Abstract
It has been widely reported that SERS can detect foodborne pathogenic bacteria, but SERS has only been limited to the detection of purified single bacteria. However, there’s few single bacterial infections in the natural state were found at the most time, and rare comparative investigations of different Gram-negative and Gram-positive bacteria by SERS exist. At the present study, we are aiming at the comparative analysis of the detection efficiency toward different foodborne pathogenic microbes by a conventional citrate-reduced gold colloid, including Escherichia coli, Salmonella enteritidis, Staphylococcus aureus and Listeria monocytogenes. The SERS spectra of samples were collected with sterile 0.5% NaCl solution as aggregating compound. First, the stability of each bacterial were detected with gold nanoparticles. Afterward, SERS signals of the four bacteria were compared at different concentrations, and it was found that Gram-positive and Gram-negative bacteria have different minimum sensitivity and detection stability. Next, the mixed bacterial SERS detection was applied to classify and identify the similar Raman spectra. Spectral data were analyzed using multivariate data analysis methods, including principal component analysis (PCA) and hierarchical cluster analysis (HCA). The SERS procedure we proposed, which utilized gold nanoparticles adsorbing on bacteria and multivariate analysis, allowed differentiation of species. The results of this pilot study demonstrate the potential of SERS to rapidly identify Gram-negative and Gram-positive foodborne pathogens.
Article Highlights
-
1.
Comparative analysis of the detection sensitivity toward different foodborne pathogenic microbes.
-
2.
SERS signal might be interrupted by the different structure of bacterial wall.
-
3.
The mixed bacterial liquid could also be effectively distinguished by SERS detection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Foodborne diseases caused by foodborne pathogenic microorganisms have become a global public health problem, unsafe food causes a vicious circle of disease and malnutrition, especially affecting infants, the elderly and patients. In recent years, the incidence rate of foodborne diseases has shown a rising trend worldwide. Foodborne diseases account for approximately 600 million cases worldwide each year, according to the World Health Organization (WHO), and 420,000 people die as a result [1]. Food safety problems caused by the contamination of common foodborne pathogenic bacteria such as Escherichia coli, Salmonella enteritidis, Staphylococcus aureus and Listeria monocytogenes often occur in the production, processing, storage and transportation of food.
Escherichia coli and S. enteritidis are Gram-negative rod (Cell walls of Gram-negative rods are more complicated than those of Gram-positive bacteria: peptidoglycan layers are thinner, sandwiching the membranes, and they contain phospholipids, lipopolysaccharides, and proteins). E. coli exists in everyone’s intestines, and generally does not cause disease, but it will cause disease when eating food contaminated by the bacteria [2]. The bacteria can pollute meat, milk and other animal-derived foods, causing serious harm and loss to the aquaculture and food industry [3]. S. enteritidis is very easy to pollute water sources, animal foods (such as meat, eggs, milk, etc.), fresh fruits and vegetables, etc., seriously threatening public health and food safety, it is one of the important zoonotic pathogens that cause foodborne diseases worldwide [4].
Listeria monocytogenes and S. aureus are belonging to Gram-positive bacteria (containing a thick layer of peptidoglycan in their cell wall). L. monocytogenes is the pathogenic bacteria of zoonosis, as facultative anaerobic bacteria, it is ubiquitous in the environment and widely exists in fresh fruits and vegetables, dairy products, meat products, aquatic products, frozen and frozen foods, ready-to-eat foods and other foods [5]. L. monocytogenes is a psychrophilic bacterium, it could still grow and reproduce at 4 °C, which increases the risk of low temperature food storage threatening human health. S. aureus is one of the most common foodborne pathogenic bacteria as a human symbiosis, once infects food, it will grow rapidly and produce protease and other endotoxins [6]. Taking food contaminated by S. aureus seriously will lead to sepsis, gastrointestinal infection, food poisoning shock syndrome and endocarditis and other diseases [7, 8].
Actions focused on accurately identifying microbes (preferably quickly and objectively), are extremely important for reducing the crisis of the foodborne pathogenic bacterial infection [9,10,11]. There are numerous methods of detection in the literature, ranging from traditional microbiology to molecular biology. Molecular biological detection, immunological method and traditional plate counting method are widely used to identify foodborne pathogens [12]. Because these methods can provide sensitive, qualitative and quantitative information on microorganisms, therefore they are considered to be the “gold standard” of pathogen identification. However, despite the accuracy of detection has improved significantly with these methods, there are limitations to these methods including high time consumption, high cost, and disagreements among observers. In light of this, it is critical to develop rapid, sensitive, and objective methods to diagnose foodborne pathogenic bacteria [13].
Surface enhanced Raman spectroscopy (SERS), an enhanced local electromagnetic field is created near the nanostructured noble metal surfaces, which is a physical method with a wide range of applications. A number of studies have demonstrated the effectiveness of SERS as an alternative method for identifying pathogenic bacteria [14]. Using SERS technology, the Raman signal intensity of target molecules can be enhanced by 104–106 times [15]. SERS signals originate primarily from the cell walls of bacteria attached to the SERS substrate [16, 17]. Thus, the difference of cell wall becomes the key point to identify different microbes by SERS. Recently, an increasing number of reports suggest that SERS including label-free and label-based SERS approaches are the technique which had been demonstrated a number of advantages for the rapid, specific and sensitive detection of pathogenic microorganisms [18]. However, less comparative analysis of the detection sensitivity of various foodborne pathogens was investigated [15, 19]. Finding out the detection sensitivity of foodborne pathogenic microorganisms may assist in the development of new strategies.
In this study, we based on a simple and efficient SERS pretreatment method, detected four foodborne pathogenic bacteria, including E. coli, S. enteritidis, S. aureus and L. monocytogenes, comparatively analyzed the detection specificity of bacterium in different batches, concentrations and mixed states, also combined with principal component analysis (PCA) and hierarchical cluster analysis (HCA), classified and identified four foodborne pathogenic microorganisms, and the feasibility of SERS for detecting four foodborne pathogenic bacteria was comprehensively evaluated. Our results provided theoretical support for the application of SERS to the efficient detection of foodborne pathogenic bacteria in a naturally manner.
2 Methods
Purification steps were not performed on any of the reagents after they were purchased. A UV–Vis spectrophotometer was used to perform ultra violet–visible spectroscopy (UV–vis) (Hash, DR6000, America). Centrifugation was performed in an Eppendorf centrifuge (Eppendorf, 5810R, Germany). The gold nanoparticles size was detected by transmission electron microscopy (Hitachi, HD-2700, Japan). The Raman spectroscopy signals were acquired by portable Raman spectrometers (Ruhai, SEED3000, China).
2.1 Gold nanoparticles preparation
Gold nanoparticles are prepared by referring to the Frens method [20]. The experimental process includes: adding polytetrafluoroethylene magnetic stirrer and 50 mL 0.01% chloroauric acid tetrahydrate (HAuCl4·4H2O, analytical reagent, 100%, Shenyang Jinke Reagent Co., China) solution into a 100 mL round-bottomed flask in turn. While the solution in the round-bottomed flask is boiling, add 0.4 mL of 1% tri-Sodium citrate dihydrate (C6H5Na3O7, analytical reagent, 100%, Shanghai Macklin Technology Co., China) solution quickly, keep heating and refluxing for 30 min to prepare gold nanoparticles. After the reaction, cooled the solution in the ice bath to 0 °C, and refrigerated in a brown reagent bottle at 0–4 °C for standby. The maximum absorption wavelength of gold nanoparticles was detected in the UV spectrophotometer, the morphology of gold nanoparticles was analyzed under the transmission electron microscope, and the best enhancement wavelength position of the substrate was detected using 1 mg L−1 phosmet (C11H12NO4PS2, analytical reagent, 100%, Shanghai Macklin Technology Co., China) solution.
2.2 Bacterial preparation
The detected bacterium E. coli (ATCC 25922), S. enteritidis (ATCC 49214), S. aureus (ATCC 6538), and L. monocytogenes (ATCC 7644) were obtained from microbiology room of Shanghai customs animal, plant and food inspection and quarantine technology center. Bacteria were inoculated in LB-broth (Sigma Aldrich, St. Louis, MI, USA), After bacterial activating, E. coli and S. enteritidis were grown at 37 °C for 14 h, S. aureus and L. monocytogenes were grown at 37 °C for 18 h to reach the stationary growth phase. 10 mL obtained bacterial solution was centrifuged and collected (4 °C, 12,000 r·min−1, 2 min) to remove the supernatant respectively. Washing the collected bacterial sediment with sterile 0.5% NaCl solution for 3 times, and left the sample for 30 min at room temperature for standby, the experimental bacterial concentration will be adjusted according to the assay requirements.
2.3 SERS measurements
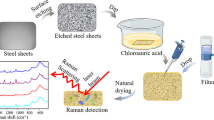
Figure 1 illustrates the steps involved in pre-processing samples before detection (Fig. 1). According to the manufacturer’s recommendation, 100% ethanol was used to calibrate the instrument. 500 µL gold nanoparticles and 300 µL bacterial solution were added into the glass liner for spectral detection, the mixture should be shaken quickly and fully before testing. The instrument parameters are set as following: laser diode with a wavelength of 785 nm as an excitation source, the laser power at the sample was 40 mW, data acquisition time was 10 s, detected 2 times for the average number of SERS.
2.4 Stability of SERS
In order to test the stability of SERS in the detection of E. coli, S. enteritidis, S. aureus and L. monocytogenes, fifteen different detection batches of four bacterial fluids were tested in this study. The concentrations of E. coli, S. enteritidis, S. aureus and L. monocytogenes were 0.5 × 107, 1 × 107, 0.25 × 107, and 0.25 × 107 CFU mL−1 in this test respectively.
2.5 PCA and HCA analysis
Since microorganisms have similar components, the Raman spectra data obtained by SERS also have strong similarity. In order to classify and identify similar Raman spectra, PCA and HCA were used to analyze the SERS profiles of four foodborne pathogens. PCA transforms the indistinguishable data into the classified visual scatter chart according to the minimum difference between the data by reducing the principal component dimension of the multivariate data, and analyzes the similarity in the comparison categories. HCA classifies the data according to the similarity, and then determines the different categories in the research data group.
2.6 Detection sensitivity measurement
Detection sensitivity was measured using a liquid phase assay. Briefly, bacteria were cultured in the LB-broth medium to stationary growth phase and adjusted to 107 CFU mL−1 with sterile 0.5% NaCl solution. In sterile glass liner, 500 µL of bacterial suspension was diluted 1/twofold with sterile 0.5% NaCl solution. Then 300 µL of gold nanoparticles were added into the glass liner and mixed. Three independent experiments were conducted in triplicate. The limit of quantification value was defined as the range between the highest concentration of the bacteria at which microbial SERS characteristic spectrum could be observed and the limit of quantification value was that no obvious SERS characteristic spectrum was observed.
2.7 SERS identification of mixed bacteria
The mixed bacterial SERS was applied to classify and identify the similar Raman spectra as the following group: L. monocytogenes and S. aureus, L. monocytogenes and S. enteritidis, L. monocytogenes and E. coli, E. coli and S. aureus, E. coli and S. enteritidis, and S. aureus and S. enteritidis. Detections were carried out in a total volume of 800 µL, containing 500 µL gold nanoparticles and 300 µL bacterial solution (150 µL bacterial A + 150 µL bacterial B). The SERS analysis was performed on all samples in triplicate.
3 Results
3.1 Characterization of the gold nanoparticles
The TEM (transmission electron microscope) image of the gold nanoparticles shows that the gold nanoparticles were uniform in size and shape, and there’s no obvious agglomeration (Fig. 2a), the solution is claret-red and transparent. Optical properties of gold nanoparticle solutions were evaluated using UV absorption spectra (Fig. 2b). The peak position and width can be used to estimate the particle size and dispersity of gold colloid. Among the synthesized four gold nanoparticles reagents, the maximum UV absorption peak is about 534, 537, 538, 540 nm respectively, and the Raman spectrum verification results of phosmet (C11H12NO4PS2) showed that 537 or 538 nm has the best enhancement effect of pure AuNPs (Fig. 2c), and the 538 nm one was selected for the next assay. The hot spot generation mechanisms have a strong analytic distance-dependence feature, a minute reduction of distance results in a significant enhancement of intensity. Only molecules on or very close to the metal surface produce the enormous field strength. Statistical analysis showed that the gold nanoparticles size distribution ranges from 55 to 65 nm, with some small molecule sizes serving an artificial hotspot. The surface has a certain roughness, which can effectively enhance the surface Plasmon resonance (SPR) effect, and is suitable for SERS substrate.
3.2 SERS spectra of different foodborne pathogenic microorganisms
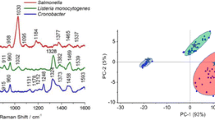
The SRES signals originated from the cell secretions, including adenine, guanine, hypoxanthine, xanthine, however, the interactions between the components of the mixture such as cell secretions and the different cell wall structure of Gram-negative or Gram-positive bacteria might interference the signal bands [21,22,23]. In this study, SERS spectra of four different bacterium E. coli, S. enteritidis, S. aureus, and L. monocytogenes were compared (Fig. 3). The overall Raman spectrum vibration peak intensity of Gram-positive bacteria is larger, this may be due to the cell wall of Gram-positive bacteria is thicker and rougher than that of gram-negative bacteria. Similar characteristic bands-pattern were found in the Gram-negative and Gram-positive bacterium. Comparing with the SERS spectra of blank control group (0.5% NaCl), it was found that the bands at 487, 598, 734, 814, 998, and 1456 cm−1 were attributed to Gram-negative bacteria, the main bands of the Gram-negative bacteria were at 665, 734, 814, 964, 1347 and 1456 cm−1. The bands assignment was summarized in Table 1. Remarkably, the strongest SERS response of Gram-positive bacteria is 734 cm−1, and that of Gram-negative bacteria is 814 cm−1, which indicates the SERS signal might be interrupted by the different structure of bacterial wall.
3.3 Stability and uniformity
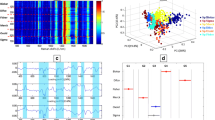
Practical applications of SERS technology rely heavily on the substrate’s stability and uniformity [29]. We collected 15 random samples of SERS spectra of four different types of bacteria to assess their uniformity (Fig. 4), and the shape and strength of the peaks of each bacterium were essentially the same, indicating the reproducibility of the enhancement effect of the substrate. Moreover, the differences in relative standard deviation (RSD) of the characteristic peak SERS intensity in Gram-negative bacteria (at 814 cm−1, 4.12% for E. coli, 4.44% for S. enteritidis) as well as that in the Gram-positive bacteria (at 734 cm−1, 5.35% for S. aureus, 5.76% for L. monocytogenes) fall within the error range for each bacterial species further suggest the stability in SERS detection for each bacteria, and confirmed the possibility of SERS application in the detection of four common foodborne pathogens.
SERS spectrum of four food borne pathogens (E. coli (a), S. enteritidis (c), S. aureus (e) and L. monocytogenes (g)) measured at 15 substrates, and Raman intensity of each bacterial at 814 cm−1 for E. coli (b), S. enteritidis (d), 734 cm−1 for S. aureus (f) L. monocytogenes (h) at 15 locations in the same basement
3.4 Detection sensitivity
The aggregation of bacteria and gold nanoparticle is mainly originated from the components of the cell wall of Gram-negative or Gram-positive bacteria which is negatively charged due to the presence of outer membrane lipopolysaccharides or teichoic acid, as well as complex molecular assemblages’ structure of the detected biological macromolecules, respectively. In order to investigate the detection sensitivity of the bacterium, four food-borne pathogenic bacteria were tested at different concentrations. Interestingly, it was found that with the concentration was growing thinner, the SERS intensity was not growing stronger, which is similar with the detection pattern of clenbuterol using different concentrations of NaCl solutions in pork [30]. For the Gram-negative bacteria, the minimum MSDC at 814 cm−1 of E. coli was 0.5 × 107 CFU mL−1 and that in S. aureus was 1 × 107 CFU mL−1. For the Gram-positive bacteria, the MSDC at 734 cm−1 of S. enteritidis was 0.25 × 107 CFU mL−1 and that in L. monocytogenes was also the same (Fig. 5). The MSDC of the Gram-positive bacteria S. enteritidis and L. monocytogenes were lower than the Gram-negative bacteria E. coli and S. aureus, which indicating that the detection sensitivity of S. enteritidis and L. monocytogenes is higher than that in the E. coli and S. aureus.
SERS spectrum of four food borne pathogens (E. coli (a), S. enteritidis (c), S. aureus (e) and L. monocytogenes (g)) measured corresponding to different bacterial solution concentrations and the trend of Raman signal intensity with bacterial solution concentration (E. coli (b), S. enteritidis (d), S. aureus (f) and L. monocytogenes (h)). The shaded areas represent the 95% confidence
3.5 PCA and HCA analysis
Due to the similar cell components of microorganisms, the Raman spectrum share a similar SERS spectrum pattern. In order to classify and identify the similar Raman spectrum, PCA and HCA were used to analyze the SERS profiles of four foodborne pathogens. As one of the most widely used unsupervised multivariate analysis methods, PCA provides an overview of spectral data [31, 32]. According to the minimum difference between the SERS data, PCA reduces the principal component dimension of the multivariate data, and analyzes the similarity in the comparison categories, transforms the indistinguishable data into the classified visual scatter chart. HCA classifies the research data firstly, then judges and analyzes the similarity and difference between the populations of the research profiles [33, 34]. PCA analysis revealed that the SERS data of pathogenic microorganisms were divided into four categories, with the spectral variance of variance 97.3% (Fig. 6a). The loadings of the model show that spectral regions 650–680, 734, 814, 950–1100, and 1315–1450 cm−1 were the main contribution region for the discrimination between these species. E. coli and S. aureus are relatively close, which is consist with the uniformity in (Fig. 4). HCA analysis showed that the tree topology can be separated into two clades, including Gram-negative and Gram-positive bacteria, E. coli and S. aureus had a similar SERS spectrum pattern, while S. enteritidis and L. monocytogenes were clustered into a big branch of Gram-positive bacteria (Fig. 6b).
3.6 SERS detection of mixed bacterial solution
Multiple bacterial SERS detection was tested using the six samples (see Sect. 2.7 for details), each food-borne pathogenic bacteria was mixed with equal amounts, and 15 duplicates were performed to represent diverse clones. PCA scored plots reveal six well-defined clusters for each sample, with PC capturing around 89.3% of the variance in the spectra (Fig. 7a). HCA analysis (Fig. 7b) revealed that six samples were divided into six categories, and there was no cross interference in each branch. The results of PCA and HCA analysis indicated that the mixed bacterial liquid could also be effectively distinguished by SERS detection.
4 Discussion
Globally, foodborne pathogens are the cause of a large number of outbreaks, diseases and deaths [35]. As a serious public health problem, foodborne pathogens with outbreaks attributed to the emergence of new pathogens as well as the reemergence of a few of old pathogens [36, 37]. There are a number of methods for identifying food borne pathogens, including phenotypic assays, PCR, and immunoassays [38]. These methods present primary limitations such as time-consuming, complex and laborious nature. A rapid and objective method of identifying foodborne bacteria can assist in designing appropriate detection strategies. In this work, SERS measurements of the application efficiency of four food borne pathogens with gold nanoparticles were comparatively detected. The results demonstrated that SERS spectra obtained from detected four food borne pathogens can be used to establish discriminatory profiles of bacterial fingerprints with good stability and special Raman peaks to different biomolecules, which was consisted with the previous report that microbes causing urinary tract infections [39].
In previous studies, most study was aiming at creating new SERS detection methods which were combined with other detection technologies [40,41,42], only a few of studies investigated the SERS detection difference of species from different genera [43, 44]. Additionally, SERS was used to differentiate bacterial pathogens based on the single stable endotoxin component, though only specific reference endotoxins were tested in the Gram-negative bacteria [12]. Our study systematically analyzed the detection stability of four food borne pathogens (two Gram-positive bacteria and two Gram-negative bacteria), and found that both Gram-positive bacteria and negative bacteria showed low RSD, these results suggested that SERS might exhibit reliable reproducibility and repeatability for further bacteria.
Recently, many studies reported that a linear relationship between the bacterial concentration and the SERS intensity were detected while the distilled water as the solvent [45, 46]. However, at the present study, SERS signals generated by the four gold nanoparticles-based bacteria didn’t exhibit a linear dependence on bacteria concentration in the NaCl solution. Interestingly, for Gram-negative bacteria, SERS intensity reduced slightly at the maximum concentration in the case of E. coli but increased in the case of S. enteritidis. In addition, for Gram-positive bacteria, the highest SERS intensity didn’t exhibit at the maximum concentration of bacteria. Previous researches confirmed that the degree of interaction between gold nanoparticles and bacteria will influence the enhancement performance [30, 47]. The degree of aggregation of gold nanoparticles would be affected by the concentration of NaCl [48]. It is believed that the electrolyte solution upsets the charge balance, resulting in a small fraction of analyst molecules adsorbing onto hot spots, thereby causing SERS to acquire a large signal [49]. For Gram-positive and Gram-negative bacteria, different components on the bacterial surface will affect the adsorption efficiency of nano-particles, thus, NaCl in the substrate acted as an aggregating compound, leads to an entirely novel pattern of SERS signal and bacterial concentration.
The advantage of SERS lies in its ability to achieve in situ chemical composition detection [50]. However, natural infection processes often do not involve a single bacterium. Comparative analysis of the detection stability, sensitivity, and discrimination ability of SERS will be helpful for new detection strategy. As observed previously in other spectroscopic techniques studies, the results obtained in this pilot study further demonstrated that, the SERS spectrum signal can be used to create discriminatory bacterial fingerprints by using biochemical information from multiple pure bacterial cultures. Furthermore, we tried to explore the gold nanoparticles-based SERS discrimination of mixed bacteria, and confirmed that SERS was able to discriminate the mixed bacteria. Nevertheless, the present experiment was only tested in an ideal concentration of 1:1 state, and it is necessary to conduct research with a higher diversity of different Gram-negative and Gram-positive bacterial species. Based on the findings in our study, along with those established in previous studies, SERS-based bacterial discrimination is supported and may be used as a guide for future experiments to expand the application of SERS for bacterial detection, as well as to optimize experimental and analytical detail procedures. However, the present work only based on the conventional citrate-reduced gold colloid method compared the SERS sensitivity toward four foodborne pathogens. For practical applications, the models able of classifying each of the bacteria strain present in mixtures separately need further research and the establishment of new SRES detection methods.
5 Conclusion
In this report, the application efficiency of gold nanoparticles-based SERS in four foodborne pathogenic microbes including the gram-negative E. coli and S. enteriditis, and the gram-positive S. aureus and L. monocytogenes was compared. All these four bacteria showed stable detection results with low RSD of the characteristic peak SERS intensity, and the minimal SERS detection concentration of the Gram-positive bacteria S. enteritidis and L. monocytogenes were lower than the Gram-negative bacteria E. coli and S. aureus, indicating the detection sensitivity of S. enteritidis and L. monocytogenes is higher than that in the E. coli and S. aureus. PCA analysis revealed that the SERS data of pathogenic microorganisms were divided into four categories, with the spectral variance of variance 97.3%, and HCA analysis showed that the tree topology can be separated into two clades, including Gram-negative and Gram-positive bacteria. In addition, multiple bacterial SERS detection revealed that the mixed bacterial liquid could also be effectively distinguished by SERS based on the PCA and HCA analysis. The results of this pilot study demonstrate the potential of SERS to rapidly identify Gram-negative and Gram-positive foodborne pathogens.
Data availability
All data are provided as figures.
Change history
23 March 2024
A Correction to this paper has been published: https://doi.org/10.1007/s42452-024-05839-9
References
World Health Organization. Food safety. 2022. https://www.who.int/news-room/fact-sheets/detail/food-safety. Accessed 19 May 2022.
Palmela C, Chevarin C, Xu Z, Torres J, Colombel JF. Adherent-invasive Escherichia coli in inflammatory bowel disease. Gut. 2017;67(3):2017–314903.
Kaper JB. Pathogenic Escherichia coli. Int J Med Micro-biol. 2005;295(6–7):355–6.
Heymans R, Vila A, Heerwaarden CA, Jansen CC, Castelijn GA, Voort M, Biesta-Peters EG. Rapid detection and differentiation of Salmonella species, Salmonella Typhimurium and Salmonella Enteritidis by multiplex quantitative PCR. PLoS ONE. 2018;13(10):0206316.
Farber JM, Peterkin PI. Listeria monocytogenes, a food-borne pathogen. Microbiol Rev. 1991;55(3):476–511.
Deurenberg RH, Stobberingh EE. The evolution of Staphylococcus aureus. Infect Genet Evol. 2008;8(6):747–63.
Castro A, Silva J, Teixeira P. Chapter 8—Staphylococcus aureus, a food pathogen: virulence factors and antibiotic resistance. In: Holban AM, Grumezescu AM, editors. Foodborne diseases, vol. 17. Handbook of food bioengineering. London: Academic Press; 2018. p. 213–38.
Martin E, Lina G, Dumitrescu O. Staphylococcus—Staphylococcus aureus. In: Batt CA, Tortorello ML, editors. Encyclopedia of food microbiology. 2nd ed. Oxford: Academic Press; 2014. p. 501–7.
Ning R, Fan J, Kong L, Jiang X, Qian Y, Du T, Zhang G, Wu W. Recent developments of droplets-based microfluidics for bacterial analysis. Chin Chem Lett. 2022;33(5):2243–52.
French K, Evans J, Tanner H, Gossain S, Hussain A. The clinical impact of rapid, direct MALDI-ToF identification of bacteria from positive blood cultures. PLoS ONE. 2016;11(12): e0169332.
Watts GS, Thornton JE, Youens-Clark K, Ponsero AJ, Hurwitz BL. Identification and quantitation of clinically relevant microbes in patient samples: comparison of three k-mer based classifiers for speed, accuracy, and sensitivity. PLoS Comput Biol. 2019;15(11):1006863.
Wu X, Zhao Y, Zughaier SM. Highly sensitive detection and differentiation of endotoxins derived from bacterial pathogens by surface-enhanced Raman scattering. Biosensors. 2021;11(7):234.
Ahmed A, Rushworth JV, Hirst NA, Millner PA. Biosensors for whole-cell bacterial detection. Clin Microbiol Rev. 2014;27(3):631–46.
Cui L, Li H-Z, Yang K, Zhu L-J, Xu F, Zhu Y-G. Raman biosensor and molecular tools for integrated monitoring of pathogens and antimicrobial resistance in wastewater. TrAC Trends Anal Chem. 2021;143: 116415.
Zhu A, Ali S, Xu Y, Ouyang Q, Wang Z, Chen Q. SERS-based Au@ Ag NPs solid-phase substrate combined with chemometrics for rapid discrimination of multiple foodborne pathogens. Spectrochim Acta Part A Mol Biomol Spectrosc. 2022;270: 120814.
Wu X, Xu C, Tripp RA, Huang YW, Zhao Y. Detection and differentiation of foodborne pathogenic bacteria in mung bean sprouts using field deployable label-free SERS devices. Analyst. 2013;138:3005–12.
Wang C, Wang J, Li M, Qu X, Zhang K, Rong Z, Xiao R, Wang S. A rapid SERS method for label-free bacteria detection using polyethylenimine-modified Au-coated magnetic microspheres and Au@Ag nanoparticles. Analyst. 2016;141(22):6226–38.
Wang K, Li S, Petersen M, Wang S, Lu X. Detection and characterization of antibiotic-resistant bacteria using surface-enhanced Raman spectroscopy. Nanomaterials. 2018;8(10):762.
Cheong Y, Kim YJ, Kang H, Choi S, Lee HJ. Rapid label-free identification of Klebsiella pneumoniae antibiotic resistant strains by the drop-coating deposition surface-enhanced Raman scattering method. Spectrochim Acta Part A. 2017;183:53–9.
Frens G. Controlled nucleation for the regulation of the particle size in monodisperse gold suspensions. Nat Phys Sci. 1973;241(105):20–2.
Fan C, Hu Z, Mustapha A, Lin M. Rapid detection of food-and water-borne bacteria using surface-enhanced Raman spectroscopy coupled with silver nanosubstrates. Appl Microbiol Biotechnol. 2011;92(5):1053–61.
Kahraman M, Keserolu K, Culha M. On sample preparation for surface-enhanced Raman scattering (SERS) of bacteria and the source of spectral features of the spectra. Appl Spectrosc. 2011;65(5):500.
Premasiri WR, Lee JC, Sauer-Budge A, Théberge R, Costello CE, Ziegler LD. The biochemical origins of the surface-enhanced Raman spectra of bacteria: a metabolomics profiling by SERS. Anal Bioanal Chem. 2016;408:4631–47.
Gillaspy AF, Worrell V, Orvis J, Roe BA, Dyer DW, Iandolo JJ. The Staphylococcus aureus NCTC 8325 genome. In: Gram-positive pathogens. New York: Wiley; 2006. p. 381–412.
Lee DG, Urbach JM, Wu G, Liberati NT, Feinbaum RL, Miyata S, Ausubel FM. Genomic analysis reveals that Pseudomonas aeruginosa virulence is combinatorial. Genome Biol. 2006;7:1–14.
Ea D, Eg E, Ov S, Na S, En E, Av E, In K. Molecular origin of surface-enhanced Raman spectra of E. coli suspensions excited at 532 and 785 nm using silver nanoparticle sols as SERS substrates. Bull Russ State Med Univ. 2018;6:25–32.
Tettelin H, Nelson KE, Paulsen IT, Eisen JA, Read TD, Peterson S, Fraser CM. Complete genome sequence of a virulent isolate of Streptococcus pneumoniae. Science. 2001;293(5529):498–506.
Qin X, Galloway-Peña JR, Sillanpaa J, Roh JH, Nallapareddy SR, Chowdhury S, Murray BE. Complete genome sequence of Enterococcus faecium strain TX16 and comparative genomic analysis of Enterococcus faecium genomes. BMC Microbiol. 2012;12(1):1–20.
Bocklitz T, Walter A, Hartmann K, Rösch P, Popp J. How to pre-process Raman spectra for reliable and stable models? Anal Chim Acta. 2011;704(12):47–56.
Guo Q, Peng Y, Zhao X, Chen Y. Rapid detection of clenbuterol residues in pork using enhanced Raman spectroscopy. Biosensors. 2022;12(10):859.
Jolliffe IT, Cadima J. Principal component analysis: a review and recent developments. Philos Trans R Soci A Math Phys Eng Sci. 2016;374(2065):20150202.
Gracie K, Lindsay D, Graham D, Faulds K. Bacterial meningitis pathogens identified in clinical samples using a SERS DNA detection assay. Anal Methods. 2015;7(4):1269–72.
Kniggendorf A-K, Gaul TW, Meinhardt-Wollweber M. Hierarchical cluster analysis (HCA) of microorganisms: an assessment of algorithms for resonance Raman spectra. Appl Spectrosc. 2011;65(2):165–73.
Lin Z, Zhao X, Huang J, Liu W, Zheng Y, Yang X, Zhang Y, La Chapelle ML, Fu W. Rapid screening of colistin-resistant Escherichia coli, Acinetobacter baumannii and Pseudomonas aeruginosa by the use of Raman spectroscopy and hierarchical cluster analysis. Analyst. 2019;144(8):2803–10.
Kirk MD, Pires SM, Black RE, Caipo M, Crump JA, Devleesschauwer B, Döpfer D, Fazil A, Fischer-Walker CL, Hald T, et al. World health organization estimates of the global and regional disease burden of 22 foodborne bacterial, protozoal, and viral diseases, 2010: a data synthesis. PLoS Med. 2015;12(12):1001921.
Akbar A, Anal AK. Food safety concerns and food-borne pathogens, Salmonella, Escherichia coli and Campylobacter. FUUAST J Biol. 2011;1(1 June):5–17.
Newell DG, Koopmans M, Verhoef L, Duizer E, Aidara-Kane A, Sprong H, Opsteegh M, Langelaar M, Threfall J, Scheutz F, et al. Food-borne diseases—the challenges of 20 years ago still persist while new ones continue to emerge. Int J Food Microbiol. 2010;139:3–15.
Berry ME, Kearns H, Graham D, Faulds K. Surface enhanced Raman scattering for the multiplexed detection of pathogenic microorganisms: towards point-of-use applications. Analyst. 2021;146(20):6084–101.
Usman M, Tang J-W, Li F, Lai J-X, Liu Q-H, Liu W, Wang L. Recent advances in surface enhanced Raman spectroscopy for bacterial pathogen identifications. J Adv Res. 2022;51:91–107.
Rahman A, Kang S, Wang W, Huang Q, Kim I, Vikesland PJ. Lectin- modified bacterial cellulose nanocrystals decorated with Au nanoparticles for selective detection of bacteria using surface-enhanced Raman scattering coupled with machine learning. ACS Appl Nano Mater. 2022;5(1):259–68.
Li JF, Zhang YJ, Ding SY, Panneerselvam R, Tian ZQ. Core-shell nanoparticle-enhanced Raman spectroscopy. Chem Rev. 2017;117(7):5002–69.
Liu H-B, Chen C-Y, Zhang C-N, Du X-J, Li P, Wang S. Functionalized AuMBA@Ag nanoparticles as an optical and SERS dual probe in a lateral flow strip for the quantitative detection of Escherichia coli O157:H7. J Food Sci. 2019;84(10):2916–24.
Dina N, Zhou H, Colniţă A, Leopold N, Szoke-Nagy T, Coman C, Haisch C. Rapid single-cell detection and identification of pathogens by using surface-enhanced Raman spectroscopy. Analyst. 2017;142(10):1782–9.
Liu Y, Zhou H, Hu Z, Yu G, Yang D, Zhao J. Label and label-free based surface-enhanced Raman scattering for pathogen bacteria detection: a review. Biosens Bioelectron. 2017;94:131–40.
Zhang H, Guan Q, Zeng P, Wu G, Hong Y, Yang W, Wang C. Ultrasensitive detection of pectin based on the decarboxylation reaction and surface-enhanced Raman spectroscopy. Anal Methods. 2022;14(23):2266–76.
Geng P, Sun S, Wang X, Ma L, Guo C, Li J, Guan M. Rapid and sensitive detection of amphetamine by SERS-based competitive immunoassay coupled with magnetic separation. Anal Methods. 2022;14(26):2608–15.
Sabur A, Havel M, Gogotsi Y. SERS intensity optimization by controlling the size and shape of faceted gold nanoparticles. J Raman Spectrosc Int J Orig Work Asp Raman Spectrosc Incl High Order Process Brillouin Rayleigh Scatt. 2008;39(1):61–7.
Bi S, Shao D, Yuan Y, Zhao R, Li X. Sensitive surface-enhanced Raman spectroscopy (SERS) determination of nitrofurazone by β-cyclodextrin-protected AuNPs/γ-Al2O3 nanoparticles. Food Chem. 2022;370: 131059.
Yaffe NR, Ingram A, Graham D, Blanch EW. A multi-component optimisation of experimental parameters for maximising SERS enhancements. J Raman Spectrosc. 2010;41(6):618–23.
Tauber JP, Matthäus C, Lenz C, Hoffmeister D, Popp J. Analysis of basidiomycete pigments in situ by Raman spectroscopy. J Biophotonics. 2018;11(6):201700369.
Acknowledgements
We appreciate Wei Han, Qiang Zhang and Xionghai Yi for their help and suggestions in this study.
Funding
This work is supported by the Startup Research Fund from the Shanghai Customs College [Grant No. 2315019A2020] and Shanghai Natural Science Foundation under Grant number 22ZR1426500. This study was made possible in part by Project ZR2022MC136 supported by Shandong Provincial Natural Science Foundation.
Author information
Authors and Affiliations
Contributions
Conceptualization, FTG; methodology, HRL, NNC, and XN; software, HRL; resources, HRL, NNC, and XN; formal analysis, HRL; investigation, HRL and NNC; writing—original draft preparation, HRL; writing—review and editing, NNC; visualization, HRL; supervision, FTG; project administration, XN; funding acquisition, HRL. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This study does not involve research on human participants and/or animals. The authors understand the informed consents and agree to all contents.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original version of this article has been revised: The Funding section has been corrected.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, H., Chen, N., Ning, X. et al. Comparative analysis of the application efficiency of gold nanoparticles-based SERS in four foodborne pathogenic microbes. Discov Appl Sci 6, 122 (2024). https://doi.org/10.1007/s42452-024-05751-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42452-024-05751-2