Luo, X. et al. Community structure and carbon metabolism functions of bacterioplankton in the Guangdong coastal zone. Mar Life Sci Technol 6, 547–561 (2024). https://doi.org/10.1007/s42995-024-00245-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42995-024-00245-x