Abstract
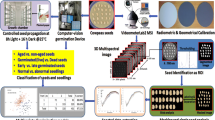
The science of seed pathology has been established since the development and application of standardized methods for assessing seed health to meet the needs of the seed industry and associated regulatory entities. Despite seed health testing being a routine operation in most countries, results of testing often vary from one laboratory to another. We evaluated computer vision using red–green–blue (RGB) imagery and machine learning algorithms to detect seed-borne fungi on common bean (Phaseolus vulgaris L.) seeds. Seeds of common bean were submitted to the standard blotter test for 7 days, followed by fungal identification using a stereo- and light microscope. A scanning electron microscope was used to confirm fungal identity. Images of seed-borne fungi were captured from a distance of approximately 5 cm. Seventeen spectral indices were derived from the RGB images. Targets of interest in the images were obtained using spatial polygons with attributes used for training six machine learning algorithms (random forest (rf), rpart, rpart1SE, rpart2, naive Bayes, and svmLinear2), with a total of five replicates per target that were identified as Aspergillus flavus, A. niger, A. ochraceus, Penicillium sp., Mucor sp, Rhizopus sp, Fusarium sp, Rhizoctonia sp., common bean tegument, and blotter paper. After a fivefold cross-validation process and a confusion matrix, the rf algorithm had the highest prediction success to detect the targets (accuracy 0.80 and Kappa 0.77, respectively). The brightness index was the most important variable in predicting targets by the rf. Using the rpart1SE algorithm, a decision tree for target identification was obtained with an accuracy of 0.70 and a Kappa value of 0.66, respectively. The rf, svmLinear2, and rpart1SE were found to be the most robust classification algorithms for predicting identification of the fungal species and other targets associated with common bean seed blotter tests using digital RGB images and indices. The use of spectral indices derived from RGB imagery has extended the training capability of algorithms, demonstrated by the importance of the variables and decision tree used for target prediction by the rf and rpart1SE algorithms, respectively.












Similar content being viewed by others
Data availability
The datasets generated during and/or analyzed in the current study are available from the corresponding author on reasonable request.
References
Alves MC, Pozza EA (2009) Scanning electron microscopy applied to seed-borne fungi examination. Microscopy Research and Technique 72:482–488
Alves MC, Pozza EA, Machado JC, Carvalho MGG (2006) Development and evaluation of an expert system for seed-borne fungi identification in the seed health analysis. Revista Brasileira De Sementes 28:176–186
Athiraja A, Vijayakumar P (2021) Banana disease diagnosis using computer vision and machine learning methods. Journal of Ambient Intelligence and Humanized Computing 12:6537–6556
Birth GS, McVey GR (1968) Measuring the color of growing turf with a reflectance spectrophotometer. Agronomy Journal 60:640–643
Benech-Arnold R, Semmartin M, Oesterheld M (2012) Seed science in the 21st century: its role in emerging economies. Seed Science Research 22:S3–S8
Boelt B, Shrestha S, Salimi Z, Jørgensen JR, Nicolaisen M, Carstensen JM (2018) Multispectral imaging–a new tool in seed quality assessment? Seed Science Research 28:222–228
Bona E, Marquetti I, Link JV et al (2017) Support vector machines in tandem with infrared spectroscopy for geographical classification of green arabica coffee. LWT-Food Science and Technology 76:330–336
Boochs F, Kupfer G, Dockter K, Kühbauch W (1990) Shape of the red edge as vitality indicator for plants. Remote Sensing 11:1741–1753
Breiman L, Friedman J, Olshen R, Stone C (1984) Cole statistics/probability series. In: Classification and regression tree. Wadsworth & Brooks, Pacific Grove, p 358
Carvalho Alves M, Pozza EA, de Bonfim Costa JC et al (2011) Adaptive neuro-fuzzy inference systems for epidemiological analysis of soybean rust. Environmental Modelling & Software 26:1089–1096
Escadafal R (1994) Soil spectral properties and their relationships with environmental parameters - examples from arid regions. In: Imaging spectrometry — a tool for environmental observations. Euro- courses: Remote Sensing, pp 71–87
Gai R, Chen N, Yuan H (2021) A detection algorithm for cherry fruits based on the improved YOLO-v4 model. Neural Computing and Applications 0123456789
Gitelson A, Stark R, Grits U et al (2002) Vegetation and soil lines in visible spectral space: a concept and technique for remote estimation of vegetation fraction. International Journal of Remote Sensing 23:2537–2562
Henrich V, Götze E, Jung A, Sandow C, Thürkow D, Gläßer C (2009) Development of an online indices database: motivation, concept and implementation. In: Proceedings of the 6th EARSel Imaging Spectroscopy SIG Workshop Innovative Tool for Scientific and Commercial Environment Applications, Tel Aviv, Israel, pp 16–18
Henrich V, Krauss G, Götze C, Sandow C (2011) The indexdatabase. Https://www.indexdatabase.de/
Huang M, Wang Q, Zhu Q et al (2015) Review of seed quality and safety tests using optical sensing technologies. Seed Science and Technology 43:337–366
Hunt ER, Daughtry CST, Eitel JUH, Long DS (2011) Remote sensing leaf chlorophyll content using a visible band index. Agronomy Journal 103(4):1090–1099. https://doi.org/10.2134/agronj2010.0395
Jensen JR (2005) Introductory Digital Image Processing: A Remote Sensing Perspective, 3rd edn. Pear- son Prentice Hall, Upper Saddle River, NJ
Kinnikar A, Desai P, Jahagirdar S (2015) Identification and detection of seed borne diseases of soybean using image processing-a survey. International Journal of Emerging Technology in Computer Science and Electronics 14:363–368
Kohavi R (1995) A study of cross-validation and bootstrap for accuracy estimation and model selection. In: IJCAI’95: Proceedings of the 14th international joint conference on artificial intelligence. Montreal, Canada, pp 1137–1145
Koklu M, Ozkan IA (2020) Multiclass classification of dry beans using computer vision and machine learning techniques. Computers and Electronics in Agriculture 174:105507
Kuhn M (2008) Building predictive models in R using the caret package. Journal of Statistical Software 28:1–26
Kuhn M (2014) Futility analysis in the cross-validation of machine learning models
Kumar R, Baloch G, Pankaj ABB, Bhatti J (2021) Fungal blast disease detection in rice seed using machine learning. International Journal of Advanced Computer Science and Applications 12:248–258
Lacaux JP, Tourre YM, Vignolles C et al (2007) Classification of ponds from high-spatial resolution remote sensing: application to rift valley fever epidemics in senegal. Remote Sensing of Environment 106:66–74
Lasso E, Thamada TT, Meira CAA, Corrales JC (2015) Graph patterns as representation of rules extracted from decision trees for coffee rust detection. In: Research conference on metadata and semantics research. Springer, pp 405–414
Liu J, Wang X (2020) Tomato diseases and pests detection based on improved Yolo V3 convolutional neural network. Frontiers in Plant Science 11:1–12
Louhaichi M, Borman MM, Johnson DE (2001) Spatially located platform and aerial photography for documentation of grazing impacts on wheat. Geocarto Int 16:65–70
Machado JC, Jaccoud Filho DS, Langerak CJ (2002) Seed-borne fungi: a contribution to routine seed health analysis. International Seed Testing Association
Madeira J, Bedidi A, Cervelle B et al (1997) Visible spectrometric indices of hematite (hm) and goethite (gt) content in lateritic soils: the application of a thematic mapper (tm) image for soil-mapping in Brasilia, brazil. International Journal of Remote Sensing 18:2835–2852
Mathieu R, Pouget M, Cervelle B, Escadafal R (1998) Relationships between satellite-based radiometric indices simulated using laboratory reflectance data and typic soil color of an arid environment. Remote Sensing of Environment 66:17–28
Medeiros AD, Capobiango NP, Silva JM et al (2020) Interactive machine learning for soybean seed and seedling quality classification. Scientific Reports 10:1–10
Miranda JR, Alves MC (2020) The use of machine learning in digital processing of satellite images applied to coffee crop. CAB Reviews: Perspectives in Agriculture, Veterinary Science, Nutrition and Natural Resources 15
Miranda JR, Alves MC, Pozza EA, Santos Neto H (2020) Detection of coffee berry necrosis by digital image processing of Landsat 8 oli satellite imagery. International Journal of Applied Earth Observation and Geoinformation 85:101983
Mohanty SP, Hughes DP, Salathé M (2016) Using deep learning for image-based plant disease detection. Frontiers in Plant Science 7:1419
Motohka T, Nasahara KN, Oguma H, Tsuchida S (2010) Applicability of green-red vegetation index for remote sensing of vegetation phenology. Remote Sensing 2:2369–2387
Neergaard P (2017) Seed Pathology. Macmillan International Higher Education
Ouhami M, Hafiane A, Es-Saady Y, Hajji ME, Canals R (2021) Computer vision, IoT and data fusion for crop disease detection using machine learning: a survey and ongoing research. Remote Sensing 13: 2486
Owomugisha G, Melchert F, Mwebaze E, Quinn JA, Biehl M (2018) Machine learning for diagnosis of disease in plants using spectral data. In: 2018 World Congress in Computer Science, Computer Engineering and Applied Computing, CSCE 2018 - Proceedings of the 2018 International Conference on Artificial Intelligence, ICAI'18, 2018. pp 9–15
Pérez-Rodríguez M, Gaiad JE, Hidalgo MJ et al (2019) Classification of cowpea beans using multielemental fingerprinting combined with supervised learning. Food Control 95:232–241
Pierozzi CG, Fujihara RT, de Souza E, S, et al (2020) Interactive key (lucid) for identification of fungi in vegetable seeds. Summa Phytopathologica 46:14–19
Ponti MP (2012) Segmentation of low-cost remote sensing images combining vegetation indices and mean shift. IEEE Geoscience and Remote Sensing Letters 10:67–70
Ponti MP, Chaves AA, Jorge FR et al (2016) Precision agriculture: using low-cost systems to acquire low-altitude images. IEEE Computer Graphics and Applications 36:14–20
Pozza EA, Maffia LA, Silva CAB et al (2018) Neuronal networks to describe epidemics of cocoa witches’ broom. Theoretical and Applied Engineering 2:1–12
Ranđelović P, Ðorđević V, Milić S et al (2020) Prediction of soybean plant density using a machine learning model and vegetation indices extracted from RGB images taken with a uav. Agronomy 10:1108
Rahman A, Cho B (2016) Assessment of seed quality using non-destructive measurement techniques: a review. Seed Science Research 26:285–305
Rego CHQ, França-Silva F, Gomes-Junior FG et al (2020) Using multispectral imaging for detecting seed-borne fungi in cowpea. Agriculture 10:361
Reudenbach C, Meyer H, Detsch F, Möller F, Nauss T, Opgenoorth L (2021) uavRst: Unmanned Aerial Vehicle R Tools. https:// www.rdocumentation.org/packages/uavRst/versions/0.5-4/
Richardson AJ, Wiegand CL (1977) Distinguishing vegetation from soil background information. Pho- Togrammetric Engineering and Remote Sensing 43:1541–1552
Richardson MJ (1990) International Seed Testing Association. In: An annotated list of seed-borne diseases, 4th edn. Zurich, Switzerland
Schneider BO, Alves MC (2020) Digital image processing and spectral data source in seed analysis. CAB Reviews 1–6
Schwartz HF, Pastor-Corrales MA (1989) Bean production problems in the tropics. CIAT, 2nd edn. Cali, Colombia
Sethy PK, Barpanda NK, Rath AK, Behera SK (2020) Image processing techniques for diagnosing rice plant disease: a survey. Procedia Computer Science 167:516–530
Sony (2002) Digital Still Camera. 33:1–44
Steinberg D (2009) CART: classification and regression trees. In: The top ten algorithms in data mining, 1st edn. Chapman; Hall/CRC, pp 193–216
Tucker CJ (1979) Red and photographic infrared linear combinations for monitoring vegetation. Remote Sensing of Environment 8:127–150
Zhu L, Spachos P, Pensini E, Plataniotis KN (2021) Deep learning and machine vision for food processing: a survey. Current Research in Food Science 4:233–249
Acknowledgements
The authors thank the Seed Pathology laboratory technician, Angela de Fátima Carvalho Santos, at the Universidade Federal de Lavras (UFLA) for her contribution to the experiments.
Author information
Authors and Affiliations
Contributions
The blotter test was prepared as routine seed health analysis in the seed pathology laboratory at UFLA. All authors processed the experimental data and designed the figures, tables, discussed the results, and drafted the manuscript. MCA captured the RGB images of the seed-borne fungi.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Pozza, E.A., de Carvalho Alves, M. & Sanches, L. Using computer vision to identify seed-borne fungi and other targets associated with common bean seeds based on red–green–blue spectral data. Trop. plant pathol. 47, 168–185 (2022). https://doi.org/10.1007/s40858-021-00485-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40858-021-00485-7