Abstract
Apis mellifera unicolor is a tropical honeybee endemic of Madagascar. Comprehensive knowledge about its mitochondrial and nuclear genetic diversity and structuration was our main purpose. Samples of worker bees were collected from 867 colonies in 76 sites in Madagascar and 1 reference population in South Africa. PCR-restriction fragment length polymorphism (RFLP) and sequencing were used to reveal variability in the COI–COII mtDNA region. Seventeen microsatellite loci were used for studying the nuclear diversity. Three PCR-RFLP profiles were observed, among which 99.4 % belonged to A1 haplotype, 0.2 % to a new A haplotype, and 0.4 % to A4 haplotype. In microsatellite analysis, moderate genetic diversity values were found for Madagascar, together with low mean number of alleles ranging from 2.47 to 3.88 compared to South Africa. Bayesian clustering assignment methods and principal component analysis (PCA) separated populations into two genetic clusters which matched with geographic areas. Several hypotheses are discussed regarding to the low genetic diversity of A. m. unicolor in its native range.
Similar content being viewed by others
1 Introduction
Apis mellifera subspecies are the most economically valuable pollinators of crop monocultures worldwide (Klein et al. 2007). In addition, honeybee contribution to floral biodiversity and conservation through pollination is estimated to affect 80 % of wild flora (Batra 1995; De la Rua et al. 2009).
Remarkable morpho-geographical differentiations are found throughout honeybee distribution areas; 28 subspecies are endemic to Africa, Europe, and Middle East (Ruttner et al. 1978; Ruttner 1988; Sheppard et al. 1997; Sheppard and Meixner 2003). Four evolutionary lineages have been described based on phenotypes and molecular traits (Garnery et al. 1992, 1993; Estoup et al. 1995; Franck et al. 2000, 2001; Alburaki et al. 2013).
In Africa, 11 subspecies are found from all lineages except from the C lineage (Hepburn and Radloff 1998; Meixner et al. 2011). The African A lineage is native to Africa and subspecies include Apis mellifera scutellata (Lepeletier 1836), Apis mellifera capensis (Eschscholtz 1822), and Apis mellifera unicolor (Latreille 1804).
A. m. unicolor is the endemic subspecies of Madagascar, a drifted tropical island, located 400 km off the East coast of Africa. Madagascar is among the five richest biodiversity hotspots of the world in terms of endemic plants and vertebrate species (Myers et al. 2000). However, less than 10 % of its original habitat still remains (Myers et al. 2000). In such an area of endemism, with over 80 % of endemic phanerogam (Ganzhorn et al. 2001), the role of pollinating insects is vital for the reproduction of these plants. Although the pollinating role of the honeybee has never been investigated in detail for these floras, A. m. unicolor is thought to play a crucial role in the pollination of endemic phanerogam species (Ruttner 1975; Ralalaharisoa-Ramamonjisoa et al. 1996).
After molecular analysis, mitotype A1 was attributed to the honeybee of Madagascar, with a surprising absence of mitochondrial polymorphism within this endemic subspecies (Franck et al. 2001). The latest molecular study using SNP analyses suggested that A. m. unicolor had a specific SNP (Whitfield et al. 2006). However, those assumptions were based on a low number of samples (<50) taken from regions of the vast territory of Madagascar. In terms of morphological and behavioral criteria, two ecotypes were described, the first from the highlands (Hauts Plateaux) and the second from the coastal area (Ruttner 1988).
Since the recent arrival of the parasitic mite Varroa destructor documented in Madagascar in 2010, severe colony losses have been observed (Rasolofoarivao et al. 2013). This situation may dramatically impact the biodiversity of Madagascar. Therefore, considering the lack of genetic data and the threat to this subspecies, it appears necessary to improve our knowledge on A. m. unicolor genetic diversity in its native area.
2 Material and methods
2.1 Area of study
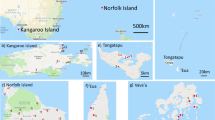
The land area and relief of Madagascar gives rise to various climatic zones. The year is characterized by two distinct seasons: the austral winter (April to November) and the austral summer (December to March). The coastal zones enjoy a warm climate, and mean annual temperatures range between 22 and 25 °C. The upland areas of the island have a more temperate climate with a mean annual temperature of 20 °C. Tropical forests mainly consist of deciduous woodland in Western Madagascar and xerophytic thorn forests in the southern region (Figure 1).
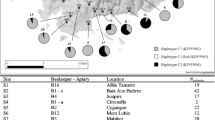
Madagascar map with ecological zonation and honeybee sampling sites (emplacements of each site are presented in red polygons). The 76 sites are spread in 6 geographic regions. Each number represents a site with correspondence in Table I.
2.2 Sampling
One adult worker honeybee per colony was sampled between August 2011 and March 2013 from 76 sites (n = 867) in Madagascar (Figure 1, SD Table I). Sampling was performed on managed colonies and 33 wild colonies (collected in Djamajar (S8) n = 6, Antsoha (S7) n = 9, Rantolava (S24) n = 3, and Tsararano (S21) n = 15). As a reference population from African lineage, samples were collected from 22 managed colonies in South Africa (one apiary from Cape region) in 2013. Honeybees were preserved in ethanol (96 %) and kept at −20 °C until molecular analysis.
2.3 DNA extraction
The six worker legs were used for DNA isolation. DNA was extracted from individual honeybees as previously described (Delatte et al. 2010). All individuals were subjected to both microsatellite and mitochondrial analyses.
2.4 Microsatellite amplification and genotyping
Microsatellite population studies were carried out using 17 loci published in Solignac et al. (2003) and combined into four different mixes (mix 1: A024, A113, Ac306, Ap055, Ap081; mix 2: A (B)124, A028, A029, A088, Ap273, Ap289; mix 3: Ap033, A035, Ap036; mix 4: A014, Ap043, Ap066). PCR reactions were performed in a 10 μL final reaction volume with a primer mix (10 μM) using Type-it Multiplex PCR Master Mix (Qiagen) kits. PCR programs were run with an initial denaturation of 94 °C for 5 min, followed by 35 cycles of denaturation at 94 °C for 30 s; annealing was 55 °C for 30 s for mix 1 and 52 °C for mixes 2–4, followed by elongation at 72 °C for 30 to 45 s. A final extension was done at 72 °C for 10 to 20 min.
Samples were further genotyped using an automated DNA sequencer (Applied Biosystems 3130XL) with capillaries. Allele sizes were scored using GeneMapper 4.0 Software. Individuals with genotype data missing for more than 40 % of all loci were excluded from statistical analysis. Small sample sizes under 10 individuals per site were not included in the nuclear genetic analysis at population level.
2.5 Microsatellite analysis
Observed (H o), expected (H e), and Nei’s 1987 unbiased expected (H n.b) heterozygosity and fixation indices (F is) (Weir and Cockerham 1984) were estimated using Genepop 4.2 (Rousset 2008) and Genetix 4.05 (Belkhir et al. 1996). All pairs of loci were tested for linkage disequilibrium using Genepop 4.2 (Rousset 2008). Deviations from Hardy–Weinberg equilibrium (HWE) were tested using a two-tailed Fisher exact test based on Markov chain (Rousset 2008). Permutation tests conducted by FSTAT (Goudet 2001) determined whether genetic diversity (H e, H n.b, H o) and F is differed significantly between geographical regions.
FreeNA (Chapuis and Estoup 2007) was used to estimate null allele frequencies. Population differentiation was quantified by calculating pairwise F ST values (Weir and Cockerham 1984) and verifying their significance through the permutational test in Genetix 4.05 (Belkhir et al. 1996). Relationships between genetic and geographic distances at all sites were tested using the Mantel’s test in Genepop 4.2 (Rousset 2008). The significance of the correlation between matrices of geographical and genetic distances among pairs of sites was tested using 1000 permutations of the data. As potential isolation by distance (ibd) patterns may not hold over the entire range (1470 km), because at some point the influence of gene flow is expected to be weak relative to the influence of genetic drift and homoplasy (Hutchison and Templeton 1999), ibd patterns were investigated at smaller spatial scales, i.e., in sites from the northern regions (500 km, n sites = 7), the central regions (370 km, n sites = 15), and southern regions of the island (380 km, n sites = 9). We further investigated the importance of scale on spatial genetic structuring by performing a hierarchical F analysis, which estimates the genetic variation found at each hierarchical level. A nested tree-level analysis of molecular variance (AMOVA, Excoffier and Lischer 2010) was performed by partitioning the total sum of squares into components representing variation between geographical regions, among sites within regions and among individuals within sites using Arlequin V3.5 (Excoffier and Lischer 2010).
Levels of population admixture were quantified using a number of Bayesian clustering procedures as implemented in Structure V2.3.4 (Pritchard et al. 2000). The number of population clusters was inferred according to Evanno et al. (2005) and the ad hoc statistic ΔK was calculated for K ranging from 1 to 10 for the full dataset comprising the reference population and 1 to 20 within the Madagascar dataset (1 million simulations and 100,000 burn-in with 10 iterations for each K). This ad hoc statistic was processed through the Structure Harvester website (http://taylor0.biology.ucla.edu/structureHarvester/). Clumpp v.1.1.2 (Jakobsson and Rosenberg 2007) was used to align the best of the five repetitions of the K. Distruct v.1.1 (Rosenberg 2004) was used to graphically display the results. A principal component analysis (PCA) was performed on the genetic data to visualize genetic differentiation among the population groups using adegenet 1.4 (Jombart 2008) in R software (Team 2005). Adegenet 1.4 was also used to check the dataset structure using an alternative clustering analysis with a discriminant analysis of principal component (DAPC).
2.6 Mitochondrial analysis
Mitochondrial DNA COI–COII region was amplified using two specific primers, E2 and H2 (Garnery et al. 1992). Amplification and PCR cycles were as described in Garnery et al. (1992). The size of the fragment amplified was visualized using 5 μL of the PCR products electrophoresed on 2 % agarose gel. PCR products were then used both for enzymatic restriction and sequencing.
Ten microliters of PCR products from Madagascar (n = 867) and South Africa (n = 22) were enzymatically digested by the DraI (Promega©) enzyme according to manufacturer recommendations. The resulting fragments were separated in 4 % agarose gel. Each enzymatic profile was scored and compared to expected sizes published (Garnery et al. 1992; Franck et al. 2001). Each profile with a different band pattern observed on gel in Madagascar was sent for sequencing (121 samples of the dominant profile, 2 per other profile). PCR products (n = 173) were sent to Macrogen© for sequencing. Of these, some were taken from wild colonies (site Tsararano S21 n = 10). DNA sequence results were aligned using MEGA 5.04 (Kumar et al. 2008) then analyzed by BLAST search on GenBank. Genetic relationships between the different sequences obtained for the A1 haplotypes from Madagascar were investigated by constructing a minimum-spanning network of the haplotypes with TCS software (Clement et al. 2000).
3 Results
3.1 Diversity indices
The amplification was successful for 710 out of 867 Malagasy and for all South African (n = 22) individuals (with at least 10 loci amplified per sample). Of the 77 sampled sites, only 33 have more than 10 individuals (SD Table I). Analysis for linkage disequilibrium showed no significant deviation from equilibrium among the 17 microsatellite marker pairs (all P > 0.05), except between loci A113 and A24 (P < 0.05).
The average number of alleles per population and over 17 markers varied between 2.47 and 3.88 in Madagascar with an overall average of 7.76 alleles/locus, and for the single reference population of South Africa, the average number was 12.57 alleles/locus (Table I).
Unbiased estimated heterozygosity (H n.b) ranged from 0.36 to 0.50 for Madagascar, and for the reference population H n.b = 0.86 (Table I). Across all loci, less than 10 % null allele was observed (Table I). Inbreeding coefficient F is ranged from heterozygote excess (outbreeding, −0.07) to heterozygote deficiency (inbreeding, 0.15) compared with HWE expectations. No difference was detected among geographical regions within Madagascar (SD Table II, Comparisons among groups, all P > 0.4).
3.2 Genetic structure
F ST estimates among pairs of sites ranged from −0.02 to 0.17, with 318 out of 496 genetic differentiation estimates being significantly different from zero (SD Table III) within Madagascar dataset. F ST estimates between the overall Malagasy populations and the reference population of South Africa were highly significant and equal to 0.34.
Within each of the northern, central, and southern parts of Madagascar (<500 km between sites), nuclear genetic differentiation between sites was positively correlated with the logarithm of geographical distance (Figure 2 a, b, c). However, at the island scale (north to south 1470 km, n sites = 32), the correlation between the two matrices was no longer significant (Mantel test, P = 0.87; Figure 2d). The analysis of molecular variance comprising only Madagascar populations showed that the main contribution to the genetic variance was variation within sites (94 %). Differences among regions (2.3 %) and among sites within regions (3.34 %) were much lower but contributed significantly to the total genetic variation (SD Table IV). When all regions were taken into account (within Madagascar), all hierarchical levels accounted for a significant part of the genetic diversity.
Relationship between logarithm of geographical distance and nuclear genetic differentiation as estimated as F ST / (1 − F ST), between a sites from the northern region only (n sites = 7, P = 0.014), b the southern region only (n sites = 9, P = 0.0027), c sites from Hauts Plateaux only (n sites = 16, 120 combinations, Mantel test P = 0.014), and d sites from all regions sampled in Madagascar (n sites = 32, 496 combinations, Mantel test P = 0.84).
Variation in allelic composition was highlighted by Structure. The optimal K = 2 was the most strongly supported in likelihood using the reference population (Figure 3), with South African individuals being far apart from the Malagasy individuals. Then, looking at the substructure intra-Madagascar populations, the optimal and strongest K population is 2 (SD Fig. 1, SD Fig. 2). The DAPC analysis corroborates the same clustering analysis using both datasets. For the full dataset (South Africa and Madagascar), three genetic clusters were found (one for South African individuals, and two closer ones within Madagascar; Figure 4).
Population structure and Euclidean distances among genetic clusters based on 17 microsatellites loci. Top: Structure bar plots (K = 2, 3, and 4) with 22 reference samples from South Africa and 710 individuals of Madagascar organized by sampling sites (S1–76). Each horizontal bar represents one individual, and sites are delimited by black lines. The height of each bar represents the probability of assignment to a genetic cluster (one color). Bottom: PCAs among individual genotypes assigned to the different clusters (K = 2, 3, and 4). Inertia percentage of each axis is indicated (using 232 variables).
DAPC based on individual genotypes of Madagascar (n = 710) and reference population of South Africa (n = 22). Dots of different colors indicate honeybee samples from different genetic clusters (K = 3). Inertia percentage of each axis, PCs eigenvalues, and discriminant factors retained are indicated.
Under the K = 2 population clustering assumption within Madagascar, a geographic structuring appears with populations from the north, northwest, and south together in cluster 1 as opposed to populations from the west and east (cluster 2). The highland populations were mostly a mix of two genetic clusters (Figures 1 and 4, SD Fig. 2).
3.3 PCR-restriction fragment length polymorphism patterns and sequences
After analysis of the Madagascar samples, three different PCR-restriction fragment length polymorphism (RFLP) profiles were detected. Two restriction profiles were congruent with ones already published. The dominant PCR-RFLP profile observed was the African A1 haplotype (99.4 %), which exhibited three lengths of fragments (∼47, 108, and 483 bp) (Franck et al. 2001). Another PCR-RFLP profile referred to as the A4 haplotype (∼47, 108, 192, and 483 bp) was found in three individuals in our sample, two from S66 and one from S23. Another notable profile was found in two individuals from S22 and S29 (∼47, 108, 150, 350 bp). The total sizes of each haplotype ranged from 638 bp for A1, 830 bp for A4, and 655 bp for the new haplotype. Within the South African samples, two different PCR-RFLP profiles were detected: A1 haplotype (n = 1) and A4 haplotype (n = 21).
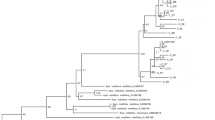
A total of 173 samples collected in 49 sites were sequenced. A total of 18 different sequences were found with variable sequence sizes belonging to the A lineage (SD Table V). The 16 sequences obtained for the A1 restriction profile have never been reported and were named A1_Mad1 to A1_Mad16 (accession numbers KF976992 to KF977009). These 16 sequences were characterized by one unit P 0 and one unit Q. The most frequent sequence, A1_Mad3, was present in all sites (n = 121, 70 %). The sequences of the two individuals exhibiting the new PCR-RFLP haplotype (in S29, S22) cluster within the A. m. unicolor group (it was subsequently named A1_Mad13) and we propose to classify it as a subtype of A1. The two other sequences (A4_Mad1 and A4_Mad2) were genetically close to the A4 haplotypes of A. m. scutellata (accession number FJ 477987 (Franck et al. 2001), similarity = 98 %) and found in two different regions of the West coast of Madagascar (R4 and R16) (SD Table V, SD Fig. 2). The A4 haplotypes were characterized by one unit P 0 and two units Q sequences.
4 Discussion
Previous PCR-RFLP analyses performed on 48 individuals from Madagascar by several authors (Garnery et al. 1992; Franck et al. 2001) detected a single restriction profile (A1). The larger sampling scale of this study in Madagascar detected a new A1 restriction profile and the occurrence of an A4 restriction profile. The sequencing approach used in our study revealed mtDNA variability with 16 new sequences. Haplotype frequencies and network analyses suggested that divergences are quite recent (SD Fig. 3), with all A1 sequences in Madagascar diverging from the predominant one through one single mutation (except for A1Mad_12). The A1 haplotype is widespread. It was found in this study in South Africa, and it has also been reported in Morocco (De la Rua et al. 2006), Algeria (Chahbar et al. 2013), Sudan (El-Niweiri and Moritz 2008), and in the Middle East (Alburaki et al. 2011). Three samples from A4 were found in the Western region of Madagascar. Two of these samples show high genetic diversity compared to A4 published sequences. A4 haplotypes might probably result from ancestral introductions to the island together with ancestral A1, as both haplotypes (belonging to AI sublineage) are commonly distributed within the different African subspecies populations (Franck et al. 2001).
The absence of haplotypes belonging to other lineages in our study implies that introductions of foreign queens are rare. This can be explained by the fact that other subspecies i) if imported/introduced were too few to be seen in our sampling, ii) are not well adapted to this environment (climate and specific endemic vegetation), or iii) are not selected by traditional Malagasy beekeepers, A. m. unicolor being easy to handle compared to other much more aggressive subspecies like the Africanized honeybee (Ruttner 1988; Winston 1992).
Madagascar populations were highly different (F ST = 0.34) to the reference population (comprising A1 and A4 haplotypes). Furthermore, A. m. unicolor has a poor allelic diversity in terms of number of alleles for each population marker (N br = 2.47 to 3.88), in comparison to our reference population (N br = 12.57) or even with other studies performed on African honeybee populations with allelic diversity ranging from 7.9 (A. m. capensis) to 9 (Apis mellifera intermissa) and even 11 (A. m. scutellata) (for 7 to 12 microsatellite loci, with most loci being the same in both studies) (Estoup et al. 1995).
The levels of heterozygosity observed in Malagasy populations were also much lower than the reference population used in this study and those reported from African populations. Across Madagascar, levels varied from 0.34 to 0.47 (H e), while the reference population H e = 0.84, and the average He ranged between 0.78 and 0.90 (A. m. intermissa to A. m. scutellata) in African populations (Franck et al. 2001). In parallel, the lowest levels ranged from 0.26 to 0.66 in west Mediterranean subspecies (Apis mellifera iberiensis and Apis mellifera siciliana, respectively) (Garnery et al. 1998).
High levels of nuclear polymorphism in African populations have been explained by i) quaternary climate changes that could be responsible for honeybee subspecies diversification and expansion in Africa (Franck et al. 2001), ii) larger effective population size (Estoup et al. 1995), allowing more alleles to be maintained, and iii) the high migratory behavior of colonies which is typical for African honeybees south of the Sahara (Hepburn and Radloff 1998; Jaffe et al. 2009). Allelic richness within populations can also be increased by introgression of foreign genes into zones with other subspecies. Due to the lack of data on the biology of the Malagasy subspecies, it is difficult to compare effective population size of A. m. unicolor to other subspecies. However, its insular situation prevents frequent natural introductions and may in part explain the low nuclear polymorphism. Both the significant pairwise F ST values observed between neighboring sites (i.e., S34–S35 only 10 km apart, SD Table III) and the significant isolation by distance patterns observed among sites 500 km apart (Figures 1 and 2) suggest that gene flow is restricted. The larger variance of F ST at longer distances (>500 km, Figure 2d) indicates that at the island scale, the influence of genetic drift is strong relative to gene flow (Hutchison and Templeton 1999) and that problems of homoplasy could be more important (Jarne and Lagoda 1996). As observed in A. m. capensis from South Africa, A. m. unicolor populations may be less mobile than other African subspecies (Estoup et al. 1995) because of the topography of the island (coastal vs. Hauts Plateaux areas) and the climatic variations between regions. Indeed this was underlined with the results of Structure indicating genetic subclustering of the observed populations into at least two major clusters. The observed genetic subclustering did not match the distribution of the two honeybee ecotypes described by Ruttner (1988). Furthermore, we found such an admixture of genetic clusters between populations from different regions and the region surrounding the capital (on the Hauts Plateaux) that those ecotypes might have been mixed in the recent past. Indeed, in Madagascar, most goods pass through the capital, central market, and free commercial exchanges, which facilitate honeybee movement. Transport routes around the island are limited but all of them lead to the capital.
Wild populations, uninfluenced by beekeeping, exist in many regions of Africa, and honeybees from natural habitats have been shown to have a higher genetic diversity than managed populations (Allsopp 2004), so more intensive studies of wild colonies in protected and wild zones of Madagascar would be interesting to confirm or not our findings on the genetic diversity of A. m. unicolor.
Several clues and hypotheses point out the fact that A. m. unicolor might be derived from a recent (in geological time) colonization event of this continental island: i) relatively low mitochondrial and nuclear genetic diversity were found on A. m. unicolor in Madagascar, compared to other subspecies of the A lineage (Estoup et al. 1995; Franck et al. 1998, 2001). ii) The hypothesis on molecular data showing A lineage split from other lineages 6 million years ago with A. m. unicolor divergence from other subspecies more recently (1 million years ago) (Han et al. 2012). iii) The prehistoric breakup of the supercontinent Gondwana which separated Madagascar from mainland Africa is dated much earlier (around 135 million years ago; Rabinowitz et al. 1983) than the first honeybee species.
Nevertheless, its morphological (two ecotypes) and behavioral differences (one of the most gentle honeybees in the world (Ruttner 1988)) from other African honeybees suggest that, such as the flora of the island, A. m. unicolor seems to have evolved in relative isolation. Furthermore, the low genetic diversity observed, if confirmed in wild and conserved areas, might also be the result of over 1400 years of high deforestation rates and habitat fragmentation on the island (Campbell 1993; Gade 1996) which has been increasing over the last 50 years (Harper et al. 2007).
References
Alburaki, M., Moulin, S., Legout, H., Alburaki, A., Garnery, L. (2011) Mitochondrial structure of eastern honeybee populations from Syria, Lebanon and Iraq. Apidologie 42, 628–641
Alburaki, M., Bertrand, B., Legout, H., Moulin, S., Alburaki, A., Sheppard, W.S., Garnery, L. (2013) A fifth major genetic group among honeybees revealed in Syria. BMC Genet. 14(1), 117
Allsopp, M.H. (2004) Cape honeybee (Apis mellifera capensis Eschscholtz) and Varroa mite (Varroa destructor Anderson & Trueman) threats to honeybees and beekeeping in Africa. Int. J. Trop. Insect Sci. 24, 87–94
Batra, S. (1995) Bees and pollination in our changing environment. Apidologie 26, 361
Belkhir, K., Borsa, P., Chikhi, L., Raufaste, N., Bonhomme, F. (1996) GENETIX 4.05, logiciel sous Windows TM pour la génétique des populations. Laboratoire génome, populations, interactions. CNRS UMR 5000, 1996–2004
Campbell, G. (1993) The Structure of Trade in Madagascar. Int. J Afr. Hist. Stud. 26, 111–148
Chahbar, N., Munõz, I., Dall’Olio, R., De la Rua, P., Serrano, J., Doumandji, S. (2013) Population structure of north african honeybees is influenced by both biological and anthropogenic factors. J. Insect Conserv. 17, 385–392
Chapuis, M.-P., Estoup, A. (2007) Microsatellite null alleles and estimation of population differentiation. Mol. Biol. Evol. 24(3), 621–631
Clement, M., Posada, D., Crandall, K.A. (2000) TCS: a computer program to estimate gene genealogies. Mol. Ecol. 9(10), 1657–1659
De la Rua, P., Galian, J., Pedersen, B.V., Serrano, J. (2006) Molecular characterization and population structure of Apis mellifera from Madeira and the Azores. Apidologie 37, 699–708
De la Rua, P., Jaffe, R., Dall’Olio, R., Munoz, I., Serrano, J. (2009) Biodiversity, conservation and current threats to European honeybees. Apidologie 40(3), 263–284
Delatte, H., Virgilio, M., Simiand, C., Risterucci, A.M., de Meyer, M., Quilici, S. (2010) Isolation and characterization of microsatellite markers from Bactrocera cucurbitae (Coquillett). Mol. Ecol. Resour. 10, 576–579
El-Niweiri, M.A., Moritz, R.F. (2008) Mitochondrial discrimination of honeybees (Apis mellifera) of Sudan. Apidologie 39, 566–573
Eschscholtz, J.F. (1822) Entomographien: Erste Lieferung. Gedruckt und verlegt bei G. Reimer 42 p.
Estoup, A., Garnery, L., Solignac, M., Cornuet, J.M. (1995) Microsatellite Variation in Honey Bee (Apis-Mellifera L) Populations - Hierarchical Genetic-Structure and Test of the Infinite Allele and Stepwise Mutation Models. Genetics 140(2), 679–695
Evanno, G., Regnaut, S., Goudet, J. (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol. Ecol. 14(8), 2611–2620
Excoffier, L., Lischer, H.E.L. (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 10(3), 564–567
Franck, P., Garnery, L., Solignac, M., Cornuet, J.M. (1998) The origin of west European subspecies of honeybees (Apis mellifera): New insights from microsatellite and mitochondrial data. Evolution 52(4), 1119–1134
Franck, P., Garnery, L., Solignac, M., Cornuet, J.M. (2000) Molecular confirmation of a fourth lineage in honeybees from the Near East. Apidologie 31(2), 167–180
Franck, P., Garnery, L., Loiseau, A., Oldroyd, B.P., Hepburn, H.R., Solignac, M., Cornuet, J.M. (2001) Genetic diversity of the honeybee in Africa: microsatellite and mitochondrial data. Heredity 86, 420–430
Gade, D.W. (1996) Deforestation and its effects in highland Madagascar. Mt. Res. Dev. 16(2), 101–116
Ganzhorn, J.U., Lowry II, P.P., Schatz, G.E., Sommer, S. (2001) The biodiversity of Madagascar: one of the world’s hottest hotspots on its way out. Oryx 35(4), 346–348
Garnery, L., Cornuet, J.M., Solignac, M. (1992) Evolutionary history of the honeybee Apis mellifera inferred from mitochondrial DNA analysis. Mol. Ecol. 1(3), 145–154
Garnery, L., Solignac, M., Celebrano, G., Cornuet, J.M. (1993) A simple test using restricted PCR-amplified mitochondrial DNA to study the genetic structure of Apis mellifera L. Experientia 49(11), 1016–1021
Garnery, L., Franck, P., Baudry, E., Vautrin, D., Cornuet, J.-M., Solignac, M. (1998) Genetic diversity of the west European honeybee (Apis mellifera mellifera and A. m. iberica) I. Mitochondrial DNA. Gen. Sel. Evol. 30, S31–S47
Goudet J. (2001) Fstat, a Program to Estimate and Test Gene Diversities and Fixation Indices Version 2.9.3., Available from http://www.unil.ch/izea/softwares/fstat.html
Han, F., Wallberg, A., Webster, M.T. (2012) From where did the Western honeybee (Apis mellifera) originate? Ecol. Evol. 2(8), 1949–1957
Harper, G.J., Steininger, M.K., Tucker, C.J., Juhn, D., Hawkins, F. (2007) Fifty years of deforestation and forest fragmentation in Madagascar. Environ. Conserv. 34(4), 325
Hepburn, H.R., Radloff, S.E. (1998) Honeybees of Africa. Springer, Berlin, Heidelberg, New York. 379 p
Hutchison, D.W., Templeton, A.R. (1999) Correlation of pairwise genetic and geographic distance measure: inferring the relative influences of gene flow and drift on distribution of genetic variability. Evolution 53, 1898–1914
Jaffe, R., Dietemann, V., Crewe, R.M., Moritz, R.F.A. (2009) Temporal variation in the genetic structure of a drone congregation area: an insight into the population dynamics of wild African honeybees (Apis mellifera scutellata). Mol. Ecol. 18(7), 1511–1522
Jakobsson, M., Rosenberg, N.A. (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23(14), 1801–1806
Jarne, P., Lagoda, P. (1996) Microsatellites, from molecules to populations and back. Trends Ecol. Evol. 11(10), 424–429
Jombart, T. (2008) adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24(11), 1403–1405
Klein, A.M., Vaissiere, B.E., Cane, J.H., Steffan-Dewenter, I., Cunningham, S.A., Kremen, C., Tscharntke, T. (2007) Importance of pollinators in changing landscapes for world crops. Proc. R. Soc. B Biol. Sci. 274(1608), 303
Kumar, S., Nei, M., Dudley, J., Tamura, K. (2008) MEGA: a biologist-centric software for evolutionary analysis of DNA and protein sequences. Brief. Bioinform. 9(4), 299–306
Latreille, P.A. (1804) Notice des espèces d’abeilles vivant en grande société, ou abeilles proprement dites, et description d’espèces nouvelles. Ann. Mus. Natl. Hist. Nat. 5, 161–178
Lepeletier, A. (1836) Histoire Naturelle des Insectes. Hymenoptères, Paris, Roret. 644 p
Meixner, M.D., Leta, M.A., Koeniger, N., Fuchs, S. (2011) The honey bees of Ethiopia represent a new subspecies of Apis mellifera—Apis mellifera simensis n. ssp. Apidologie 42(3), 425–437
Myers, N., Mittermeier, R.A., Mittermeier, C.G., Da Fonseca, G.A., Kent, J. (2000) Biodiversity hotspots for conservation priorities. Nature 403, 853–858
Pritchard, J.K., Stephens, M., Donnelly, P. (2000) Inference of population structure using multilocus genotype data. Genetics 155(2), 945–959
Rabinowitz, P.D., Coffin, M.F., Falvey, D. (1983) The separation of Madagascar and Africa. Science 220, 67–69
Ralalaharisoa-Ramamonjisoa Z., Ralimananai H., Lobreau-Callen D. (1996) Comportement de butinage d’Apis mellifera var Unicolor (HYMENOPTERA, APIDAE) dans divers biotopes. Biogéographie de Madagascar, 517–522.
Rasolofoarivao, H., Clemencet, J., Raveloson-Ravaomanarivo, L.H., Razafindrazaka, D., Reynaud, B., Delatte, H. (2013) Spread and strain determination of Varroa destructor (Acari: Varroidea) in Madagascar since its first description in 2010. Exp. Appl. Acarol. 60(4), 521–530
Rosenberg, N.A. (2004) DISTRUCT: a program for the graphical display of population structure. Mol. Ecol. Notes 4(1), 137–138
Rousset, F. (2008) Genepop‘007: a complete reimplementation of the Genepop software for Windows and Linux. Mol. Ecol. Resour. 8, 103–106
Ruttner, F. (1975) Races of bees. In: Sons Dadant (ed.) The Hive and the Honeybee, pp. 19–38. Dadant, publ, Hamilton
Ruttner, F. (1988) Biogeography and taxonomy of honeybees. Springer, Berlin, Germany. 284 p
Ruttner, F., Tassencourt, I., Louveaux, J. (1978) Biometrical-statistical analysis of the geographic variability of Apis mellifera L. Apidologie 9, 363–381
Sheppard, W.S., Meixner, M.D. (2003) Apis mellifera pomonella, a new honeybee subspecies from Central Asia. Apidologie 34(4), 367–376
Sheppard, W., Arias, M., Grech, A., Meixner, M. (1997) Apis mellifera ruttneri, a new honeybee subspecies from Malta. Apidologie 28(5), 287–293
Solignac, M., Vautrin, D., Loiseau, A., Mougel, F., Baudry, E., Estoup, A., Garnery, L., et al. (2003) Five hundred and fifty microsatellite markers for the study of the honeybee (Apis mellifera L.) genome. Mol. Ecol. Notes 3(2), 307–311
Team R.D.C. (2005) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Weir, B.S., Cockerham, C.C. (1984) Estimating F-statistics for the analysis of population structure. Evolution 38(6), 1358–1370
Whitfield, C.W., Behura, S.K., Berlocher, S.H., Clark, A.G., Johnston, J.S., Sheppard, W.S., Smith, D.R., et al. (2006) Thrice out of Africa: ancient and recent expansions of the honeybee, Apis mellifera. Science 314, 642–645
Winston M. (1992) Killer bees. The Africanized honeybee in the Americas. Harvard University Press, 162 p.
Acknowledgments
We thank Ravelomanana A. for mapping sites with GIS and Simiand C. for technical help in the laboratory. We thank the following people for their help with data collection: Rousse P., Porphyre V. (and QualiREG networking), Borsa C., Mandirola V., Andrianaivoariseta N., Razafindrazaka D., Cattel J., Chesnais Q., ADEFA, and FENAM. We are grateful to the Malagasy beekeepers who participated in the study. We would like to thank Garnery L. for the fruitful discussions related to those genetic results. This work is part of the Ph.D. of Rasolofoarivao H., recipient of a grant of CIRAD-AIRD-Sud. Field work had been partly funded by CIRAD, the Enlargement and sustainability of the Plant Protection Network supported by the European Union, the French government, and the Région Réunion and the Département of la Réunion. In addition, we would like to thank the editor (M. Meixner) and the anonymous referees for their remarks that greatly improved our manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Manuscript editor: Marina Meixner
Diversité génétique de l’abeille endémique de Madagascar: Apis mellifera unicolor (Hymenoptera: Apidae)
Microsatellite / lignée africaine / ADNmt / aire de répartition / haplotype
Genetische Diversität der endemischen Honigbiene von Madagaskar, Apis mellifera unicolor (Hymenoptera: Apidae)
Apis mellifera unicolor / Mikrosatelliten / Afrikanische Lineage / genetische Diversität / mtDNA
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 1211 kb)
Rights and permissions
About this article
Cite this article
Rasolofoarivao, H., Clémencet, J., Techer, M.A. et al. Genetic diversity of the endemic honeybee: Apis mellifera unicolor (Hymenoptera: Apidae) in Madagascar. Apidologie 46, 735–747 (2015). https://doi.org/10.1007/s13592-015-0362-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13592-015-0362-1