Abstract
Introduction
RPGR ORF15 is an exon present almost exclusively in the retinal transcript of RPGR. It is purine-rich, repetitive and notoriously hard to sequence, but is a hotspot for mutations causing X-linked retinitis pigmentosa.
Methods
Long-read nanopore sequencing on MinION and Flongle flow cells was used to sequence RPGR ORF15 in genomic DNA from patients with inherited retinal dystrophy. A flow cell wash kit was used on a MinION flow cell to increase yield. Findings were confirmed by PacBio SMRT long-read sequencing.
Results
We showed that long-read nanopore sequencing successfully reads through a 2 kb PCR-amplified fragment containing ORF15. We generated reads of sufficient quality and cumulative read-depth to detect pathogenic RP-causing variants. However, we observed that this G-rich, repetitive DNA segment rapidly blocks the available pores, resulting in sequence yields less than 5% of the expected output. This limited the extent to which samples could be pooled, increasing cost. We tested the utility of a MinION wash kit containing DNase I to digest DNA fragments remaining on the flow cell, regenerating the pores. Use of the DNase I treatment allowed repeated re-loading, increasing the sequence reads obtained. Our customised workflow was used to screen pooled amplification products from previously unsolved inherited retinal disease (IRD) in patients, identifying two new cases with pathogenic ORF15 variants.
Discussion
We report the novel finding that long-read nanopore sequencing can read through RPGR-ORF15, a DNA sequence not captured by short-read next-generation sequencing (NGS), but with a more reduced yield. Use of a flow cell wash kit containing DNase I unblocks the pores, allowing reloading of further library aliquots over a 72-h period, increasing yield. The workflow we describe provides a novel solution to the need for a rapid, robust, scalable, cost-effective ORF15 screening protocol.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Long-read DNA sequencing with an ONT nanopore sequencer successfully reads across the frequently mutated and notoriously hard to sequence ORF15 region of the RPGR gene, but with low yield. |
Yield was increased using a flow cell wash kit. |
The method described allows simultaneous sequencing of up to 24 samples in a single experiment, providing a rapid cost-effective protocol. |
1 Introduction
Approaches to DNA sequencing have advanced significantly since the landmark report of Sanger sequencing in 1977 [1, 2]. The prevailing technology, next generation sequencing (NGS), which uses sequencing-by-synthesis chemistry to generate short (approximately 150-bp) sequence reads, has increased the accessibility of genetic testing and the number of genes that can be concurrently analysed in a single assay. In recent years, population-scale sequencers have enabled whole genome sequencing (WGS), allowing the UK 100,000 Genomes Project [3], and other national large-scale sequencing programmes, to be completed. Due to its capability to deliver large volumes of highly accurate sequence data at relatively low cost, short-read NGS has become the dominant technology for determining a molecular diagnosis in patients with rare genetic diseases.
However, short-read DNA sequencing has several widely reported limitations. Generic enrichment PCR conditions can lead to non-uniform or absent coverage [4, 5], de novo assembly and haplotype phasing is rarely possible [6], structural variants can prove difficult to detect [7] and the characterisation of repetitive sequence remains challenging [8]. It is likely that these issues underlie many of the approximately half of cases with a suspected Mendelian disease that remain undiagnosed following whole-exome (WES) short-read sequencing [9, 10]. More recently, applications showcasing the diagnostic utility of long-read sequencing have emerged. Third generation single molecule sequencing platforms, such as the Sequel and Revio instruments (Pacific Biosciences), in addition to the nanopore range of devices (Oxford Nanopore Technologies; ONT), can generate long reads (> 10 kb) at a rapidly increasing rate and scale [9, 11,12,13]. This is facilitating the investigation of so-called dark and camouflaged genomic loci, which have remained refractory to short-read analyses, either due to informatic difficulties (e.g. an inability to determine an unambiguous mapping position) or wet-laboratory processes that relate to their underlying genomic architecture (e.g. the high GC content of some first exons). These studies are increasing our understanding of the frequency and complexity of structural variants, and enabling improved analysis of challenging genomic regions [14,15,16].
Retinitis pigmentosa (RP) is the most prevalent inherited retinal disease (IRD) [17], with dominant, recessive and X-linked inheritance patterns described. X-linked retinitis pigmentosa (XLRP) is generally the more severe form and accounts for up to 20% of patients [18]. The majority of pathogenic variants causing XLRP are in the retinitis pigmentosa GTPase regulator (RPGR) gene [19, 20], which has multiple isoforms. Over 60% of disease-causing variants in RPGR are in the notoriously hard-to-sequence open reading frame 15 (ORF15) exon and the ORF15-containing isoform is the predominant transcript expressed in the retina (NM_001034853.2). ORF15 contains a 999 bp low-complexity region (chrX:38,145,048-38,145,046, GRCh37/hg19), 98.3% of which is made up of the purines adenine and guanine. The nucleotide sequence consists of an imperfect tandem array of ~ 27 bp repeats with a consensus sequence GAGGAGGAAGGAGAAGGGGAGGGGGAA. This encodes a 333 amino acid protein domain, 90% of which consists of glutamic acid and glycine residues, consisting of imperfect repeats of EEEGEGEGE [21]. This sequence is thought to be responsible for the high mutability and reduction in replication fidelity observed in this region [22]. Standard short-read NGS captures the outer extremities of the exon but is unable to comprehensively characterise the repetitive central region. Although WGS performs better than WES in most GC-rich areas [23], this is not the case for ORF15. It has been suggested that the super helical tension caused by the repeats leads to the formation of hairpins and other complex structures that cause instability and polymerase slippage or arrest [24, 25]. This region has been identified as a hotspot for disease-causing variations [21], the most prevalent of which are small deletions that create a frameshift in the encoded protein [19, 20, 26].
A scalable, high-throughput, reliable approach is therefore required to screen this exon. Here, we assess the viability of long-read nanopore sequencing as a screening strategy for the identification of pathogenic mutations in RPGR-ORF15, from PCR-amplified ORF15 DNA. We found that it can be read using a MinION sequencer (Oxford Nanopore Technologies), but flow cell pores became rapidly blocked. Use of a MinION wash kit containing DNase I to digest any remaining library fragments reactivated the pores and enabled the flow cell to be re-loaded. This increased the number of sequence reads that were mapped to the ORF15 locus.
2 Methods and Materials
2.1 Patient Recruitment
Patients were recruited prospectively at Ophthalmology clinics in St James’s University Hospital, Leeds, and other UK centres. Informed written consent was obtained using a protocol that followed the precepts of the Declaration of Helsinki and was approved by the Leeds East Research Ethics Committee (Project reference 17/YH/0032). Genomic DNA was extracted from blood using standard protocols.
2.2 Transcript
All sequence variants are numbered based on transcript NM_001034853.2.
2.3 Short-Read Exome Sequencing (WES)
Targeted enriched libraries were prepared using the SureSelectXT Human All Exon V6 kit (Agilent Technologies, Santa Clara, CA, USA) and sequenced with a paired-end protocol on a HiSeq 3000 Sequencer (Illumina, Little Chesterford, UK). The quality control of the raw sequence data, base quality scores, GC content and duplications were checked using java based FastQC software (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/). Sequence adaptors were removed with Trim Galore (http://www.bioinformatics.babraham.ac.uk/projects/trim_galore/). Sequences were then aligned against the reference genome (hg19/GRCh37) using the Burrows–Wheeler Aligner BWA (v0.7.12-r1.39) [27]. SAM files were converted to BAM files with SAMtools then sorted by Picard tools (v2.5.0) (https://broadinstitute.github.io/picard/), which were also used to remove PCR duplicates. BAM files were realigned locally around the indels using the Genome Analysis Tool Kit GATK (https://gatk.broadinstitute.org/hc/en-us) (v3.5) [28, 29]. The GATK HaplotypeCaller function was used to call small indels and single nucleotide variants (SNVs) in genomic variant call format (g.VCF). The variant list was then annotated using Variant Effect Predictor (VEP) software [30].
2.4 PCR Amplification
Two PCR reactions were performed to generate ORF15 amplification products that were uniquely indexed on a per-sample basis. Pre-indexing PCR: a first PCR was carried out using ORF15 specific primers tailed with universal sequencing tags. The PCR reaction mix consisted of 1 µl of genomic DNA (20–50 ng/µl), 0.8 µl of 5 mM dNTPs (Invitrogen, Paisley, UK), 0.2 µl Phusion DNA polymerase [New England Biolabs (NEB), Ipswich, MA, USA], 4 µl GC buffer (NEB), 0.6 µl DMSO (Invitrogen), 1 µl each of 0.5 µM forward (TTTCTGTTGGTGCTGATATTGCTGATGAAGTGGAAACTGACCA) and reverse (ACTTGCCTGTCGCTCTATCTTCTGTCTGACTGGCCATAATCG) primers (universal sequencing tags are underlined) (ThermoFisher Scientific, Waltham, MA, USA) and 11.4 µl nuclease-free water. Thermocycling conditions are detailed in Online Supplementary Material (OSM) Table S1. Indexing PCR: to sequence multiple samples in a single flow cell, unique indexing barcodes were added to each sample during a second-round PCR. The pre-indexed PCR amplification products were purified using AMPure XP beads (Beckman Coulter, Indianapolis, IN, USA), then quantified by Qubit fluorometer (ThermoFisher). Molarity was calculated using the NEBiocalculator (https://nebiocalculator.neb.com/#!/ssdnaamt). A total of 100–200 fmol of each pre-indexed amplicon was adjusted to 24 µl with nuclease-free water then combined with 25 µl Long Amp Taq 2X master mix (NEB) and 1 µl of barcode reagent from kit EXP-PBC096 (ONT, Oxford, UK). Thermocycling conditions are recorded in OSM Table S2.
2.5 Library Preparation
Barcoded amplification products were pooled in equimolar quantities to a total mass of 5 µg. From this solution, 1 µg of DNA was aliquoted and end-repaired by combining 3.5 µl FFPE DNA repair buffer (NEB), 2 µl FFPE DNA repair mix (NEB), 3.5 µl Ultra™ II end-prep reaction buffer (NEB) and 3 µl Ultra™ II end-prep enzyme mix (NEB), made up with nuclease-free water in a total reaction volume of 60 µl. The reaction was incubated at 20 °C for 5 min then 65 °C for 5 min. After cleaning with AMPure XP beads, sequencing adapters were ligated to the double-stranded amplimers. The reaction comprised 60 µl of PCR amplimers, 25 µl Ligation Buffer (ONT), 10 µl Quick T4 DNA Ligase (NEB) and 5 µl Adapter Mix (ONT). The reaction was incubated for 10 min at room temperature then cleaned up using AMPure XP beads; the beads were washed twice with 250 µl short fragment buffer (ONT). The pellet was eluted in 15 µl elution buffer (ONT) then quantified using a Qubit fluorometer (ThermoFisher) to enable library molarity to be calculated.
2.6 Long-Read Sequencing
Long-read sequencing was carried out on two types of nanopore flow cell. Flongle sequencing: a separate sequencing library was created for each sample, using half volumes of the above-described end-prep and ligation reactions. A Flongle flow cell (R.9.4.1) was next prepared for sequencing by loading 120 µl priming mix, 3 µl of Flush Tether (FT) (ONT) and 117 µl of Flush Buffer (FB) (ONT). A total of 3–20 fmol of the library was then combined with 15 µl of sequencing buffer (SQB) (ONT) and 10 µl of loading beads (LB) (ONT) prior to loading onto the flow cell. A 24-h Flongle sequencing run was initiated using MinKNOW software (v.3.6.0; ONT). MinION sequencing: 800 µl of MinION flowcell (R9.4.1 FLO-MIN106D) priming mix (30 µl of Flush Tether (ONT) well mixed into a vial of Flush Buffer (ONT)) was loaded into the flow cell priming port. A total of 50 fmol of the eluted library was made up to 12 µl using nuclease-free water mixed with 37.5 µl of sequencing buffer (ONT) and 25.5 µl of loading beads (ONT), then loaded into the flow cell via the SpotOn port in a dropwise fashion. The MinION sequencer was run for 72 h using MinKNOW software (v.3.6.5; ONT). Use of flow cell wash kit with MinION sequencing: when ORF15 amplification products were initially sequenced, pores were observed to be rapidly blocked, resulting in the production of relatively few reads. In subsequent runs a flow cell wash kit (WSH003) (ONT) was used to reactivate pores and boost instrument yields. The sequencer was run as described, but paused after 4 h. A total of 2 µl of wash solution (ONT) was mixed with 398 µl of diluent to make a wash mix. Liquid was withdrawn from the waste port and discarded before 400 µl of the wash mix was loaded into the priming port and left for 60 min. This was then removed from the waste port, more priming mix was loaded into the priming port, then more library was loaded into the SpotON port.
2.7 Nanopore Sequence Analysis
Base calling and sample demultiplexing to convert the raw data from fast5 to FASTQ format was performed using Guppy (v.6.4.2; https://nanoporetech.com) with the super-high accuracy model. This included the detection of mid-strand adapters and barcodes, primers and read splitting. NanoFilt (v.2.2.0; https://github.com/wdecoster/nanofilt) was used to remove low-quality reads (Q score ≥ 10) and perform length-based filtering (minimum 1819 bp, maximum 2019 bp) [31]. Processed reads were next aligned to the human reference genome (build hg19) using minimap2 (v.2.16; https://github.com/lh3/minimap2 [32]) prior to being converted to BAM format and sorted by alignment coordinate using samtools (v.1.9; https://github.com/samtools/samtools [33]). Variant calling was performed using Clair3 (v.0.1; https://github.com/HKU-BAL/Clair3) in a singularity container with the pre-trained nanopore-specific model “r941_prom_sup_g5014”. NanoStat (v.1.1.2; https://github.com/wdecoster/nanostat [31]) was used to calculate read metrics and statistics. BAM files were visualised using the Integrative Genomics Viewer (IGV; v.2.16.0.; https://software.broadinstitute.org/software/igv/).
2.8 Reference Laboratory Sanger Sequencing of ORF15
Four primer pairs (RPGR_Ex15-1F/ RPGR_Ex15-1R, RPGR_Ex15-2F/ RPGR_Ex15-2R, RPGR_Ex15-3F/ RPGR_Ex15-3R and RPGR_Ex15-4F/ RPGR_Ex15-4R) were used to sequence the ORF15 region of RPGR. The sequence-specific primers are listed in OSM Table S3. All of the RPGR exon ORF15 primers were tailed with N13 tags (forward: GTAGCGCGACGGCCAGT and reverse: CAGGGCGCAGCGATGAC). The PCR mix used for primer pairs RPGR_Ex15-1F/RPGR_Ex15-1R and RPGR_Ex15-4F/RPGR_Ex15-4R consisted of 10 µl GoTaq master mix (Promega, Madison, Wisconsin, USA), 2 µl primer mix (final concentration 500 nM), 2 µl of genomic DNA and 6 µl nuclease-free water. The PCR mix used for primer pairs RPGR_Ex15-2F/RPGR_Ex15-2R and RPGR_Ex15-3F/RPGR_Ex15-3R comprised 2 µl of 10× PCR buffer minus MgCl2 (Invitrogen), 1 µl of 20 mM dNTP mix, 0.5 µl of 50 mM MgCl2, 1.20 µl primer mix (final concentration 300 nM), 0.20 µl Platinum Taq DNA polymerase (Invitrogen), 2 µl of genomic DNA and 13.10 µl nuclease-free water. Thermocycling conditions for these PCR reactions are recorded in OSM Table S4. Sanger sequencing reaction mixes are recorded in OSM Table S5. The sequencing run was performed on an ABI 3730 Genetic Analyzer (Applied Biosystem) and the sequences produced were analysed on sequence scanner software (v2.0; Applied Biosystem).
2.9 Variant Verification by Pacific Biosciences (PacBio) Sequencing
Long-range PCR amplification was performed for 48 samples using different combinations of barcoded forward (CAGTAGAAAAGCCAGACAGTTACATG) and barcoded reverse (GTATATTCCTGTTTCCTAAAGCTGCC) primers. The full primer list is given in OSM Table S6. The PCR reaction was performed by mixing 1 µl of genomic DNA (30–50 ng/µl), 4 µl of GC buffer (final concentration 1×) (NEB), 0.8 µl of 5 mM dNTPs (final concentration 200 µM) (Invitrogen), 1 µl of each primer (final concentration 0.5 µM each), 0.2 µl Phusion High-Fidelity DNA polymerase (final concentration 0.4 units/20 µl PCR reaction) (NEB) and nuclease-free water up to 20 µl total volume. No additives were added. A positive control and a negative control with no gDNA were included for each pair of barcoded primers. Thermocycling conditions are recorded in OSM Table S7. Long-read sequencing was carried out using a Sequel (PacBio, California) following the manufacturer’s instructions. The generated subreads were converted to circular consensus sequences (ccs) using the command-line tool ccs (v.4.2.0) with default parameters (a minimum of three full-length subreads were required to generate a ccs). Sequence reads were next aligned to the human reference genome (build hg19) using minimap2 (v.2.16) before being converted to BAM format and sorted by alignment coordinate using samtools (v.1.9). Variant calling was performed using Clair3 (v.0.1) with the PacBio-specific pre-trained “HiFi” model. The PacBio amplicon overlapped all MinION generated target nucleotides. The comparative analysis included a review of the variant call files in addition to manual inspection of the alignment BAM files using the IGV (v.2.16.0).
3 Results
3.1 MinION Long-Read Sequencing of ORF15
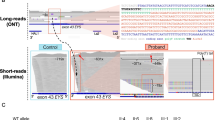
A 1919 bp DNA fragment containing ORF15 was amplified from genomic DNA from five males with RP, using the pre-indexing PCR protocol. One patient was hemizygous for the RPGR-ORF15 pathogenic variant c.2426_2427delAG, p.(Glu809Glyfs*25) and the remaining four were at that time unsolved following previous analyses of targeted or exome enriched NGS datasets. It is well documented that conventional short-read NGS approaches perform poorly when they are used to sequence this exon [19, 22, 34, 35]. Figure 1A shows an example reference-based alignment at the ORF15 locus, generated using short-reads obtained from a HiSeq 3000 (Illumina, Inc.). While there is sufficient read depth at the extremities of the ORF15 exon, there are no alignments spanning the central ~ 900 bp repetitive region.
Sequencing RPGR ORF15. A Short-read next generation sequencing at the RPGR ORF15 locus. Hybridisation capture enrichment was performed prior to sequencing on an Illumina HiSeq 3000. Aligned sequence reads are viewed using the integrative genomic viewer (IGV). There is an absence of mapped reads across the central region. B Long-read sequencing alignment at the RPGR ORF15 locus generated using a nanopore MinION sequencer. The male RP patient is hemizygous for the two base-pair deletion c.2426_2427delAG (NM_001034853.2), p.(Glu809Glyfs*25) (ChrX: 38145827_38145828delCT (hg19). C Sanger sequencing electropherogram generated by the Manchester Reference Laboratory confirms the absence of a two base-pair sequence at the dashed vertical line
Ligation-based library preparation was performed on ORF15 amplification products, and 3–20 fmol of each library was loaded on individual Flongle flow cells. Visual examination of the aligned sequence reads using the IGV confirmed that the nanopore workflow is capable of generating full-length RPGR-ORF15 sequences. Unique sequences flanking the repeat enabled the long reads to be anchored to the target locus, generating sufficient read coverage across the highly repetitive ORF15 sequence to enable mutation detection. Identification of the previously reported ORF15 pathogenic variant c.2426_2427delAG, p.(Glu809Glyfs*25) is demonstrated in Fig. 1B. Run yields obtained were between 9 and 56 Mb, corresponding to read counts of between 6.41K and 34.04K, as detailed in OSM Table S8. The c.2426_2427delAG variant was confirmed by Sanger sequencing (Fig. 1C) using a specialised ORF15 sequencing protocol developed by the Manchester reference laboratory (see “Materials and Methods”).
While Flongle sequencing of ORF15 was successful, yields were between 1 and 5% of the conservatively anticipated 1 Gb sequencer output. Furthermore, it was evident from cumulative read traces that, by contrast to a typical Flongle sequencing run, throughput from the flow cell after loading the ORF15 amplimer slowed rapidly, within minutes. Figure 2A shows the cumulative read trace of a sequencing run targeting a non-repetitive region spanning 11.5 kb of the ALMS1 gene, with no known difficulty for conventional DNA sequence analysis. A proportion of pores remained open and able to generate significant numbers of new reads after 24 h (Fig. 2B). In contrast, when the ORF15 amplimer was sequenced, the cumulative read count plateaued within the first hour (Fig. 2C) and pore availability declined rapidly within 35 min of loading (Fig. 2D).
Cumulative read counts and pore availability plots for Flongle and MinION long-read sequencing of RPGR ORF15, with and without the use of a flow cell wash kit. A Cumulative read count plot for Flongle sequencing of an 11.5 kb PCR amplimer containing the ALMS1 gene. B Pore availability over time for the Flongle run plotted in A. The plot shows that pores were available over a 24-h period, with a slow decline over that time. C Cumulative read count plot for Flongle sequencing of a 1.9 kb PCR amplimer containing RPGR ORF15. Pore availability dropped rapidly within the first hour and reads produced declined to almost none within three hours. D Pore availability for the Flongle run shown in C, demonstrating that pores rapidly became “unavailable” over the first hour of sequencing, resulting in a dramatic decrease in data acquisition. E Cumulative read count plot for MinION sequencing of the same RPGR ORF15-containing amplimer in five pooled, tagged samples, with application of flow cell wash buffer after 3 and 24 h. After each wash, the rate at which reads were acquired recovered to near the original starting rate, then rapidly declined again over the first hour. As a result, throughput was more than doubled from the point of the first wash. F Pore availability for the MinION run plotted in E, with two wash treatments. After washing, pores recovered from “unavailable” to the “single pore” state, increasing the rate of data acquisition, though they then rapidly dropped over the next hour. G Cumulative read count plot for MinION sequencing of the RPGR ORF15 amplicon in 12 pooled samples, with nine washing steps over a two-day period. Throughput rebounds after every washing step but this effect declines progressively over the course of the run. H Pore availability for the MinION sequencing run of exon ORF15 plotted in G, which included nine treatments to reactivate the pores. After every wash, the pores recovered from the “unavailable” state to the “single pore” state, increasing the rate of data acquisition. I Colour key showing the pore status during sequencing in the nanopore runs shown in B, D, F and H
Several post-amplification clean-up protocols were investigated to determine whether contaminants were blocking the pores. These included post PCR clean ups using AMPure XP beads, manual gel extraction of the PCR product and automated size separation using the Pippin Prep System (Sage Science, Beverly, MA, USA). However, no improvement was obtained (data not shown). We therefore hypothesised that the reduced throughput of the ORF15 amplimer may be a consequence of the formation of secondary structures within the repetitive ORF15 sequence, such that these structures then progressively blocked the flow cell pores until no further reads could be generated.
3.2 Use of a Flow Cell Wash Kit to Increase Sequencer Yield
To address this possibility, a flow cell wash kit (WSH003, ONT) containing DNase I was used. Wash kits are intended to facilitate reuse of MinION flow cells by digesting, and therefore removing, any residual DNA from the flow cell pores before a different library is loaded. We hypothesised that application of a nuclease wash treatment would clear the pores and allow reloading of a further aliquot of the same ORF15 library, thereby increasing yield and cumulative read count at the target locus. However, the nuclease wash can only be used on the MinION flow cell, not the lower-throughput Flongle, because opening and resecuring the Flongle flow cell cover is not a supported procedure. At this point, experiments were therefore switched to use MinION flow cells.
When an ORF15 amplimer library was run on a MinION flow cell, we observed the expected rapid decline in the cumulative read count (Fig. 2E) and pore availability (Fig. 2F), over a period of 2–3 h. Use of the DNase I wash led to an immediate rebound in cumulative read output and pore availability, but this declined within a similar timeframe, requiring a further DNase I nuclease treatment. The resultant output, though still well below the manufacturer’s expected yield, was considerably increased as a result of washing and reloading. Finally, to establish the likely limit of the rewashing protocol, we ran multiple aliquots of an ORF15 amplimer library on a single MinION flow cell over a period of 3 days, washing and reloading nine times. A cumulative read count trace and pore availability plot for this run are shown in Fig. 2G, H, respectively. Pore availability continued to rebound after each wash but declined over the course of the run until little benefit was gained from further reloading.
3.3 Screening ORF15 in Untested Cases
Our customised ORF15 workflow was applied to a further 49 individuals, and the initial five were re-analysed. Clinical and genotyping details for all 54 screened individuals can be found in OSM Table S9. These included 30 males and 1 female with unsolved RP and 4 males with unsolved macular disease. However, over the course of this work, ten of these cases were subsequently solved and marked as obligate negative in OSM Table S9. In addition, four males and two females with RP and one male with macular disease, each carrying known ORF15 pathogenic variants [two with c.3334C>T, p.(Gln1112Ter), two with c.2426_2427delAG, p.(Glu809Glyfs*25) and three with c.2405_2406delAG, p.(Glu802Glyfs*32)], were included. Lastly, 12 unaffected individuals were tested as controls to assess population variation. Libraries from each DNA sample were indexed and combined in pools of up to 24 cases, then sequenced on a MinION flow cell, with multiple nuclease washes performed. Per-sample raw read counts ranged between 5716 and 97,596 (mean: 27,412) with processed read counts (i.e., those remaining following read length, quality score and target site filtering) being reduced to between 81 and 17,346 (mean: 2635). All previously known variants were observed, and we identified two new cases of RP caused by the ORF15 variants c.2041_2042delAA, p.(Lys681Glyfs*2), ChrX:38146210_38146211del (hg19) and c.2323_2324delAG, p.(Arg775Glufs*59), ChrX:38145933_38145934del (hg19). Both variants have been reported previously on the ClinVar database as either likely pathogenic (c.2041_2042delAA; accession number: VCV000865836.2) or pathogenic (c.2323_2324delAG; accession number: VCV000438144.18), respectively. We also identified a heterozygous pathogenic nonsense mutation (c.2074G>T p.(Gly692*), ChrX:38146178C>A (hg19)) in a carrier female. We note that the single molecule reads enabled us to determine this variant was arranged in cis with the other identified variants in this patient. All pathogenic variants, both those included as controls and those newly identified in this study, were first identified by the variant caller then confirmed by manual inspection using the IGV.
In addition, several benign single nucleotide variants (SNVs) and in-frame deletions and duplications were observed, in both cases and controls. For 48 of the 54 analysed patients, these variants were verified using a PacBio generated long-read dataset (individual sample variants, and the outcome of these comparative analyses are detailed in OSM Table 9). For single nucleotide variants we obtained 100% concordance between the two datasets; for this class of variant the assay was therefore 100% sensitive and specific. For insertion/deletion variants, all non-reference events were “identified” by the automated variant caller (Clair3). However, for the MinION dataset, a 21-bp duplication was incorrectly resolved as a single (c.2939dup), rather than 21 nucleotide, duplication [c.2919_2939dup (p.(Gly977_Glu983dup)]. Manual inspection of the aligned sequence reads revealed it to be a multi-nucleotide insertion, which was “correctly” resolved from the Pacific Biosciences dataset. To assess inter-run comparability, 20 samples were analysed twice by MinION sequencing; there was complete concordance between the variants identified in these samples.
4 Discussion
Use of standard PCR and Sanger sequencing to amplify the RPGR-ORF15 locus presents a technical challenge. This is thought to be due to polymerase slippage or arrest caused by hairpins and other complex structures in the ORF15 repetitive region. The repetitive sequence, together with the presence of common polymorphic indels, means that sequence alignment is also difficult. Previous studies have used a range of different mutation detection approaches since ORF15 was reported as a mutation hotspot [21]. These include direct Sanger sequencing [36], cloning the PCR product and then Sanger sequencing [37, 38] and direct sequencing of the repetitive part of RPGR-ORF15 with nested sequencing primers [39]. Short-read NGS, using sequencing-by-synthesis chemistry, results in poor depth-of-coverage over the highly repetitive region [34]. An NGS-based approach using a de novo assembly pipeline has been developed, which reportedly overcomes the limitations of the traditional pipeline but required considerable optimisation to reduce the number of false positive calls [40].
In this study we successfully screened RPGR-ORF15 for disease causing variants using a novel approach, long-range PCR target enrichment combined with long-read nanopore sequencing, in cases with unsolved RP and macular disease. The target locus was amplified using a two-step PCR which incorporated per-sample barcodes prior to sequencing on a MinION flow cell. During the run, sequencing pores become “unavailable”, possibly due to secondary structures formed by the ORF15 repetitive sequence; this reduced the number of reads generated to less than 5% of those expected. We therefore repurposed a flow cell wash kit containing DNase I, originally designed to allow flow cells to be reused [41], and used it to digest any remaining DNA fragments and unclog the pores. We then reloaded either a further aliquot of the same library, or a freshly prepared library. We demonstrated that DNase I treatment restores pores to an “active” state, resulting in higher per-run yields and cumulative read depth at the target ORF15 locus. The benefits of this approach became progressively more limited over time; after 72-h and nine cycles of washing and reloading, output declined to a point where few further reads could be obtained. This was probably due to both the natural deterioration of the membrane-embedded pores and frequent washing and reloading steps throughout the run. Nevertheless, we speculate that our workflow is likely to be of value to investigators aiming to sequence other similarly intractable genomic regions.
In comparison to WES, our novel method allowed us to screen the entire length of the ORF15 exon and resulted in a depth of coverage that allowed detection of three previously verified pathogenic variants, two further pathogenic variants in previously unsolved male cases and a pathogenic heterozygous nonsense mutation in a carrier female. Single molecule reads in this latter case allowed us to ascertain that the additional variants identified in this patient were arranged in cis. For single nucleotide variants, no false positive calls were detected, and the false negative rate (when compared with the PacBio generated dataset) was also zero, demonstrating that the sensitivity of this approach is high. We note that the number of reported false positives is lower than the de novo assembly workflow reported by Maggi et al. (2020), but acknowledge that our study size is limited and further testing of our workflow would be beneficial. For insertion variants the discrepancy between automatically resolving a single- and 21-nucleotide duplication highlights the ongoing utility of manually scrutinising aligned sequence reads in this region of complex genomic architecture.
One recognised limitation of our workflow is the requirement to perform PCR-based target-enrichment. This can lead to polymerase slippage across low-complexity repeats, and biased amplification of parental alleles (although this concern is mitigated when analysing hemizygous male cases). Two additional long-read target enrichment strategies are being developed which avoid PCR amplification. CRISPR/Cas9 workflows enable specific cleavage sites to be generated in bulk genomic DNA, prior to the ligation of instrument-specific sequencing adapters [42]. ReadUntil sequencing (also known as adaptive sampling) using the ReadFish software package, allows nanopore devices to selectively reject off-target sequences from the pore, in real-time, by reversing the voltage across individual nanopores [43, 44]. However, both workflows suffer from relatively low yield and on-target read depth, which may further exacerbate the already much reduced run yields obtained from nanopore sequencing of ORF15.
In conclusion, we demonstrate the novel finding that long-read nanopore sequencing can read through the region of RPGR-ORF15 that is refractory to analysis by short read NGS. Furthermore, we show that repurposing of a flow cell wash kit intended to facilitate flow cell reuse, which contains DNase I and unblocks the pores, allowing researchers to increase yield by reloading further aliquots of the library over a 72-h period. The workflow described here also permits the sequencing of indexed pooled libraries, from up to 24 individuals, on a single MinION flow cell, providing a rapid cost-effective screening protocol for this notoriously hard-to-sequence mutation hotspot. This new approach may be of value in analysing other similarly hard-to-sequence DNA regions and suggests that widespread adoption of long-read sequencing in a diagnostic setting may lead to improved coverage of so-called dark and camouflaged genomic regions.
References
Sanger F, Nicklen S, Coulson AR. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA. 1977;74(12):5463–7.
Van Dijk EL, Jaszczyszyn Y, Naquin D, Thermes C. The third revolution in sequencing technology. Trends Genet. 2018;34(9):666–81.
Turnbull C, Scott RH, Thomas E, Jones L, Murugaesu N, Pretty FB, et al. The 100,000 Genomes Project: bringing whole genome sequencing to the NHS. BMJ. 2018;361:k1687.
Manase D, D’Alessandro LC, Manickaraj AK, Al Turki S, Hurles ME, Mital S. High throughput exome coverage of clinically relevant cardiac genes. BMC Med Genom. 2014;7:67.
Mandelker D, Schmidt RJ, Ankala A, McDonald Gibson K, Bowser M, Sharma H, et al. Navigating highly homologous genes in a molecular diagnostic setting: a resource for clinical next-generation sequencing. Genet Med. 2016;18(12):1282–9.
Snyder MW, Adey A, Kitzman JO, Shendure J. Haplotype-resolved genome sequencing: experimental methods and applications. Nat Rev Genet. 2015;16(6):344–58.
Kosugi S, Momozawa Y, Liu X, Terao C, Kubo M, Kamatani Y. Comprehensive evaluation of structural variation detection algorithms for whole genome sequencing. Genome Biol. 2019;20(1):117.
Karimzadeh M, Ernst C, Kundaje A, Hoffman MM. Umap and Bismap: quantifying genome and methylome mappability. Nucleic Acids Res. 2018;46(20): e120.
Wenger AM, Peluso P, Rowell WJ, Chang P-C, Hall RJ, Concepcion GT, et al. Accurate circular consensus long-read sequencing improves variant detection and assembly of a human genome. Nat Biotechnol. 2019;37(10):1155–62.
Frésard L, Montgomery SB. Diagnosing rare diseases after the exome. Cold Spring Harb Mol Case Stud. 2018;4(6): a003392.
Lu H, Giordano F, Ning Z. Oxford Nanopore MinION sequencing and genome assembly. Genom Proteom Bioinform. 2016;14(5):265–79.
Pugh J. The current state of nanopore sequencing. Methods Mol Biol. 2023;2632:3–14.
Peng C, Zhang H, Ren J, Chen H, Du Z, Zhao T, et al. Analysis of rare thalassemia genetic variants based on third-generation sequencing. Sci Rep. 2022;12(1):9907.
Ebbert MT, Jensen TD, Jansen-West K, Sens JP, Reddy JS, Ridge PG, et al. Systematic analysis of dark and camouflaged genes reveals disease-relevant genes hiding in plain sight. Genome Biol. 2019;20(1):1–23.
Watson CM, Crinnion LA, Lindsay H, Mitchell R, Camm N, Robinson R, et al. Assessing the utility of long-read nanopore sequencing for rapid and efficient characterization of mobile element insertions. Lab Invest. 2021;101(4):442–9.
Huddleston J, Eichler EE. An incomplete understanding of human genetic variation. Genetics. 2016;202(4):1251–4.
Daiger SP, Bowne SJ, Sullivan LS. Perspective on genes and mutations causing retinitis pigmentosa. Arch Ophthalmol. 2007;125(2):151–8.
Churchill JD, Bowne SJ, Sullivan LS, Lewis RA, Wheaton DK, Birch DG, et al. Mutations in the X-linked retinitis pigmentosa genes RPGR and RP2 found in 8.5% of families with a provisional diagnosis of autosomal dominant retinitis pigmentosa. Invest Ophthalmol Vis Sci. 2013;54(2):1411–6.
Li J, Tang J, Feng Y, Xu M, Chen R, Zou X, et al. Improved diagnosis of inherited retinal dystrophies by high-fidelity PCR of ORF15 followed by next-generation sequencing. J Mol Diagn. 2016;18(6):817–24.
Tuupanen S, Gall K, Sistonen J, Saarinen I, Kämpjärvi K, Wells K, et al. Prevalence of RPGR-mediated retinal dystrophy in an unselected cohort of over 5000 patients. Transl Vis Sci Tech. 2022;11(1):6.
Vervoort R, Lennon A, Bird AC, Tulloch B, Axton R, Miano MG, et al. Mutational hot spot within a new RPGR exon in X-linked retinitis pigmentosa. Nat Gen. 2000;25(4):462–6.
Chiang JP, Lamey TM, Wang NK, Duan J, Zhou W, McLaren TL, et al. Development of high-throughput clinical testing of RPGR ORF15 using a large inherited retinal dystrophy cohort. Invest Ophthalmol Vis Sci. 2018;59(11):4434–40.
Nash BM, Ma A, Ho G, Farnsworth E, Minoche AE, Cowley MJ, et al. Whole genome sequencing, focused assays and functional studies increasing understanding in cryptic inherited retinal dystrophies. Int J Mol Sci. 2022;23(7):3905.
Holder IT, Wagner S, Xiong P, Sinn M, Frickey T, Meyer A, et al. Intrastrand triplex DNA repeats in bacteria: a source of genomic instability. Nucleic Acids Res. 2015;43(21):10126–42.
De Bustos A, Cuadrado A, Jouve N. Sequencing of long stretches of repetitive DNA. Sci Rep. 2016;6(1):1–7.
Neidhardt J, Glaus E, Lorenz B, Netzer C, Li Y, Schambeck M, et al. Identification of novel mutations in X-linked retinitis pigmentosa families and implications for diagnostic testing. Mol Vis. 2008;14:1081.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25(14):1754–60.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20(9):1297–303.
DePristo MA, Banks E, Poplin R, Garimella KV, Maguire JR, Hartl C, et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Gen. 2011;43(5):491–8.
McLaren W, Pritchard B, Rios D, Chen Y, Flicek P, Cunningham F. Deriving the consequences of genomic variants with the Ensembl API and SNP Effect Predictor. Bioinformatics. 2010;26(16):2069–70.
De Coster W, D’Hert S, Schultz DT, Cruts M, Van Broeckhoven C. NanoPack: visualizing and processing long-read sequencing data. Bioinformatics. 2018;34(15):2666–9.
Li H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics. 2018;34(18):3094–100.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The sequence alignment/map format and SAMtools. Bioinformatics. 2009;25(16):2078–9.
Wang J, Zhang VW, Feng Y, Tian X, Li F-Y, Truong C, et al. Dependable and efficient clinical utility of target capture-based deep sequencing in molecular diagnosis of retinitis pigmentosa. Invest Ophthalmol Vis Sci. 2014;55(10):6213–23.
Huang X-F, Wu J, Lv J-N, Zhang X, Jin Z-B. Identification of false-negative mutations missed by next-generation sequencing in retinitis pigmentosa patients: a complementary approach to clinical genetic diagnostic testing. Genet Med. 2015;17(4):307–11.
Breuer DK, Yashar BM, Filippova E, Hiriyanna S, Lyons RH, Mears AJ, et al. A comprehensive mutation analysis of RP2 and RPGR in a North American cohort of families with X-linked retinitis pigmentosa. Am J Hum Genet. 2002;70(6):1545–54.
Zhang Q, Acland GM, Wu WX, Johnson JL, Pearce-Kelling S, Tulloch B, et al. Different RPGR exon ORF15 mutations in Canids provide insights into photoreceptor cell degeneration. Hum Mol Genet. 2002;11(9):993–1003.
Ebenezer ND, Michaelides M, Jenkins SA, Audo I, Webster AR, Cheetham ME, et al. Identification of novel RPGR ORF15 mutations in X-linked progressive cone-rod dystrophy (XLCORD) families. Invest Ophthalmol Vis Sci. 2005;46(6):1891–8.
Bader I, Brandau O, Achatz H, Apfelstedt-Sylla E, Hergersberg M, Lorenz B, et al. X-linked retinitis pigmentosa: RPGR mutations in most families with definite X linkage and clustering of mutations in a short sequence stretch of exon ORF15. Invest Ophthalmol Vis Sci. 2003;44(4):1458–63.
Maggi J, Roberts L, Koller S, Rebello G, Berger W, Ramesar R. De novo assembly-based analysis of rpgr exon orf15 in an indigenous african cohort overcomes limitations of a standard next-generation sequencing (NGS) data analysis pipeline. Genes (Basel). 2020;11(7):800.
Lipworth S, Pickford H, Sanderson N, Chau KK, Kavanagh J, Barker L, et al. Optimized use of Oxford Nanopore flowcells for hybrid assemblies. Microb Genom. 2020;6(11):mgen000453.
Gilpatrick T, Lee I, Graham JE, Raimondeau E, Bowen R, Heron A, et al. Targeted nanopore sequencing with Cas9-guided adapter ligation. Nat Biotechnol. 2020;38(4):433–8.
Miller DE, Sulovari A, Wang T, Loucks H, Hoekzema K, Munson KM, et al. Targeted long-read sequencing identifies missing disease-causing variation. Am J Hum Genet. 2021;108(8):1436–49.
Payne A, Holmes N, Clarke T, Munro R, Debebe BJ, Loose M. Readfish enables targeted nanopore sequencing of gigabase-sized genomes. Nat Biotechnol. 2021;39(4):442–50.
Acknowledgements
The authors would like to thank the patients who consented to be involved in and support this research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval
Informed written consent was obtained using a protocol that followed the precepts of the Declaration of Helsinki and was approved by the Leeds East Research Ethics Committee (Project reference 17/YH/0032).
Conflict of interest
Authors Samar Yahya, Christopher M Watson, Ian Carr, Martin McKibbin, Laura A Crinnion, Morag Taylor, Hope Bonin, Tracy Fletcher, Mohammed E El-Asrag, Manir Ali, Carmel Toomes and Chris F Inglehearn declare that they have no conflicts of interest that might be relevant to the contents of this manuscript.
Funding
This work was funded by RP Fighting Blindness and Fight for Sight UK (RP Genome Project GR586) and by a scholarship from the Saudi Arabian Cultural Bureau and King Abdulaziz University to S.Y. The open access fee was paid from the scholarship funding. No other sources of financial assistance were used to conduct the study or to assist with the preparation of the manuscript. Funding organisations had no role in the design and conduct of this research.
Data availability
The data that support the findings of this study are available on request from the corresponding author.
Code availability statement
Not applicable.
Author contributions
Samar Yahya contributed to study design, acquired, analysed and interpreted data and wrote the first draft of the paper. Christopher Watson contributed to study design, acquired, analysed and interpreted data and worked on early drafts of the paper. Ian Carr acquired, analysed and interpreted data and commented on the paper draft. Martin McKibbin acquired, analysed and interpreted data and commented on the paper draft. Laura Crinnion acquired data and commented on the paper draft. Morag Taylor acquired data and commented on the paper draft. Hope Bonin acquired data and commented on the paper draft. Tracey Fletcher acquired data and commented on the paper draft. Mohammed El-Asrag analysed data and commented on the paper draft. Manir Ali contributed to study design and commented on the paper draft. Carmel Toomes contributed to study design, carried out data analysis and worked on early drafts of the paper. Chris Inglehearn contributed to study design, acquired, analysed and interpreted data and worked on early drafts of the paper. All authors read and approved the final manuscript.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Yahya, S., Watson, C.M., Carr, I. et al. Long-Read Nanopore Sequencing of RPGR ORF15 is Enhanced Following DNase I Treatment of MinION Flow Cells. Mol Diagn Ther 27, 525–535 (2023). https://doi.org/10.1007/s40291-023-00656-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40291-023-00656-z