Abstract
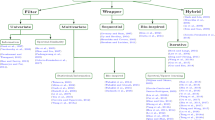
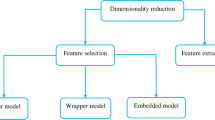
Autism is not just the consequence of mutations in single genes but across many genes. The dynamic changes in genes are well expressed by gene expression microarray data. The potential role of researchers still persists in finding the most prominent genes that influence autism risk. In this work, microarray data collected has 146 samples of autism and a control group with 54,613 gene expression features. The features and samples suffer from dimensionality issues to train the machine learning model. To reduce the dimension of features without loss of generality, four feature selection methods are used to select 100 prominent genes each with statistical significance. The feature selection model is concatenated to the neural network classifier model for classification. The neural network model is configured with proper adjustments of hyper parameters. Each feature selection method is combined with a neural network model for the analysis of prominent genes to better the classification of autism. The metrics used to access the classification are accuracy and loss. The neural network model is trained for ten folds and tested for untrained data. The optimized framework shows an accuracy of 78.6% with a minimum loss of 0.694 for the untrained data. The optimized model is identified with a trade-off between accuracy and loss.







Similar content being viewed by others
References
Kaliyappan K, Palanisamy M, Govindarajan R, Duraiyan J. Microarray and its applications. J Pharm Bioallied Sci. 2012;4:310.
Latkowski T, Osowski S. Gene selection in autism—comparative study. Neurocomputing. 2017;250:37–44.
Hameed SS, Hassan R, Muhammad FF. Selection and classification of gene expression in autism disorder: use of a combination of statistical filters and a GBPSO-SVM algorithm. PLoS ONE. 2017;12:1–25.
Liu S, et al. Feature selection of gene expression data for cancer classification using double RBF-kernels. BMC Bioinform. 2018;19:1–14.
Journal I, Factor I. biomed research international (J Biomed Biotechnol). Comput Math Methods Med. 2015;2015:2–4.
Wagner RF. From medical images to multiple-biomarker microarrays. Med Phys. 2007;34:4944–51.
Asif M, Martiniano HF, Vicente AM, Couto FM. Identifying disease genes using machine learning and gene functional similarities, assessed through Gene Ontology. PLoS ONE. 2018;13:1–15.
Latkowski T, Osowski S. Data mining for feature selection in gene expression autism data. Expert Syst Appl. 2015;42:864–72.
Wang L, Audenaert P, Michoel T. High-dimensional bayesian network inference from systems genetics data using genetic node ordering. Front Genet. 2019;10:1–13.
Roopa BS, Manjunatha Prasad R. Concatenating framework in ASD analysis towards research progress. In 1st international conference on advanced technologies in intelligent control, environment, computing and communication engineering. ICATIECE 2019; 2019. p. 269–71. https://doi.org/10.1109/ICATIECE45860.2019.9063816
Roopa BS, Prasad RM. Identification of best fit learning models based on calibration for better classification of autism. Int J Eng Adv Technol. 2020;9:2090–4.
Krishnan A, et al. Genome-wide prediction and functional characterization of the genetic basis of autism spectrum disorder. Nat Neurosci. 2016;19:1454–62. https://doi.org/10.1038/nn.4353
Courchesne E, Pierce K. Brain overgrowth in autism during a critical time in development: implications for frontal pyramidal neuron and interneuron development and connectivity. Int J Dev Neurosci. 2005;23:153–70.
Giorgia Canali, Marta Garcia, Bruno Hivert, Delphine Pinatel, Aline Goullancourt, et al. Genetic variants in autism-related CNTNAP2 impair axonal growth of cortical neurons. Human Molecular Genetics, Oxford University Press (OUP), pp.1941–1954. 2018. https://doi.org/10.1093/hmg/ddy102.hal-01963618.
Rylaarsdam L, Guemez-Gamboa A. Genetic causes and modifiers of autism spectrum disorder. Front Cell Neurosci. 2019. https://doi.org/10.3389/fncel.2019.00385.
Tost H, et al. A common allele in the oxytocin receptor gene (OXTR) impacts prosocial temperament and human hypothalamic-limbic structure and function. Proc Natl Acad Sci USA. 2010;107:13936–41.
Wilson DS. Benign Application of Knowledge through Evolutionary Theory. In: Madhavan G, Oakley B, Kun L, editors. Career Development in Bioengineering and Biotechnology. Series in Biomedical Engineering. New York, NY: Springer; 2008. https://doi.org/10.1007/978-0-387-76495-5_67.
Vissers LELM, et al. De novo variants in CNOT1, a central component of the CCR4-NOT complex involved in gene expression and RNA and protein stability, cause neurodevelopmental delay. Am J Hum Genet. 2020;107:164–72.
Winkler GS, Mulder KW, Bardwell VJ, Kalkhoven E, Timmers HTM. Human Ccr4-Not complex is a ligand-dependent repressor of nuclear receptor-mediated transcription. EMBO J. 2006;25:3089–99.
Kruszka P, et al. A CCR4-NOT transcription complex, subunit 1, CNOT1, variant associated with holoprosencephaly. Am J Hum Genet. 2019;104:990–3.
Alter MD, et al. Autism and increased paternal age related changes in global levels of gene expression regulation. PLoS ONE. 2011;6(2): https://doi.org/10.1371/journal.pone.0016715.
Chen Y, Dougherty ER, Bittner ML, Meltzer P, Trent J. Microarray Image Analysis and Gene Expression Ratio Statistics. In: Zhang W, Shmulevich I, editors. Computational and Statistical Approaches to Genomics. Boston, MA: Springer; 2006. https://doi.org/10.1007/0-387-26288-1_1.
Technically Speaking: Why We Use Random Sampling in Reading ResearchBy: Adam Reeger, M.S., Ariel M. Aloe, Ph.D., Posted on: November 12 2019.
Jordan J. Neural networks: training with backpropagation (2017).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the topical collection “Data Science and Communication” guest edited by Kamesh Namudri, Naveen Chilamkurti, Sushma S J and S. Padmashree.
Rights and permissions
About this article
Cite this article
Roopa, B.S., Manjunatha Prasad, R. A Selection of an Optimal Framework Identifying the Prominent Autism Risk Gene Biomarkers from Gene Expression Data Using Neural Network. SN COMPUT. SCI. 2, 241 (2021). https://doi.org/10.1007/s42979-021-00559-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42979-021-00559-y