Abstract
Salicylic acid (SA) is a phytohormone required for plant growth and defense signaling. There are two major SA biosynthesis pathways in plants: the isochorismate synthase (ICS) pathway and the phenylalanine ammonia-lyase (PAL) pathway. It has been demonstrated in several plant species, including the model plant Arabidopsis, that SA is derived predominantly from the ICS pathway. Here, we employed the CRISPR/Cas9 system to generate ICS knockout mutants in rice (Oryza sativa L.). The Osics mutants display severe growth defects, and are completely devoid of phylloquinone, an isochorismate-derived product. The growth defects of Osics can be rescued through exogenous application of 1,4-dihydroxy-2-naphthoic acid (NA), a precursor of phylloquinone. Remarkably, the basal SA levels are not altered in the Osics mutants. Our findings support a role of OsICS in the biosynthesis of phylloquinone, and imply that SA biosynthesis in rice may occur through an alternative route other than the ICS pathway.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Salicylic acid (SA) not only functions as a pivotal phytohormone that mediates plant defense responses (Ding and Ding 2020), but also contributes to various aspects of plant growth and development, including photosynthesis, transpiration, seed germination, thermogenesis, drought resistance, and senescence (Khalvandi et al. 2021; Uzunova and Popova 2000; Vicente and Plasencia 2011; Vlot et al. 2009). Previous work acknowledges the existence of both the isochorismate synthase (ICS) and phenylalanine ammonia-lyase (PAL) pathways for SA biosynthesis in plants, in both cases SA originates from chorismate in the plastid (Lefevere et al. 2020). However, the exact SA biosynthesis process still remains largely unknown in lots of plant species. Dissecting the mechanism underlying plant SA biosynthesis will further our understanding of plant immunity and development, and may open avenues for developing novel strategies for crop improvement.
Genetic analysis of SA-deficient mutants has revealed that SA is predominantly synthesized through the ICS pathway in Arabidopsis and some other plant species, similar to the pathway described in Pseudomonas bacteria (Mercado-Blanco et al. 2001; Muller et al. 1996; Serino et al. 1995). The ICS pathway has been fully elucidated in Arabidopsis, in which chorismate is converted into isochorismate by ICS in plastid, and the resultant compound is exported to the cytosol to produce SA (Rekhter et al. 2019; Torrens-Spence et al. 2019). The ICS pathway accounts for over 90% of SA biosynthesis induced by pathogens or ultraviolet light in Arabidopsis (Garcion et al. 2008). The remaining 10% SA is believed to originate from the PAL pathway where chorismate undergoes a multistep enzymatic conversion into phenylalanine (Phe) and PAL converts Phe into trans-cinnamic acid, a precursor of SA (Yalpani et al. 1993). Interestingly, a recent isotopic tracing study in Arabidopsis challenged the notion that SA is synthesized from Phe (Wu et al. 2023), suggesting the presence of a PAL-independent pathway.
The ICS pathway is also predominant in some other plants such as tomato (Uppalapati et al. 2007), maize (Djamei et al. 2011), and tobacco (Catinot et al. 2008). However, the relative contribution of the ICS pathway to SA biosynthesis varies among different plant species. In soybean (Glycine max), both the ICS and PAL pathways are crucial for pathogen-induced SA biosynthesis, as knocking down either pathway halts SA production and compromises pathogen resistance (Shine et al. 2016). In addition, the ICS pathway is also implicated in the biosynthesis of phylloquinone (known as vitamin K1), an essential component for electron transfer in photosystem I (PSI) (Furt et al. 2010; Yuan et al. 2009). In a barley (Hordeum vulgare) ics mutant, phylloquinone is deficient while the basal SA level remains unchanged, suggesting that SA in barley likely synthesized through an ICS-independent pathway (Qin et al. 2019).
While being one of the most important staple crops, rice (Oryza sativa L.) also serves as a widely-used model monocot. Notably, rice has relatively high basal SA levels, which surpass those observed in other model plants such as Arabidopsis and tobacco, as well as in other cereal species like wheat and maize (Klessig et al. 2016; Pál et al. 2014). In this study, we generated Osics mutants in rice using CRISPR/Cas9 system and found that these Osics plants are seedling lethal and deficient in phylloquinone. However, the SA levels in the Osics mutants remained unchanged, indicating that SA in rice is biosynthesized through an ICS-independent pathway.
Results
Characterization of ICS in rice
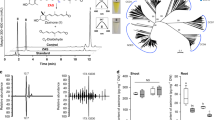
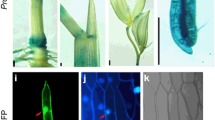
Unlike Arabidopsis, which harbors two ICS homologs, the rice Nipponbare genome contains a single-copy gene of ICS (Os09g036150). A maximum likelihood (ML) phylogenetic tree was constructed using the deduced amino acid sequences of OsICS and sequences of ICS proteins from other plant species, and OsICS clearly clusters together with other monocot ICSs (Fig. 1A). OsICS is expressed in the root, stem, and leaf of rice, with the highest expression level observed in the leaf (Fig. 1B). Analysis of the OsICS using the online tool (https://predictprotein.org/) predicted a chloroplastic localization. To confirm the subcellular localization of OsICS, OsICS was fused in frame with green fluorescent protein (GFP) and then transiently expressed in rice protoplasts. Confocal microscopy imaging of the transformed cells revealed that OsICS-GFP fluorescence colocalized with chlorophyll fluorescence (Fig. 1C), supporting that the OsICS protein is localized in the chloroplast, where the chorismate substrate is present.
OsICS knockout mutants exhibit severe growth defects. A Phylogenetic analysis of ICS proteins from rice and other plant species. The OsICS is highlighted in red. EntC, Escherichia coli enterobactin-specific isochorismate synthase; ICS, isochorismate synthase. B The expression levels of OsICS in root, stem, and leaf of wild-type (WT) plants at the seedling stage. Data are represented as mean ± SD of three biological replicates. C Subcellular localization of the OsICS protein in rice protoplasts. D OsICS knockout mutants. sgRNAs with NGG and NAG are designed as PAMs. The PAM sequences are indicated in red. The black line represents the genome sequence, and black boxes represent exons. Sequences of the sgRNA-induced OsICS mutations in the individual mutant lines are shown. E Phenotypes of 15-day-old (left) and 20-day-old (right) seedlings of the wild-type plants and Osics mutants. Scale bar, 5 cm. F, G Plant height (F) and fresh weight (G) of the 15-day-old wild-type and Osics seedlings. H Chlorophyll contents, including total chlorophyll (Chlt), chlorophyll a (Chla), and chlorophyll b (Chlb) of leaves from the 15-day-old wild-type and Osics seedlings. FW fresh weight. Data are represented as mean ± SD of three biological replicates. **P < 0.01, ***P < 0.001, ****P < 0.0001 (two-tailed Student’s t test)
OsICS knockout mutants exhibit severe growth defects
We used the CRISPR/Cas9 system to generate rice mutants of OsICS. Initially, we designed a single-guide RNA (sgRNA1) targeting the second exon of OsICS with a canonical NGG PAM (Fig. 1D). Due to the high editing efficiency of the sgRNA1, Osics mutants obtained were all homozygous and seedling lethal. Consequently, these homozygous rice ics mutant materials could not be reproduced for further study. Therefore, we redesigned sgRNAs with NAG PAM (Fig. 1D), facilitating the generation of heterozygous mutants (Meng et al. 2018). The sgRNAs, sgRNA2 and sgRNA3, targeting exon 3 and exon 10 of OsICS, resulted in two independent heterozygous lines with a 1 bp and 4 bp deletion, respectively (Fig. 1D). Homozygous Osics mutants (named as Osics-1 and Osics-2) were segregated and utilized for subsequent analyses.
Osics-1 and Osics-2 both exhibited yellow leaves and dwarf at 15 days after germination (Fig. 1E, left). Consistent with their phenotypes, the accumulation of chlorophyll, as well as fresh weight and plant height of the two Osics mutants, were significantly lower than those of wild-type plants (Fig. 2F–H). Finally, the seedlings became wilted and died (Fig. 1E, right). These results support that OsICS plays crucial roles in rice growth and development.
OsICS is required for phylloquinone, but not salicylic acid biosynthesis. A, B Contents of total SA (A) and phylloquinone (B) in 15-day-old wild-type plants and Osics mutants. Data are represented as mean ± SD of three biological replicates. ns indicates no statistical significance determined by two-tailed Student’s t test (p > 0.05). ND not detected. FW fresh weight. C Phenotypes of Osics mutants were rescued by exogenous application of 1,4-dihydroxy-2-naphthoic acid (NA). Rice plants were grown in Kimura B nutrient solution supplemented with either 0.1 mM NA or 0.1% dimethyl sulfoxide (mock) after seed germination. Scale bar, 5 cm. D Gene ontology enrichment analysis was performed on the differentially expressed genes (DEGs) in Osics-2 compared with the wild-type plants. Bar charts showing the representative terms. BP biological process, MF molecular function, CC cellular component. E Heatmap showing the differential expression of genes encoding photosystem I and photosystem II related proteins. Gene expression levels are represented by a heatmap using the log2-transformed FPKM values. The accession number for the indicated genes are listed in Table S1
The Osics mutants are phylloquinone-deficient, but have unaltered basal SA levels
In addition to catalyzing the production of SA, ICS also plays a role in the biosynthesis of phylloquinone, another isochorismate-derived end product present in many plants. In the Arabidopsis ics1ics2 double mutant, phylloquinone is entirely devoid, while retaining a low, but detectable residual level of SA (Garcion et al. 2008). To assess the content of phylloquinone and total SA in the homozygous Osics mutants, we conducted high performance liquid chromatography-tandem mass spectrometry (HPLC–MS/MS) analyses to precisely measure these two compounds. Both Osics-1 and Osics-2 mutants accumulated approximately 10 µg of total SA per gram of fresh weight, a level similar to that of wild-type plants (Fig. 2A). In contrast, phylloquinone was undetectable in Osics-1 and Osics-2 mutants (Fig. 2B). To confirm the involvement of ICS in phylloquinone biosynthesis, we exogenously supplied the mutant plants with 1,4-dihydroxy-2-naphthoic acid (NA), a precursor for phylloquinone. The Osics mutants were effectively rescued by NA, exhibiting green leaves and viable seedlings (Fig. 2C). These data indicate that the isochorismate synthesis is completely blocked in the Osics mutants and that basal SA is not derived from the ICS pathway in rice.
Disruption of OsICS compromised the expression of photosynthesis-associated genes
To elucidate the potential impacts of disrupting ICS in rice, we carried out RNA-seq analysis of a mutant line (Osics-2) and wild-type plants to identify differentially expressed genes (DEGs) between them. Gene ontology (GO) enrichment of the DEGs showed that processes related to photosynthesis, photosynthesis/light reaction, and pigment metabolism within the biological processes (BP) category were highly enriched and down-regulated in the Osics-2 mutant (Fig. 2D). Within the molecular function (MF) category, the majority of DEGs were associated with ATP binding and chlorophyll binding, which are crucial for light absorption and electron transfer during photosynthesis (Fig. 2D). Similarly, DEGs in the cellular components (CC) category were predominantly observed in chloroplast and photosystem (Fig. 2D). These data suggest that the process of photosynthesis is greatly affected in the Osics-2 mutant. We further checked DEGs associated with photosystem I and photosystem II. The expression of genes encoding the major subunits of photosystem I (PsaD, PsaF, PsaG, PsaH, PsaK, PsaL, and PsaN) exhibited obvious down-regulation in the Osics-2 mutant (Fig. 2E). Similarly, PsbO, PsbP, PsbQ, PsbR, PsbR3, PsbW, PsbX, PsbY, Psb27, and Psb28 family genes of photosystem II were also down-regulated in the Osics-2 mutant (Fig. 2E). Overall, our results demonstrate that disrupting ICS in rice results in compromised expression of photosynthesis-associated genes, supporting the function of ICS in phylloquinone biosynthesis.
Discussion
Biosynthesis of SA is fundamental for understanding of its function and regulation. SA was originally proposed to be synthesized through the PAL pathway in plants, as indicated by studies based on radiolabeled compounds in cucumber plants (Meuwly et al. 1995). However, in Arabidopsis, a second pathway, known as the ICS pathway which is similar to that described in some Pseudomonas bacteria, was discovered (Wildermuth et al. 2001). The existence of the ICS pathway gained strong support from investigations involving ics mutants in Arabidopsis. In addition to its deficiency in pathogen-induced SA accumulation (Wildermuth et al. 2001), the ics1ics2 double mutants exhibited a 80% decrease in total SA levels under basal conditions and a 95% decrease after UV induction in leaves (Garcion et al. 2008), which underscores the significance of the ICS pathway in biosynthesis of both basal and UV-induced SA in Arabidopsis. The indispensability of the ICS enzyme for SA production in response to biotic and abiotic stresses has been demonstrated in Nicotiana benthamiana, providing further evidence for the critical role of the ICS pathway in SA biosynthesis (Catinot et al. 2008; Shibata et al. 2010). It was also demonstrated in several other plant species that SA is primarily derived from the ICS pathway (Garcion et al. 2008). In this study, we successfully identified a functional ICS gene in rice and generated Osics mutants. These Osics mutants exhibited a yellowish leaf phenotype and were lethal at the seedling stage (Fig. 1), resembling the phylloquinone-deficient mutants pha, aae14, and abc4 in Arabidopsis (Gross et al. 2006; Kim et al. 2008; Shimada et al. 2005). Importantly, no detectable phylloquinone was observed in the Osics mutants (Fig. 2), indicating that this ICS protein is solely responsible for the isochorismate production in rice. We measured the total SA content in these Osics mutants and found that the total SA content remained unchanged (Fig. 2). This observation provides genetic evidence for the existence of an ICS-independent pathway for SA production in rice. The biosynthesis of SA in rice appears similar to that in barley, in which SA is synthesized through an ICS-independent pathway as well (Qin et al. 2019). These observations provide evidence that the production of SA exclusively via an ICS-independent route is likely widespread in monocots. However, the pathway through which SA is synthesized in rice still remains for further investigation, with the PAL pathway being the most likely candidate.
It has been reported that some plants predominantly employ the PAL pathway for SA biosynthesis. For example, in the PAL1-silenced plants of Aegilops variabilis, basal SA levels in the roots decreased by 75%. In contrast, the SA levels were not changed in the AevICS-silenced plants (Zhang et al. 2021), suggesting that SA is primarily synthesized through the PAL pathway in Ae. Variabilis. In the case of rice, which has very high basal SA levels, its SA biosynthesis has been previously presumed to be through a PAL-dependent pathway (Xu et al. 2023). Nine genes have been annotated as PALs in rice, with PAL1-7 co-localizing with disease resistance quantitative trait loci (QTLs), indicating their involvement in plant defense (Tonnessen et al. 2015). Among these, PAL6 has been shown to play a role in SA accumulation. The rice PAL6 T-DNA insertion line exhibited great reduction in PAL activities and a decrease in SA levels (Duan et al. 2014). In addition, SA accumulation in rice was shown to be dependent on OsAIM1, a β-oxidase responsible for the production of benzoic acid (BA), which may serve as a precursor of SA in the PAL pathway. Knocking out OsAIM1 leads to decreased SA levels (Xu et al. 2017, 2023). These observations underscore the significance of the PAL pathway for SA biosynthesis in rice. However, the regulation of SA biosynthesis in rice may differ from other plant species due to the high basal SA levels. Further research should be extended on identification of key SA biosynthesis enzymes in rice and on better understanding the multifaceted functions of SA in this major staple crop.
Materials and methods
Plant materials and growth conditions
Rice (Oryza sativa ssp. japonica cv. Nipponbare) seeds were germinated at 37 °C under dark conditions. After germination, rice plants were grown in Kimura B nutrient solution (Coolaber, China) in a growth chamber with 85% relative humidity under 13 h/11 h light/dark cycle at 28 °C/24 °C day/night temperatures. For NA treatment experiment, 5-day-old Osics plants were supplied with 0.1 mM NA or buffer (0.1% dimethyl sulfoxide) and the nutrient solution was renewed every 3 days.
Phylogenetic analysis of ICS
Multiple sequence alignments of ICS proteins from different species were performed using ClustalW with default parameters. The maximum likelihood (ML) phylogenetic tree was constructed using MEGA 11.0. Accession number of ICS proteins are listed in Table S2.
CRISPR/Cas9 editing vector construction and rice genetic transformation
The CRISPR v2.0 (http://crispr.hzau.edu.cn/CRISPR2/) was used for designing sgRNA. A pair of complimentary oligos corresponding to each sgRNA were synthesized commercially, and then were annealed and inserted at the BsaI site of the vector pHUE411 (Xing et al. 2014). The construct was then introduced into rice embryogenic calli via Agrobacterium tumefaciens as described (Hiei et al. 1994). The edited plants were identified with PCR. The primers used are listed in Table S3.
Chlorophyll, SA and phylloquinone measurements
Chlorophyll content was determined as described, with minor modifications (Alam et al. 2022; Li et al. 2022). Fresh leaves (approximately 30 mg) were ground to powder with liquid nitrogen and extracted with 1.8 mL of 80% acetone. After centrifuged at 10,000 rpm for 5 min, the resulting supernatants were measured with spectrophotometric scanning at 663 nm and 645 nm for chlorophyll a (Chla) and chlorophyll b (Chlb), respectively. For SA and phylloquinone measurements, the leaves (approximately 0.1 g) of 15-day-old rice seedlings were collected and ground into powder in liquid nitrogen. Extraction and measurement of SA and phylloquinone were performed as previously described (Glauser et al. 2014; Lohmann et al. 2006).
Subcellular localization of OsICS
OsICS was fused in frame with GFP by cloning the coding sequence without the stop codon of OsICS into the pJIT163-ubi-GFP vector (Wang et al. 2014). The pJIT163-Ubi-GFP (control) and pJIT163-Ubi-OsICS-GFP constructs were transformed into rice protoplasts as described (Shan et al. 2014). The fluorescence images were obtained with a Leica TCS SP8 laser scanning confocal microscope.
RNA-seq and quantitative RT-PCR assays
Total RNA was extracted using a RiboPure kit (Invitrogen, USA). RNA-seq was conducted using the Illumina NovaSeq platform, and DEGs were detected by edgeR with a threshold absolute value of |log2(fold change)|≥ 1 and Padj (adjusted P-value) ≤ 0.05 (Robinson et al. 2010). GO enrichment analysis of DEGs was performed using Gene Ontologies (https://www.geneontology.org/). For quantitative RT-PCR (qRT-PCR), 1 µg of total RNA was reverse-transcribed into first-strand cDNA using Superscript Reverse Transcriptase III (Invitrogen, USA). qRT-PCR was performed using the SYBR Premix Ex Taq™ kit (Takara, Japan). Ubiqutin was used as the internal control. Normalized expression levels were determined using the 2−ΔCt method. The primers are listed in Table S3.
Data availability
All data generated in this study are available in the paper.
References
Alam MS, Kong J, Tao R, Ahmed T, Alamin M, Alotaibi SS, Abdelsalam NR, Xu JH (2022) CRISPR/Cas9 mediated knockout of the OsbHLH024 transcription factor improves salt stress resistance in rice (Oryza sativa L.). Plants-Basel 11:1184. https://doi.org/10.3390/plants11091184
Catinot J, Buchala A, Abou-Mansour E, Métraux JP (2008) Salicylic acid production in response to biotic and abiotic stress depends on isochorismate in Nicotiana benthamiana. FEBS Lett 582:473–478. https://doi.org/10.1016/j.febslet.2007.12.039
Ding P, Ding Y (2020) Stories of salicylic acid: a plant defense hormone. Trends Plant Sci 25:549–565. https://doi.org/10.1016/j.tplants.2020.01.004
Djamei A, Schipper K, Rabe F, Ghosh A, Vincon V, Kahnt J, Osorio S, Tohge T, Fernie AR, Feussner I, Feussner K, Meinicke P, Stierhof YD, Schwarz H, Macek B, Mann M, Kahmann R (2011) Metabolic priming by a secreted fungal effector. Nature 478:395–398. https://doi.org/10.1038/nature10454
Duan L, Liu H, Li X, Xiao J, Wang S (2014) Multiple phytohormones and phytoalexins are involved in disease resistance to Magnaporthe oryzae invaded from roots in rice. Physiol Plant 152:486–500. https://doi.org/10.1111/ppl.12192
Furt F, Oostende C, Widhalm JR, Dale MA, Wertz J, Basset GJ (2010) A bimodular oxidoreductase mediates the specific reduction of phylloquinone (vitamin K1) in chloroplasts. Plant J 64:38–46. https://doi.org/10.1111/j.1365-313X.2010.04305.x
Garcion C, Lohmann A, Lamodière E, Catinot J, Buchala A, Doermann P, Métraux JP (2008) Characterization and biological function of the ISOCHORISMATE SYNTHASE2 gene of Arabidopsis. Plant Physiol 147:1279–1287. https://doi.org/10.1104/pp.108.119420
Glauser G, Vallat A, Balmer D (2014) Hormone profiling. In: Sanchez-Serrano, J., Salinas, J. (eds) Arabidopsis Protocols. Methods Mol Biol 1062:597–608. https://doi.org/10.1007/978-1-62703-580-4_31
Gross J, Cho WK, Lezhneva L, Falk J, Krupinska K, Shinozaki K, Seki M, Herrmann RG, Meurer J (2006) A plant locus essential for phylloquinone (vitamin K1) biosynthesis originated from a fusion of four eubacterial genes. J Biol Chem 281:17189–17196. https://doi.org/10.1074/jbc.M601754200
Hiei Y, Ohta S, Komari T, Kumashiro T (1994) Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J 6:271–282. https://doi.org/10.1046/j.1365-313x.1994.6020271.x
Khalvandi M, Siosemardeh A, Roohi E, Keramati S (2021) Salicylic acid alleviated the effect of drought stress on photosynthetic characteristics and leaf protein pattern in winter wheat. Heliyon 7:e05908. https://doi.org/10.1016/j.heliyon.2021.e05908
Kim HU, van Oostende C, Basset GJC, Browse J (2008) The AAE14 gene encodes the Arabidopsis o-succinylbenzoyl-CoA ligase that is essential for phylloquinone synthesis and photosystem-I function. Plant J 54:272–283. https://doi.org/10.1111/j.1365-313X.2008.03416.x
Klessig DF, Tian M, Choi HW (2016) Multiple targets of salicylic acid and its derivatives in plants and animals. Front Immunol 7:206. https://doi.org/10.3389/fimmu.2016.00206
Lefevere H, Bauters L, Gheysen G (2020) Salicylic acid biosynthesis in plants. Front Plant Sci 11:521987. https://doi.org/10.3389/fpls.2020.00338
Li S, Lin D, Zhang Y, Deng M, Chen Y, Lv B, Li B, Lei Y, Wang Y, Zhao L, Liang Y, Liu J, Chen K, Liu Z, Xiao J, Qiu JL, Gao C (2022) Genome-edited powdery mildew resistance in wheat without growth penalties. Nature 602:455–460. https://doi.org/10.1038/s41586-022-04395-9
Lohmann A, Schöttler MA, Bréhélin C, Kessler F, Bock R, Cahoon EB, Dörmann P (2006) Deficiency in phylloquinone (vitamin K1) methylation affects prenyl quinone distribution, photosystem I abundance, and anthocyanin accumulation in the Arabidopsis AtmenG mutant. J Biol Chem 281:40461–40472. https://doi.org/10.1074/jbc.M609412200
Meng X, Hu X, Liu Q, Song X, Gao C, Li J, Wang K (2018) Robust genome editing of CRISPR-Cas9 at NAG PAMs in rice. Sci China Life Sci 61:122–125. https://doi.org/10.1007/s11427-017-9247-9
Mercado-Blanco J, van der Drift KM, Olsson PE, Thomas-Oates JE, van Loon LC, Bakker PA (2001) Analysis of the pmsCEAB gene cluster involved in biosynthesis of salicylic acid and the siderophore pseudomonine in the biocontrol strain Pseudomonas fluorescens WCS374. J Bacteriol 183:1909–1920. https://doi.org/10.1128/jb.183.6.1909-1920.2001
Meuwly P, Molders W, Buchala A, Metraux JP (1995) Local and systemic biosynthesis of salicylic acid in infected cucumber plants. Plant Physiol 109:1107–1114. https://doi.org/10.1104/pp.109.3.1107
Muller R, Dahm C, Schulte G, Leistner E (1996) An isochorismate hydroxymutase isogene in Escherichia coli. FEBS Lett 378:131–134. https://doi.org/10.1016/0014-5793(95)01436-5
Pál M, Kovács V, Szalai G, Soós V, Ma X, Liu H, Mei H, Janda T (2014) Salicylic acid and abiotic stress responses in rice. J Agron Crop Sci 200:1–11. https://doi.org/10.1111/jac.12037
Qin Y, Torp AM, Glauser G, Pedersen C, Rasmussen SK, Thordal-Christensen H (2019) Barley isochorismate synthase mutant is phylloquinone-deficient, but has normal basal salicylic acid level. Plant Signal Behav 14:1671122. https://doi.org/10.1080/15592324.2019.1671122
Rekhter D, Lüdke D, Ding Y, Feussner K, Zienkiewicz K, Lipka V, Wiermer M, Zhang Y, Feussner I (2019) Isochorismate-derived biosynthesis of the plant stress hormone salicylic acid. Science 365:498–502. https://doi.org/10.1126/science.aaw1720
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140. https://doi.org/10.1093/bioinformatics/btp616
Serino L, Reimmann C, Baur H, Beyeler M, Visca P, Haas D (1995) Structural genes for salicylate biosynthesis from chorismate in Pseudomonas aeruginosa. Mol Gen Genet 249:217–228. https://doi.org/10.1007/bf00290369
Shan Q, Wang Y, Li J, Gao C (2014) Genome editing in rice and wheat using the CRISPR/Cas system. Nat Protoc 9:2395–2410. https://doi.org/10.1038/nprot.2014.157
Shibata Y, Kawakita K, Takemoto D (2010) Age-related resistance of Nicotiana benthamiana against hemibiotrophic pathogen Phytophthora infestans requires both ethylene- and salicylic acid-mediated signaling pathways. Mol Plant Microbe Interact 23:1130–1142. https://doi.org/10.1094/mpmi-23-9-1130
Shimada H, Ohno R, Shibata M, Ikegami I, Onai K, Ohto M, Takamiya K (2005) Inactivation and deficiency of core proteins of photosystems I and II caused by genetical phylloquinone and plastoquinone deficiency but retained lamellar structure in a T-DNA mutant of Arabidopsis. Plant J 41:627–637. https://doi.org/10.1111/j.1365-313X.2004.02326.x
Shine MB, Yang JW, El-Habbak M, Nagyabhyru P, Fu DQ, Navarre D, Ghabrial S, Kachroo P, Kachroo A (2016) Cooperative functioning between phenylalanine ammonia lyase and isochorismate synthase activities contributes to salicylic acid biosynthesis in soybean. New Phytol 212:627–636. https://doi.org/10.1111/nph.14078
Tonnessen BW, Manosalva P, Lang JM, Baraoidan M, Bordeos A, Mauleon R, Oard J, Hulbert S, Leung H, Leach JE (2015) Rice phenylalanine ammonia-lyase gene OsPAL4 is associated with broad spectrum disease resistance. Plant Mol Biol 87:273–286. https://doi.org/10.1007/s11103-014-0275-9
Torrens-Spence MP, Bobokalonova A, Carballo V, Glinkerman CM, Pluskal T, Shen A, Weng J-K (2019) PBS3 and EPS1 complete salicylic acid biosynthesis from isochorismate in Arabidopsis. Mol Plant 12:1577–1586. https://doi.org/10.1016/j.molp.2019.11.005
Uppalapati SR, Ishiga Y, Wangdi T, Kunkel BN, Anand A, Mysore KS, Bender CL (2007) The phytotoxin coronatine contributes to pathogen fitness and is required for suppression of salicylic acid accumulation in tomato inoculated with Pseudomonas syringae pv. tomato DC3000. Mol Plant Microbe Interact 20:955–965. https://doi.org/10.1094/mpmi-20-8-0955
Uzunova AN, Popova LP (2000) Effect of salicylic acid on leaf anatomy and chloroplast ultrastructure of barley plants. Photosynthetica 38:243–250. https://doi.org/10.1023/a:1007226116925
Vicente MR-S, Plasencia J (2011) Salicylic acid beyond defence: its role in plant growth and development. J Exp Bot 62:3321–3338. https://doi.org/10.1093/jxb/err031
Vlot AC, Dempsey DMA, Klessig DF (2009) Salicylic acid, a multifaceted hormone to combat disease. Annu Rev Phytopathol 47:177–206. https://doi.org/10.1146/annurev.phyto.050908.135202
Wang Y, Cheng X, Shan Q, Zhang Y, Liu J, Gao C, Qiu JL (2014) Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat Biotechnol 32:947–951. https://doi.org/10.1038/nbt.2969
Wildermuth MC, Dewdney J, Wu G, Ausubel FM (2001) Isochorismate synthase is required to synthesize salicylic acid for plant defence. Nature 414:562–565. https://doi.org/10.1038/35107108
Wu J, Zhu W, Zhao Q (2023) Salicylic acid biosynthesis is not from phenylalanine in Arabidopsis. J Integr Plant Biol 65:881–887. https://doi.org/10.1111/jipb.13410
Xing HL, Dong L, Wang ZP, Zhang HY, Han CY, Liu B, Wang XC, Chen QJ (2014) A CRISPR/Cas9 toolkit for multiplex genome editing in plants. BMC Plant Biol 14:327. https://doi.org/10.1186/s12870-014-0327-y
Xu L, Zhao H, Ruan W, Deng M, Wang F, Peng J, Luo J, Chen Z, Yi K (2017) ABNORMAL INFLORESCENCE MERISTEM1 functions in salicylic acid biosynthesis to maintain proper reactive oxygen species levels for root meristem activity in rice. Plant Cell 29:560–574. https://doi.org/10.1105/tpc.16.00665
Xu L, Zhao H, Wang J, Wang X, Jia X, Wang L, Xu Z, Li R, Jiang K, Chen Z, Luo J, Xie X, Yi K (2023) AIM1-dependent high basal salicylic acid accumulation modulates stomatal aperture in rice. New Phytol 238:1420–1430. https://doi.org/10.1111/nph.18842
Yalpani N, Leon J, Lawton MA, Raskin I (1993) Pathway of salicylic acid biosynthesis in healthy and virus-inoculated tobacco. Plant Physiol 103:315–321. https://doi.org/10.1104/pp.103.2.315
Yuan Y, Chung JD, Fu X, Johnson VE, Ranjan P, Booth SL, Harding SA, Tsai CJ (2009) Alternative splicing and gene duplication differentially shaped the regulation of isochorismate synthase in Populus and Arabidopsis. Proc Natl Acad Sci U S A 106:22020–22025. https://doi.org/10.1073/pnas.0906869106
Zhang H, Huang Q, Yi L, Song X, Li L, Deng G, Liang J, Chen F, Yu M, Long H (2021) PAL-mediated SA biosynthesis pathway contributes to nematode resistance in wheat. Plant J 107:698–712. https://doi.org/10.1111/tpj.15316
Funding
This work was supported by the National Key R&D Program of China (2023ZD0407305), National Natural Science Foundation of China (32230088), Open Project of State Key Laboratory of Plant Genomics (SKLPG2016A-32), Biological Breeding Engineering of Shanxi Agricultural University (YZGC094).
Author information
Authors and Affiliations
Contributions
J-LQ and JW conceived the study. ZW, GY and DZ conducted the experiments. ZW, GY and J-LQ analyzed the data. JW, GY and J-LQ wrote the manuscript. All authors read and approved the final version for publication.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, Z., Yang, G., Zhang, D. et al. Isochorismate synthase is required for phylloquinone, but not salicylic acid biosynthesis in rice. aBIOTECH (2024). https://doi.org/10.1007/s42994-024-00166-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42994-024-00166-4