Abstract
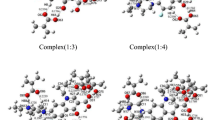
Molecular simulations are widely used to model molecularly imprinted polymers(MIPs) in order to enhance their adsorption and selectivity. In this study, chloramphenicol(CAP) and acrylamide(AM) were used as the template and functional monomer, respectively, and pentaerythritol triacrylate(PETA), ethylene glycol dimethacrylate (EGDMA), and trimethylolpropane trimethylacrylate(TRIM) were used as cross-linking agents. The ωB97XD/6-31G(d,p) density functional theory method was employed to simulate binding sites, binding energy, the number of hydrogen bonds, the imprinted molar ratio, which produced the most stable complex, and the interaction mechanism. The cross-linking agent was optimized based on the binding energy. The atoms in molecules theory were used to study the nature of the imprinting effects. The theoretical calculations revealed that CAP and AM formed ordered complexes via hydrogen bonding interactions when the molar ratio between CAP and AM was 1:7 using TRIM as the cross-linking agent. The CAP-AM complex(molar ratio 1:7) had the most stable structure, the largest number of hydrogen bonds, and the smallest ΔE. The experimental results indicate that the CAP-MIPs formed perfect microspheres with an average particle size of 314 nm. Scatchard plot analysis showed that the CAP-MIPs had only one type of binding site over the studied concentration ranges. The dissociation equilibrium constant and maximum apparent adsorption capacities were 1887.35 mg/L(5.84 mmol/L) and 155.56 mg/g(0.482 mmol/g), respectively.
Similar content being viewed by others
References
Chen H. X., Ying J., Chen H., Huang J. L., Liao L., Chromatogr., 2008, 68, 629
Yusof N. A., Rahman S. K. A., Hussein M. Z., Ibrahim N. A., Polymers, 2013, 5, 1215
Douny C., Widart J., Pauw E. D., Maghuin-Rogister G., Scippo M. L., Food Anal. Methods, 2013, 6, 1458
Xu Z. X., Gao H. J., Zhang L. M., Chen X. Q., Qian X. G., J. Food Sci., 2015, 76, R69
Liang W. X., Hu H. W., Guo P. R., Ma Y. F., Li P. Y., Zheng W. R., Zhang M. J., Sci. Food Agr., 2014, 94, 1409
Cheong W. J., Yang S. H., Ali F. D., J. Sep. Sci., 2013, 36, 60
Zhang Y., Qian L., Yin W., He B., Liu F. M., Hou C. J., Huo D. Q., Fa H. B., Chem. Res. Chinese Universities, 2016, 32(5), 725
Huang Y. X., Lian H. T., Sun X.Y., Liu B., Chem. Res. Chinese Universities, 2011, 27(1), 28
Wulff G., Liu J. Q., Acc. Chem. Res., 2012, 45, 239
Liang D. D., Wang Y., Li S.Y., Li Y. Q., Zhang M. L., Li Y., Tian W. S., Liu J. B., Tang S. S., Li B., Jin R. F., Int. J. Mol. Sci., 2016, 17, 1750
Liu J. B., Shi Y., Tang S. S., Jin R. F., Struc. Chem., 2016, 7, 897
Liu J. B., Wang Y., Su T. T., Li B., Tang S. S., Jin R. F., Struc. Chem., 2016, 7, 1135
Liu J. B., Wang Y., Tang S. S., Gao Q., Jin R. F., New J. Chem., 2017, 41, 13370
Turiel E., Martínesteban A., Comput. Theor. Chem., 2017, 1108, 76
Yu H., Koide H., Urakami T., Kanazawa H., Kodama T., Oku N., Shea K. J., Comput. Theor. Chem., 2017, 1117, 130
Zhu R., Zhao W. H., Zhai M. J., Wei F. D., Cai Z., Sheng N., Hu Q., Anal. Chim. Acta, 2010, 658, 209
Wang Y. D., Wang E. L., Dong H., Liu F., Wu Z. M., Li H., Wang Y., Adsorpt. Sci. Technol., 2014, 32, 321
Chen H. Y., Ding L., Liu M. L., Chem. J. Chinese Universities, 2015, 36(1), 67
Frisch M. J., Trucks G. W., Schlegel H. B., Scuseria G. E., Robb M. A., Cheeseman J. R., Scalmani G., Barone V., Mennucci B., Petersson G. A., Nakatsuji H., Caricato M., Li X., Hratchian H. P., Izmaylov A. F., Bloino J., Zheng G., Sonnenberg J. L., Hada M., Ehara M., Toyota K., Fukuda R., Hasegawa J., Ishida M., Nakajima T., Honda Y., Kitao O., Nakai H., Vreven T., Montgomery J. A. Jr., Peralta J. E., Ogliaro F., Bearpark M., Heyd J. J., Brothers E., Kudin K. N., Staroverov V. N., Kobayashi R., Normand J., Raghavachari K., Rendell A., Burant J. C., Iyengar S. S., Tomasi J., Cossi M., Rega N., Millam J. M., Klene M., Knox J. E., Cross J. B., Bakken V., Adamo C., Jaramillo J., Gomperts R., Stratmann R. E., Yazyev O., Austin A. J., Cammi R., Pomelli C., Ochterski J. W., Martin R. L., Morokuma K., Zakrzewski V. G., Voth G. A., Salvador P., Dannenberg J. J., Dapprich S., Daniels A. D., Farkas Ö., Foresman J. B., Ortiz J. V., Cioslowski J., Fox D. J., Gaussian 09(Revision A.02), Gaussian, Inc., Wallingford CT, 2009
Liu J. B., Wang G. Y., Tang S. S., Gao Q., Liang D. D., Jin R. F., J. Sep. Sci., 2019, 42, 769
Achary K. R., Gowda D. S. S., Post M., Mat. Sci. Eng. C: Mater., 2013, 33, 189
Steiner T., Angew. Chem. Int. Ed., 2002, 41, 49
Jeffrey G. A., Crystallogr. Rev., 2003, 9, 135
Steiner T., Crystallogr. Rev., 2003, 9, 177
Lipkowski P., Grabowski S. J., Robinson T. L., Leszczynski J., J. Phys. Chem. A, 2004, 108, 10865
Rozas I., Alkorta A. I., Elguero J., J. Am. Chem. Soc., 2000, 122, 11154
Author information
Authors and Affiliations
Corresponding author
Additional information
Supported by the National Natural Science Foundation of China(No.21563002) and the Science and Technology Development Program Project of Jilin Province, China(No.JJKH20170299KJ).
Rights and permissions
About this article
Cite this article
Liu, J., Zhao, W., Tang, S. et al. Theoretical Design and Adsorption Properties of Molecularly Imprinted Polymers Obtained from Chloramphenicol and Acrylamide. Chem. Res. Chin. Univ. 36, 915–920 (2020). https://doi.org/10.1007/s40242-019-9267-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40242-019-9267-2