Abstract
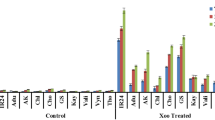
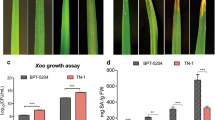
Bacterial blight of rice caused by Xanthomonas oryzae pv. oryzae (Xoo) is an important disease of rice causing significant yield losses. Currently, there are ten highly diverse pathotypes of Xoo prevalent in Punjab state of India. Breeding for and deploying host plant resistance is the most economical approach for the management of this disease. But pathogen variability coupled with uncertainties in prevailing weather conditions due to climate change poses serious threat to the durability of R genes and makes the plant vulnerable to attack by the pathogen. During the current study, biochemical basis of resistance of two R genes, Xa4 and Xa7 was studied at different temperature regimes and five pathotypes of Xoo viz. PbXo-4, PbXo-7, PbXo-8, PbXo-10 and Sgr-1001. Level of resistance in Xa4 and Xa7 genes was observed to be highly correlated with the expression of three defense related enzymes viz. peroxidase, polyphenol oxidase and phenylalanine ammonia lyase. It was further observed that the expression of Xa4 and Xa7 was influenced by temperature as Xa4 showed increased effectiveness at lower temperature while Xa7 was more effective at high temperature. The differential pattern of lesion length progression on these near isogenic lines in response to different temperature regimes also suggests the level of quantitative resistance imparted by these genes.
Similar content being viewed by others
References
Anonymous (2015) Package of practices for kharif crops. Punjab Agricultural University, Ludhiana, pp 1–24
Arfaoui A, Hadram EI, A, Mabrouk Y, Sifi B, Boudabous A, EI Hadrami I, Daayy F, Cherif M, (2007) Treatment of chickpea with Rhizobium isolates enhances the expression of phenylpropanoid defense related genes in response to infection by Fusarium oxysporum f.sp. ciceris. Plant Physiol Biochem 45:470–479
Baker B, Zambryski P, Staskawicz B, Dinesh-Kumar SP (1997) Signaling in plant-microbe interactions. Science 276:726–733
Bastin M, Unluer O (1972) Effects of actinomycetes-D on the formation of enzymes in Jerusalem artichoke tuber slices. Phytopathol 94:693–705
Burdon JJ, Thrall PH, Ericson AL (2006) The current and future dynamics of disease in plant communities. Annu Rev Phytopathol 44:19–39
Burrell MM, Rees TAP (1974) Metabolism of phenylalanine and tyrosine by rice leaves infected by Pyricularia oryzae. Physiol Plant Pathol 4:497–508
Busungu C, Taura S, Sakagami JI, Ichitani K (2016) Identification and linkage analysis of a new rice bacterial blight resistance gene from XM14, a mutant line from IR24. Breed Sci 66:636–645
Chakraborty S (2005) Potential impact of climate change on plant-pathogen interactions. Aust Plant Pathol 34:443–448
Chakraborty S, Panga IB, Lupton J, Hart L, Room PM, Yates D (2000) Production and dispersal of Colletotrichum gloeosporioides spores on Stylosanthes scabra under elevated CO2. Environ Pollut 108:317–326
Chen S, Wang C, Yang J, Chen B, Wang W, Su J, Feng A, Zeng L, Zhu X (2020) Identification of the novel bacterial blight resistance gene Xa46(t) by mapping and expression analysis of the rice mutant H120. Sci Rep 10:12642. https://doi.org/10.1038/s41598-020-69639-y
Chittoor JM, Leach JE, White FF (1997) Differential induction of peroxidase gene family during infection of rice by Xanthomonas oryzae pv. oryzae. Mol Plant Microbe Interact 10:861–871
Cottyn B, Mew TW (2004) Bacterial blight of rice. In: Sci EPC (ed) Goodman RM. Marcel Dekker, New York, pp 79–83
Crowl TA, Crist TO, Parmenter RR, Belovsky G, Lugo AE (2008) The spread of invasive species and infectious disease as drivers of ecosystem change. Front Ecol Environ 6:238–246
Debnath S, Satya P, Sahab C (2013) Pathotype characterization of Xanthomonas oryzae pv. oryzae isolates of West Bengal and evaluation of resistance genes of bacterial blight of rice (Oryza sativa L.). J Crop Weed 9:198–200
Dhkal M, Hunjan MS, Kaur H, Kaur R (2016) Biochemical basis of bacterial leaf streak resistance in maize. Indian Phytopath 69:373–380
Dossa GS, Oliva R, Maiss E, Wydra K (2015) High temperature enhances the resistance of cultivated African rice, Oryza glaberrima, to bacterial blight. Plant Dis 100:380–387
Dossa GS, Quibod I, Atienza-Grande G, Olive R, Maiss E, Vera Cruz C, Wydra K (2020) Rice pyramided line IRBB67 (Xa4/ Xa7) homeostasis under combined stress of high temperature and bacterial blight. Sci Rep 10:683. https://doi.org/10.1038/s41598-020-57499-5
Eastburn DM, McElrone AJ, Bilgin DD (2011) Influence of atmospheric and climatic change on plant-pathogen interactions. Plant Pathol 60:54–69
Garrett K, Dendy S, Frank E, Rouse M, Travers S (2006) Climate change effects on plant disease: genomes to ecosystems. Annu Rev Phytopathol 44:489–509
Gautam RK, Singh PK, Sakthivel K, Srikumar M, Kumar N, Kumar K, Singh AK, Roy SD (2015) Analysis of pathogenic diversity of the rice bacterial blight pathogen (Xanthomonas oryzae pv. oryzae) in the Andaman Islands and identification of effective resistance genes. J Phytopathol 163:423–432
Grulke NE (2011) The nexus of host and pathogen phenology: Understanding the disease triangle with climate change. New Phytol 189:8–11
Hunjan MS (2012) Pathogenic diversity and molecular characterization of Xanthomonas oryzae pv. oryzae [(Ishiyama) Swings et al.] causing bacterial blight of rice in Punjab. Ph.D. dissertation. Punjab Agricultural University, Ludhiana, India.
Hutin M, Sabot F, Ghesquiere A, Koebnik R, Szurek B (2015) A knowledge-based molecular screen uncovers a broad-spectrum OsSWEET14 resistance allele to bacterial blight from wild rice. Plant J 84:694–703
Jiang N, Yan J, Liang Y, Shi Y, He Z, Wu Y, Zeng Q, Liu X and Peng J (2020) Resistance Genes and their Interactions with Bacterial Blight/Leaf Streak Pathogens (Xanthomonas oryzae) in Rice (Oryza sativa L.)—an Updated Review. Rice 13:3 https://doi.org/https://doi.org/10.1186/s12284-019-0358-y
Kaji R, Ogawa T (1995) Identification of the located chromosome of the resistance gene, Xa-7, to bacterial leaf blight in rice. Breed Sci 45(Suppl. 1):79
Kauffman HE, Reddy APK, Hsieh SPY, Merca SD (1973) An improved technique for evaluating resistance of rice varieties to Xanthomonas oryzae pv. oryzae. Pl Dis Rep 57:537–541
Kim SM, Reinke RF (2019) A novel resistance gene for bacterial blight in rice, Xa43(t) identified by GWAS, confirmed by QTL mapping using a bi-parental population. PLoS ONE 14:e0211775
Kim SM, Suh JP, Qin Y, Noh TH, Reinke RF, Jena KK (2015) Identification and fine-mapping of a new resistance gene, Xa40, conferring resistance to bacterial blight races in rice (Oryza sativa L.). Theor Appl Genet 128:1933–1943
Kumar A, Hunjan MS, Kaur H, Dhillon HK, Singh PP (2017) Biochemical responses associated with resistance to bacterial stalk rot caused by Dickeya zeae in maize. J Phytopath 165:822–832
Li L, Stiffen JC (2002) Overexpression of polyphenol oxidase in transgenic tomato plants results in enhanced bacterial disease resistance. Planta 215:239–247
Liang Xiao X, Ping Li, Zheng A (2013) In vitro selection of sheath blight resistance germplasms in rice. Afr J Microbiol Res 7:4422–4429
Lore JS, Vikal Y, Hunjan MS, Goel RK, Bharaj TS, Raina GL (2011) Genotypic and pathotypic diversity of Xanthomonas oryzae pv. oryzae, the cause of bacterial blight of rice in Punjab state of India. J Phytopath 159:479–487
Lore JS, Mangat GS, Jain J, Hunjan MS, Kaur R, Singh K, Zaidi NW, Singh US and Vera Cruz CM (2016) Pathotyping and host-plant resistance to bacterial blight of rice in north-western region of India: A success story (Abstract). The 5th International Conference on Bacterial Blight of Rice, The Bellevue Manila, Alabang, Philippines, October 17–19, 2016. pp 61
Nandakumar R, Babu S, Viswanthan R, Raguchander T, Samiyappan R (2001) Induction of systemic resistance in rice against sheath blight disease by Pseudomonas flourescens. Soil Biol Biochem 3:603–612
Neelam K, Mahajan R, Gupta V, Bhatia D, Gill BK, Komal R, Lore JS, Mangat GS, Singh K (2019) High-resolution genetic mapping of a novel bacterial blight resistance gene xa-45(t) identified from Oryzaglaberrima and transferred to Oryzasativa. TheorAppl Genet 133:689–705
Noda T, Li C, Li J, Ochiai H, Ise K, Kaku H (2001) Pathogenic diversity of Xanthomonas oryzae pv. oryzae strains from Yunnan Province, China. Jpn Agric Res Q 35:97–103
Ou SH (1985) Rice diseases. 2nded, Commonwealth Agricultural Bureaux, Kew, Surrey, United Kingdom pp380.
Pal TK, Bhattacharaya S, Chakraborty K (2011) Induction of systemic resistance in rice by leaf extract of Cymbopogan citrus and Ocimum sanctum against sheath blight disease. Arch Appl Sci Res 3:392–400
Parihar PS, Prakash O, Punetha H (2012) Investigation on defensive enzymes activity of Brassica juncea genotypes during pathogenesis of Alternaria blight. Nat Sci 10:63–68
Santigo R, De Armas R, Legaz M, Vicente C (2009) Changes in phenolic acids content, phenylalanine ammonia lyase and peroxidase activities in sugarcane leaves induced by elicitors isolated from Xanthomonas albilineans. Aust J Plant Pathol 38:357–365
Seetharaman K, Babu RM, Sajeena A, Ebenezar EG, Raja AS, Biji KR, Prakas MS (2004) Induction of phenols in rice bacterial leaf blight resistance. J Ecobiol 16:53–56
Shannon LM, Kay E, Lew JY (1966) Peroxidase isozymes from horeseradish roots. Isolation and physical properties. J Biol Chem 241:2166–2172
Sharma N, Kaur C, Singh J, Gupta AK (2012) Biochemical responses associated with bacterial blight of rice (Oryza sativa). Arch Phytopathol Plant Protect 45:47–61
Sun X, Yang Z, Wang S, Zhang Q (2003) Identification of a 47 kb DNA fragment containing Xa4, a locus for bacterial blight resistance in rice.Theor Appl Genet 106:683–87
Tahsili J, Sharifi M, Safaie N, Esmaeilzadeh BS, Behmanesh M (2014) Induction of lignans and phenolic compounds in cell culture of Linum album by culture filtrate of Fusarium graminearum. J Plant Interact 14:412–417
Vanitha SC, Niranjana SR, Umesha S (2009) Role of phenylalanine ammonia lyase and polyphenol oxidase in host resistance to bacterial wilt of tomato. J Phytopath 157:552–557
Velazhahan R, Jayarajb J, Khabbaza S, Kumara RS, Muthukrishnan S (2006) Xanthomonas oryzae pv. oryzae infection triggers accumulation of phenolics, defense-related enzymes and thaumatin-like proteins in rice leaves. Arch Phytopath Plant Protect 39:329–339
Vera Cruz CM, Bai J, On I, Leung H, Nelson RJ, Mew TW, Leach JE (2000) Predicting durability of a disease resistance gene based on an assessment of the fitness loss and epidemiological consequences of avirulence gene mutation. PNAS 97:13500–13505
Webb KM, Ona I, Bai J, Garrett KA, Mew T, Vera Cruz CM, Leach JE (2010) A benefit of high temperature: increased effectiveness of a rice bacterial blight disease resistance gene. New Phytol 185:568–576
Acknowledgements
The present research was supported by Department of Science and Technology, Government of India through ‘Promotion of University Research and Scientific Excellence’ grant.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hunjan, M.S., Kamboj, I., Lore, J.S. et al. Expression of defense related enzymes in rice near isogenic lines IRBB4 and IRBB7 challenged with Xanthomonas oryzae pv. oryzae at elevated temperature. Indian Phytopathology 74, 33–43 (2021). https://doi.org/10.1007/s42360-020-00304-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42360-020-00304-0