Abstract
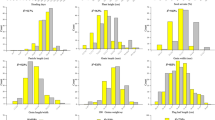
Success of crop improvement program depends on systematic exploitation of genetic diversity. Improved understanding on the genetic basis of traits contributing to yield and stress tolerance is necessary to accelerate development of resilient crop varieties. In this study, a subset of 102 diverse rice accessions was assembled after analysing population structure (K = 8) and removal of admixtures from a larger set of IRRI 3 K panel. The constructed subset showed adequate diversity in yield related traits. Genome wide association analysis using the genome wide SNP markers identified a total of 42 SNPs showing significant association with major yield traits. Out of the identified SNPs, 20 SNPs were found to be present in QTL or genes reported previously for yield traits. Remaining 22 loci were found to be novel and needs validation. Elite genetic stocks with increased yield potential will permit us to dissect out the physiological and molecular basis of spikelet number per panicle in rice and thereby accelerate yield enhancement in rice through haplotype based breeding.




Similar content being viewed by others
References
Abbai, R., Singh, V. K., Nachimuthu, V. V., Pallavi, S., Ramchander, S., Abhilash, K. V., Singh, A.K., Singh, U.M., Varshney, R.K., & Kumar, A. (2019). Haplotype analysis of key genes governing grain yield and quality traits across 3K RG panel reveals scope for the development of tailor-made rice with enhanced genetic gains. Plant Biotechnology Journal. https://doi.org/10.1111/pbi.13087.
Abdurakhmonov, I. Y., & Abdukarimov, A. (2008). Application of association mapping to understanding the genetic diversity of plant germplasm resources. International Journal of Plant Genomics. https://doi.org/10.1155/2008/574927
Alexander, D. H., Shringarpure, S. S., Novembre, J., & Lange, K. (2015). Admixture 1.3 software manual.
Allen, T. T. (2019). Software overview and methods review: Minitab. In T.T. Allen (Ed.), Introduction to engineering statistics and lean six sigma (pp. 575–600). London: Springer.
Asimit, J., & Zeggini, E. (2010). Rare variant association analysis methods for complex traits. Annual Review of Genetics, 44, 293–308. https://doi.org/10.1146/annurev-genet-102209-163421
Bhandari, A., Sandhu, N., Bartholome, J., Cao-Hamadoun, T.-V., Ahmadi, N., Kumari, N., & Kumar, A. (2020). Genome-wide association study for yield and yield related traits under reproductive stage drought in a diverse indica-aus rice panel. Rice, 13(1), 1–22. https://doi.org/10.1186/s12284-020-00406-3
Bradbury, P. J., Zhang, Z., Kroon, D. E., Casstevens, T. M., Ramdoss, Y., & Buckler, E. S. (2007). TASSEL: Software for association mapping of complex traits in diverse samples. Bioinformatics, 23(19), 2633–2635. https://doi.org/10.1093/bioinformatics/btm308
Browning, B. L., & Browning, S. R. (2016). Genotype imputation with millions of reference samples. The American Journal of Human Genetics, 98(1), 116–126. https://doi.org/10.1016/j.ajhg.2015.11.020
FAOSTAT. (2019). Food and agricultural organisation. http://www.fao.org/faostat/en/#data/QC. Accessed 17 May, 2021, 2021
Gonzalez, D., Bowen, A. J., Carroll, T. S., & Conlan, R. S. (2007). The transcription corepressor LEUNIG interacts with the histone deacetylase HDA19 and mediator components MED14 (SWP) and CDK8 (HEN3) to repress transcription. Molecular and Cellular Biology, 27(15), 5306–5315. https://doi.org/10.1128/MCB.01912-06
https://www.indiastat.com/data/agriculture/rice/data-year/all-years
Korte, A., & Farlow, A. (2013). The advantages and limitations of trait analysis with GWAS: A review. Plant Methods, 9(1), 1–9. https://doi.org/10.1186/1746-4811-9-29
Kushwaha, U. K. S., Mangal, V., Bairwa, A. K., Adhikari, S., Ahmed, T., Bhat, P., Yadav, A., Dhaka, N., Prajapati, D. R., & Gaur, A. (2017). Association mapping, principles and techniques. J Biol Environ Eng, 2(1), 1–9.
Li, G., Zhang, H., Li, J., Zhang, Z., & Li, Z. (2021). Genetic control of panicle architecture in rice. The Crop Journal. https://doi.org/10.1016/j.cj.2021.02.004
Lin, Y. L., Wu, D. H., Wu, C. C., & Huang, Y. F. (2021). Explore the genetics of weedy traits using rice 3K database. Botanical Studies, 62(1), 1–16. https://doi.org/10.1186/s40529-020-00309-y
Ling, S., Chen, C., Wang, Y., Sun, X., Lu, Z., Ouyang, Y., & Yao, J. (2015). The mature anther-preferentially expressed genes are associated with pollen fertility, pollen germination and anther dehiscence in rice. BMC Genomics, 16(1), 1–17. https://doi.org/10.1186/s12864-015-1305-y
Ray, D. K., Mueller, N. D., West, P. C., & Foley, J. A. (2013). Yield Trends Are Insufficient to Double Global Crop Production by 2050. PLoS ONE. https://doi.org/10.1371/journal.pone.0066428.
Ren, M., Huang, M., Qiu, H., Chun, Y., Li, L., Kumar, A., Fang, J., Zhao, J., He, H., & Li, X. (2021). Genome-wide association study of the genetic basis of effective tiller number in rice. Rice, 14(1), 1–13. https://doi.org/10.1186/s12284-021-00495-8
RGP, K. (2014). The 3000 rice genomes project. Gigascience, 3(1), 7. https://doi.org/10.1186/2047-217X-3-7
Sakai, H., Lee, S. S., Tanaka, T., Numa, H., Kim, J., Kawahara, Y., Wakimoto, H., Yang, C. C., Iwamoto, M., & Abe, T. (2013). Rice annotation project database (RAP-DB): an integrative and interactive database for rice genomics. Plant and Cell Physiology, 54(2), e6–e6. https://doi.org/10.1093/pcp/pcs183
Sang, J., Zou, D., Wang, Z., Wang, F., Zhang, Y., Xia, L., Li, Z., Ma, L., Li, M., & Xu, B. (2020). IC4R-2.0: rice genome reannotation using massive RNA-seq data. Genomics, Proteomics and Bioinformatics, 18(2), 161–172. https://doi.org/10.1016/j.gpb.2018.12.011
Subedi, S. R., Sandhu, N., Singh, V. K., Sinha, P., Kumar, S., Singh, S., Ghimire, S. K., Pandey, M., Yadaw, R. B., & Varshney, R. K. (2019). Genome-wide association study reveals significant genomic regions for improving yield, adaptability of rice under dry direct seeded cultivation condition. BMC Genomics, 20(1), 1–20. https://doi.org/10.1186/s12864-019-5840-9
Tian, C., Gregersen, P. K., & Seldin, M. F. (2008). Accounting for ancestry: Population substructure and genome-wide association studies. Human Molecular Genetics, 17(R2), R143–R150. https://doi.org/10.1093/hmg/ddn268
Turner, S. D. (2014). qqman: An R package for visualizing GWAS results using QQ and manhattan plots. Biorxiv. https://doi.org/10.1101/005165
Wang, J., & Zhang, Z. (2021). GAPIT Version 3: Boosting power and accuracy for genomic association and prediction. Genomics, Proteomics and Bioinformatics. https://doi.org/10.1016/j.gpb.2021.08.005
Zhao, K., Tung, C. W., Eizenga, G. C., Wright, M. H., Ali, M. L., Price, A. H., Norton, G. J., Islam, M. R., Reynolds, A., & Mezey, J. (2011). Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nature Communications, 2(1), 1–10. https://doi.org/10.1038/ncomms1467
Acknowledgements
Authors are thankful to IRRI-South Asia Breeding Hub, Hyderabad for providing the valuable genetic materials. Authors also thank ICAR-NASF for the funding support.
Funding
Funding was provided by Indian Council of Agricultural Research (GTR 8030).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sundaramoorthy, M., Ramasamy, S.P., Rajagopalan, V.R. et al. Pilot scale genome wide association mapping identified novel loci for grain yield traits in rice. Plant Physiol. Rep. 27, 11–21 (2022). https://doi.org/10.1007/s40502-021-00641-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40502-021-00641-w