Abstract
Sunflower is important for food, feed and ornamentals, faces rising demand for quality oil. Identifying resilient high-yielding genotypes is essential. Multi-environmental trials and integrated approaches to evaluate genotype by environment interactions (GEI) enhance accuracy and comprehensiveness of breeding programs. This study assess the representativeness and discriminative power of tested environments; classify mega-environments for genotype recommendations and evaluate how GGE and BLUP methods identify stable and superior sunflower genotypes. The experiment used a randomized complete block design with three replications across six locations for two seasons. The pooled analysis of variance for GEI on seed yield showed significant variability (p < 0.05) with mean values from 1171 to 2564 kg ha−1, indicating inconsistent genotypes performance across environments. The study identified E11, E2, and E6 as key discriminating and representative environments and grouped test locations into four mega-environments. Genotypes G6, G1, and G4 are the most desirable based on stability and seed yield. E11 and G6 are identified as ideal environment and genotype, respectively. GGE biplot and BLUP methods consistently identify stable and ideal sunflower genotypes. However, GGE stability indices differ somewhat in assessing genotypes stability possibly due to low number of genotypes, testing environment, and year in the dataset. The study confirmed that simultaneous consideration of GGE and BLUP models for GEI analysis helps to select more stable and adaptable sunflower genotypes across environments. It also clarifies GEI pattern, identifying mega-environments, and aiding genotype selection in sunflower production. However, validating these mega-environments and exploring the genetic basis of adaptability and stability remains essential.
Article Highlights
-
Explored how genetics and environment interact to influence sunflower yield.
-
Identified optimal environments for precise genotype recommendations.
-
Assessed the efficiency of GGE and BLUP models for robust genotype selection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Sunflower (Helianthus annuus L.) is a herbaceous annual plant belonging to the family Asteraceae, grown for its seeds. It is a predominantly cross-pollinated diploid (2n = 2x = 34) species with a genome size of 3.6 GB [1]. It is the most important source of vegetable oil next to palm, soybean, and canola oil [2]. Global production of sunflower seed exceeded 54.2 million metric tons in 2022 [3]. Sunflower seed is widely used for edible oil production as a protein source for humans and animal, and as an ornamental plant in gardens. The residual cake after oil extraction is becoming a high protein meal for livestock [4]. Its oil is also recognized for its medicinal properties, aiding in infection recovery [5]. Sunflower seeds enhance human immunity with micronutrients like zinc, selenium and iron. Its oil is rich in tocopherols, sterols and antioxidants; reducing blood cholesterol [6]. Additionally, it is utilized in soap making, paint and varnishes [2]. Due to its ability to grow in diverse agro-ecological conditions, high oil quality compared to other major oilseeds and multiple advantage; sunflower has emerged as an oil seed crop of the future by enduring environmental changes on the global level. Even though, there is ample potential and growing interest, limited acreage and very low national productivity which is below yield potential is observed in Ethiopia. The lack of high-yielding and adapted cultivars in the target environment is a key challenge for the low productivity of sunflower in the country [7]. Sunflower breeding focused more on the development of stable and high-yielding varieties in target environments by confirming superiority and stability across contrasting environments. The ability of the genotype responses to environmental conditions is determined by the genetic makeup of individual [8, 9].
Every breeding program conducts multi-environmental trials to identify superior varieties suitable for specific environments. The significance of GEI has direct implications for breeders' decisions, influencing how genotypes perform across different environments [8]. Proper handling and analysis of multi-environmental data are crucial either to ignore, reduce, or exploit the interaction effect [9]. Identifying stable and high-yielding genotypes across environments is essential, as it allows for the exploitation of crossover GEI [8, 10]. A significant GEI, particularly for quantitative traits like sunflower seed yield, can substantially hinder efforts to select superior genotypes [9]. Breeding lines, including those for sunflowers, lack consistency in responding to environmental fluctuations, greatly influencing crop production and complicating breeding efforts. Therefore, various numerical and graphical methods are employed to identify high-yield and stable genotypes across numerous environments.
To draw valid conclusions regarding high seed yield, stable, and adaptable sunflower cultivars, proper statistical analysis is essential. Parametric stability measurements and non-parametric or qualitative statistical models have been broadly utilized and reported to determine the degree of GEI in sunflower. These approaches have been documented in previous experiments conducted by Luque [11], Grunvald [12], Grunvald [13] and Chakraborty [14]. Additionally, multivariate methods have been employed to assess sunflower seed yield stability. Baraki [15] reported the use of AMMI and BLUP models, while Cherinet [16], Ahmed [17], Ma’ali [18] and Ansarifard [19] utilized GGE models. The choice of GGE and BLUP methods over other stability statistics is based on their robustness, flexibility and ability to effectively handle complex GEI pattern [9, 20]. GGE analysis, in particular, stands out as a versatile and superior approach compared to other multivariate statistics due to its ability to provide graphical representation of GEI through biplot. This graphical representation allows interpretation of genotypes performance across different environments and facilitate the visualization of crossover interactions [20,21,22]. Such visual analysis aids breeders in the determination of stable and responsive genotypes. Moreover, the flexibility of the GGE model accommodates non-linear responses to environmental variation, making the method suitable for capturing more complex GEI patterns that may not be adequately addressed by traditional stability approaches [23, 24]. This flexibility allows for a more accurate assessment of genotype performance in diverse environments. Additionally, the method increases the precision of stability assessment by incorporating spatial information when field experiments exhibit spatial variability. GGE is efficient for mega-environment determination; for assessing cultivars mean and stability, and to identify discriminating and representative environments [23,24,25]. It is more successful than parametric and nonparametric univariate statistics in terms of GEI data analysis because it examines the complicated genotype effects and interactions using graphical method simultaneously. It also provides solution by combining mean seed yield and stability by transforming into criterion to evaluate graphically [23]. In multi-environmental trials trait like seed yield in sunflower is complex and affected by genetic, environmental and genetic by environment interactions which makes complicate the selection and identification of superior and stable genotypes; Therefore, statistical models like GGE is more powerful and essential to evaluate the genetic potential of genotypes and recommend for wide adaptation.
The genetic values of genotypes were evaluated using BLUP by accounting for both genetic and environmental variability. It provides more precise estimate of genotype performance compared to other traditional stability analysis approaches, which often focus solely on GEI [26]. The method leads to more reliable results because it uses sound statistical principles by ensuring unbiased estimation of genetic values and robust inference about genotype performance. Additionally, the use of mixed linear model in the analysis allows for the incorporation of genotype and environment random effects, enabling the simultaneous evaluation of genotype performance across multiple environments [9]. The study aims to examine the pattern of genotype by environment interaction (GEI) of sunflower seed yield using BLUP and GGE biplot analysis. The representativeness and discriminative power of the target locations are also highlighted and mega-environments identification were employed for effective genotype recommendation. Furthermore, the level of association between BLUP and GGE models in identifying stable and superior sunflower genotypes was determined.
2 Material and methods
2.1 Descriptions of testing sites and experimental design
The geographic positions of experimental sites were presented in Fig. 1. The experiment was carried out in six locations for two consecutive years (2017 and 2018) under rain-fed conditions. Detailed descriptions of the test locations including altitude, rainfall, and soil properties were provided in Table 1.
The experimental genotypes were derived from the national sunflower improvement program through population improvement using head to row selection scheme. The head to row selection was employed on cultivated sunflower genotypes with individual heads being carefully selected and evaluated under controlled situation and superior heads were further promoted to preliminary trial followed by multilocation yield trials. The evaluation criterias include agronomic traits such as shorter plant height, larger head size, earliness and uniformity; tolerance to the most common sunflower disease and insect pests; yield potential and yield related traits; and oil content. To identify superior and stable genotypes eight outstanding sunflower genotypes interms of desired characters were selected and promoted from early stage yield trial to multilocation yield trial and tested in representative sunflower growing areas of Ethiopia along with one locally adapted (Russian black) and one standard check (Oissa) varieties. The material utilized in the experiment along with their corresponding data analysis codes were outlined in the Table 2. The experiment was conducted in a randomized complete block design (RCBD) with three replications. Each plot consisted of six rows 4 m in length with 75 cm between rows and 25 cm within row. During planting, a seed rate of 10 kg and a fertilizer of 23 kg nitrogen and 23 kg P2O5 per hectare were used. Other agronomic practices such as thinning, hoeing, weeding, disease and insect pest controls were conducted as per the recommendations.
2.2 Data collection and statistical data analysis
The harvested four middle rows or plot yield was converted to yield per hectare in kilograms for data analysis. The data were subjected to combined analyses of variance (ANOVA) by utilizing linear mixed-effect model (LMM) to partition the total variation [27, 28]. We considered seed yield as response variable and genotype as fixed effect. Environment, replication with in environment and genotype-environment interaction (GEI) effects were treated as random effects to capture the inherent variability across growing conditions. The significance of random effects was assessed using likelihood ratio test (LRT) by fitting the model to maximum likelihood (ML). But the significance of fixed effect (genotypes) was determined using F-test. This comprehensive approach allowed determining the impact of variabilities for sunflower yield trial [27, 28]. To simplify the assessment, each year at each location was treated as a separate and distinct individual environment. The differences between means of the genotypes were compared by least significant difference (LSD) test. Before conducting stability analysis the significance of GEI was checked.
GGE biplot analysis was conducted to classify mega environments, interaction and yield stability of multi-environment trial data. The principal component 1 and 2 were used to construct the biplot using environment-centered yield data to singular value decomposition (SVD) [29] of the first two principal components to fit the GGE biplot model as proposed by Yan [20, 29].
where, Yij is the trait mean for genotype i in environment j; µ is the grand mean; βj is the main effect of environment j; µ + βj is the mean yield across all genotypes in environment j; λ1 and λ2 are the singular values (SV) for the first and second principal components (PC1 and PC2), respectively; ξi1 and ξi2 are eigenvectors of genotype i for PC1 and PC2, respectively; η1j and η2j are eigenvectors of environment j for PC1 and PC2; and εij is the residual associated with genotype i in environment j [20, 29].
Genotype-focused and environment-focused scaling was implemented to compare genotypes and environments, respectively. Symmetric scaling was also performed to visualize the which-won-where pattern of the sunflower multi-environment trial yield data [20, 29]. The GGE Distance (GGED) and GGE instability index (GGEIN) can be calculated by the GGE biplot model [30]. For simultaneous identification of high-yielding, stable, and adapted sunflower genotypes, the best linear unbiased prediction (BLUP) parameters such as harmonic mean of genotypic values (HMGV) to evaluate genotypes adaptability, the relative performance of genotypic values (RPGV) and the harmonic mean of RPGV (HMRPGV) stability parameter were evaluated as proposed by De Resende [31].
where, E is the number of environments; GVij is the genetic value of ith genotype in jth environment; where GVij = uj + gi + geij, µj is the mean seed yield in the jth environment, and gi is the BLUP value of ith genotype, and geij is the BLUP value of the interaction between ith genotype and jth environment. The data analysis was performed using R software version 4.3.2 lmer4 package [32] for combined ANOVA analysis and Metan package [33] for GGE and BLUP analysis.
3 Result and discussion
3.1 Analysis of variance
The analysis of variance (ANOVA) for genotype and environment revealed significant level of variability at p < 0.01 and p < 0.001 respectively. While the GEI effect was significant at p < 0.05 on seed yield performance of the tested sunflower genotypes (Table 3). The significance of GEI indicates that genotypes performance varies across environments, This illustrates that some genotypes may perform better or worse depending on specific environmental condition, which implies the importance of due attention for both genetic and environmental factors in sunflower breeding programs. The analysis of variance result delivered understanding into the source of variations [34], enhances the predictive accuracy of the GGE and BLUP analyses and guiding in identifying genotype recommendations and breeding decisions for specific or broad adaptability across environments [35]. The present finding was in agreement with the previous reports [16, 36, 37] which underscore the importance of environmental studies for sunflower variety selection.
3.2 Sunflower seed yield performance in different environments
The environment means in this study ranged from 1171 kgha−1 (E4/kulumsa2018) to 2564 kg ha−1 (E11/kulumsa2017), with a grand mean of 1805 kg ha−1 (Table 4). Only E3, E4, E5, E7, and E12 out of the examined environments displayed below-average mean values. The overall mean across 12 environments showed that, the highest mean yield was recorded by genotype 8 (G8), which had a yield of 2032.6 kg ha−1. G1 and G4 had mean yields of 2021.1 and 1930.4 kg ha−1 respectively. The local (G10) and standard check (G9) exhibited the lowest average seed yield (1615.6 kg ha−1 and 1636.9 kg ha−1 respectively than other genotypes. This finding underscores that they have limited yield potential and need to replace these varieties with better performing ones to enhance yield, adaptability and overall profitability of the producers. Chakraborty [14] emphasized the importance of selecting superior genotypes for stable performance across different environmental conditions. Reports in sunflower germplasms revealed that the significance of genetic resource in breeding programs with desirable traits showing high heritability and genetic advance indicating the potential of improving seed yield [38, 39]. This highlights the need to drive effort in breeding research to develop and promote sunflower varieties with enhanced seed yield potential for sustainable and profitable production in Ethiopia and beyond. From the present study, inconsistent seed yield ranks across environments were observed; for instance, the high-yielding genotype G8 performed best in the fifth environment (E5) but worst rank in three other environments. G1 ranked first in five environments but exhibited poor rank in environments E7 and E8. This shows the genotypic reaction of genotype performances was not consistent across the test locations. Therefore, this differential response pattern of genotypes yield ranking across environments revealed a cross-over interaction [40]. The study clearly shows that there is significant difference in seed yield between environments and years (Table 4); this may be because of a large geographic area with diverse weather patterns, as well as edaphic and other environmental factors that affect sunflower production. This means that a significant contribution from the environment and GEI is estimated. Ansarifard [19] and his colleagues conducted a study on GEI in sunflower genotypes using GGE biplot and they conclude similar finding with the present result, which confirm the instability of some genotypes across different environments. Similar findings on sunflower seed yield stability and environment interaction were reported [16, 41,42,43,44]. These disparate expressions of genotypes over the environments also known as genotype-environment interaction complicate the selection of sunflower genotype for seed yield under diverse environmental conditions. Hence, recognizing the better consistent genotypes across varied locations, quantifying the interaction effect based on statistical approaches, and taking advantage of positive interaction help to minimize the consequence. The change of genotype rank across environments justified the use of robust statistical methods to visualize and interpret the genotype by environment data which helps to identify stable and widely adapted genotypes, representative and mega environments identification. To analyze such parameters; GGE biplot analysis is a potential candidate and has become a popular data visualization for genotype by environment data [23]. Besides achieving high yield performance, genetic stability is crucial factor in enhancing sunflower production. The result of this study demonstrated the importance, impact and variation in performance between environments and GEI showing how climate change influences sunflower production year to year and location to location. The performance difference among environments and GEI has an implication for breeding program; requiring the selection of genotypes that are well suited and consistence across diverse environments [22]. This could be achieved through conducting trials in divers location over multiple years, utilize suitable stability analysis, and considering the concept of mega-environment identification [44, 45]. In addition the result underlines the importance of environmental characterization for sunflower production in Ethiopia and beyond. Comparable results were reported on sunflower multi-environments trials [13, 14, 16, 19].
3.3 Test environment evaluation
3.3.1 Environment discrimination and representativeness
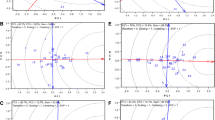
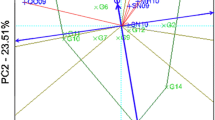
To evaluate the discriminating ability and representativeness of environments for sunflower seed yield (Fig. 2), we follow environment centered model Yan [20, 29] by setting the parameters as centering = 2, scaling = 0, and SVP = 2. The model allows capturing discriminative power of each environments and its representativeness across different genotypes. Principal component 1 and 2 explained 54.25 percent of the entire variance which showed a poor explanation of variability than the previous report [16, 44]. This reduced explanation of variation by the axis is the result of complex Genotype interaction with environment [45]. Similar variations were reported for other crops [46]. Figure 2 explained the discriminating power and representativeness of the test environments. The environment vectors length shows the capability of the environment’s to differentiate the genotypes and the length of the vectors visualized by the concentric circles; the longer victor showed the more discriminating power [20, 21]. Therefore, the biplot validates that E8, E11, and E2 are the most discriminatory environments respectively because of their long vector. But E5, E7, E1, and E4 have very short vectors with very low discriminating power; they provide little or no information about the evaluated sunflower genotypes compared to those with longer vectors. The biplot result supports the seed yield value presented in Table 4 for E5, E7, E1, and E4 in order; that most of the genotypes exhibited below the average mean value than other environments. Therefore, these environments should not be used as test locations for sunflower yield trials. Another interesting feature within the biplot is the concentric circles located on the abscissa representing the average-environment axis (AEA) which helps to visualize the magnitude of environment vectors extending from the origin. Furthermore, the measure of the angle between two vectors following the AEA helps to inspect the relationship between environments and ideal environment [45]. The biplot analysis (Fig. 2) identified E11 followed by E2 and E6 as the most suitable environments interms of representing target environments (representativeness) as well as exhibiting genotypic difference (discriminating ability) for selecting superior sunflower genotypes. This is because they form small (acute) angles between AEA and their victors [21]. Environments with long vectors but large angles with AEA such as E8 are regarded as lower representatives than small-angle environments. They are not considered as ideal in selecting superior sunflower genotypes but are useful in discarding unstable genotypes. Moreover, E5 and E7; E1 and E4; and E12 and E10 are regarded as redundant environments for each other which produced similar information about the genotypes and can be dropped in future sunflower evaluations. Target and discriminate environment identification using GGE biplot was reported in other crops including sunflower [36, 47, 48].
3.3.2 Ranking and ideal environment identification
In order to illustrate the distance between each testing environment from the most favorable environment the concentric circles were employed as best visualizing tools in GGE biplot. In the biplot a more desirable environment is located closer to the favorable environment or close to the center [20]. Hence, the study identified E11 as the optimal environment and E10, E9, and E12 environments considered as desirable environments because they are close to favorable environment E11. However, E8 followed by E5 and E7 are considered as the worst environments since they are very far from the center (Fig. 3).
3.3.3 Mega environment identification
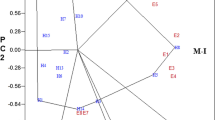
The Which-won-where and mega environment identification analysis helps to elucidated the presence of crossovers, separate mega environment and identify genotypes adapted to specific locations [20, 29]. Five sunflower genotypes namely G1, G8, G9, G5, and G7 were helped to draw the shape as a vertices and placing the remaining genotypes within the polygon (Fig. 4). This implies that the vertex genotypes are positioned distant from the biplot origin and they are outstanding in those specific environments, either positively or negatively [20]. Genotypes G4 and G6 were located proximate to the center; hence they were not responsive to environments and would rank the same or show stable performance in all test locations. The vertical line originating from the ballot’s origin partition into five sectors (Fig. 4). The vertex genotype is the highest yielder in all environments falling in the sector [49]. Therefore, G1 was superior genotype in E6, E9 and E10; G12 and G5 are winning in E5 and E7; G7 was paramount genotype in E3 and G8 is the highest yielder in environments E1, E2, E4, E8, E11, and E12. In addition G8 and G1 have similar performance in E12. However, no environments belonged to genotype G9. Even though further investigations will be required, the causes for lack of environmental representation for G9 might be triggered by the genotype performance limited to very specific environments, its unique performance characteristics or the limitations in the environmental representation of the study [20, 25]. Similar result in sunflower genotypes was previously reported by Ansarifard [19]. Moreover, the biplot revealed the presence of interaction effect and the existence of mega-environments. The crossover interaction in the biplot indicates that different genotypes excel in different environments, identified mega-environments and grouping similar test environments. Reports on various oil crops like sunflower [16, 19]; sesame [50]; canola [51]; Cotton [52] and groundnuts [53] utilized biplot analysis in delineating stable genotypes for specific environments and categorize environments into distinct mega-environments and sub-mega environments.
The study identifies four mega-environments (ME) and one sub-mega-environment (SME). The first mega environment (ME1) includes E5 and E7. This group consists of environments with moderate altitudes and varying rainfall. The Aridisols in E5 categorized as typically dry and nutrient poor [54]; Whereas E7 soil type is more fertile and better moisture retention [55]. The exclusive presence of G5 in this environment suggests its strong adaptability to these contrasting soil conditions. Breeding should focus for drought tolerance and efficient nutrient uptake in G5 to maintain productivity across these varied environments. The second group ME2 contains E3; ME2 characterized by high altitude and moderate rainfall with fertile soil type [56]. The presence of multiple genotypes in this environment indicates their broader adaptability to these conditions. Breeding of ME2 should focus on traits performance in the given altitude to ensure genotypes competitive yields under these conditions. The third ME3 consisted of E6, E9 and E10; it is described by moderate to high altitudes and varying rainfall, with fertile black soil [55] and less fertile Aridisols [56]. The presence of G1 indicates strong specific adaptation to both fertile moisture retained soils and environment with moderate water availability. Breeding efforts for this environment focus on maintaining the traits that offer high performance with particular focus traits for soil fertility and moisture retention. The fourth mega environment ME4 contains E1, E2, E4, E8, and E11; but it was further subdivided into two sub-mega-environments (SME). Closer similarity between E1 and E4 was observed than the other three environments because of their proximity to the origin and smaller angle between them. This revealed their less discriminating and representing power of the target environment as explained by Yan [21] and Weikai [49]. Therefore, they are considered as separate sub-mega-environment. G4 and G6 are best-performed genotypes in environments E1 and E4. Besides, E2, E8, and E11 had long vectors and acute angles; as explained by Yan [21] and Weikai [49] this environments are more discriminating and more descriptive of the target region. Therefore, they clustered as a separate subgroup into sub-mega-environment. Conversely, E12 is found in between G1 and G8 which shows that it has similar genotypic responses to both sectors, and it is not very discriminating or representative of either sector. Therefore, E12 is a common environment that helps to compare the performance of genotypes across the two mega-environments, which is useful as a reference environment [21]. ME4 was characterized by high altitude with diverse rainfall and soil types. The soil types in this location offer fertile conditions but differ in water retention capacity [54,55,56,57]. Breeding in this group should focus in enhancing versatile traits in G4 and G6 for stability and focus on traits that optimize high performance in specific conditions for G8. The absence of specific environments for G9 and G10 suggest that they may have broader adaptability preference not captured within the current testing environments. Therefore, further testing in additional locations is important to determine their optimal growing conditions. The present finding underscores the importance of selecting genotypes for specific environments and highlights the need for targeted breeding programs to optimize performance.
3.4 Genotype evaluation using GGE biplot
3.4.1 Mean and stability of genotypes
The visualization of the mean and stability of 10 sunflower genotypes was presented in Fig. 5. Ideal genotypes were identified by applying genotype centered singular value partitioning (SVP) and drawing an AEC in the biplot. As a criterion superior yield and relatively stable performance across the tested locations were considered as ideal as stated by Yan [21, 29]. The biplot analysis indicates that G8 was a high mean yielder followed by genotypes G6 and G1. In the mean verse stability analysis, the average environment coordinates (AEC) abscissa and the AEC ordinate represented by x and y-axis’s respectively in the biplot. The arrow on the AEC abscissa axis points towards the superior genotypes, allowing ranking of genotypes based on their mean performance [20, 21]. Whereas, the average environment is defined by PC1 and PC2 and represented by an arrow examines the mean performance and the consistency of the genotype in any direction of the abscissa [49]. As general rule genotypes with the shortest projection with the abscissa revealed better stability but the longest projection is seen to indicate more fluctuation or instability [20]. Therefore, G6 is the more consistence and good yielder genotype, followed by G1. However, G8 and G9 showed a long projection which implies greater instability than the tested genotypes. Genotype G4 was located near the abscissa and short projection onto the AEC ordinate. This indicates that its rank was highly consistent across environments within the mega-environment. Therefore, G6, G1, and G4 respectively are the desired sunflower genotypes in terms of stability and mean seed yield.
3.4.2 Evaluation of genotypes relative to ideal genotypes
Ideal genotypes are those located in close proximity to the center of the concentric circle. Hence, the biplot in Fig. 6 reflected that the proximity of G6 to the center, making it the most ideal and desirable genotype. Other genotypes such as G1, followed by G8 and G4, which are located near G6 are considered as desirable genotypes. However, genotypes G10 and G9 are the worst, as they are located distant from the center. In any breeding program, high yielder and more stable across all environments may be considered as ideal genotypes [30] and assessing other genotypes in relation to an ideal genotype assists in evaluating the respective roles of stability and seed yield in identifying the best varieties[58].
The GGED and GGEIN indexes, obtained through GGE biplot analysis are essential measures for selecting adapted and stable genotypes by estimating genotypic response to environmental variations. It ultimately improves breeding efficiency to enhance genotype selection and improve productivity [21, 49, 58]. Based on the GGED value, genotypes G3, G4, and G7 resulted in lower values whereas, genotypes G2, G8, and G9 exhibit the highest value (Table 5). The greater the GGED distance indicates the genotypes are more distinct in terms of their performance across environments [21]. Genotypes with smaller values are considered more stable across environments [49]. Therefore, genotype G3 and G4 perform consistently under varying environmental conditions and are considered as more stable and adaptive than other genotypes. However, genotype G8 and G9 are considered as less stable and more responsive to specific environmental conditions. This indicated that they respond differently to environmental variations and there is a clear distinction in their adaptability across environments, and they may recommend for specific adaptations [30]. GGEIN helps to understand adaptability and genotypes interaction with environments. Its values ranged from 0 to 1 and if the GGEIN value closer to 1 the genotype become more unstable across diverse environments [21, 30, 49, 58]. Hence, genotypes G1, G3, and G7 showed better adaptability and constancy across the tested environments. In contrast, genotypes G5, G9, and G10 are considered as less adapted and unstable genotypes. GGED and GGEIN estimates are valuable for sunflower breeders in selecting genotypes that exhibit wide adaptation or specific adaptation to certain environments. Based on the two parameters, G3 and G7 are identified interms of adaptability and constancy which are recommended for wider production across environments because of their low yield performance fluctuation under diverse environments. GGED and GGEIN indexes are important approaches in target breeding programs in selecting targeted traits either for specific or wide adaptability. We confirmed that the GGEIN value identify high performing stable sunflower genotypes like G1 and G3 for broad adaptation while environment specific genotypes G5 and G9 selected with GGED. This result was somehow similar with biplot analysis displayed above in identifying G9 as specific adapted genotype. The use of both indexes provides robust framework for evaluating the performance and stability of genotypes [59, 60] to make informed decisions to enhance crop productivity and adaptability across diverse environments [21, 45]. The importance the two parameters in identification of adapted cotton genotypes were reported by Baxevanos [30].
3.5 Best linear unbiased prediction based stability indices
The BLUP-based scores for mean yield ranking and the predicted mean values of sunflower genotypes for seed yield were presented in (Table 6 and Fig. 7). Based on the result G1, G8, and G4 were recognized as highly stable and high-yielding genotypes, but G9 and G10 were unstable and low-yielding genotypes (Table 6). The BLUP stability parameters under the study were almost the same with sunflower seed yield genotype ranking, this suggests that the genotypes had similar performance across those parameters indicating the BLUP models has strong association with seed yield. Comparable findings were reported in other crops such as soybean [61]; Jatropha [62]; finger millet [63] and lentil [64]. Even though all the BLUP parameters are designed to detect difference between genotypes in multi-environmental experiments, they have their own unique characteristics. As highlighted by Piepho [26] and Crossa [65]; the mean seed yield and stability of genotypes were considered simultaneously to estimate the harmonic mean of genotypic values (HMGV) which aims to identify stable genotypes. The harmonic mean of RPGV (HMRPGV) estimates stability, adaptability, and mean performances simultaneously, while the relative performance of genotypic values (RPGV) is considered as adaptability index. Figure 7 showed the higher mean values represented by blue and the lower with red circles. Among the analyzed genotypes five revealed a higher value than the predicted mean of which G8, G1, and G4 showing the highest predicted mean value compared to other sunflower genotypes. The BLUP estimates fill the gaps of other stability measures like GGE by predicting the average performance of genotypes in a mixed model with high efficiency and accuracy [66]. The GEI study based on the BLUP model was reported for other crops like canola [67] soybean [57] and potato [68].
3.6 Relation between stability statistics
The study provides excellent understanding of using the biplot and BLUP statistical methods in sunflower multi-environment trial. Both methods identified G1, G4, G6, and G8 as desirable genotypes based on their positive interaction across diverse environments and seed yield performance. However, the result of GGE stability indices varied from the biplot and the BLUP results in selecting stable and high yielder sunflower genotypes. This may be because of the indexes requirement of a big data set interms number of experimental years, testing environments and number of evaluated genotypes for accurate estimation of genotype-by-environment interactions and obtain reliable conclusions [21, 65]. The result for identifying superior sunflower genotypes were consistence between the two approaches, though they have their own pros and cons both methods complement each other in future sunflower research. The BLUP method exploit genotypes performance by interpreting the genotypic values but not group or provide information regarding testing environments but GGE biplot did that. Therefore, for any GEI analysis, simultaneous consideration of both techniques helps to select more stable and adaptable sunflower genotypes across wider environments will increases analysis accuracy and decision-making. To exploit the integration of GGE and BLUP models in stability analysis results were reported for rapeseed [69], cowpea [70], and lentil [64].
4 Conclusions
The study aimed to assess the representativeness and discriminative power of the test environments, group mega environments for genotype recommendations and evaluate how the combination of GGE and BLUP methods identify stable and superior sunflower genotypes. Our findings revealed significant variability in the representativeness and discriminative power of the tested environments as highlighted by ANOVA. This variability emphasizes the importance of environmental studies and enhances predictive accuracy of GGE and BLUP analyses. Notably the principal component 1 and 2 analyzed by SVD explained 54.25 percent of the variation for sunflower seed yield, providing insight into the underlying factors influencing yield variations. Genotype G8 recorded the highest seed yield while the checks varieties G9 and G10 consistently showed the lowest yield and instability by most of the tested parameters underlining cultivar replacement. The study confirmed that genotype performance was inconsistence across test environments, necessitates the use of robust statistical methods to recognize superior and constant sunflower genotypes. Environment E11 appeared as the most representativeness and discriminative environments for selecting stable and superior genotypes. Four mega-environments were identified for effective genotype recommendations. Genotype G6 identified as the most stable and ideal genotype, while G1, G4 and G8 showing high mean values and better stability, offering breeders promising candidates for specific and wide adaptation.
Except GGE stability indices, GGE biplot and BLUP methods consistently identify stable and ideal sunflower genotypes. This result underscore their reliability in breeding programs enables breeders to confidently select genotypes with superior performance across diverse environmental conditions which enhance the efficiency of sunflower cultivation and breeding efforts. Understanding the GEI pattern and employing recommended mega environments are significant for informed variety selection, contributing in recognizing outstanding and stable sunflower varieties. Future research should validate identified mega-environments over multiple years and explore the genetics of adaptability and stability in sunflower genotypes. Expanding testing environments to include various conditions and incorporating genomic tools can enhance our understanding of sunflower stability and develop more effective breeding strategies. In summary this study contributes to advancing sunflower production by elucidating GEI patterns, identifying mega-environments, and facilitating genotype selection, ultimately enhancing productivity in sunflower cultivation and potentially other crops. To conclude, this study advances sunflower production by clarifying GEI patterns, identifying mega-environments, and facilitating genotype selection, ultimately enhancing productivity in sunflower cultivation and potentially others.
Data availability
The data used in this study are not publicly available due to confidentiality. However, the data used in this study are available upon request from the corresponding author.
References
Badouin H, Gouzy J, Grassa CJ, Murat F, Staton SE, Cottret L, et al. The sunflower genome provides insights into oil metabolism, flowering and Asterid evolution. Nature. 2017;546:148–52. https://doi.org/10.1038/nature22380.
Rauf S, Jamil N, Ali Tariq S, Khan M, Kausar M. Progress in modification of sunflower oil to expand its industrial value. J Sci Food Agric. 2017;97:1997–2006. https://doi.org/10.1002/jsfa.8214.
Food and Agriculture Organization of the United Nations. FAOSTAT statistical database; 2022. Retrieved July 17, 2024, from http://www.fao.org/faostat/en/#home
Pal, D. Sunflower (Helianthus annuus L.) seeds in health and nutrition. In: Nuts and seeds in health and disease prevention. London: Academic Press, 2011;1097–1105.
Jayawardena R, Sooriyaarachchi P, Chourdakis M, Jeewandara C, Ranasinghe P. Enhancing immunity in viral infections with special emphasis on COVID-19: a review. diabetes and metabolic syndrome. Clin Res Rev. 2020;14:367–82.
Rauf, S. Breeding strategies for sunflower (Helianthus annuus L.) genetic improvement. In: Advances in plant breeding strategies: industrial and food crops. Springer, Cham, 2019; 637–673.
Misteru Tesfaye and Birhanu Mengistu. Sunflower research: current status and future prospects in Ethiopia. Int J Agric Innov Res. 2021;Volume 9, Issue 5, ISSN (Online) 2319–1473.
Leon ND, Jannink J-L. Edwards, J.W.; Kaeppler, S.M. Introduction to a special issue on genotype by environment interaction. Crop Sci. 2016;56:2081–2089.
Bernardo, R. Breeding for quantitative traits in plants. Stemma Press: Woodbury, MN, USA;2020.
Saltz JB, Bell AM, Flint J, Gomulkiewicz R, Hughes KA, Keagy J. Why does the magnitude of genotype-by-environment interaction vary? Ecol Evol. 2018;8:6342–53.
Luquez JE, Aguirrezabal LAN, Agüero ME, Pereyra VR. Stability and adaptability of cultivars in non‐balanced yield trials. comparison of methods for selecting ‘high oleic’sunflower hybrids for grain yield and quality. J Agron Crop Sci 2002;188(4):225–34. https://doi.org/10.1046/j.1439-037X.2002.00562.x
Grunvald AK, Carvalho CGPD, Oliveira ACBD, Andrade CADB. Adaptability and stability of sunflower genotypes in Central Brazil. Pesquisa Agropecuaria Brasileira. 2008;43:1483–93.
Grunvald AK, Carvalho CGP, Amabile RF, Godinho V, de Oliveira ACB, Ramos NP. Adaptability and stability of conventional and high oleic sunflower genotypes cultivated in Central Brazil. Genom. Quant. Genet. 2014;8:7–15. ISSN 2157-9903.
Chakraborty NR, Lakshman SS, Debnath S, Rahimi M. Yield stability and economic heterosis analysis in newly bred sunflower hybrids throughout diverse agro-ecological zones. BMC Plant Biol. 2022;22(1):579. https://doi.org/10.1186/s12870-022-03983-1.
Baraki F, Gebregergis Z, Belay Y, Teame G, Gebremedhin Z, Berhe M, Gebregergs G. Identification of adaptable sunflower (Helianthus annuus L.) genotypes using yield performance and multiple-traits index. Heliyon, 2023; https://doi.org/10.1016/j.heliyon.2024.e29405.
Alem C, Worku A, Mekonnen M, Asres T, Fentie D, Mihiretu E, Esmael J. GGE stability analysis of seed yield in sunflower genotypes (Helianthus annuus L.) in Western Amhara Region, Ethiopia. Int J Plant Breed Genet. 2016;10:104–9. https://doi.org/10.3923/ijpbg.2016.104.109.
Ahmed MA, Abdelsatar MA, Attia MA, Abeer AA. GGE biplot analysis of Line by tester for seed yield and its attributes in sunflower. RUDN J. Agron. Anim. Ind. 2019;14(4):374–89. https://doi.org/10.22363/2312-797X-2019-14-4-374-389
Ma’ali S, Makgoga W, Erasmus J, Swanepoel S. Genotype-by-environment interaction and yield stability of sunflower hybrids across production environments in South Africa. S Afr J Plant Soil .36(4):271–8. https://doi.org/10.1080/02571862.2018.1558461.
Ansarifard I, Mostafavi K, Khosroshahli M, Reza Bihamta M, Ramshini H. A study on genotype–environment interaction based on GGE biplot graphical method in sunflower genotypes (Helianthus annuus L.). Food Sci. Nutr. 2020;8(7), 3327–34. https://doi.org/10.1002/fsn3.1610
Yan W, Hunt AL, Sheng Q, Szlavnics Z. Cultivar evaluation and mega-environment investigation based on GGE-biplot. Crop Sci. 2000;40:596–605. https://doi.org/10.2135/cropsci2000.403597x.
Yan W, Kang MS. GGE biplot analysis: a graphical tool for breeders, geneticists, and agronomists (1st ed.) CRC Press. 2003;https://doi.org/10.1201/9781420040371
Yan W, Kang MS, Ma B, Woods S, Cornelius PL. GGE biplot vs. AMMI analysis of genotype-by-environment data. Crop Sci. 2007;47:643–53. https://doi.org/10.2135/cropsci2006.06.0374.
Badu-Apraku B, Fakorede B, Akinwale R, Annor B, Adewale S, Toyinbo J, et al. Application of the GGE biplot as a statistical tool in the breeding and testing of early and extra-early maturing maize in Sub-Saharan Africa. Crop Breed Genet Genome. 2020;2(3):e200012. https://doi.org/10.20900/cbgg20200012
Angelini J, Faviere GS, Bortolotto EB, Arroyo L, Valentini GH, Domingo Lucio Cervigni G. Biplot pattern interaction analysis and statistical test for crossover and non-crossover genotype-by-environment interaction in peach. Sci Hortic. 2019;252:298–309. https://doi.org/10.1016/j.scienta.2019.03.024
Ding M, Tier B, Yan W. Application of GGE biplot analysis to evaluate genotypes (G), environments (E), and G × E interaction on P. radiate: A case study. In: Proceedings of the Australasian Forestry Genetic Conference Breeding for Wood Quality. Apr 11–14; Hobart, Australia; 2007.
Piepho HP, Möhring J, Melchinger AE, Büchse A. BLUP for phenotypic selection in plant breeding and variety testing. Euphytica. 2008;161:209–28. https://doi.org/10.1007/s10681-007-9449-8.
Crossa J, Gauch HG Jr, Zobel RW. Additive main effects and multiplicative interaction analysis of two international maize cultivar trials. Crop Sci. 1990;30(3):493–500.
West BT, Welch KB, Galecki AT. Linear mixed models: a practical guide using statistical software. Chapman and Hall/CRC;2022.
Yan W. Singular-value partitioning in biplot analysis of multienvironment trial data. Agron J. 2002;94:990–6. https://doi.org/10.2134/agronj2002.9900.
Baxevanos D, Goulas C, Tzortzios S, et al. Interrelationship among and repeatability of seven stability indices estimated from commercial cotton (Gossypium hirsutum L.) variety evaluation trials in three Mediterranean countries. Euphytica. 2008;161:371–82. https://doi.org/10.1007/s10681-007-9586-0.
De Resende MDV. Métodos Estatísticos Ótimos na Análise de Experimentos de Campo. Ostra: INFOTECA-E. Colombo: Embrapa Florestas Documentos; 2004.
Bates, D. lme4: Linear mixed‐effects models using Eigen and S4. R package version, 1, 1; 2016.
Olivoto T, Lúcio ADC. metan: An R package for multi-environment trial analysis. Methods Ecol Evol. 2020;11(6):783–9. https://doi.org/10.1111/2041-210X.13384.
Melo RCD, Coimbra JLM, Trevisani N, Guidolin AF, Kavalco SAF. Consideration of the appropriate variation sources of the statistical model and their impacts on plant breeding. Acta Sci Agron. 2018;41(1):39791. https://doi.org/10.4025/actasciagron.v41i0.39791.
Akter A, Hasan MJ, Kulsum U, Rahman MH, Khatun M, Islam MR. GGE biplot analysis for yield stability in multi-environment trials of promising hybrid rice (Oryza sativa L.). Bangladesh Rice J. 2015;19(1):1–8.
Pourdad SS, Moghaddam MJ. Study on seed yield stability of sunflower inbred lines through GGE biplot. Helia. 2013;36(58):19–28. https://doi.org/10.2298/HEL1358019P.
Dos Santos IG, Carneiro VQ, Sant’anna IDC, Cruz CD, Carvalho C, Borba Filho AL, Alves AD. Factor analysis and GGE biplot for environmental and genotypic evaluation in sunflower trials. Funct Plant Breed J. 2019;1(2):555. https://doi.org/10.35418/2526-4117/v1n2a3.
Umarani E, Saritha A, Ramanjaneyulu AV. Estimation of genetic parameters in sunflower (Helianthus annuus L.) germplasm lines for yield and its contributing traits. Int J Environ Clim Change. 2022;12(12):936–42.
Abu M. Evaluation of sunflower genotypes using principal component analysis. Int J Genet Genom. 2022;10(1):32–6. https://doi.org/10.11648/j.ijgg.20221001.15.
Yan W, Cornelius PL, Crossa J, Hunt LA. Two types of GGE biplots for analyzing multi-environment trial data. Crop Sci. 2001;41:656–63. https://doi.org/10.2135/cropsci2001.413656x.
Mrdja J, Crnobarac J, Radić V, Miklič V. Sunflower seed quality and yield in relation to environmental conditions of production region. Helia. 2012;35(57):123–34. https://doi.org/10.2298/hel1257123m.
Khomari A, Mostafavi K, Mohammadi A. Grain yield stability analysis and genotype-environment interaction study in sunflower (Helianthus annuus L.) cultivars; 2016.
Radić V, Balalić I, Miladinov Z, Ćirić M, Vasiljević M, Jocić S, Marjanović-Jeromela A. Genotype × environment interaction of some traits in sunflower (Helianthus annuus L.) lines. Appl Ecol Environ Res. 2020;18(1):1707–19. https://doi.org/10.15666/aeer/1801_17071719.
Shah MH, Rauf S, Nazir S, Ortiz R, Naveed A, Fatima S. Stability analyses of sunflower (Helianthus annuus L.) hybrids for oleic acid and yield traits under multi location trials in Pakistan, Italian. J Agron. 2023;18(1):55. https://doi.org/10.4081/ija.2023.2079.
Yan W, Tinker NA. An integrated biplot analysis system for displaying, interpreting and exploring genotype × environment interaction. Crop Sci. 2005;45:1004–16. https://doi.org/10.2135/cropsci2004.0076.
Ghaffar M, Asghar MJ, Shahid M, et al. Estimation of G × E Interaction of lentil genotypes for yield using AMMI and GGE biplot in Pakistan. J Soil Sci Plant Nutr. 2023;23:2316–30. https://doi.org/10.1007/s42729-023-01182-x.
Brankovic GR, Balalic IM, Zoric MZ, Miklic VJ, Jocic SB, Surlan-momirovic GG. Characterization of sunflower testing environments in Serbia. Turk J Agric Forestry. 2012; 36(3):2. https://doi.org/10.3906/tar-1106-45
Mousavi SMN, Hejazi P, Khalkhali SKZ. Study on stability of grain yield sunflower cultivars by AMMI and GGE biplot in Iran. Mol Plant Breed. 2016;7(2):1–6. https://doi.org/10.5376/mpb.2016.07.0002.
Yan W, Tinker NA. Biplot analysis of multi-environment trial data: principles and applications. Can J Plant Sci. 2006;86(3):623–45. https://doi.org/10.4141/P05-169.
Daba C, Ayana A, Zeleke H, Wakjira A. Genotype x environment interactions for seed yield in Sesame in western Ethiopia. East Afr J Sci. 2015;9(2):85–96.
Shojaei SH, Mostafavi K, Ghasemi SH, Bihamta MR, Illés Á, Bojtor C, Mousavi SMN. Sustainability on different canola (Brassica napus L.) cultivars by GGE biplot graphical technique in multi-environment. Sustainability. 2023;15(11):8945.
Yehia W. Investigate stability and genotypes X environments association in Egyptian cotton genotypes under normal and drought condition utilizing GGE-biplot model. Asian J Adv Agric Res. 2022;20(2):18–31. https://doi.org/10.9734/AJAAR/2022/v20i2393.
Olanrewaju OS, Oyatomi O, Babalola OO, Abberton M. GGE biplot analysis of genotype × environment interaction and yield stability in bambara groundnut. Agronomy. 2021;11(9):1839.
United States Department of Agriculture. Soil taxonomy: a basic system of soil classification for making and interpreting soil surveys (2nd ed.). Natural Resources Conservation Service. 1999; Available online: https://www.nrcs.usda.gov/Internet/FSE_DOCUMENTS/nrcs142p2_051232.pdf
Saraswat VK, Dadhwal VK (Eds.) (2003). Vertisols of tropical and temperate regions of Asia. Science Publishers, Inc
IUSS Working Group WRB. "World Reference Base for Soil Resources 2014, update 2015." International soil classification system for naming soils and creating legends for soil maps. World Soil Resources Reports No. 106. Food and Agriculture Organization of the United Nations, Rome. 2015. Available online: https://www.researchgate.net/publication/285828913_World_Reference_Base_for_Soil_Resources_2014_update_2015
FAO. Guidelines for soil description (4th ed.). Food and Agriculture Organization of the United Nations. 2006; Available online: http://www.fao.org/3/y5263e/y5263e.pdf
Sousa MBE, Damasceno-Silva KJ, Rocha MDM, De Menezes N, Júnior JÂ, Lima LRL. Genotype by environment interaction in cowpea lines using GGE biplot method. Rev Caatinga. 2018;31:64–71.
Pour-Aboughadareh A, Barati A, Koohkan SA, Jabari M, Marzoghian A, Gholipoor A, Kheirgo M. Dissection of genotype-by-environment interaction and yield stability analysis in barley using AMMI model and stability statistics. Bull Natl Res Centre. 2022;46(1):19.
Pour-Aboughadareh A, Khalili M, Poczai P, Olivoto T. Stability indices to deciphering the genotype-by-environment interaction (GEI) effect: an applicable review for use in plant breeding programs. Plants. 2022;11(3):414.
Freiria GH, Gonçalves LSA, Furlan FF, Fonseca Junior NDS, Lima WF, Prete CEC. Statistical methods to study adaptability and stability in breeding lines of food-type soybeans. Bragantia. 2018;77:253–64.
Alves RS, Peixoto LA, Teodoro PE, Silva LA, Rodrigues EV, Resende MDV, et al. Selection of Jatropha curcas families based on temporal stability and adaptability of genetic values. Ind Crops Prod. 2018;119:290–3. https://doi.org/10.1016/j.indcrop.2018.04.029.
Anuradha N, Patro TSSK, Singamsetti A, Sandhya Rani Y, Triveni U, Nirmala Kumari A, Govanakoppa N, Lakshmi Pathy T, Tonapi VA. Comparative study of AMMI- and BLUP-based simultaneous selection for grain yield and stability of finger millet [Eleusine coracana (L.) Gaertn.] genotypes. Front Plant Sci. 2022;6(12):786839. https://doi.org/10.3389/fpls.2021.786839.
Hossain MA, Sarker U, Azam MG, Kobir MS, Roychowdhury R, Ercisli S, Ali D, Oba S, Golokhvast KS. Integrating BLUP, AMMI, and GGE models to explore GE interactions for adaptability and stability of winter lentils (Lens culinarisedik.). Plants. 2023;12(11):2079. https://doi.org/10.3390/plants12112079.
Crossa J, Pérez-Rodríguez P, Cuevas J, Montesinos-López O, Jarquín D, De Los Campos G, Varshney RK. Genomic selection in plant breeding: methods, models, and perspectives. Trends Plant Sci. 2017;22(11):961–75. https://doi.org/10.1016/j.tplants.2017.08.011.
Nardino M, Baretta D, Carvalho IR, Olivoto T, Follmann DN, Szareski VJ, Ferrari M, de Pelegrin AJ, Konflanz VA, de Souza VQ. Restricted maximum likelihood/best linear unbiased prediction (REML/BLUP) for analyzing the agronomic performance of corn. Afr J Agric Res. 2016;11(48):4864–72.
Zhang H, Berger JD, Herrmann C. Yield stability and adaptability of canola (Brassica napus L.) in multiple environment trials. Euphytica. 2017;213:155. https://doi.org/10.1007/s10681-017-1948-7.
Sood S, Bhardwaj V, Kumar V, Gupta VK. BLUP and stability analysis of multi-environment trials of potato varieties in sub-tropical Indian conditions. Heliyon. 2020. https://doi.org/10.1016/j.heliyon.2020.e05525.
Hu X. A comprehensive comparison between ANOVA and BLUP to valuate location-specific genotype effects for rape cultivar trials with random locations. Field Crop Res. 2015;179:144–9. https://doi.org/10.1016/j.fcr.2015.04.023.
Santos AD, Ceccon G, Teodoro PE, Correa AM, Alvarez RDCF, Silva JFD, Alves VB. Adaptability and stability of erect cowpea genotypes via REML/BLUP and GGE Biplot. Bragantia. 2016;75:299–306.
Acknowledgements
The authors thank Ethiopian Institute of Agricultural Research for financial support and the High and Midland Oilseeds Research Program staff members and colaborative research centers for their tremendous effort in trial management and data collection.
Plant guidelines
The plant material utilized in study was obtained from cultivated source, and all procedures were conducted in strict compliance with regulations and guidelines governing the cultivation of sunflower. No plant material was collected from wild populations.
Funding
This work was supported by Ethiopian Institute of Agricultural Research, crop research directorate with grant number EIAR-2017-18.
Author information
Authors and Affiliations
Contributions
All authors contribute to the experiment by developing proposals, trial management and data collection. Statistical data analysis and first draft preparation was taken part by the corresponding author Birhanu Mengistu. The co-author Mohammed Abu commented on the draft of the manuscript and provide feedback on the reviewers comment.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Aboye, B.M., Edo, M. Exploring genotype by environment interaction in sunflower using genotype plus genotype by environment interaction (GGE) and best linear unbiased prediction (BLUP) approaches. Discov Appl Sci 6, 431 (2024). https://doi.org/10.1007/s42452-024-06136-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42452-024-06136-1