Abstract
Characterization of the Cryphonectria parasitica population was initially done by a phenotypical assessment of 40 in vitro grown isolates obtained from 52 healing cankers collected from eight important chestnut-growing provinces of Turkey. The results of Bavendamm test, often correlated indirectly to hypovirulence, suggested 31 possibly hypovirulent and 9 virulent isolates. PCR tests amplified two regions of ORFs A and B of Cryphonectria hypovirus 1 (CHV-1) from 36 of 40 isolates. The PCR test confirmation was more sensitive than the Bavendamm test. Partial ORFA sequencing revealed 36 CHV-1 haplotypes belonging to Italian subtype (I), with all hypovirulent isolates being of EU-1 vc type. The CHV-1 from 10 native EU-1 isolates were first transferred to six European vc type testers, EU-2, EU-3, EU-5, EU-7, EU-26, and EU-44, having heteroallelism at one vic locus. The presence of the vic locus difference generally reduces virus transmission. The easiest and highest frequency virus transfer was obtained by vic4 and vic6 allelic differences, while the differences vic2 and vic7 made the transfer more challenging. Finally, in this study we successfully transferred CHV-1 to an EU-1 isolate obtained from the Bursa province to an EU-12 European tester isolate.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chestnut, Castanea sativa Mill., is not only a forest tree comprising 81.232 ha pristine forest (OGM 2021) in Turkey, but with the production of 76.045 tons (FAOSTAT D 2023), is also an important fruit commodity. One of the biological constraints for its production is chestnut blight, a fungal disease caused by Cryphonectria parasitica (Murrill) M.E. Barr. The disease occurs in all chestnut-growing countries. The agent was first noted in Turkey in 1967 (Akdoğan and Erkam 1968; Delen 1975) and has spread in all parts of Turkey since then (Çeliker and Onoğur 2001; Erincik et al. 2008; Akıllı et al. 2009; FAO 2014).
Cryphonectria parasitica affects all the above-ground parts of the trees, causing cankers on the branches and on the stems, eventually producing severe damage to the trees. Control of the disease by cultural and chemical methods is not feasible since the pathogen infects the barks, and preventing the infection is not an easy task.
In the second half of the twentieth century, natural healing of the cankers was reported originally from Europe and this phenomenon was attributed to infections by a virus, Cryphonectria hypovirus 1 (CHV-1), that reduces the virulence of its fungal host (Grente 1965; Choi and Nuss 1992; Heiniger and Rigling 1994; Milgroom and Cortesi 2004). Cryphonectria parasitica can be infected by CHV-1, a cytoplasmic RNA mycovirus in the family Hypoviridae, which can cause hypovirulence (Anagnostakis 1977; Hilman et al. 2004). After the biological control potential of hypoviruses was discovered in Italy in the 1950s, it became a common tool for disease control. Since then, various mycoviruses, including CHV-1, CHV-2, CHV-3, and CHV-4, have been identified for chestnut blight control in North America; however, CHV-4 does not cause hypovirulence. To trigger biological control when no natural hypovirulence exists, the hypovirulent strain of C. parasitica is applied to the active cankers and the virus is expected to infect the resident canker-causing strain and reduce its virulence. For this reason, the hypovirulent strain have to be vegetatively compatible (vc) with the canker-causing one.
Cryphonectria parasitica exhibits a diversity of vc types in Europe, and this variation changes according to the countries. In Turkey, a few vc types of C. parasitica were previously determined (Akıllı et al. 2009; Mangıl et al. 2018), with EU-1 and EU-12 being the most common (Akıllı Şimşek et al. 2019). Anyway, natural hypovirulence was found only in EU-1 vc types.
Vegetative incompatibility has an important role in restricting the horizontal transmission of the virus (Nuss 1992; Cortesi et al. 2001). Cortesi et al. (2001) found significant differences in virus transmission isolates investigating the effect of allelic variation for six vic loci in C. parasitica.
CHV-1 population was previously studied in detail in the Black Sea region of Turkey, and the hypovirus was found only in EU-1 vc type of C. parasitica, with subtype I being prevalent and subtype F2 appearing in lesser amounts (Akıllı et al. 2013). Another study, comprising the western Anatolia and Black Sea regions, was done more recently, and a laboratory for biological control of chestnut canker was established in Bolu (Akıllı Şimşek et al. 2019).
Anyway, further studies on vc types present in Turkey are needed to extend biological control studies of chestnut canker. The aims of this study were (i) to characterize CHV-1, naturally occurring in important chestnut-growing provinces of Turkey, (ii) to transfer CHV-1 to vc types of EU-2, EU-3, EU-5, EU-7, EU-26, EU-44, which are either present or likely to be present, and, most importantly, to EU-12 vc type which does not display natural hypovirulence in Turkey.
Materials and methods
Fungal isolates and determination of the hypovirulent isolates
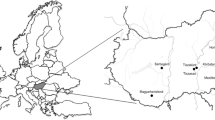
The 40 C. parasitica isolates used in the study were obtained from 52 healing cankers collected from eight provinces (Bursa, Düzce, Çanakkale, Kastamonu, Samsun, Sinop, Yalova, and Zonguldak) (Fig. 1). Vegetative-compatibility types of 40 isolates used in this study were determined with the classical method and multiplex PCR based on vic loci as previously described (Çakar et al. 2023).
Bavendamm test
A total of 40 white or cream-colored isolates were submitted to phenol oxidase (laccase) test. The test was performed by growing the isolates on an agar medium containing tannic acid (Bavendamm’s medium), as described by Rigling et al. (1989). Bavendamm test was carried out with two replications, and isolates with weak color change were considered as possible hypovirulent. A CHV-1-free EU-1 tester isolate was used as a virulent control.
Determination of the presence of Cryphonectria hypovirus 1
Total RNA isolation
For total RNA isolation, hypovirulent C. parasitica isolates were cultivated on PDA at 25 °C for 7 days. Approximately 50 mg of mycelium with conidia were scraped from the agar surface and treated with liquid nitrogen. RNA extraction was performed using NucleoZOL (Macherey-Nagel, Dueren, Germany) according to the manufacturer’s guidelines and confirmed by agarose gel electrophoresis. The resultant RNA was measured with a DS-11 FX + spectrophotometer (Denovix Inc., Wilmington, DE, USA) and stored at − 80 °C till used.
cDNA synthesis and amplifying ORFA and ORFB
The first strand cDNA synthesis kit (Eurx Ltd, Gdańsk, Poland) was employed to reverse transcribe 1 µg total RNA. Reverse transcription was carried out in 20 µl of reaction mixture containing 1 µg total RNA, 4 µl 5× cDNA Buffer, 1 µl mix RT (reverse transcriptase), 1 µl oligo (dT), 1 µl random hexamers, 12 µl nuclease-free water. The thermal cycling conditions of the reaction for cDNA were 10 min at 20 °C, 40 min at 50 °C, 5 min at 85 °C, and finally, 10 min at 4 °C.
PCR to amplify ORFA and ORFB regions of CHV-1 was carried out using hvep1/2 primers (Gobbin et al. 2003) and 12 F/12R primers (Feau et al. 2004), respectively. PCR amplification was conducted in 50 µl of reaction mixture. The thermal cycling conditions were as follows: 2 min at 94 °C, 40 cycles of 1 min at 94 °C, 1.5 min at 50 °C and 2 min at 72 °C, and 8 min at 72 °C for a final extension. The PCR products were fractionated by electrophoresis in a 1% agarose gel, stained with ethidium bromide, and visualized under UV.
Sequencing and phylogenetic analysis of CHV-1
ORFA region of CHV-1 isolates was chosen for sequencing. The amplified ORFA products were bi-directionally sequenced by Macrogen Inc. (Seoul, Korea) with hvep1/2 primers. The resultant sequences were edited, and consensus sequences were assembled by the SeqMan and MegAlign modules of DNASTAR software version 7.1.0 (DNASTAR Inc., Madison, Wisconsin, USA). The generated ORFA sequences were deposited in the GenBank nucleotide database (https://www.ncbi.nlm.nih.gov/genbank/, accessed on 10 Feb 2023).
For phylogenetic analyses, the ORFA sequences from this study and additional reference sequences obtained from GenBank (Supplementary Table ST1) were aligned using the MAFFT v.7 online interface (Katoh et al. 2019; http://mafft.cbrc.jp/alignment/server/, accessed on 10 Feb 2023). Maximum likelihood (ML) phylogenetic tree was inferred for ORFA datasets using the command-line version of IQ-TREE 1.6.7 (Nguyen et al. 2015) with an ultrafast bootstrap approximation approach (UFBoot2) implemented with 1000 replicates (Hoang et al. 2018). The CIPRES Science Gateway V 3.3 was used for the analyses (https://www.phylo.org/, accessed on 10 Feb 2023).
Transfer of the hypovirus to various vc types
Virus transmission between vc types, which have heteroallelism at one vic locus compared to EU-1 vc, was performed by placing a mycelial plug from a virus-infected donor and virus-free recipient isolates 2–3 mm apart on a PDA plate and incubating for 10 days at 25 °C in the dark as previously described (Anagnostakis and Day 1979; Cortesi et al. 2001). Pairs of hypovirus-infected and hypovirus-free isolates are co-cultured on PDA. Virus-infected donor isolates from only one vc type were paired with virus-free recipient isolates from six other vc types (EU-2, EU-3, EU-5, EU-7, EU-26, EU-44). Virus transmission between incompatible isolates was tested in four plates for each comparison with three repetitions (total of 12 replications). Transfer of the hypovirus was first observed by color changes of the cultures at far edges from the inoculation points. Then horizontal virus transmission was verified by PCR amplification of the ORFA and ORFB regions of Cryphonectria hypovirus 1.
Hypovirulent BU06 (EU-1) was used as a source for CHV-1 transfer to the EU-12 vc tester isolate. Initially, the strain EU-3 (with vic6 difference from EU-1) was previously transformed by the BU06 strain, which was utilized as the donor. Then the virus in EU-3 was gradually transferred to European vc testers EU-30, EU-29, and EU-12 vc type, respectively. The virus transmission was confirmed by PCR tests.
Statistical analysis
To establish the significance of the variations in the values among transmission rates with donor isolates and vic loci, an analysis of variance (ANOVA) was employed, followed by Tukey’s Honestly Significant Difference (HSD) at P ≤ 0.05 with JMP® 16 software (SAS Institute Inc, Cary, NC, USA).
Results
Characteristics of the hypovirulent isolates and their distribution
All 40 isolates grown on PDA medium with methionine and biotin displayed either whitish or creamy phenotype (Table 1). Most isolates (31 out of 40) produced weak color change on the Bavendamm test media. Thirty-six isolates resulted CHV-1 infected as they produced 391 bp and 741 bp DNA bands generated by hvep1 and hvep2 primers coding ORFA and 12 F and 12R primers for ORFB regions, respectively, in RT-PCR tests from the total RNA (Fig. 2), while CHV-1 was not determined in remain four isolates. The GenBank accession numbers of the CHV-1 ORFA nucleotide sequences generated in this study are listed in Table 2. Four of the all 40 isolates were CHV-1 negative in RT-PCR tests.
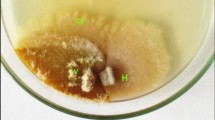
Agarose gel electrophoreses of RT-PCR products of ORFA (1–8) and ORFB (9–16) from some of Cryphonectria parasitica hypovirus 1 (CHV-1) infected Cryphonectria parasitica isolates. C. parasitica isolates are: 1, 9: KS04; 2, 10: AM03; 3, 11: Si02; 4, 12: KS01; 5, 13: BL13; 6, 14: BL01; 7, 15: BL09; and 8, 16: Si04. M: 100 bp DNA ladder (Solis BioDyne, Tartu, Estonia). Primers used were hvep1/hvep2 and 12 F/12R, respectively
Subtypes of Cryphonectria hypovirus 1
Cryphonectria hypovirus 1 subtypes were determined by constructing an ML phylogenetic tree using the sequences of the ORFA region of the Turkish 36 isolates and sequences representatives of various CHV-1 subtypes (Fig. 3). As shown in Fig. 3, all the isolates from this study grouped with reference subtype I isolates forming a cluster with high bootstrap values. Sequences from this study were most similar to European CHV-1 subtype I, but none was identical to previously sequenced CHV-1 isolates listed in Table 2. Nevertheless, some of isolates sequenced in this study shared 100% nucleotide sequence identity among themselves (BL08, BL14 and YL10; YL01 and AM06; AM02 and AM05; CK01, Si03 and Si03).
Transmission of Cryphonectria hypovirus 1 to virulent isolates of Cryphonectria parasitica belonging to different vc types
Ten hypovirulent isolates from Samsun (AM03, AM04), Düzce (BL01, BL03), Bursa (BU06), Çanakkale (CK01), Kastamonu (KS01), Sinop (Si02), Yalova (YL09), and Zonguldak (ZN04), all having EU-1 vc type with CHV-1 subtype I, were used for virus transmission by dual inoculation of the hypovirulent isolate with the virulent tester EU vc types. When compatible, hypovirulence was transmitted on the far edges of the tester isolate (Fig. 4). Hypovirus transmission was initially checked by visual observation of colony morphology and color and confirmed by RT-PCR assays with the same primer pairs (hvep1/hvep2 and 12 F/12R) used in previous phases of this study.
Transmission of the hypovirus to the virulent tester EU isolates showed variation based on the vic allelic differences. In the experiments involving ten CHV-1 infected isolates and six different vic genotypes of C. parasitica, all but one donor isolates successfully transferred the hypovirus to all recipients, EU-3, EU-5, EU-26, and EU-44 genotypes of the fungus, but one of them did not transfer the same virus to EU-2 and EU-7 vc types (Table 1).
Differences in vic loci have been found to be important for horizontal virus transmissions. A statistically significant result has also been found among donor isolates (Table 3). Donor isolate had a significant effect on the virus transmission rate of isolates (P < 0.001); also, there was a significant effect of EU vc tester isolates (P < 0.001) on the transmission rate (Table 3).
Transfer of CHV-1 to the mentioned European vc types was also verified by PCR of the ORFA and ORFB regions of the EU-3, EU-5, and EU-26 vc types (Supplementary Fig. S1).
Transfer of CHV-1 to EU-12 vc type was done by using hypovirulent EU-3 previously transfected by hypovirulent BU06 strain, as outlined in Table 4.
Discussion
Chestnut canker is known to occur in Turkey since 1967 and to induce severe disease in all chestnut-growing areas of the country (Akdoğan and Erkam 1968; Delen 1975; Coşkun et al. 1999; Çeliker and Onoğur 2001; Gürer et al. 2001; Erincik et al. 2008; Akıllı et al. 2009). Since the other control methods are not very practical or feasible, biological control using hypovirulent fungal strains infected by a particular type of viruses called hypoviruses has been widely studied and used in the world (Anagnostakis 1982; MacDonald and Fulbright 1991; Robin et al. 2000; Milgroom and Cortesi 2004; Heiniger and Rigling 2009; FAO 2014; Şimşek et al. 2019; Çakar et al. 2020). In practice, hypovirulent C. parasitica strains are applied to the peripheries of the active cankers, so, in case of compatibility, that they can transfer the hypovirus (CHV-1) to the virulent strain. Thus, CHV-1 infection reduces the virulence of the fungal host, and the tree eventually recovers gradually (Choi and Nuss 1992). To achieve success in biological control, the vc types of the hypovirulent and virulent strains must be compatible since the hypovirus transfer of occurs by hyphal anastomosis. For this reason, for successful application of CHV-1, vc types of the local C. parasitica population are first determined and then properly selected hypovirulent strains are applied. Vc types of virulent and hypovirulent C. parasitica isolates have been found in almost all of the chestnut-growing areas of Turkey, represented by two EU-1 and EU-12, with EU-1 being more prevalent (Çeliker and Onoğur 2001; Gürer et al. 2001; Erincik et al. 2011).
Natural hypovirulence is mainly occurring along the Black Sea region of Turkey but not in the Aegean region, which is the main good-quality chestnut-producing part of Turkey (Çakar et al. 2021). Both, EU-1 and EU-12 vc types are distributed at almost equal rates (Akıllı Şimşek et al. 2019; Erincik et al. 2008, 2011), whereas some new vc types, such as EU-2, EU-3, EU-5, and EU-7, with one vic allelic difference from EU-1, were reported only in some locations (Akıllı et al. 2009; Daldal 2018; Mangıl and Erincik 2018; Hatipoğlu 2019).
Thirty-six hypovirulent C. parasitica isolates obtained from eight provinces used to study the population diversity of CHV-1 based on partial ORF-A sequencing resulted to be subtype I, which is in agreement with what previously reported (Akıllı et al. 2013). Subtype I was also reported as the most effective and common biological control agent in Turkey and elsewhere (Robin et al. 2010; Akıllı et al. 2013; Ringling and Prospero 2018). This is not unexpected given that CHV-1 subtype I is the most prevalent in Europe while other subtypes have a much more restricted distribution. Indeed, compared to other CHV-1 subtypes, subtype I of the virus has a stronger propensity for propagation and establishment, which may contribute to its greater invasiveness (Robin et al. 2010).
CHV-1 transfer from EU-1 to some other vc types having one allelic difference showed variation concerning efficiency. Transfer to vc types EU-3, EU-5, and EU-26, which have allelic differences at vic6, vic4, and vic1 loci, respectively, was done easily; the number of transfers from 10 donor isolates was 10, 10, and 10, respectively. Transfer of the virus to vc types having one allelic difference at vic2 and vic7, such as EU-2 and EU-7, did not happen in the first attempt but was finally accomplished after many repetitions. Difficulty for the virus transfer to fungal isolates belonging to these vc types was also emphasized by Cortesi et al. (2001) and Choi et al. (2012). Transfer of the virus from vc type EU-29 to EU-12, which has one allelic difference at vic7 locus (Table 4), was achieved only after a total of 30 repeats/attempts. Hypovirus transfer from a native EU-1 hypovirulent isolate to the EU-12 virulent European tester isolate was done for the first time in Turkey, and the biological control studies against EU-12 vc type could be facilitated by the availability and use of this isolate.
The restrictive effect of heteroallelism at vic loci on virus transmission, observed in previous studies (Cortesi et al. 2001; Papazova et al. 2008), has also been confirmed in this study. Additionally, previous published data suggest that there is an effect of hypovirulent isolates on transmission. QingChao et al. (2009) reported that different hypovirulent isolates used as donors had different virus transmission capabilities.
In conclusion, it has been determined that vic loci and the allelic variations in these loci have an impact on virus transmission. The hypovirus transmission from a native EU-1 hypovirulent isolate to the EU-12 virulent European tester isolate was successfully done for the first time for a local Turkish isolate. This has been an important finding for biological control studies with next step of validating these results in the open field.
References
Akdoğan S, Erkan E (1968) Dikkat! Kestane kanseri görüldü. Tomurcuk 1:4–5 (in Turkish)
Akıllı S, Katırcıoğlu YZ, Maden S (2009) Vegetative compatibility types of Cryphonectria parasitica, chestnut blight agent. Black Sea Region Pathol 39(6):390–396. https://doi.org/10.1111/j.1439-0329.2009.00601.x
Akıllı S, Ulubaş-Serçe Ç, Katırcıoğlu YZ, Maden S, Rigling D (2013) Characterisation of hypovirulent isolates of the chestnut blight fungus, Cryphonectria parasitica from the Marmara and Black Sea regions of Turkey. Eur J Plant Pathol 135:323–334. https://doi.org/10.1007/s10658-012-0089-z
Akıllı Şimşek S, Katircioğlu YZ, Borst N, Çakar D, Prospero S, Rigling D, Maden S (2019) Identification and characterisation of hypovirus-infected Cryphonectria parasitica isolates from biological control plots in İzmir, Kütahya, and Sinop. Turk J Agric for 43(6):527–537. https://doi.org/10.3906/tar-1901-82
Anagnostakis S (1977) Vegetative incompatibility in Endothia Parasitica. Exp Mycol 1:306–316. https://doi.org/10.1016/S0147-5975(77)80006-6
Anagnostakis SL (1982) Biological control of chestnut blight. Science 215(4532):466–471. https://doi.org/10.1126/science.215.4532.466
Anagnostakis SL, Day PR (1979) Hypovirulence conversion, in Endothia Parasitica. Phytopathology 69:1226–1229. https://doi.org/10.1094/Phyto-69-1226
Çakar D, Akıllı Şimşek A, Katırcıoğlu YZ, Maden S (2020) Biological control of chestnut canker caused by Cryphonectria Parasitica (Murr.) by hypovirulent strains at selected orchards in İzmir and Aydın provinces of Turkey. Current Researches in Science and Mathematics Sciences, 1st edn. Ivpe Cetinje, Montenegro, pp 12–39
Çakar D, Akıllı S, Can T, Katircioğlu YZ, Maden S (2021) Determination of Vc types of chestnut canker agent Cryphonectria parasitica and evulation of hyphovirulence in chestnut forest areas in Bolu and İzmir Regional directorates of Forestry. IJAWS 7(1):41–55. https://doi.org/10.24180/ijaws.818343
Çakar D, Özer G, Akıllı Şimşek S, Maden S (2023) Determination of vc and mating types of Cryphonectria parasitica isolates by multiplex PCR and their genetic diversity in 13 chestnut-growing provinces of Turkey. https://doi.org/10.1111/efp.12813. For Pathol e12813
Çeliker NM, Onoğur E (2001) Evaluation of hypovirulent isolates of Cryphonectria parasitica for the biological control of chestnut blight. Snow Landsc Res 76:378–382
Choi GH, Nuss DL (1992) Hypovirulence of chestnut blight fungus conferred by an infectious viral cDNA. Science 257(5071):800–803. https://doi.org/10.1126/science.1496400
Choi GH, Dawe AL, Churbanov A, Smith ML, Milgroom MG, Nuss DL (2012) Molecular characterization of vegetative incompatibility genes that restrict hypovirus transmission in the chestnut blight fungus Cryphonectria parasitica. Genetics 190(1):113–127. https://doi.org/10.1534/genetics.111.133983
Cortesi P, McCulloch CE, Song H, Lin H, Milgroom MG (2001) Genetic control of horizontal virus transmission in the chestnut blight fungus, Cryphonectria parasitica. Genetics 159(1):107–118. https://doi.org/10.1093/genetics/159.1.107
Çoşkun H, Turchetti T, Maresi G, Santagada A (1999) Preliminary investigations into Cryphonectria parasitica (Murr) Barr isolates from Turkey. Phytopathol Mediterr 38:101–110
Daldal M, Erincik Ö, Wall JR (2018) Geographical distribution of vegetative compatibility and mating types of Cryphonectria parasitica in İzmir, Manisa and Denizli provinces in western Turkey. Pathol 48(5):e12444. https://doi.org/10.1111/efp.12444
Delen N (1975) Distribution and the biology of chestnut blight (Endothia Parasitica) (Murrill) Anderson and Anderson. J Turkish Phytopath 4:93–113
Deng Q, Ye Y, Miao M, Fang Q, Li T, Wang K (2009) The horizontal transmission of Cryphonectria hypovirus 1 (CHV1) is affected by virus strains. Sci Bull 54(17):3053–3060. https://doi.org/10.1007/s11434-009-0368-z
Erincik O, Doken MT, Acikgoz S, Ertan E (2008) Characterisation of Cryphonectria parasitica isolates collected from Aydin province in Turkey. Phytoparasitica 36:249–259. https://doi.org/10.1007/BF02980771
Erincik Ö, Özdemir Z, Durdu ÖF, Döken MT, Açıkgöz S (2011) Diversity and spatial distribution of vegetative compatibility types and mating types of Cryphonectria parasitica in the Aydın Mountains, Turkey. Eur j Plant Pathol 129:555–566. https://doi.org/10.1007/s10658-010-9719-5
FAO (2014) Management of chestnut blight and for improving forest health and vitality (2012–2014), Report TCP/TUR/615676
FAOSTAT D (2023) Food and Agriculture Organization Statistical Database. Available online: http://www.fao.org/faostat/en/#data/QC/. Accessed 21 Jan 2023
Gobbin D, Hoegger PJ, Heiniger U, Rigling D (2003) Sequence variation and evolution of Cryphonectria hypovirus 1 (CHV-1) in Europe. Virus Res 97(1):39–46. https://doi.org/10.1016/s0168-1702(03)00220-x
Grente J (1965) Les formes hypovirulentes d’Endothia Parasitica et les espoirs de lutte contre le chancre du chataignier. CR Acad Agric France 51:1033–1037
Gürer M, Turchetti T, Biagioni P, Maresi G (2001) Assessment and characterisation of Turkish hypovirulent isolates of Cryphonectria parasitica (Murr) Barr. Phytopathol Mediterr 40(3):265–275. https://doi.org/10.14601/Phytopathol_Mediterr-1612
Hatipoğlu E (2019) Detection of distribution of vegetative incompatibility genes (vic) of Cryphonectria parasitica, population in the Eastern Black Sea region. Master’s thesis. Aydın Menderes University
Heiniger U, Rigling D (1994) Biological control of chestnut blight in Europe. Annu Rev Phytopathol 32:581–599. https://doi.org/10.1146/annurev.py.32.090194.003053
Heiniger U, Rigling D (2009) Application of the Cryphonectria hypovirus (CHV-1) to control the chestnut blight, experince from Switzerland. Acta Hortic 815:233–245. https://doi.org/10.17660/ActaHortic.2009.815.31
Hoang DT, Chernomor O, von Haeseler A, Minh BQ, Vinh LS (2018) UFBoot2: improving the ultrafast bootstrap approximation. Mol Biol Evol 35:518–522. https://doi.org/10.1093/molbev/msx281
Katoh K, Rozewicki J, Yamada KD (2019) MAFFT online service: multiple sequence alignment, interactive sequence choice and visualisation. Brief Bioinform 20:1160–1166. https://doi.org/10.1093/bib/bbx108
Lin H, Lan X, Liao H, Parsley TB, Nuss DL, Chen B (2007) Genome sequence, full-length infectious cDNA clone, and mapping of viral double-stranded RNA accumulation determinant of hypovirus CHV1-EP721. J Virol 81:1813–1820. https://doi.org/10.1128/JVI.01625-06
MacDonald WL, Fulbright DW (1991) Biological control of chestnut blight: use and limitations of transmissible hypovirulence. Plant Dis 75(7):656–661. https://doi.org/10.1094/PD-75-0653
Mangıl E, Erincik Ö (2018) Sexual reproduction contributes to vegetative compatibility type diversity in the population of Cryphonectria parasitica in the East Black Sea region of Turkey. Australas Plant Pathol 47:301–310. https://doi.org/10.1007/s13313-018-0556-x
Milgroom GM, Cortesi P (2004) Biological control of chestnut blight with hypovirulence; a critical analysis. Annu Revi Phytopathol 42:311–338. https://doi.org/10.1146/annurev.phyto.42.040803.140325
Nguyen LT, Schmidt HA, Von Haeseler A (2015) IQTREE: a fast and effective stochastic algorithm for estimating maximum likelihood phylogenies. Mol Biol Evol 32:268–274. https://doi.org/10.1093/molbev/msu300
Nuss DL (1992) Biological control of chestnut blight–an example of virus-mediated attenuation of fungal pathogenesis. Microbiol Rev 56:561–576. https://doi.org/10.1128/mr.56.4.561-576.1992
OGM (2021) Orman Genel Müdürlüğü, Türkiye Orman Varlığı (in Turkish)
Rigling D, Heiniger U, Hohl HR (1989) Reduction of laccase activity in dsRNA-containing hypovirulent strains of Cryphonectria (Endothia) parasitica. Phytopathology 79:219–223. https://doi.org/10.1094/Phyto-79-219
Rigling D, Borst N, Cornejo C, Supatashvili A, Prospero S (2018) Genetic and phenotypic characterisation of Cryphonectria hypovirus 1 from Eurasian Georgia. Viruses 10(12):687. https://doi.org/10.3390/v10120687
Robin C, Anziani C, Cortesi P (2000) Relationship between biological control, incidence of hypovirulence, and diversity of vegetative compatibility types of Cryphonectria parasitica in France. Phytopathology 90(7):730–737. https://doi.org/10.1094/PHYTO.2000.90.7.730
Robin C, Lanz S, Soutrenon A, Rigling D (2010) Dominance of natural over released biological control agents of the chestnut blight fungus Cryphonectria parasitica in south-eastern France is associated with fitness-related traits. Biol Control 53(1):55–61. https://doi.org/10.1016/j.biocontrol.2009.10.013
Acknowledgements
This research was produced as part of the PhD thesis of Deniz Çakar. The authors appreciate funding of this study by Çankırı Karatekin University Research Fund (Project no: FF210621D05). We thank the Western Black Sea Forestry Research Institute and the General Directory of Forestry. We would also like to thank Dr. Daniel Rigling and Dr. Paolo Cortesi for sharing European tester strains. We would also like to thank Editor Dr. Sead Sabanadzovic for his contribution to the article. The help of Aslı Özcan on the design of the map in Fig. 1 is also appreciated. We also thank Dr. Ali Çelik for help with the process of sequencing for isolates.
Funding
This work was supported by Çankırı Karatekin University Research Fund (Project no: FF210621D05).
Open access funding provided by the Scientific and Technological Research Council of Türkiye (TÜBİTAK).
Author information
Authors and Affiliations
Contributions
DÇ, Methodology, Investigation, Experimental Work, Field Study, Molecular Analysis, Formal Analysis, Writing, GÖ, Methodology, Molecular Analysis, Formal Analysis, Writing, SAŞ, Conceptualization, Methodology, Investigation, Writing, SM, Conceptualization, Writing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Çakar, D., Özer, G., Şimşek, S.A. et al. Characterisation of Cryphonectria hypovirus 1 strains in Turkey and their transmission to various vegetative compatibility types of Cryphonectria parasitica. J Plant Pathol (2024). https://doi.org/10.1007/s42161-024-01691-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42161-024-01691-3