Abstract
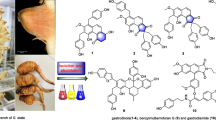
Four new 14(13 → 12)-abeolanostane triterpenoids featuring extended π-conjugated systems, kadcoccitanes E–H (1–4), were obtained from the stems of Kadsura coccinea through using a HPLC − UV-guided approach. Their structural and configurational determination was accomplished through extensive spectroscopic analysis coupled with quantum chemical calculations. Kadcoccitanes E–H were tested for their cytotoxic activities against five human tumor cell lines (HL-60, A-549, SMMC-7721, MDA-MB-231, SW-480) but none of them exhibited activities at the concentration 40 μM.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The plant family Schisandraceae is composed of two genera, Schisandra and Kadsura. Plant species within this family have attracted much attention for serving as an eminent source of triterpenoids with diverse structures and various bioactivities [1, 2]. Triterpenoids from the Schisandraceae family can be categorized into three major groups on the basis of their different carbon frameworks: lanostanes, cycloartanes, and schinortriterpenoids [1, 2]. Their fascinating architectures have aroused great interest among organic synthetic chemists and a few molecules have been successfully totally synthesized [3].
Kadsura coccinea is a vine plant grown widely in southern China. It possesses reputed medicinal value and has been utilized as effective remedies for gastropathy, rheumatic arthritis, etc. In the last decades, considerable efforts in phytochemical research on K. coccinea have led to the characterization of many lanostane triterpenoids and more than 10 kinds of novel scaffolds [4,5,6,7,8,9], including the recently reported neokadcoccitane A [10] possessing an aromatic ring D and an extended π-conjugated systems (EPCS). The EPCS moieties are not commonly present in lanostane triterpenoids from the Schisandraceae family, but can be found in a few cycoartane triterpenoids. For example, kadlongilactones A and B were both found to possess an α,β,γ,δ-unsaturated lactone moiety and characterized as potent inhibitors of K562 cells [11]. To explore more structurally and biologically fascinating triterpenoids from K. coccinea, further phytochemical investigations were carried out on stems collected in Jingzhou County, Hunan Province. During this research, several unusual chromatographic peaks in HPLC with maximum ultraviolet absorption (UV) wavelengths at 300 ~ 380 nm were observed, suggesting the existence of triterpenoids with EPCS. Subsequent UV-guided isolation afforded kadcoccitanes E–H (1–4), four undescribed 14(13 → 12)-abeolanostane triterpenoids featuring α,β,γ,δ-unsaturated aldehyde or α,β,γ,δ,ε,ζ-unsaturated aldehyde/ketone. Herein, the isolation and structure elucidation of kadcoccitanes E–H (1–4), along with their cytotoxicity evaluation were described.
2 Results and discussion
Kadcoccitane E (1), a light yellow amorphous solid, was assigned the molecular formula of C30H42O4 through its HRESIMS ([M − H]– m/z 465.3014, calcd 465.3010), indicating ten degrees of unsaturation (DOUs). The 1H NMR spectrum (Table 1) exhibited resonances for two olefinic protons (δH 5.90, m; δH 6.86, d, J = 2.1 Hz), an aldehyde group (δH 10.33, s), a doublet methyl group (δH 1.10, d, J = 6.8 Hz), and five singlet methyl groups (δH 0.99, 1.05, 1.13, 1.18, and 2.07). The 13C NMR and DEPT spectra (Table 2) exhibited 30 carbon signals, including six methyls, eight methylenes, seven methines (two olefinic and one aldehyde), and nine nonprotonated carbons (one carboxyl, one carbonyl, four olefinic, and three quaternary carbons). Apart from the six DOUs occupied by carboxyl, carbonyl, aldehyde and olefinic groups, the remaining four ones uncovered that 1 possessed a tetracyclic architecture.
Detailed analysis of the NMR spectra of 1 indicated that it was a 14(13 → 12)-abeolanostane triterpenoid featuring 6/6/5/6-fused ring system. The HMBC correlations (Fig. 2) from H3-19 (δH 1.13) to C-1 (δC 35.7), C-5 (δC 52.5), C-9 (δC 169.6), and C-10 (δC 38.4), from H3-29 (δH 1.05) to C-3 (δC 214.8), C-4 (δC 47.9), C-5, and C-30 (δC 26.3), and from H3-30 (δH 1.18) to C-3, C-4, C-5, and C-29 (δC 21.9), in combination with the 1H–1H COSY correlations (Fig. 2) of H-1β (δH 2.06)/H-2β (δH 2.71) and H-6β (δH 1.50)/H-7α (δH 1.34)/H-8 (δH 2.55) revealed the existence of typical rings A and B in kadcoccitane E (1). In addition, the HMBC correlations from H-11 (δH 6.86) to C-8 (δC 52.0), C-9, C-10, C-12 (δC 173.8), and C-14 (δC 45.9), from H3-28 (δH 0.99) to C-8, C-12, C-14, and C-15 (δC 31.6), and from H-18 (δH 10.33) to C-13 (δC 128.7) and C-17 (δC 36.3), in combination with the 1H–1H COSY correlations of H-16α (δH 1.81)/H-17 (δH 2.88) demonstrated that a six-membered ring with C-12/C-13 double bond and an aldehyde group at C-13 fused with a five-membered ring, which revealed the existence of rings C and D in kadcoccitane E. Furthermore, the HMBC correlations from H3-21 (δH 1.10) to C-17 and C-22 (δC 32.9), from H-24 (δH 5.90) to C-22, C-23 (δC 28.9), C-26 (δC 170.6), and C-27 (δC 21.6), and from H3-27 (δH 2.07) to C-24 (δC 142.0), C-25 (δC 129.1), and C-26, in combination with the 1H–1H COSY correlations of H-17/H-20 (δH 2.42)/H-22b (δH 1.30)/H-23a (δH 2.84)/H-24 revealed the existence of typical side chain (C-20–C-27) in kadcoccitane E (1). Thus, the planar structure of 1 could be established as shown in Fig. 1.
With respect to the stereochemistry of 1, the C-24/C-25 double bond was assigned Z geometry due to the H-24/H3-27 correlation in the ROESY spectrum (Fig. 2). Besides, correlations of H3-30/H-5α (δH 1.49), H-2β/H3-29, H3-19/H-2β, H-8/H3-19, H-15β (δH 1.69)/H-8, H3-28/H-15α (δH 1.78), H-17/H3-28 indicated that both H-8 and H3-19 adopted β-orientations, while both H-17 and H3-28 adopted α-orientations. According to these deductions, as well as biosynthetic considerations, the absolute configuration of 1 was determined as 5R, 8S, 10S, 14S, 17R, and 20R.
Kadcoccitane F (2) was obtained as a light yellow amorphous solid and its molecular formula was determined as C30H40O4 by its HRESIMS data ([M − H]– m/z 463.2852, calcd 463.2854). The 1H NMR spectrum (Table 1) displayed signals for three olefinic protons (δH 5.72, dt, J = 6.5, 2.1 Hz; δH 5.84, brs; δH 7.08, s), an aldehyde group (δH 10.37, s), a doublet methyl group (δH 1.10, d, J = 6.8 Hz), and five singlet methyl groups (δH 1.07, 1.16, 1.17, 1.22, and 2.05). The 13C NMR and DEPT spectra (Table 2) showed 30 carbon resonances, including six methyls, seven methylenes, seven methines (three olefinic and one aldehyde), and ten nonprotonated carbons (one carboxyl, one carbonyl, five olefinic, and three quaternary carbons). Apart from the seven DOUs generated by carboxyl, carbonyl, aldehyde and olefinic groups, the remaining four ones manifested the tetracyclic system of 2. Analysis of the NMR spectra of 2 indicated that its structure was closely similar to that of 1, except that 2 possessed one additional double assumed to be located in ring B. This deduction could be supported by HMBC correlations (Fig. 3) from H-7 (δH 5.72) to C-5 (δC 51.2), C-6 (δC 23.9), C-9 (δC 163.6), and C-14 (δC 44.2), as well as 1H–1H COSY correlations (Fig. 3) of H-5 (δH 1.82)/H-6β (δH 2.15)/H-7. Upon carefully analyzing the ROESY spectrum of 2, in combination with comparisons between NMR data of 2 and 1, and biosynthetic considerations, the absolute configuration of 2 was determined as 5R, 10S, 14S, 17R, and 20R.
Kadcoccitane G (3) was isolated as a light yellow amorphous solid and its molecular formula was determined as C30H40O4, with eleven DOUs based on its HRESIMS data ([M – H]– m/z 463.2854, calcd 463.2854). The 1H NMR spectrum (Table 1) showed signals for four olefinic protons (δH 4.96, d, J = 2.4 Hz; δH 5.09, d, J = 2.4 Hz; δH 5.93, brs; δH 6.31, s), a doublet methyl group (δH 0.95, d, J = 6.7 Hz), and five singlet methyl groups (δH 1.09, 1.12, 1.26, 1.36, and 2.12). Totally 30 carbon signals could be observed in the 13C NMR and DEPT spectra (Table 2), including six methyls, eight methylenes (one olefinic), five methines (two olefinic), and eleven nonprotonated carbons (one carboxyl, two carbonyl, five olefinic, and three quaternary carbons). Apart from the seven DOUs generated by carboxyl, carbonyl and olefinic groups, the remaining four ones disclosed the existence of four rings in its structure.
Further analysis of the 1D and 2D NMR spectra of 3 indicated that it possessed the same scaffold as kadcoccitanes E and F (1 and 2). The HMBC correlations (Fig. 4) from H-11 (δH 6.31) to C-8 (δC 144.6), C-9 (δC 169.7) and C-14 (δC 54.0), from H3-28 (δH 1.36) to C-8, C-12 (δC 168.5), C-14, and C-15 (δC 31.2), and from H2-18 (δH 4.96, 5.09) to C-12, C-13 (δC 146.4) and C-17 (δC 50.7), in combination with the 1H–1H COSY correlations (Fig. 4) of H-5 (δH 2.39)/H-6β (δH 2.68), H-15α (δH 2.48)/H-16α (δH 1.83)/H-17 (δH 2.21) uncovered the existence of an α,β,γ,δ,ε,ζ-unsaturated ketone system distributed in rings B–D. The ROESY correlations (Fig. 4) suggested that 3 possessed identical relative configuration as that of 2. Finally, the absolute configuration of 3 was determined as 5R, 10S, 14R, 17R, and 20R through biosynthetic considerations.
Kadcoccitane H (4) was obtained as a yellow amorphous solid and had the molecular formula of C30H40O4 as determined by its HRESIMS data ([M + H]+ m/z 465.2999, calcd 465.2999). The 1H NMR spectrum (Table 1) showed signals for two olefinic protons (δH 5.98, t, J = 7.7 Hz; δH 6.30, s), a doublet methyl group (δH 1.06, d, J = 6.9 Hz), and six singlet methyl groups (δH 1.10, 1.12, 1.24, 1.38, 2.02 and 2.12). The 13C NMR and DEPT spectra (Table 2) exhibited 30 carbon signals, including seven methyls, seven methylenes, four methines (two olefinic), and twelve nonprotonated carbons (one carboxyl, two carbonyl, six olefinic, and three quaternary carbons). Apart from the seven DOUs occupied by carboxyl, carbonyl and olefinic groups, the remaining four ones revealed that 4 had a tetracyclic structure.
An in-depth analysis of the 1D and 2D NMR spectra of 4 indicated that its structure resembled closely to that of 3, and only one variation could be observed. The exocyclic C-13/C-18 double bond in 3 became C-13/C-17 double bond in 4, which could be revealed by HMBC correlations (Fig. 5) from H3-18 (δH 2.02) to C-12 (δC 169.1), C-13 (δC 123.2) and C-17 (δC 145.5).
The relative configuration of rings A–D in 4 was elucidated to be the same as that in 3 through analysis of the ROESY spectrum (Fig. 5) and NMR data comparison. However, it is difficult to determine the configuration of C-20 which was located on the conformationally flexible side chain as well as on an allyl position. Thus, (5R*, 10S*, 14R*, 20R*)-4 (4a) and (5R*, 10S*, 14R*, 20S*)-4 (4b) were subjected to GIAO NMR calculations at mPW1PW91-SCRF/6–31 + G(d,p)//B3LYP-D3/6-31G(d) level of theory. Then, considering the flexibility of compound 4, DP4 + analysis based on random conformational amplitudes [12] was employed to distinguish 4a and 4b. As a result, 4a and 4b got a probability of 67.1% and 32.9%, respectively (Fig. S53). In addition, as the calculated chemical shifts of the α,β-unsaturated carboxyl moiety usually endure relatively large errors, which is also the case in the present study, an additional DP4 + analysis was undertaken using partial data in which 1H and 13C chemical shifts/shielding tensors in C-24–C-27 moiety were excluded from experimental/calculated data. As a result, 4a and 4b got a probability of 66.1% and 33.9%, respectively (Fig. S54). Thus, the relative configuration of 4 was determined as 5R*,10S*,14R*, and 20R*. Subsequent TDDFT ECD calculation on (5R,10S,14R,20R)-4 (4aA) afforded a theoretical curve which matched with the experimental spectrum very well (Fig. 6a). Then, the dominant conformer 4a-1 was subjected to molecular orbital (MO) analysis, and the π → π* transition from HOMO to LUMO MOs contributed significantly to the Cotton effect around 350 nm (Fig. 6b and c). Finally, the absolute configuration of 4 was determined as 5R, 10S, 14R, and 20R.
a Experimental ECD spectrum of 4 (black). Calculated ECD spectra (shift = + 8 nm) of (5R,10S,14R,20R)-4 (4aA) (red) and ent-4aA (blue). b Calculated ECD spectrum (curve) of conformer 4a-1 with rotatory strength (bar), and the key transitions in three important excited states (table). c Key MOs of conformer 4a-1
The potential cytotoxic activities of kadcoccitanes E–H (1–4) against five human tumor cell lines, HL-60 (leukemia), A-549 (lung cancer), SMMC-7721 (liver cancer), MDA-MB-231 (breast cancer), and SW-480 (colon cancer) were tested, but no remarkable activities were observed (Table 3).
3 Experimental
3.1 General experimental procedures
General experimental procedures can be found in the Supplementary Material (part 1).
3.2 Plant material
The source of the plant material is the same as described in ref 9 or can be found in the Supplementary Material (part 2), but the investigated parts in the present research are stems, instead of roots in ref 9.
3.3 Extraction and isolation
The detailed procedures for extraction and isolation can be found in in the Supplementary Material (part 3).
3.4 Characteristic data of compounds 1–4
Kadcoccitane E (1): light yellow amorphous powder; [α] 23D − 73.57 (c 0.107, MeOH); UV(MeOH) λmax (log ε): 195 (4.16), 217 (4.02), 265 (3.57), 304 (4.13) nm; ECD (MeOH) λmax (Δε): 195 (+ 8.00), 223 (− 15.29), 303 (+ 12.01), 343 (− 8.88) nm; IR (KBr) νmax 3407, 2951, 1707, 1658, 1605, 1205, 1132, 869, 577 cm−1; negative HRESIMS m/z 465.3014 [M − H]– (calcd for C30H41O4, 465.3010).
Kadcoccitane F (2): light yellow amorphous powder; [α] 25D − 51.09 (c 0.092, MeOH); UV(MeOH) λmax (log ε): 195 (4.17), 272 (3.53), 337 (3.94) nm; ECD (MeOH) λmax (Δε): 202 (+ 4.30), 229 (− 4.93), 256 (+ 2.36), 322 (+ 5.08), 368 (− 4.93) nm; IR (KBr) νmax 3427, 2962, 2933, 2872, 1706, 1652, 1381, 1204, 581 cm−1; negative HRESIMS m/z 463.2852 [M − H]– (calcd for C30H39O4, 463.2854).
Kadcoccitane G (3): light yellow amorphous powder; [α] 19D + 103.68 (c 0.086, MeOH); UV(MeOH) λmax (log ε): 195 (4.25), 270 (3.20), 318 (3.90) nm; ECD (MeOH) λmax (Δε): 195 (+ 5.48), 217 (− 2.60), 337 (+ 5.16) nm; IR (KBr) νmax 3430, 2964, 2935, 2872, 1707, 1645, 1457, 1392, 1245, 596 cm−1; negative HRESIMS m/z 463.2854 [M − H]– (calcd for C30H39O4, 463.2854).
Kadcoccitane H (4): yellow amorphous powder; [α] 20D –275.76 (c 0.198, MeOH); UV(MeOH) λmax (log ε): 199 (4.04), 224 (4.19), 285 (2.95), 368 (4.01) nm; ECD (MeOH) λmax (Δε): 195 (–15.23), 247 (+ 24.69), 358 (–6.89) nm; IR (KBr) νmax 3428, 2963, 2931, 1708, 1638, 1394, 1191, 1100, 601, 575 cm−1; positive HRESIMS m/z 465.2999 [M + H]+ (calcd for C30H41O4, 465.2999).
For 1H NMR data of kadcoccitanes E–H (1–4), see Table 1. For their 13C NMR data, see Table 2.
3.5 The cytotoxicity assay
The cytotoxicity assay has been described previously [13].
Data availability
The data that support the findings of this study are openly available in the Science Data Bank at https://doi.org/10.57760/sciencedb.j00144.00004 (Under review).
References
Xiao WL, Li RT, Huang SX, Pu JX, Sun HD. Triterpenoids from the Schisandraceae family. Nat Prod Rep. 2008;25:871–91.
Shi YM, Xiao WL, Pu JX, Sun HD. Triterpenoids from the Schisandraceae family: an update. Nat Prod Rep. 2015;32:367–410.
Yang P, Li J, Sun L, Yao M, Zhang X, Xiao WL, et al. Elucidation of the structure of pseudorubriflordilactone B by chemical synthesis. J Am Chem Soc. 2020;142:13701–8.
Liang CQ, Shi YM, Luo RH, Li XY, Gao ZH, Li XN, et al. Kadcoccitones A and B, two new 6/6/5/5-fused tetracyclic triterpenoids from Kadsura coccinea. Org Lett. 2012;14:6362–5.
Liang CQ, Shi YM, Li XY, Luo RH, Li Y, Zheng YT, et al. Kadcotriones A-C: tricyclic triterpenoids from Kadsura coccinea. J Nat Prod. 2013;76:2350–4.
Hu ZX, Shi YM, Wang WG, Li XN, Du X, Liu M, et al. Kadcoccinones A-F, new biogenetically related lanostane-type triterpenoids with diverse skeletons from Kadsura coccinea. Org Lett. 2015;17:4616–9.
Liang CQ, Shi YM, Wang WG, Hu ZX, Li Y, Zheng YT, et al. Kadcoccinic acids A-J, triterpene acids from Kadsura coccinea. J Nat Prod. 2015;78:2067–73.
Yang YP, Jian YQ, Liu YB, Ismail M, Xie QL, Yu HH, et al. Triterpenoids from Kadsura coccinea with their anti-inflammatory and inhibited proliferation of rheumatoid arthritis-fibroblastoid synovial cells activities. Front Chem. 2021;9: 808870.
Xu HC, Hu K, Sun HD, Puno PT. Four 14(13→12)-abeolanostane triterpenoids with 6/6/5/6-fused ring system from the roots of Kadsura coccinea. Nat Prod Bioprospect. 2019;9:165–73.
Xu HC, Hu K, Shi XH, Tang JW, Li XN, Sun HD, et al. Synergistic use of NMR computation and quantitative interproton distance analysis in the structural determination of neokadcoccitane A, a rearranged triterpenoid featuring an aromatic ring D from Kadsura coccinea. Org Chem Front. 2019;6:1619–26.
Pu JX, Xiao WL, Lu Y, Li RT, Li HM, Zhang L, et al. Kadlongilactones A and B, two novel triterpene dilactones from Kadsura longipedunculata possessing a unique skeleton. Org Lett. 2005;7:5079–82.
Zanardi MM, Marcarino MO, Sarotti AM. Redefining the impact of Boltzmann analysis in the stereochemical assignment of polar and flexible molecules by NMR calculations. Org Lett. 2020;22:52–6.
Xia JN, Hu K, Su XZ, Tang JW, Li XN, Sun HD, et al. Discovery of ent-kaurane diterpenoids, characteristic metabolites of Isodon species, from an endophytic fungal strain Geopyxis sp. XY93 inhabiting Isodon parvifolia. Fitoterapia. 2022;158:105160.
Acknowledgements
We thank Service Center for Bioactivity Screening, State Key Laboratory of Phytochemistry and Plant Resources in West China, Kunming Institute of Botany for bioactivity screening. This project was supported financially by the Natural Science Foundation of Yunnan Province (No. 202101AT070188).
Funding
Natural Science Foundation of Yunnan Province (202101AT070188).
Author information
Authors and Affiliations
Contributions
PTP conceived and designed the research; QQZ, KH carried out the experiment and wrote the manuscript; PTP, HDS and KH supervised the whole study and critically reviewed the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
No conflict of interest is declared.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
13659_2023_376_MOESM1_ESM.pdf
Supplementary file1 It includes general experimental procedures; plant material; extraction and isolation; 1D NMR, 2D NMR, HRESIMS, UV, ECD, IR spectra, and OR of compounds 1–4, as well as computational methods and data for compound 4.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, QQ., Hu, K., Sun, HD. et al. Four new lanostane triterpenoids featuring extended π-conjugated systems from the stems of Kadsura coccinea. Nat. Prod. Bioprospect. 13, 12 (2023). https://doi.org/10.1007/s13659-023-00376-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13659-023-00376-1