Abstract
Objectives
To introduce bibliometric features of Iranian documents on microbiota and to provide descriptive information about retrieved documents related to the medical sciences and documents utilizing molecular techniques for microbiota detection.
Methods
This is a descriptive bibliometric study of all Iranian documents on microbiota in any language that were indexed in Scopus before 7 September 2019. We assessed the research performance through statistical analysis of the bibliometric indicators, including number of publications, citations, institutions and journals activities, co-citations and bibliographic couplings, and network analysis of co-authorships, countries’ collaborations, terms and keywords.
Results
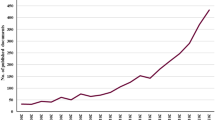
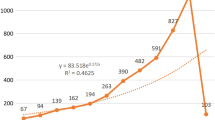
We extracted 425 relevant documents, 260 of which pertain to the medical sciences. The most focused microbiota modulating interventions and diseases in 33 clinical trials are ‘synbiotics’ (n = 8) and ‘probiotics’ (n = 8), and ‘Obesity’ (n = 3) and ‘non-alcoholic fatty liver disease’ (n = 3), respectively. During the last decade, Iranian microbiota publications have increasingly grown with a constant upward slope, particularly in the area of medical sciences after 2016. Citation counting reveals that originals and reviews have been cited 4221 times, with an average 10.76 citations and H-index of 34. The most significant performance in publishing Iranian microbiota documents belongs to ‘Tehran University of Medical Sciences’ as the active institution (n = 89 publications) and the supporting sponsor (n = 19), ‘Microbial Pathogenesis’ as the productive journal (n = 12), ‘Seidavi A’ as the most authorships (n = 19), and ‘the United States’ as the collaborative country (n = 46).
Conclusions
The qualitative and quantitative information of this study will be a practical guidance for future study planning and policy-decision making.






Similar content being viewed by others
Data availability
Not applicable.
References
Sokolov-Mladenović S, Cvetanović S, Mladenović I. R&D expenditure and economic growth: EU28 evidence for the period 2002–2012. Econ Res-Ekonomska istraživanja. 2016;29(1):1005–20.
Sharifi V, Rahimi Movaghar A, Mohammadi M, Goodarzi R, Izadian E, Farhoudian A et al. Analysis of mental health research in the Islamic Republic of Iran over 3 decades: a scientometric study. 2008.
Ataie-Ashtiani B. Chinese and Iranian scientific publications: fast growth and poor ethics. Sci Eng Ethics. 2017;23(1):317–9.
Akhondzadeh S. Iranian science shows world's fastest growth: ranks 17th in science production in 2012. Avicenna J Med Biotechnol. 2013;5(3):139.
Agarwal A, Durairajanayagam D, Tatagari S, Esteves SC, Harlev A, Henkel R, et al. Bibliometrics: tracking research impact by selecting the appropriate metrics. Asian J Androl. 2016;18(2):296–309.
Li Y, Zou Z, Bian X, Huang Y, Wang Y, Yang C, et al. Fecal microbiota transplantation research output from 2004 to 2017: a bibliometric analysis. PeerJ. 2019;7:e6411.
Pritchard A. Statistical bibliography or bibliometrics. J Doc. 1969;25(4):348–9.
Turnbaugh PJ, Ley RE, Hamady M, Fraser-Liggett CM, Knight R, Gordon JI. The human microbiome project. Nature. 2007;449(7164):804.
Baudoin L, Sapinho D, Maddi A, Miotti L. Scientometric analysis of the term “microbiota” in research publications (1999-2017): A second youth of a century-old concept. FEMS Microbiol Lett. 2019;366(12).
The unseen life of the soil. Science. 1927;65(1695):x–x. https://doi.org/10.1126/science.65.1695.0x.
Crawford JJ, Shankle RJ. Application of newer methods to study the importance of root canal and oral microbiota in endodontics. Oral Surg Oral Med Oral Pathol. 1961;14(9):1109–23.
Gibbons R, Socransky S, Sawyer S, Kapsimalis B, MacDonald J. The microbiota of the gingival crevice area of man—II: the predominant cultivable organisms. Arch Oral Biol. 1963;8(3):281–9.
Socransky S, Gibbons R, Dale A, Bortnick L, Rosenthal E, Macdonald J. The microbiota of the gingival crevice area of man—I: Total microscopic and viable counts and counts of specific organisms. Arch Oral Biol. 1963;8(3):275–80.
Mann S, Masson FM, Oxford A. Effect of feeding aureomycin to calves upon the establishment of their normal rumen microflora and microfauna. Br J Nutr. 1954;8(3):246–52.
Baldwin R, Wood W, Emery R. Conversion of glucose-C14 to propionate by the rumen microbiota. J Bacteriol. 1963;85(6):1346–9.
Mraz O, Cerny L, editors. Functional Composition+ Main Metabolites of Rumen Microflora in Cattle with Definite Food Intake. FOLIA MICROBIOLOGICA; 1964: INST MICROBIOLOGY, VIDENSKA 1083, PRAGUE 4 142 20, CZECH REPUBLIC.
Peterson J, Garges S, Giovanni M, McInnes P, Wang L, Schloss JA, et al. The NIH human microbiome project. Genome Res. 2009;19(12):2317–23.
Regan NHMPATlpngLPJLAMPDDXRFLBRRPMK. A review of 10 years of human microbiome research activities at the US National Institutes of Health, Fiscal Years 2007–2016. Microbiome. 2019;7:1–19.
O’Mahony SM, Clarke G, Borre Y, Dinan T, Cryan J. Serotonin, tryptophan metabolism and the brain-gut-microbiome axis. Behav Brain Res. 2015;277:32–48.
Kamada N, Chen GY, Inohara N, Núñez G. Control of pathogens and pathobionts by the gut microbiota. Nat Immunol. 2013;14(7):685–90.
Lemon KP, Armitage GC, Relman DA, Fischbach MA. Microbiota-targeted therapies: an ecological perspective. Sci Transl Med. 2012;4(137):137rv5-rv5.
Collado MC, Cernada M, Baüerl C, Vento M, Pérez-Martínez G. Microbial ecology and host-microbiota interactions during early life stages. Gut Microbes. 2012;3(4):352–65.
Bar-Ilan J. Citations to the “Introduction to informetrics” indexed by WOS, Scopus and Google Scholar. Scientometrics. 2010;82(3):495–506.
Van Eck N, Waltman L. Software survey: VOSviewer, a computer program for bibliometric mapping. Scientometrics. 2009;84(2):523–38.
Team S. Science of science (Sci2) tool. Indiana University and SciTech Strategies. 2009.
Nazemi M, Amini M, Salehi R. Difference in quantities of bifidobacteria from the intestinal microflora of individuals with type 2 diabetes and healthy individuals from Iran. Journal of Isfahan Medical School. 2013;31(247):1216–1225.
Nasrollahzadeh D, Malekzadeh R, Ploner A, Shakeri R, Sotoudeh M, Fahimi S, et al. Variations of gastric corpus microbiota are associated with early esophageal squamous cell carcinoma and squamous dysplasia. Sci Rep. 2015;5:8820.
Heidarian F, Noormohammadi Z, Aghdaei HA, Alebouyeh M. Relative abundance of streptococcus spp. and its association with disease activity in inflammatory bowel disease patients compared with controls. Arch Clin Infect Dis. 2017;12(2) e57291.
Nabizadeh E, Jazani NH, Bagheri M, Shahabi S. Association of altered gut microbiota composition with chronic urticaria. Ann Allergy Asthma Immunol. 2017;119(1):48–53.
Navab-Moghadam F, Sedighi M, Khamseh ME, Alaei-Shahmiri F, Talebi M, Razavi S, et al. The association of type II diabetes with gut microbiota composition. Microb Pathog. 2017;110:630–6.
Sedighi M, Razavi S, Navab-Moghadam F, Khamseh ME, Alaei-Shahmiri F, Mehrtash A, et al. Comparison of gut microbiota in adult patients with type 2 diabetes and healthy individuals. Microb Pathog. 2017;111:362–9.
Zamani S, Shariati SH, Zali MR, Aghdaei HA, Asiabar AS, Bokaie S, et al. Detection of enterotoxigenic Bacteroides fragilis in patients with ulcerative colitis. Gut Pathog. 2017;9(1):53.
Ghavami SB, Rostami E, Sephay AA, Shahrokh S, Balaii H, Aghdaei HA, et al. Alterations of the human gut Methanobrevibacter smithii as a biomarker for inflammatory bowel diseases. Microb Pathog. 2018;117:285–9.
Rezasoltani S, Aghdaei HA, Dabiri H, Sepahi AA, Modarressi MH, Mojarad EN. The association between fecal microbiota and different types of colorectal polyp as precursors of colorectal cancer. Microb Pathog. 2018;124:244–9.
Rezasoltani S, Sharafkhah M, Aghdaei HA, Mojarad EN, Dabiri H, Sepahi AA, et al. Applying simple linear combination, multiple logistic and factor analysis methods for candidate fecal bacteria as novel biomarkers for early detection of adenomatous polyps and colon cancer. J Microbiol Methods. 2018;155:82–8.
Al-Bayati L, Fasaei BN, Merat S, Bahonar A. Longitudinal analyses of gut-associated bacterial microbiota in ulcerative colitis patients. Arch Iran Med (AIM). 2018;21(12).
Tavasoli S, Alebouyeh M, Naji M, Shakiba majd G, Shabani Nashtaei M, Broumandnia N, et al. The association of the intestinal oxalate degrading bacteria with recurrent calcium kidney stone formation and hyperoxaluria: a case-control study. BJU Int. 2019;125(1):133–143.
Mohammadzadeh N, Kalani BS, Bolori S, Azadegan A, Gholami A, Mohammadzadeh R, et al. Identification of an intestinal microbiota signature associated with hospitalized patients with diarrhea. Acta Microbiol Immunol Hung. 2019:1–14.
Heidarian F, Alebouyeh M, Shahrokh S, Balaii H, Zali MR. Altered fecal bacterial composition correlates with disease activity in inflammatory bowel disease and the extent of IL8 induction. Curr Res Transl Med. 2019;67(2):41–50.
Moossavi S, Engen PA, Ghanbari R, Green SJ, Naqib A, Bishehsari F, et al. Assessment of the impact of different fecal storage protocols on the microbiota diversity and composition: a pilot study. BMC Microbiol. 2019;19(1):145.
Nami Y, Haghshenas B, Khosroushahi AY. Molecular identification and probiotic potential characterization of lactic acid Bacteria isolated from human vaginal microbiota. Adv Pharm Bull. 2018;8(4):683–95.
Mousavi SH, Mehrara S, Barzegari A, Ostadrahimi A. Correlation of gut microbiota profile with body mass index among school age children. Iran Red Crescent Med J. 2018;20(4):e58049 (In Press).
Ejtahed H-S, Tito RY, Siadat S-D, Hasani-Ranjbar S, Hoseini-Tavassol Z, Rymenans L, et al. Metformin induces weight loss associated with gut microbiota alteration in non-diabetic obese women: a randomized double-blind clinical trial. Eur J Endocrinol. 2018;180(3):165–176(aop).
Payahoo L, Khajebishak Y, Alivand MR, Soleimanzade H, Alipour S, Barzegari A, et al. Investigation the effect of oleoylethanolamide supplementation on the abundance of Akkermansia muciniphila bacterium and the dietary intakes in people with obesity: a randomized clinical trial. Appetite. 2019;141:104301.
Laffin MR, Tayebi Khosroshahi H, Park H, Laffin LJ, Madsen K, Kafil HS, et al. Amylose resistant starch (HAM-RS2) supplementation increases the proportion of Faecalibacterium bacteria in end-stage renal disease patients: microbial analysis from a randomized placebo-controlled trial. Hemodial Int. 2019;23(3):343–347.
Ahmadi S, Nagpal R, Wang S, Gagliano J, Kitzman DW, Soleimanian-Zad S, et al. Prebiotics from acorn and sago prevent high-fat-diet-induced insulin resistance via microbiome–gut–brain axis modulation. J Nutr Biochem. 2019;67:1–13.
Hosseinifard E-S, Morshedi M, Bavafa-Valenlia K, Saghafi-Asl M. The novel insight into anti-inflammatory and anxiolytic effects of psychobiotics in diabetic rats: possible link between gut microbiota and brain regions. Eur J Nutr. 2019;58(01):1–15.
Ghazifard A, Kasra-Kermanshahi R, Far ZE. Identification of thermophilic and mesophilic bacteria and fungi in Esfahan (Iran) municipal solid waste compost. Waste Manag Res. 2001;19(3):257–61.
Seidavi A, Mirhosseini SZ, Shivazad M, Chamani M, Sadeghi AA. The development and evaluation of a duplex polymerase chain reaction detection of Bifidobacterium spp. and Lactobacillus spp. in duodenum, jejunum, ileum and cecum of broilers. J Rapid Meth Aut Microbiol. 2008;16(1):100–12.
Ghiyasi M, Rezaei M, Sayyahzadeh H, Firouzbakhsh F, Attar A. Effects of prebiotic (Fermacto) in low protein diet on some blood parameters and intestinal microbiota of broiler chicks. Ital J Anim Sci. 2008;7(3):313–20.
Jajarm H, Jahanbin A, Mokhber N, Gooyandeh S, Mansourian A, Beitollahi J. Effects of persica mouthwash on oral microbiota of cleft lip and palate patients during fixed orthodontic treatment. J Appl Sci. 2009;9:1593–6.
Henderson G, Cox F, Ganesh S, Jonker A, Young W, Collaborators GRC, et al. Rumen microbial community composition varies with diet and host, but a core microbiome is found across a wide geographical range. Sci Rep. 2015;5:14567.
Llewellyn MS, Boutin S, Hoseinifar SH, Derome N. Teleost microbiomes: the state of the art in their characterization, manipulation and importance in aquaculture and fisheries. Front Microbiol. 2014;5:207.
Davari S, Talaei SA, Alaei H. Probiotics treatment improves diabetes-induced impairment of synaptic activity and cognitive function: behavioral and electrophysiological proofs for microbiome–gut–brain axis. Neuroscience. 2013;240:287–96.
Bishehsari F, Mahdavinia M, Vacca M, Malekzadeh R, Mariani-Costantini R. Epidemiological transition of colorectal cancer in developing countries: environmental factors, molecular pathways, and opportunities for prevention. World J Gastroenterol: WJG. 2014;20(20):6055–72.
Vaghef-Mehrabany E, Alipour B, Homayouni-Rad A, Sharif S-K, Asghari-Jafarabadi M, Zavvari S. Probiotic supplementation improves inflammatory status in patients with rheumatoid arthritis. Nutrition. 2014;30(4):430–5.
Rastmanesh R. High polyphenol, low probiotic diet for weight loss because of intestinal microbiota interaction. Chem Biol Interact. 2011;189(1–2):1–8.
Homayouni A, Bastani P, Ziyadi S. Mohammad-Alizadeh-Charandabi S, Ghalibaf M, Mortazavian AM et al. effects of probiotics on the recurrence of bacterial vaginosis: a review. J Low Genit Tract Dis. 2014;18(1):79–86.
Huang X, Fan X, Ying J, Chen S. Emerging trends and research foci in gastrointestinal microbiome. J Transl Med. 2019;17(1):67.
Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C, et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature. 2010;464(7285):59.
Yao H, Wan J-Y, Wang C-Z, Li L, Wang J, Li Y, et al. Bibliometric analysis of research on the role of intestinal microbiota in obesity. PeerJ. 2018;6:e5091.
Cho YA, Kim J. Effect of probiotics on blood lipid concentrations: a meta-analysis of randomized controlled trials. Medicine. 2015;94(43) e1714.
Sarao LK, Arora M. Probiotics, prebiotics, and microencapsulation: a review. Crit Rev Food Sci Nutr. 2017;57(2):344–71.
Mahmoudi R, Fakhri O, Farhoodi A, Kaboudari A, Rahimi SF. A review on probiotic dairy products as functional foods reported from Iran. Int J Food Nutr Saf. 2015;6(1):2.
Ejtahed H-S, Angoorani P, Soroush A-R, Atlasi R, Hasani-Ranjbar S, Mortazavian AM, et al. Probiotics supplementation for the obesity management; a systematic review of animal studies and clinical trials. J Funct Foods. 2019;52:228–42.
Roberfroid M, Gibson GR, Hoyles L, McCartney AL, Rastall R, Rowland I, et al. Prebiotic effects: metabolic and health benefits. Br J Nutr. 2010;104(S2):S1–S63.
Ejtahed H-S, Soroush A-R, Siadat S-D, Hoseini-Tavassol Z, Larijani B, Hasani-Ranjbar S. Targeting obesity management through gut microbiota modulation by herbal products: a systematic review. Complement Ther Med. 2018;42:184–204.
Anderson A, McNaught C, Jain P, MacFie J. Randomised clinical trial of synbiotic therapy in elective surgical patients. Gut. 2004;53(2):241–5.
Ejtahed H-S, Tabatabaei-Malazy O, Soroush A-R, Hasani-Ranjbar S, Siadat S-D, Raes J, et al. Worldwide trends in scientific publications on association of gut microbiota with obesity. Iran J Basic Med Sci. 2019;22(1):65–71.
Turnbaugh PJ, Hamady M, Yatsunenko T, Cantarel BL, Duncan A, Ley RE, et al. A core gut microbiome in obese and lean twins. Nature. 2009;457(7228):480.
Ley RE, Turnbaugh PJ, Klein S, Gordon JI. Microbial ecology: human gut microbes associated with obesity. Nature. 2006;444(7122):1022.
Ley RE, Bäckhed F, Turnbaugh P, Lozupone CA, Knight RD, Gordon JI. Obesity alters gut microbial ecology. Proc Natl Acad Sci. 2005;102(31):11070–5.
Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature. 2006;444(7122):1027.
Integrative H. The Integrative human microbiome project. Nature. 2019;569(7758):641.
Kulkarni AV, Aziz B, Shams I, Busse JW. Comparisons of citations in web of science, Scopus, and Google scholar for articles published in general medical journals. Jama. 2009;302(10):1092–6.
Mongeon P, Paul-Hus A. The journal coverage of web of science and Scopus: a comparative analysis. Scientometrics. 2016;106(1):213–28.
Acknowledgements
Not applicable.
Funding
This study was funded by the Endocrinology and Metabolism Research Institute, Tehran University of Medical Sciences, Tehran, Iran.
Author information
Authors and Affiliations
Contributions
H.A., H.D., H.E., F.R. and P.P. conceived of the presented idea. S.D.S., B.L. and P.P. developed the theory and H.A. performed the computations. N.F., A.S., S.A.B, and S.H.R. verified the analytical methods. H.E. investigated the molecular technique aspect of retrieved articles, H.E., H.A., and H.D. contributed to the interpretation of the results and S.D.S supervised the findings of this work. H.D. and H.A. wrote the manuscript in consultation with H.E. and N.F. All authors discussed the results, provided critical feedback, helped shape the research and analysis, and contributed to the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Consent for publication
Not applicable.
Ethics approval and consent to participate
Not applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 1319 kb)
Rights and permissions
About this article
Cite this article
Aazami, H., DehghanBanadaki, H., Ejtahed, HS. et al. The landscape of microbiota research in Iran; a bibliometric and network analysis. J Diabetes Metab Disord 19, 163–177 (2020). https://doi.org/10.1007/s40200-020-00488-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40200-020-00488-2