Abstract
Background
People living with HIV (PLWH) are aging and experience age-related physiological changes and comorbidities. Atorvastatin is a widely prescribed lipid-lowering agent metabolized by cytochrome P450 (CYP) 3A4, whose hepatocyte uptake is facilitated by organic anion transporting polypeptide (OATP) 1B1/1B3. Inhibition or induction of this enzyme and hepatic transporter can increase or decrease atorvastatin exposure, respectively.
Objective
This study aimed to describe the pharmacokinetic profile of atorvastatin and its major metabolite, and to evaluate drug–drug interactions (DDIs) with antiretrovirals (ARVs).
Methods
The atorvastatin pharmacokinetic profile was best described by a two-compartment model with first-order absorption and elimination. Metabolite concentrations were described by considering both linear metabolism from atorvastatin and presystemic metabolism. The influence of demographic and clinical covariates on drug and metabolite pharmacokinetics was assessed using NONMEM®. Model-based simulations were performed to evaluate the magnitude of DDIs with ARVs.
Results
Full pharmacokinetic profiles (98 atorvastatin + 62 o-OH-atorvastatin concentrations) and sparse concentrations (78 and 53 for atorvastatin and o-OH-atorvastatin, respectively) were collected in 59 PLWH. Interindividual variability was high. The coadministration of boosted ARVs decreased atorvastatin clearance by 58% and slowed down o-OH-atorvastatin formation by 88%. Atorvastatin clearance increased by 78% when coadministered with CYP3A4 inducers. Simulations revealed a 180% increase and 44% decrease in atorvastatin exposure (area under the curve) in the presence of ARVs with inhibiting and inducing properties, respectively.
Conclusion
This study showed an important interindividual variability in atorvastatin pharmacokinetics that remains largely unexplained after the inclusion of covariates. Since boosted ARVs double atorvastatin exposure, the initial dosage might be reduced by half, and titrated based on individual clinical targets.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Our findings highlight the high variability in atorvastatin pharmacokinetics, which is partially explained by drug–drug interactions with antiretroviral (ARV) treatments. |
Simulations revealed a 180% increase and 44% decrease in atorvastatin exposure (area under the curve) in the presence of ARVs with inhibiting and inducing properties, respectively. |
The present model provides a rationale for the selection of initial atorvastatin dosage, taking into account the associated ARV regimen. |
1 Introduction
People living with HIV (PLWH) live longer and experience age-related physiological changes and comorbidities, notably cardiovascular diseases. Polypharmacy is frequent in elderly PLWH, leading to an increased risk for drug–drug interactions (DDIs), which may harm this vulnerable population. Antiretroviral drugs (ARVs) are among the therapeutic agents with the highest potential for DDIs. Protease inhibitors (PIs) and non-nucleoside reverse transcriptase inhibitors (NNRTIs) can indeed inhibit and/or induce cytochrome P450 (CYP) isoforms [1] as well as drug transporters [2].
Atorvastatin is a widely prescribed lipid-lowering agent that undergoes extensive first-pass metabolism [3]. It is predominantly metabolized by CYP3A4 into two active metabolites: the major ortho-hydroxy atorvastatin (o-OH-atorvastatin) and the minor para-hydroxy atorvastatin (p-OH-atorvastatin). Both atorvastatin and its active metabolites can undergo lactonization and thus exist in equilibrium with their respective inactive lactone forms. A study has suggested that most of the acid metabolites present in human plasma results from an interconversion of lactone metabolites [4]. It has been reported that about 70% of the HMG-CoA reductase inhibition is attributable to o-OH-atorvastatin and p-OH-atorvastatin, while the lactone forms are inactive [3]. Nevertheless, the latter may be incriminated for statin-induced myotoxicity [5, 6].
Importantly, the organic anion transporting polypeptide (OATP1B1/1B3) facilitates the entry of atorvastatin in the liver (i.e. the site of action) [7]. PIs inhibit OATP1B1 in addition to CYP3A4 and are therefore expected to substantially increase atorvastatin exposure, both by inhibiting the entry of the statin in the liver and by further inhibiting its biotransformation. According to the summary of product characteristics, atorvastatin exposure could increase by three- to fourfold in the presence of ritonavir-boosted darunavir [8]. This interaction can lead to serious adverse effects, such as rhabdomyolysis [9]. The current recommendations indicate to initiate atorvastatin at a low dosage in the presence of boosted darunavir and not to exceed a daily dose of 20 mg. However, formal DDI studies have not been performed, particularly in the elderly, leading to a lack of knowledge on the magnitude of DDIs.
To date, several studies have evaluated the factors influencing atorvastatin pharmacokinetics (PK). The effect of age is controversial, with some authors reporting an age-related increase in atorvastatin exposure [10,11,12], while others did not find any significant influence [13, 14]. One non-compartmental PK study showed an effect of sex (11% decrease in area under the curve [AUC] in women) on atorvastatin disposition [11]. Moreover, population PK studies indicate a body weight-related decrease in atorvastatin clearance [13], an influence of liver enzymes (aspartate aminotransferase [AST] and lactate dehydrogenase) on atorvastatin disposition [14, 15], and an effect of polymorphisms in the intestinal breast cancer resistance protein (BCRP) on atorvastatin bioavailability [16]. However, to our knowledge, no study investigated the effect of ARVs on atorvastatin disposition in a real-life setting.
The aims of this observational study were to develop a population PK model for atorvastatin and its major active metabolite in aging PLWH, and to quantify the effect of ARVs and other covariates on their disposition.
2 Material and Methods
2.1 Data Collection
Full PK investigations were performed in the framework of a study evaluating DDIs in PLWH enrolled in the Swiss HIV Cohort Study (SHCS), as described elsewhere [17]. All study participants gave written informed consent before entering the study. The study protocol was reviewed and approved by the Ethics Committee of Vaud and northwest/central Switzerland (CER-VD 2018-00369) and registered in ClinicalTrials.gov (NCT03515772). In addition, sparse plasma samples were collected at the patient’s biannual cohort visits (SHCS project #815), at unselected times after the last drug intake. Undetectable atorvastatin plasma levels during the SHCS follow-up visits, suggestive of non-adherence to treatment, were excluded from the analysis. In addition, samples with non-reliable time information (i.e. time of blood sampling or last dose intake) were excluded from the analysis. Information on concurrent comedications (HIV and non-HIV medications), bodyweight, sex, age and liver function tests (AST and alanine aminotransferase [ALT]) were also available.
2.2 Analytical Method
Blood samples were collected in EDTA-containing tubes, immediately placed at + 4 °C. Shortly afterwards, blood samples were centrifuged and the plasma was stored at − 80 °C until analysis. Atorvastatin, o-OH-atorvastatin and p-OH-atorvastatin concentrations were determined by ultra-high-performance liquid chromatography coupled with tandem mass spectrometry (UHPLC–MS/MS) using a multiplex method developed and validated purposely for this research project [18]. The assay showed appropriate repeatability and intermediate precision for the quantification of atorvastatin and its two active metabolites o-OH-atorvastatin and p-OH-atorvastatin (coefficient of variation [CV] 2.1–13.4% and 4.2–13.4%, respectively) and trueness (98.4–110.8%). Lower limits of quantification (LLOQs) were 0.3 ng/mL for atorvastatin and p-OH-atorvastatin, and 0.5 ng/mL for o-OH-atorvastatin. In addition, darunavir and ritonavir plasma concentrations were measured using previously published LC-MS/MS methodology [19].
2.3 Model-Based Pharmacokinetic Analysis
Population PK analyses were performed using the non-linear mixed-effect modeling (NONMEM®) program (version 7.4.2, ICON Development Solutions, Ellicott City, MD, USA). PsN v4.2.0 was used for automation of various model development and evaluation methods, Pirana v2.9.2 was used to structure and document model development, and R v3.6.1 (1.2.1335) was used for data management, statistical analysis and graphical output [20, 21].
2.3.1 Base Model
Since a substantial proportion (42%) of p-OH-atorvastatin concentrations were below the LLOQ (BQL), population PK modeling was pursued for atorvastatin and its major measurable active metabolite o-OH-atorvastatin. o-OH-atorvastatin concentrations were mainly BQL over the full PK (i.e. ‘rich’) investigations and were distributed throughout the dosing interval. The population PK analysis was first performed using the samples collected during the full PK investigations for the parent atorvastatin, and subsequently using all available samples for the parent atorvastatin and the metabolite, assuming linear metabolism and integrating the first-pass effect of atorvastatin. Administered doses, atorvastatin and o-OH-atorvastatin plasma concentrations were converted into nanomoles (nmol) and nanomoles per liter (nmol/L), respectively, for the analyses of drug and metabolite data. Pharmacodynamic properties of atorvastatin and its active metabolites are generally considered equivalent [22] and the sum of both substances was defined as the ‘active moiety’. With the exception of PLWH who reported missing atorvastatin doses during the last week before their blood intake, steady state was assumed for all PLWH.
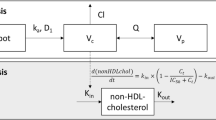
A stepwise procedure was used to find the model that adequately fitted the data. The two-compartment model for atorvastatin, with an additional compartment for o-OH-atorvastatin, schematically depicted in Fig. 2, was finally retained for data description. Presystemic metabolism was modeled by estimating a proportional coefficient (FRator-oOH) between total atorvastatin and o-OH-atorvastatin absorption rate constants (ka). This parameter allowed the description of atorvastatin presystemic metabolism by both considering the fraction of the dose directly converted into metabolite and by adjusting the o-OH-atorvastatin ka. The total ka was fixed to the value estimated during the analysis of rich parent PK data (2.59 h−1), and (1 − FRator-oOH) × ka and FRator-oOH × ka are the atorvastatin and o-OH-atorvastatin ka—k12 and k13, respectively. Owing to identifiability problems, both compounds were assumed to have the same apparent volume of distribution. Since atorvastatin was administered orally, apparent PK parameters were estimated. Exponential errors were used to describe between-subject variability for all PK parameters, with the exception of FRator-oOH. Individual FRator-oOH were constrained to vary between 0 and 1 by using the logit of FRator-oOH, and its interindividual variability was calculated as previously reported [23, 24]. Finally, several error models (i.e. proportional, additive and mixed) were compared to describe the residual variability for both drug and metabolite. The correlation between atorvastatin and its metabolite concentration measurements was tested using the L2 function in NONMEM®.
2.3.2 Covariate Model
The analysis of each covariate was sequentially examined using a stepwise insertion/deletion approach. First, correlation between post hoc individual estimates of the PK parameters and the covariates of interest were visually inspected. Potentially influential covariates were then incorporated sequentially into the model using linear or non-linear functions as appropriate. Categorical variables (sex and comedications, classified as the presence or absence of a boosted regimen or CYP3A4 inducers [25]) were coded as 0 and 1, and continuous covariates (age, weight, AST and ALT) were centered on their median value. Missing values for weight, AST and ALT were imputed to the population median value. Darunavir and ritonavir area under the concentration-time curves from zero to 24 h (AUC24) were calculated using previously published population PK models [26]. Non-competitive interaction models including darunavir and ritonavir AUC24 on CLator and FRator-oOH were tested using linear, power or exponential functions.
2.3.3 Model Selection and Parameter Estimation
Atorvastatin and metabolite concentrations were fitted by using the first-order conditional method (FOCE) with interaction using the ADVAN5 subroutine. BQL plasma levels for sparse data were excluded from the analysis. BQL concentrations for rich PK data were treated using the M6 approach, replacing BQL by LLOQ/2 at the first BQL observation and ignoring the following ones [27, 28]. The model was selected based on the likelihood ratio test (based on the reduction of the objective function value [ΔOFV]), visual inspection of diagnostic plots, and evaluation of estimates of population fixed and random parameters, along with their precision. Since an ΔOFV between two models approximates a Chi-square distribution, a decrease in the OFV > 3.84 (p < 0.05) for one additional parameter was considered statistically significant in the model building and the forward inclusion of covariates. During the backward deletion step, a covariate was retained in the final multivariate model if its deletion from the full model led to a 6.63-point increase in the objective function (p < 0.01, 1 degree of freedom).
2.4 Model Evaluation
A sensitivity analysis was carried out to assess the possible leverage effect on significant covariates due to potential outlier concentrations. Population parameters obtained when excluding data were compared with those obtained using the full dataset. In addition, 2000 datasets were generated by resampling from the original dataset for the evaluation of the final model by the bootstrap method implemented in PsN [20]. Stratification was based on the presence of boosted ARVs, CYP3A4 inducers and on rich PK sampling. Mean parameter values with their 95% confidence intervals (CI95%) were compared with the original model estimates. Finally, prediction- and variability-corrected visual predictive checks (pvcVPCs) were also performed on the final PK model with variability using the PsN-Toolkit and the R package Xpose4, to visually compare observed concentrations with 5th, 50th and 95th prediction percentiles [20, 29, 30].
2.5 Model-Based Simulations
Simulations of 1000 individuals with different ARV regimens based on the final model with between-subject variability were conducted to derive the average AUC24 with 95% prediction intervals (PI95%) for atorvastatin, o-OH-atorvastatin and the active moiety (AUCator, AUCo-OH, AUCactive moiety = AUCator + AUCo-OH).
3 Results
3.1 Data
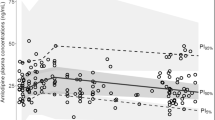
Nine PLWH contributed to 98 atorvastatin and 62 o-OH-atorvastatin plasma concentrations collected in a rich sampling design. In addition, 78 sparse atorvastatin and 53 sparse o-OH-atorvastatin concentrations from 55 PLWH were included in the analysis. Overall, 176 atorvastatin and 115 o-OH-atorvastatin plasma concentrations were available from 59 PLWH who were receiving atorvastatin at a dose ranging from 5 to 40 mg once daily. Characteristics of the study population are presented in Table 1. The median (range) of samples available per study individual was 11 (10–11) for PLWH included in the full PK study, and 1 (1–2) for PLWH whose samples were collected during the SHCS follow-up visits. Plasma concentration measurements varied from 0.3 to 106 ng/mL (0.5–190 nmol/L), and from 0.5 to 24 ng/mL (0.9–42 nmol/L), for atorvastatin and o-OH-atorvastatin, respectively. None of the atorvastatin concentrations and 38% (n = 69) of o-OH-atorvastatin levels were below their respective LLOQs. Figure 1 represents concentration-time profiles for atorvastatin and o-OH-atorvastatin according to concurrent ARV drugs.
Observed atorvastatin (left) and o-OH-atorvastatin concentrations (right) [log-scale] versus time after dose. Concentrations were standardized for a daily dose of 10 mg. Concentrations for PLWH receiving neither CYP3A4 inhibitors nor CYP3A4 inducers (grey circles), or receiving boosted ARVs (yellow squares) or strong CYP3A4 inducers (blue triangles) are shown. Plasma drug concentrations of PLWH who participated in the pharmacokinetic study with rich sampling are joined by black lines. PLWH people living with HIV, CYP cytochrome P450, ARVs antiretrovirals
In addition, darunavir and ritonavir plasma concentrations were available for PLWH receiving such ARV regimens concomitantly to atorvastatin.
3.2 Base and Covariate Model
Atorvastatin full PK profiles were best described by a two-compartment model with first-order absorption and elimination. The addition of a second compartment to describe atorvastatin disposition significantly improved the fit (variation in OFV, ∆OFV = − 113; p < 0.001) and the individual plots of rich PK data. The ka was estimated at 2.59 h−1 and was fixed to this value for subsequent model development to allow precise estimation of the other model parameters during the analysis of the full dataset. The model presented in Fig. 2 adequately described atorvastatin and o-OH-atorvastatin data. The inclusion of the factor FRator-oOH describing presytemic metabolism of atorvastatin improved the description of the data. Residual variabilities on atorvastatin and o-OH-atorvastatin were satisfactorily described using proportional and mixed-error models, respectively. The additive part of the metabolite error model was estimated at 0.44 nmol/L. Parameter estimates and between-subject variability (CV%) of the base PK model were a ka fixed to 2.59 h−1 (239%), an FRator-oOH of 11% (131%), an atorvastatin clearance (CLator) of 204 L/h (94%), a central volume of distribution of atorvastatin and o-OH-atorvastatin (Vcator=Vcmet) of 3170 L (137%), a peripheral volume of distribution (Vpator) of 591 L, an intercompartmental clearance (Q) of 104 L/h, a metabolic rate constant (k23) of 0.0096 h−1, and a metabolite clearance (CLo-OH-ator) of 118 L/h. Drug and metabolite concentrations were found to be correlated (59%).
Compartmental model used to describe atorvastatin and o-OH-atorvastatin plasma concentration-time profiles. k12 absorption rate constant from depot to the atorvastatin compartment, k13 absorption rate constant from depot to the o-OH-atorvastatin compartment, ka total absorption rate constant, FRator-oOH proportional coefficient between total atorvastatin and o-OH-atorvastatin absorption rate constants, k20 atorvastatin elimination rate constant, k30 o-OH-atorvastatin elimination rate constant, CLator apparent atorvastatin clearance, CLo-OH-ator apparent o-OH-atorvastatin clearance, Vcator apparent atorvastatin volume of distribution, Vcmet apparent o-OH-atorvastatin volume of distribution, Vpator peripheral atorvastatin volume of distribution, Q intercompartmental clearance
Due to substantial eta shrinkage on FRator-oOH (46%) in the base model, the graphic exploration was interpreted cautiously between the parameter estimate and covariates. Since visual inspection of exploratory plots did not reveal an effect of sex, weight, AST and ALT on atorvastatin and o-OH-atorvastatin PK, these covariates were not tested in the model.
The coadministration of boosted ARVs decreased atorvastatin clearance by 58% and logit(FRator-oOH) by 225% (ΔOFV < − 26; p < 0.001). Consequently, o-OH-atorvastatin was formed 88% slower in the presence of boosted ARVs than in the absence of such a regimen. Atorvastatin clearance increased by 78% in PLWH receiving CYP3A4 inducers (ΔOFV = − 21; p < 0.001). The inclusion of covariates decreased original between-subject variability on CLator and FRator-oOH, by 34 and 46%, respectively. Aging did not significantly influence FRator-oOH or CLator and was not retained in the final model (ΔOFV > − 0.4; p > 0.1). The parameter estimates of the final model and their precisions are presented in Table 2. Interaction models integrating darunavir or ritonavir AUC24 on atorvastatin clearance or FRator-oOH did not improve the description of the data. Diagnostic plots for the final model are shown in electronic supplementary Fig. S1. While the model seemed to adequately describe the absorption phase, a bias remained for very high atorvastatin concentrations, as shown on the observed concentrations versus population predictions plot. This bias was entirely due to a single individual, who was the only one treated with ritonavir-boosted darunavir concomitantly with atorvastatin at a dose of 40 mg once daily. Reassuringly, this extremely high atorvastatin dose is not recommended by the US product label of darunavir in patients treated with ritonavir-boosted darunavir and is therefore not expected to be usually encountered in clinical practice.
3.3 Model Evaluation
The sensitivity analysis performed while removing one individual with extremely high atorvastatin concentrations (11 blood samplings) did not reveal any significant influence on the estimated PK parameters, but the effect of boosted regimens on the logit(FRator-oOH) parameter decreased by 15%. o-OH-atorvastatin was formed 83% slower in the presence of boosted ARVs than in the absence of such a regimen when removing this individual. This effect was not considered significantly different compared with the results obtained with the full dataset (the o-OH-atorvastatin formation rate was reduced by 88% in the presence of boosted ARVs) and this individual was therefore maintained in the dataset.
The final model parameters, together with their bootstrap estimations, are displayed in Table 2. The model was considered reliable since the parameters were within the bootstrap CI95% and differed less than 9% from the bootstrap estimations, with the exception of between-subject variability on ka and FRator-oOH, which were 13% and 25%, respectively. CI95% of the peripheral volume was very large, despite good precision of its estimate (27%). In addition, while the bootstrap was stratified by rich sampling data, including several drug concentrations in the absorption phase, CI95% of the between-subject variability on the ka remains very wide. This reflects the difficulty of the model in accurately describing the highly variable absorption phase.
Finally, predictive performance of the model was supported by the results of the pvcVPCs (Fig. 3).
Prediction- and variability-corrected visual predictive check of the final model of atorvastatin (left) and o-OH-atorvastatin (right). Open circles represent prediction- and variability-corrected observed plasma concentration; black solid and dashed lines represent the median and PI90% of the observed data; shaded surfaces represent the model-predicted 90% confidence interval of the simulated median and PI90%; horizontal black lines are the LLOQ of atorvastatin (0.54 nmol/L) and o-OH-atorvastatin (0.87 nmol/L). In the lower panel, shaded areas represent the PI90% of the simulated (shaded surface). BQL data and close circles show the fraction of observed BQL data. LLOQ lower limit of quantification, BQL below the LLOQ, PI90% 90% prediction interval
3.4 Simulations
Model-based simulations were performed to estimate and compare AUCator, AUCo-OH and the sum of both, the AUCactive moiety, for individuals receiving atorvastatin 10 mg with different ARV regimens (Fig. 4). The simulated average AUCator indicated a 180% increase in atorvastatin exposure in PLWH receiving boosted ARVs compared with those receiving ARVs devoid of interaction potential with atorvastatin. Conversely, AUCo-OH decreased by 12% in PLWH treated with boosted regimens. In total, AUCactive moiety increased by 110% in PLWH receiving boosted ARVs compared with PLHW receiving ARVs without interaction potential. Following coadministration of CYPA4 inducers, AUCator and AUCactive moiety decreased by 44 and 31%, respectively, compared with PLWH receiving non-interacting ARV treatments.
Simulated estimates of AUC24 for atorvastatin, o-OH-atorvastatin and the active moiety in PLWH receiving ARVs not interacting with atorvastatin (grey boxes), receiving boosted ARVs (yellow boxes), or CYP3A4 inducers (blue boxes). AUC area under the concentration–time curve, PLWH people living with HIV, ARVs antiretrovirals, CYP cytochrome P450
In PLWH receiving both boosted regimens and CYP3A4 inducers, AUCator and AUCactive moiety increased by 61 and 21%, respectively.
4 Discussion
Our study provides a description of the population PK profile of atorvastatin and o-OH-atorvastatin, and quantifies the magnitude of DDIs with ARVs in real-life situations. Although parameter estimates widely differ between published population PK analyses, the reported PK parameters estimated in the present study were generally in fair concordance with overall reported values [13, 14].
The present model revealed large interindividual variability in atorvastatin PK, notably during the absorption phase, known to be affected by multiple factors. First, food has been reported to decrease atorvastatin peak concentration (Cmax) and increase time to Cmax (Tmax) [3]. Although all full PK samples were obtained under standardized conditions, this parameter was not controlled for samples collected during the follow-up visits. Since, atorvastatin is exposed to intestinal CYP3A4 during the absorption phase, CYP3A4 inhibitors and inducers may further contribute to the observed important variability in this parameter. In our PK model, this has been captured by integrating the effect of boosted regimens (all CYP3A4 inhibitors) on the absorption parameter FRator-oOH. Finally, several transporters are involved in the disposition of atorvastatin and its metabolites. Genetic polymorphisms can affect the intrinsic activity and/or expression of transporters and the observed variability in atorvastatin absorption could therefore be explained by the genetic background [10, 31]. Shitara et al. showed that OATP could play a significant role in atorvastatin absorption [32]. In addition, ABCG2 and SLCOB polymorphisms have been shown to affect atorvastatin Cmax, with no effect on elimination half-life [33,34,35], supporting an effect of transporter genotypes on atorvastatin mainly during the absorption phase. However, the lack of genotyping data in our study prevented the estimation of such an effect. In our model, due to the complexity of the absorption phase, ka was fixed to the value obtained during the analysis, using atorvastatin rich PK data to obtain a reasonable value of Tmax. Predicted Tmax values ranged from 0.5 to 3.7 h, with a median of 1.3 h, in accordance with the manufacturer’s data [36]. Studies also reported Tmax values varying from 0.5 to 2 h [37,38,39,40,41]. In addition, the ka value of 2.59 h−1 is in the range of values reported in published population PK models, varying from 0.2 to 3.5 h−1.
The present study also identified large between-subject variability in atorvastatin clearance and central volume of distribution. Although non-compartmental analyses showed an effect of age on atorvastatin disposition [10, 11], the majority of previously published population PK analyses did not report any significant influence [13, 14, 16], while one of the studies found an effect in men only [12]. In our study, this association did not reach statistical significance, although visual inspection of the plots evaluating the effect of age on atorvastatin clearance suggested a slight decrease in clearance for PLWH older than 60 years of age. This absence of age effect could result from the narrow interquartile range of age (58–71 years) in our population.
This model allowed for the evaluation of the impact of DDIs that are encountered in clinical practice. The dual inhibition of cytochromes and transporters is expected to substantially increase atorvastatin exposure [42]. Inhibition of the hepatic uptake transporter OATP1B1 is expected to reduce the entry of atorvastatin in the liver, whereas inhibition of hepatic BCRP and P-gp decreases the hepatobiliary excretion of atorvastatin. A previously published study demonstrated that inhibition of hepatic transporters of atorvastatin might yield to DDIs with the same magnitude as enzyme inhibition [38], with potential occurrence of atorvastatin toxicity. Indeed, adverse effects such as rhabdomyolysis appeared to be at least partially related to atorvastatin plasma concentrations [43, 44], and several cases of rhabdomyolysis have been reported with the simultaneous administration of moderate or strong CYP3A4 inhibitors [45,46,47,48]. Studies suggested that myotoxicity may be related to either atorvastatin lactone or hydroxylated metabolites, or both [6, 39]. However, regardless of the actual incriminated species, its formation critically depends on the disposition and the circulating concentrations of the parent statin, which keeps a determinant interest. No atorvastatin target plasma trough concentrations have been clearly established to avoid toxicity, but caution is needed when co-prescribing enzyme and transporter inhibitors with atorvastatin.
The magnitude of DDIs with atorvastatin differs between boosted regimens. Atorvastatin AUC was shown to be increased by 822% when coadministered with ritonavir-boosted atazanavir, while atorvastatin AUC increased by 200–300% and 700–800% when coadministered with ritonavir-boosted darunavir or ritonavir-boosted tipranavir, respectively [37]. Differences in the magnitude of DDIs have been attributed to differences in the ability of PIs to inhibit OATP1B1 [49]. In our study, the lack of data prevented us from differentiating the effect of different boosted regimens on atorvastatin and o-OH-atorvastatin exposure. Model-based simulations revealed a 2.8-fold increase in AUCator when coadministered with boosted regimens that were mostly boosted darunavir. This result is in good agreement with the manufacturer’s data reporting a three- to fourfold increase in atorvastatin AUC when coadministered with ritonavir-boosted darunavir. In addition, another study showed that atorvastatin AUC increased by 290% in cases of coadministration of cobicistat-boosted darunavir [50]. To our knowledge, no study has reported the effect of boosted regimens on the active moiety, which is modulated by the decrease in o-OH-atorvastatin exposure. Our results demonstrated that PLWH receiving concomitantly boosted regimens and atorvastatin at a daily dose of 10 mg obtained an atorvastatin exposure 29% lower than PLWH receiving atorvastatin alone at a daily dose of 40 mg. This result is slightly different, with manufacturer’s data reporting a difference of 15% [8]. Conversely, AUCator and AUCactive moiety were 44 and 31% lower, respectively, in PLWH receiving CYP3A4 inducers compared with PLWH receiving ARVs not involved in DDIs with atorvastatin. This is in perfect agreement with studies reporting a moderate magnitude of DDIs between atorvastatin and NNRTIs [51, 52]. Finally, the inducing effect of NNRTIs on CYP3A4 partly compensates for the magnitude of DDIs with boosted ARVs. Of interest, when coadministered together, the inhibitory effect of boosted regimens was shown to be stronger than the concurrent inducing effect of NNRTIs, as evidenced by the fact that AUCator and AUCactive moiety increased by 61 and 21%, respectively.
This study has some limitations. First, the small sample size prevented us from differentiating the effect of different boosted regimens. However, atorvastatin PK data in PLWH are limited in the literature and this work aims to expand the current knowledge on DDIs in a real-life setting. In addition, the effect of boosted ARVs on the magnitude of DDIs could have been slightly attenuated as among the 80 concentrations obtained in PLWH treated with boosted regimens, 12 (15%) were also influenced by CYP3A4 inducers.
Despite these limitations, our study is the first to describe atorvastatin and o-OH-atorvastatin disposition by considering the first-pass and presystemic metabolism. The availability of rich PK data with concentrations collected in the absorption phase allowed for a satisfactory description of the entire concentration-time profile of atorvastatin and its metabolite. In addition, data collected in a real-life setting evidenced the high between-subject variability, which is partly explained by DDIs.
5 Conclusions
The present study showed an important between-subject variability in atorvastatin PK that remained largely unexplained after the inclusion of covariates. Since atorvastatin exposure doubles in the presence of boosted ARV drugs, the initial dosage might be reduced by half, and titrated based on clinical risk factors and targets.
References
Barry M, Mulcahy F, Merry C, Gibbons S, Back D. Pharmacokinetics and potential interactions amongst antiretroviral agents used to treat patients with HIV infection. Clin Pharmacokinet. 1999;36(4):289–304.
Moss D, Siccardi M, Marzolini C. Mechanisms of drug interactions II: transport proteins. In: Pai M, Kiser J, Gubbins P, Rodvold K, editors. Drug interactions in infectious diseases: mechanisms and models of drug interactions. Infectious Disease. Cham: Humana Press; 2018. p. 49–85.
Lennernas H. Clinical pharmacokinetics of atorvastatin. Clin Pharmacokinet. 2003;42(13):1141–60.
Jacobsen W, Kuhn B, Soldner A, Kirchner G, Sewing KF, Kollman PA, et al. Lactonization is the critical first step in the disposition of the 3-hydroxy-3-methylglutaryl-CoA reductase inhibitor atorvastatin. Drug Metab Dispos. 2000;28(11):1369–78.
Skottheim IB, Gedde-Dahl A, Hejazifar S, Hoel K, Asberg A. Statin induced myotoxicity: the lactone forms are more potent than the acid forms in human skeletal muscle cells in vitro. Eur J Pharm Sci. 2008;33(4–5):317–25.
Hermann M, Bogsrud MP, Molden E, Asberg A, Mohebi BU, Ose L, et al. Exposure of atorvastatin is unchanged but lactone and acid metabolites are increased several-fold in patients with atorvastatin-induced myopathy. Clin Pharmacol Ther. 2006;79(6):532–9.
Wu X, Whitfield LR, Stewart BH. Atorvastatin transport in the Caco-2 cell model: contributions of P-glycoprotein and the proton-monocarboxylic acid co-transporter. Pharm Res. 2000;17(2):209–15.
Janssen-Cilag. Prezista Summary of Product Characteristics. Janssen-Cilag; 2012. https://www.ema.europa.eu/en/documents/product-information/prezista-epar-product-information_en.pdf
Neuvonen PJ, Niemi M, Backman JT. Drug interactions with lipid-lowering drugs: mechanisms and clinical relevance. Clin Pharmacol Ther. 2006;80(6):565–81.
DeGorter MK, Tirona RG, Schwarz UI, Choi YH, Dresser GK, Suskin N, et al. Clinical and pharmacogenetic predictors of circulating atorvastatin and rosuvastatin concentrations in routine clinical care. Circ Cardiovasc Genet. 2013;6(4):400–8.
Gibson DM, Bron NJ, Richens A, Hounslow NJ, Sedman AJ, Whitfield LR. Effect of age and gender on pharmacokinetics of atorvastatin in humans. J Clin Pharmacol. 1996;36(3):242–6.
Schwartz JB, Verotta D. Population analyses of atorvastatin clearance in patients living in the community and in nursing homes. Clin Pharmacol Ther. 2009;86(5):497–502.
Knebel W, Gastonguay MR, Malhotra B, El-Tahtawy A, Jen F, Gandelman K. Population pharmacokinetics of atorvastatin and its active metabolites in children and adolescents with heterozygous familial hypercholesterolemia: selective use of informative prior distributions from adults. J Clin Pharmacol. 2013;53(5):505–16.
Narwal R, Akhlaghi F, Asberg A, Hermann M, Rosenbaum SE. Development of a population pharmacokinetic model for atorvastatin acid and its lactone metabolite. Clin Pharmacokinet. 2010;49(10):693–702.
Macwan JS, Sam WJ, Gohh R, Akhlaghi F. Development of a complex combined parent-metabolite population pharmacokinetic model for atorvastatin acid and its lactone metabolite: implication of renal transplantation. J Pharmacokinet Pharmacodyn. 2013;40:S73–4.
Tsamandouras N, Guo Y, Wendling T, Hall S, Galetin A, Aarons L. Modelling of atorvastatin pharmacokinetics and the identification of the effect of a BCRP polymorphism in the Japanese population. Pharmacogenet Genomics. 2017;27(1):27–38.
Courlet P, Stader F, Guidi M, Saldanha SA, Stoeckle M, Cavassini M, et al. Pharmacokinetic profiles of boosted darunavir, dolutegravir and lamivudine in aging patients enrolled in the Swiss HIV Cohort Study. Aids. 2020;34(1):103–8.
Courlet P, Spaggiari D, Desfontaine V, Cavassini M, Alves Saldanha S, Buclin T, et al. UHPLC-MS/MS assay for simultaneous determination of amlodipine, metoprolol, pravastatin, rosuvastatin, atorvastatin with its active metabolites in human plasma, for population-scale drug-drug interactions studies in people living with HIV. J Chromatogr B Anal Technol Biomed Life Sci. 2019;1125:121733.
Fayet A, Beguin A, Zanolari B, Cruchon S, Guignard N, Telenti A, et al. A LC-tandem MS assay for the simultaneous measurement of new antiretroviral agents: raltegravir, maraviroc, darunavir, and etravirine. J Chromatogr B Anal Technol Biomed Life Sci. 2009;877(11–12):1057–69.
Lindbom L, Pihlgren P, Jonsson EN. PsN-Toolkit—a collection of computer intensive statistical methods for non-linear mixed effect modeling using NONMEM. Comput Methods Programs Biomed. 2005;79(3):241–57.
Keizer RJ, van Benten M, Beijnen JH, Schellens JH, Huitema AD. Pirana and PCluster: a modeling environment and cluster infrastructure for NONMEM. Comput Methods Programs Biomed. 2011;101(1):72–9.
European Medicine Agency. Atorvastatin, summary of product characteristics. Available at: https://www.ema.europa.eu/en/documents/referral/lipitor-article-30-referral-annex-iii_en.pdf.
Vandenberghe F, Guidi M, Choong E, von Gunten A, Conus P, Csajka C, et al. Genetics-based population pharmacokinetics and pharmacodynamics of risperidone in a psychiatric cohort. Clin Pharmacokinet. 2015;54(12):1259–72.
Bouzom F, Laveille C, Merdjan H, Jochemsen R. Use of nonlinear mixed effect modeling for the meta-analysis of preclinical pharmacokinetic data: application to S 20342 in the rat. J Pharm Sci. 2000;89(5):603–13.
US FDA. Drug Development and Drug Interactions: Table of Substrates, Inhibitors and Inducers. Available at: https://www.fda.gov/drugs/developmentapprovalprocess/developmentresources/druginteractionslabeling/ucm093664.htm.
Arab-Alameddine M, Lubomirov R, Fayet-Mello A, Aouri M, Rotger M, Buclin T, et al. Population pharmacokinetic modelling and evaluation of different dosage regimens for darunavir and ritonavir in HIV-infected individuals. J Antimicrob Chemother. 2014;69(9):2489–98.
Bergstrand M, Karlsson MO. Handling data below the limit of quantification in mixed effect models. AAPS J. 2009;11(2):371–80.
Beal SL. Ways to fit a PK model with some data below the quantification limit. J Pharmacokinet Pharmacodyn. 2001;28(5):481–504.
Bergstrand M, Hooker AC, Wallin JE, Karlsson MO. Prediction-corrected visual predictive checks for diagnosing nonlinear mixed-effects models. AAPS J. 2011;13(2):143–51.
Jonsson EN, Karlsson MO. Xpose: an S-PLUS based population pharmacokinetic/pharmacodynamic model building aid for NONMEM. Comput Methods Programs Biomed. 1999;58(1):51–64.
Niemi M. Transporter pharmacogenetics and statin toxicity. Clin Pharmacol Ther. 2010;87(1):130–3.
Shitara Y, Maeda K, Ikejiri K, Yoshida K, Horie T, Sugiyama Y. Clinical significance of organic anion transporting polypeptides (OATPs) in drug disposition: their roles in hepatic clearance and intestinal absorption. Biopharm Drug Dispos. 2013;34(1):45–78.
Woo HI, Kim SR, Huh W, Ko JW, Lee SY. Association of genetic variations with pharmacokinetics and lipid-lowering response to atorvastatin in healthy Korean subjects. Drug Des Devel Ther. 2017;11:1135–46.
Keskitalo JE, Zolk O, Fromm MF, Kurkinen KJ, Neuvonen PJ, Niemi M. ABCG2 polymorphism markedly affects the pharmacokinetics of atorvastatin and rosuvastatin. Clin Pharmacol Ther. 2009;86(2):197–203.
Li R, Barton HA. Explaining ethnic variability of transporter substrate pharmacokinetics in healthy asian and caucasian subjects with allele frequencies of OATP1B1 and BCRP: a mechanistic modeling analysis. Clin Pharmacokinet. 2018;57(4):491–503.
Swiss Medic. Information sur le médicament. Available at: http://www.swissmedicinfo.ch.
Pham PA, la Porte CJ, Lee LS, van Heeswijk R, Sabo JP, Elgadi MM, et al. Differential effects of tipranavir plus ritonavir on atorvastatin or rosuvastatin pharmacokinetics in healthy volunteers. Antimicrob Agents Chemother. 2009;53(10):4385–92.
Lau YY, Huang Y, Frassetto L, Benet LZ. effect of OATP1B transporter inhibition on the pharmacokinetics of atorvastatin in healthy volunteers. Clin Pharmacol Ther. 2007;81(2):194–204.
Kantola T, Kivisto KT, Neuvonen PJ. Effect of itraconazole on the pharmacokinetics of atorvastatin. Clin Pharmacol Ther. 1998;64(1):58–65.
Bullman J, Nicholls A, Van Landingham K, Fleck R, Vuong A, Miller J, et al. Effects of lamotrigine and phenytoin on the pharmacokinetics of atorvastatin in healthy volunteers. Epilepsia. 2011;52(7):1351–8.
Backman JT, Luurila H, Neuvonen M, Neuvonen PJ. Rifampin markedly decreases and gemfibrozil increases the plasma concentrations of atorvastatin and its metabolites. Clin Pharmacol Ther. 2005;78(2):154–67.
Chauvin B, Drouot S, Barrail-Tran A, Taburet AM. Drug-drug interactions between HMG-CoA reductase inhibitors (statins) and antiviral protease inhibitors. Clin Pharmacokinet. 2013;52(10):815–31.
Ucar M, Mjorndal T, Dahlqvist R. HMG-CoA reductase inhibitors and myotoxicity. Drug Saf. 2000;22(6):441–57.
Heerey A, Barry M, Ryan M, Kelly A. The potential for drug interactions with statin therapy in Ireland. Ir J Med Sci. 2000;169(3):176–9.
Hsiao SH, Chang HJ, Hsieh TH, Kao SM, Yeh PY, Wu TJ. Rhabdomyolysis caused by the moderate CYP3A4 inhibitor fluconazole in a patient on stable atorvastatin therapy: a case report and literature review. J Clin Pharm Ther. 2016;41(5):575–8.
Nandy A, Gaini S. Severe rhabdomyolysis as complication of interaction between atorvastatin and fusidic acid in a patient in lifelong antibiotic prophylaxis: a dangerous combination. Case Rep Med. 2016;2016:4705492.
Kahri J, Valkonen M, Backlund T, Vuoristo M, Kivisto KT. Rhabdomyolysis in a patient receiving atorvastatin and fluconazole. Eur J Clin Pharmacol. 2005;60(12):905–7.
Mah Ming JB, Gill MJ. Drug-induced rhabdomyolysis after concomitant use of clarithromycin, atorvastatin, and lopinavir/ritonavir in a patient with HIV. AIDS Patient Care STDs. 2003;17(5):207–10.
Annaert P, Ye ZW, Stieger B, Augustijns P. Interaction of HIV protease inhibitors with OATP1B1, 1B3, and 2B1. Xenobiotica. 2010;40(3):163–76.
Custodio JM, West S, SenGupta D, Zari A, Humeniuk R, Ling KH, et al. Evaluation of the drug-drug interaction potential between cobicistat-boosted protease inhibitors and statins [abstract no. O_04]. In: 18th international workshop on clinical pharmacology of antiviral therapy; 14–16 June 2017; Chicago.
Schöller-Gyüre M, Kakuda TN, De Smedt G, Woodfall B, Bollen S, Peeters M, Vandermeulen K, Hoetelmans RM, et al. Pharmacokinetic interaction between the non-nucleoside reverse transcriptaseinhibitors (NNRTI) TMC125 and atorvastatin in HIV-negative volunteers. In: IAS Conference on HIV Pathogenesis, Treatment and Prevention. Sydney:NSW;2007.
Gerber JG, Rosenkranz SL, Fichtenbaum CJ, Vega JM, Yang A, Alston BL, et al. Effect of efavirenz on the pharmacokinetics of simvastatin, atorvastatin, and pravastatin: results of AIDS Clinical Trials Group 5108 Study. J Acquir Immune Defic Syndr. 2005;39(3):307–12.
Acknowledgements
The authors would like to thank all patients who agreed to participate in the studies, as well as the following persons for their invaluable help and implication in blood sample collection, as study physicians, study coordinators and study nurses: Anne-Sophie Brunel, Benjamin Viala, Chiara Saracci, Dan Lebowitz, Katharine Darling, Deolinda Alves, Vreneli Waelti Da Costa, Alexandra Mitouassiwou-Samba, Valérie Sormani (Lausanne Center), and Manuel Battegay, Marcel Stoeckle, Irena Ferati, Kerstin Asal, Rebekka Plattner, Reinhild Harant, Silke Purschke, Vanessa Grassedonio, Vreni Werder (Basel Center).
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Funding
This work was supported by two Swiss National Science Foundation grants (Grant SNF No. 324730-165956 to LAD (Lausanne) and Grant No. 324730-166204 to CM (Basel), as well as Grants from the OPO and Isaac Dreyfus Foundations to CM (Basel). The SHCS #815 study received the 2017 SHCS Abbvie Award (awarded to LAD and CM).
Conflict of interest
Perrine Courlet, Laurent A. Decosterd, Susana Alves Saldanha, Felix Stader, Thierry Buclin, Catia Marzolini, Chantal Csajka, and Monia Guidi have no conflicts of interest to declare. Matthias Cavassini has, through his institution, received research grants from ViiV and Gilead, and offered expert testimony for Abbvie, MSD, Gilead, and Sandoz. Marcel Stoeckle received advisory board fees from Gilead, ViiV, MSD, Sandoz and Mepha, as well as grants for conferences from Gilead and MSD, however these were unrelated to the present study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
40262_2020_876_MOESM1_ESM.pdf
Supplementary material 1 (PDF 137 kb) Supplementary file S1: Goodness-of-fit plots of atorvastatin (1) and o-OH-atorvastatin (2) final population pharmacokinetic model. Loess smooth curves are shown in grey. (a) Observed concentrations versus individual predictions; the line of identity is shown in black. (b) Observed concentrations versus population predictions; the line of identity is shown in black. (c) CWRES versus population predictions; the ordinate value zero line is shown in black. (d) CWRES versus time postdose; the ordinate value zero line is shown in black. CWRES conditional weighted residuals
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Courlet, P., Decosterd, L.A., Alves Saldanha, S. et al. Influence of Drug–Drug Interactions on the Pharmacokinetics of Atorvastatin and Its Major Active Metabolite ortho-OH-Atorvastatin in Aging People Living with HIV. Clin Pharmacokinet 59, 1037–1048 (2020). https://doi.org/10.1007/s40262-020-00876-0
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40262-020-00876-0