Abstract
Purpose
The prevalence of diabetes is increasing worldwide. The associations between the lipid profile and glycated hemoglobin (HbA1c), fasting glucose, and diabetes remain unclear, so we aimed to perform a cohort study and a two-sample Mendelian randomization (MR) study to investigate the causality between blood lipid profile and HbA1c, fasting glucose, and diabetes.
Methods
A total of 25,171 participants from the Taiwan Biobank were enrolled. We applied a cohort study and an MR study to assess the association between blood lipid profile and HbA1c, fasting glucose, and diabetes. The summary statistics were obtained from the Asian Genetic Epidemiology Network (AGEN), and the estimates between the instrumental variables (IVs) and outcomes were calculated using the inverse-variance weighted (IVW) method. A series of sensitivity analyses were performed.
Results
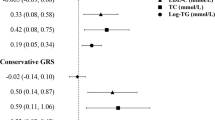
In the cohort study, high-density lipoprotein cholesterol (HDL-C) was negatively associated with HbA1c, fasting glucose, and diabetes, while the causal associations between HDL-C and HbA1c (βIVW = − 0.098, p = 0.003) and diabetes (βIVW = − 0.594, p < 0.001) were also observed. Furthermore, there was no pleiotropy effect in this study using the MR-Egger intercept test and MR-PRESSO global test.
Conclusions
Our results support the hypothesis that a genetically determined increase in HDL-C is causally related to a reduction in HbA1c and a lower risk of diabetes.





Similar content being viewed by others
Data availability
Summary data of blood lipid profiles GWAS were used in our study is available at the AGEN consortium website (https://blog.nus.edu.sg/agen/summary-statistics/lipids/).
Abbreviations
- MR:
-
Mendelian randomization
- IVW:
-
Inverse-variance weighted
- HbA1c:
-
Glycated hemoglobin
- HDL-C:
-
High-density lipoprotein cholesterol
- LDL-C:
-
Low-density lipoprotein cholesterol
- RCTs:
-
Randomized controlled trials
- IVs:
-
Instrumental variables
- SNPs:
-
Single nucleotide polymorphisms
- WHR:
-
Waist-hip ratio
- BMI:
-
Body mass index
- PRESSO:
-
Pleiotropy residual sum and outliers
- MBE:
-
Mode-based estimate
References
International Diabetes Federation diabetes atlas Tenth edition. 2021
Saeedi P, Petersohn I, Salpea P, Malanda B, Karuranga S, Unwin N et al (2019) Global and regional diabetes prevalence estimates for 2019 and projections for 2030 and 2045: results from the International Diabetes Federation Diabetes Atlas, 9(th) edition. Diabetes Res Clin Pract 157:107843
Diagnosis and classification of diabetes mellitus (2013) Diabetes Care 36(Suppl 1):S67-74
Siqueira ISL, Alves Guimarães R, Mamed SN, Santos TAP, Rocha SD, Pagotto V et al (2020) Prevalence and risk factors for self-report diabetes mellitus: a population-based study. Int J Environ Res Public Health 17:6497
Malta DC, Bernal RTI, Iser BPM, Szwarcwald CL, Duncan BB, Schmidt MI (2017) Factors associated with self-reported diabetes according to the 2013 National Health Survey. Rev Saude Publ 51:12s
Bertoldi AD, Kanavos P, França GV, Carraro A, Tejada CA, Hallal PC et al (2013) Epidemiology, management, complications and costs associated with type 2 diabetes in Brazil: a comprehensive literature review. Global Health 9:62
Zheng Y, Ley SH, Hu FB (2018) Global aetiology and epidemiology of type 2 diabetes mellitus and its complications. Nat Rev Endocrinol 14:88–98
Gudjinu HY, Sarfo B (2017) Risk factors for type 2 diabetes mellitus among out-patients in Ho, the Volta regional capital of Ghana: a case–control study. BMC Res Notes 10:324
Temneanu OR, Trandafir LM, Purcarea MR (2016) Type 2 diabetes mellitus in children and adolescents: a relatively new clinical problem within pediatric practice. J Med Life 9:235–239
Khan HA, Sobki SH, Khan SA (2007) Association between glycaemic control and serum lipids profile in type 2 diabetic patients: HbA1c predicts dyslipidaemia. Clin Exp Med 7:24–29
Laverdy OG, Hueb WA, Sprandel MC, Kalil-Filho R, Maranhão RC (2015) Effects of glycemic control upon serum lipids and lipid transfers to HDL in patients with type 2 diabetes mellitus: novel findings in unesterified cholesterol status. Exp Clin Endocrinol Diabetes 123:232–239
Wang S, Ji X, Zhang Z, Xue F (2020) Relationship between lipid profiles and glycemic control among patients with type 2 diabetes in Qingdao, China. Int J Environ Res Public Health 17:5317
Drew BG, Duffy SJ, Formosa MF, Natoli AK, Henstridge DC, Penfold SA et al (2009) High-density lipoprotein modulates glucose metabolism in patients with type 2 diabetes mellitus. Circulation 119:2103–2111
Zhu XW, Deng FY, Lei SF (2015) Meta-analysis of Atherogenic Index of Plasma and other lipid parameters in relation to risk of type 2 diabetes mellitus. Prim Care Diabetes 9:60–67
Davis PJ, Liu M, Sherman S, Natarajan S, Alemi F, Jensen A et al (2018) HbA1c, lipid profiles and risk of incident type 2 Diabetes in United States Veterans. PLoS ONE 13:e0203484
Liu J, van Klinken JB, Semiz S, van Dijk KW, Verhoeven A, Hankemeier T et al (2017) A Mendelian randomization study of metabolite profiles, fasting glucose, and type 2 diabetes. Diabetes 66:2915–2926
Agarwal T, Lyngdoh T, Dudbridge F, Chandak GR, Kinra S, Prabhakaran D et al (2020) Causal relationships between lipid and glycemic levels in an Indian population: a bidirectional Mendelian randomization approach. PLoS ONE 15:e0228269
Schmidt AF, Swerdlow DI, Holmes MV, Patel RS, Fairhurst-Hunter Z, Lyall DM et al (2017) PCSK9 genetic variants and risk of type 2 diabetes: a mendelian randomisation study. Lancet Diabetes Endocrinol 5:97–105
Fall T, Xie W, Poon W, Yaghootkar H, Mägi R, Knowles JW et al (2015) Using genetic variants to assess the relationship between circulating lipids and type 2 diabetes. Diabetes 64:2676–2684
White J, Swerdlow DI, Preiss D, Fairhurst-Hunter Z, Keating BJ, Asselbergs FW et al (2016) Association of lipid fractions with risks for coronary artery disease and diabetes. JAMA Cardiol 1:692–699
Zhu Z, Zheng Z, Zhang F, Wu Y, Trzaskowski M, Maier R et al (2018) Causal associations between risk factors and common diseases inferred from GWAS summary data. Nat Commun 9:224
Yuan S, Larsson SC (2020) An atlas on risk factors for type 2 diabetes: a wide-angled Mendelian randomisation study. Diabetologia 63:2359–2371
Soremekun O, Karhunen V, He Y, Rajasundaram S, Liu B, Gkatzionis A et al (2022) Lipid traits and type 2 diabetes risk in African ancestry individuals: a Mendelian Randomization study. EBioMedicine 78:103953
Haase CL, Tybjærg-Hansen A, Nordestgaard BG, Frikke-Schmidt R (2015) HDL cholesterol and risk of type 2 diabetes: a mendelian randomization study. Diabetes 64:3328–3333
Marott SC, Nordestgaard BG, Tybjærg-Hansen A, Benn M (2016) Components of the Metabolic syndrome and risk of type 2 diabetes. J Clin Endocrinol Metab 101:3212–3221
Pan W, Sun W, Yang S, Zhuang H, Jiang H, Ju H et al (2020) LDL-C plays a causal role on T2DM: a Mendelian randomization analysis. Aging (Albany NY) 12:2584–2594
Lee K, Lim CY (2019) Mendelian randomization analysis in observational epidemiology. J Lipid Atheroscler 8:67–77
Smith GD, Ebrahim S (2003) “Mendelian randomization”: can genetic epidemiology contribute to understanding environmental determinants of disease? Int J Epidemiol 32:1–22
Chen CH, Yang JH, Chiang CWK, Hsiung CN, Wu PE, Chang LC et al (2016) Population structure of Han Chinese in the modern Taiwanese population based on 10,000 participants in the Taiwan Biobank project. Hum Mol Genet 25:5321–5331
Lin WY, Liu YL, Yang AC, Tsai SJ, Kuo PH (2020) Active cigarette smoking is associated with an exacerbation of genetic susceptibility to diabetes. Diabetes 69:2819–2829
Diagnosis and classification of diabetes mellitus (2006) Diabetes Care 29(Suppl 1):S43–S48
Gillett MJ (2009) International Expert Committee report on the role of the A1c assay in the diagnosis of diabetes. Diabetes Care 32(7):1327–1334
Daniel L. Hartl AGC (2007) Principles of population genetics. 4th edn
Hellwege JN, Keaton JM, Giri A, Gao X, Velez Edwards DR, Edwards TL (2017) Population stratification in genetic association studies. Curr Protoc Hum Genet. 95:1.22.1-3.3
Zhang F, Zhang L, Deng HW (2009) A PCA-based method for ancestral informative markers selection in structured populations. Sci China C Life Sci 52:972–976
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D (2006) Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38:904–909
Hartwig FP, Davies NM, Hemani G, Davey SG (2016) Two-sample Mendelian randomization: avoiding the downsides of a powerful, widely applicable but potentially fallible technique. Int J Epidemiol 45:1717–1726
Network AGE. 2017.
Nowak C, Ärnlöv J (2018) A Mendelian randomization study of the effects of blood lipids on breast cancer risk. Nat Commun 9:3957
Beeghly-Fadiel A, Khankari NK, Delahanty RJ, Shu XO, Lu Y, Schmidt MK et al (2020) A Mendelian randomization analysis of circulating lipid traits and breast cancer risk. Int J Epidemiol 49:1117–1131
Luo M, Sun M, Wang T, Zhang S, Song X, Liu X et al (2023) Gut microbiota and type 1 diabetes: a two-sample bidirectional Mendelian randomization study. Front Cell Infect Microbiol 13:1163898
Zhang LP, Zhang XX (2022) Relationship between lipids and sleep apnea: Mendelian randomization analysis. World J Clin Cases 10:11403–11410
Anderson CA, Pettersson FH, Clarke GM, Cardon LR, Morris AP, Zondervan KT (2010) Data quality control in genetic case–control association studies. Nat Protoc 5:1564–1573
Burgess S, Thompson SG (2011) Avoiding bias from weak instruments in Mendelian randomization studies. Int J Epidemiol 40:755–764
Pierce BL, Ahsan H, Vanderweele TJ (2011) Power and instrument strength requirements for Mendelian randomization studies using multiple genetic variants. Int J Epidemiol 40:740–752
Fortier I, Raina P, Van den Heuvel ER, Griffith LE, Craig C, Saliba M et al (2017) Maelstrom research guidelines for rigorous retrospective data harmonization. Int J Epidemiol 46:103–105
Jung S (2013) Structural equation modeling with small sample sizes using two-stage ridge least-squares estimation. Behav Res Methods 45:75–81
Burgess S, Scott RA, Timpson NJ, Davey Smith G, Thompson SG (2015) Using published data in Mendelian randomization: a blueprint for efficient identification of causal risk factors. Eur J Epidemiol 30:543–552
Burgess S, Davies NM, Thompson SG (2016) Bias due to participant overlap in two-sample Mendelian randomization. Genet Epidemiol 40:597–608
Burgess S, Davey Smith G, Davies NM, Dudbridge F, Gill D, Glymour MM et al (2019) Guidelines for performing Mendelian randomization investigations. Wellcome Open Res 4:186
Burgess S, Dudbridge F, Thompson SG (2016) Combining information on multiple instrumental variables in Mendelian randomization: comparison of allele score and summarized data methods. Stat Med 35:1880–1906
Burgess S, Butterworth A, Thompson SG (2013) Mendelian randomization analysis with multiple genetic variants using summarized data. Genet Epidemiol 37:658–665
Bowden J, Davey Smith G, Burgess S (2015) Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int J Epidemiol 44:512–525
Hartwig FP, Davey Smith G, Bowden J (2017) Robust inference in summary data Mendelian randomization via the zero modal pleiotropy assumption. Int J Epidemiol 46:1985–1998
Burgess S, Foley CN, Allara E, Staley JR, Howson JMM (2020) A robust and efficient method for Mendelian randomization with hundreds of genetic variants. Nat Commun 11:376
Huang D, Lin S, He J, Wang Q, Zhan Y (2022) Association between COVID-19 and telomere length: a bidirectional Mendelian randomization study. J Med Virol 94:5345–5353
Hemani G, Zheng J, Elsworth B, Wade KH, Haberland V, Baird D et al (2018) The MR-Base platform supports systematic causal inference across the human phenome. Elife 7
Dragioti E, Gerdle B, Larsson B (2019) Longitudinal associations between anatomical regions of pain and work conditions: a study from The SwePain Cohort. Int J Environ Res Public Health 16:2167
Westerlund H, Kivimäki M, Singh-Manoux A, Melchior M, Ferrie JE, Pentti J et al (2009) Self-rated health before and after retirement in France (GAZEL): a cohort study. Lancet 374:1889–1896
Diagnosis and classification of diabetes mellitus (2014) Diabetes Care 37(Suppl 1):S81-90
Cai X, Zhang Y, Li M, Wu JH, Mai L, Li J et al (2020) Association between prediabetes and risk of all cause mortality and cardiovascular disease: updated meta-analysis. BMJ 370:m2297
Cai X, Liu X, Sun L, He Y, Zheng S, Zhang Y et al (2021) Prediabetes and the risk of heart failure: a meta-analysis. Diabetes Obes Metab 23:1746–1753
Gordon SM, Hofmann S, Askew DS, Davidson WS (2011) High density lipoprotein: it’s not just about lipid transport anymore. Trends Endocrinol Metab 22:9–15
Rahmoun MN, Ghembaza CE, El-Amine GM (2019) Lipid profile in type 2 patients with diabetes from Tlemcen: A Western Algerian population. Diabetes Metab Syndr 13:1347–1351
von Eckardstein A, Sibler RA (2011) Possible contributions of lipoproteins and cholesterol to the pathogenesis of diabetes mellitus type 2. Curr Opin Lipidol 22:26–32
Fazio S, Linton MF (2013) Killing two birds with one stone, maybe: CETP inhibition increases both high-density lipoprotein levels and insulin secretion. Circ Res 113:94–96
Drew BG, Rye KA, Duffy SJ, Barter P, Kingwell BA (2012) The emerging role of HDL in glucose metabolism. Nat Rev Endocrinol 8:237–245
Chapman MJ, Le Goff W, Guerin M, Kontush A (2010) Cholesteryl ester transfer protein: at the heart of the action of lipid-modulating therapy with statins, fibrates, niacin, and cholesteryl ester transfer protein inhibitors. Eur Heart J 31:149–164
Association AD (2020) 2. Classification and diagnosis of diabetes: standards of medical care in diabetes—2021. Diabetes Care 44:S15–S33
Sacks DB (2011) A1C versus glucose testing: a comparison. Diabetes Care 34:518–523
Little RR, Sacks DB (2009) HbA1c: how do we measure it and what does it mean? Curr Opin Endocrinol Diabetes Obes 16:113–118
Funding
This work was supported by grants from the Ministry of Science and Technology, Taiwan [MOST 107-2314-B-037-086-MY3 and MOST 110-2314-B-037-048 MY3]. The funding agencies had no role in the study design, data collection, data analysis, data interpretation, or writing of the report.
Author information
Authors and Affiliations
Contributions
YCL and TNW contributed to the conception and design. TNW contributed to the acquisition of data. YCL and HPT contributed to statistical analysis and interpretation. YCL and TNW drafted the manuscript for important intellectual content. TNW is the guarantor of this work and, as such, had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Corresponding author
Ethics declarations
Conflict of interests
The authors have declared that they have no conflict of interest.
Ethical approval
Ethical approval for the study was granted by the Institutional Review Board of Kaohsiung Medical University and Chung-Ho Memorial Hospital and the Ethics and Governance Committee (EGC) of the Taiwan Biobank.
Informed consent
Written informed consent of participation was obtained from all participants when joining TWB and the personal information of each participant was fully encrypted for protection.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lin, YC., Tu, HP. & Wang, TN. Blood lipid profile, HbA1c, fasting glucose, and diabetes: a cohort study and a two-sample Mendelian randomization analysis. J Endocrinol Invest 47, 913–925 (2024). https://doi.org/10.1007/s40618-023-02209-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40618-023-02209-x